

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0159.10
(353 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
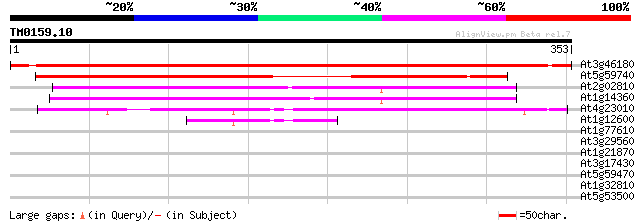
Score E
Sequences producing significant alignments: (bits) Value
At3g46180 unknown protein 559 e-160
At5g59740 protein serine /threonine kinase - like protein 369 e-102
At2g02810 unknown protein 142 3e-34
At1g14360 putative UDP-galactose transporter MSS4 135 3e-32
At4g23010 unknown protein 82 4e-16
At1g12600 hypothetical protein 50 2e-06
At1g77610 unknown protein 37 0.012
At3g29560 hypothetical protein 32 0.40
At1g21870 glucose 6 phosphate/phosphate translocator, putative 32 0.40
At3g17430 unknown protein 29 4.4
At5g59470 unknown protein 28 5.8
At1g32810 hypothetical protein 28 5.8
At5g53500 unknown protein 28 9.9
>At3g46180 unknown protein
Length = 347
Score = 559 bits (1441), Expect = e-160
Identities = 270/353 (76%), Positives = 310/353 (87%), Gaps = 6/353 (1%)
Query: 1 MAESPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHS 60
MAE S + + + KLWK +FA+SGIM+TLV YG+LQEKIMRVPYG +KEYFKHS
Sbjct: 1 MAEPDSVNEAKE----KKKKLWKAVFAISGIMLTLVIYGLLQEKIMRVPYGLKKEYFKHS 56
Query: 61 LFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPV 120
LFLVFCNR+TTSAVSA +LLASKK LDPVAP+YKYCL+SV+NILTTTCQYEALKYVSFPV
Sbjct: 57 LFLVFCNRLTTSAVSAAALLASKKVLDPVAPVYKYCLISVTNILTTTCQYEALKYVSFPV 116
Query: 121 QTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGREN 180
QTLAKCAKMIPVM+WGT+IMQK+Y+G DY++AFLVTLGCSVFIL+PAG+DISPY++GREN
Sbjct: 117 QTLAKCAKMIPVMVWGTLIMQKKYRGFDYLVAFLVTLGCSVFILFPAGDDISPYNKGREN 176
Query: 181 TVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIP 240
TVWGV LM GYLG DGFTSTFQDKLF+GY+MEIHNQIFYTT+CS ILS TGLILQGHL+P
Sbjct: 177 TVWGVSLMVGYLGFDGFTSTFQDKLFKGYNMEIHNQIFYTTICSSILSFTGLILQGHLLP 236
Query: 241 AIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWF 300
A++FV RH DC FDIALLSTVATASQFFISYTIRTFGALTFA IMTTRQL SIMLSC+WF
Sbjct: 237 AVDFVSRHRDCLFDIALLSTVATASQFFISYTIRTFGALTFAAIMTTRQLASIMLSCIWF 296
Query: 301 SHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDSIPLVQSGDSNNLKDNP 353
SHPLSWEQ IG+VIVFGSLYAK+F +K +K ++ +P + ++ LK NP
Sbjct: 297 SHPLSWEQCIGSVIVFGSLYAKTFVKKKSEKPPAAQELP--RDEEAQPLKGNP 347
>At5g59740 protein serine /threonine kinase - like protein
Length = 295
Score = 369 bits (947), Expect = e-102
Identities = 191/297 (64%), Positives = 216/297 (72%), Gaps = 49/297 (16%)
Query: 17 RDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSA 76
++NKLWKG+FAVSGIM TLV YGVLQEKIMRVPYG KE+FKHSLFLVFCNR+TTSAVSA
Sbjct: 12 KENKLWKGVFAVSGIMSTLVIYGVLQEKIMRVPYGVNKEFFKHSLFLVFCNRLTTSAVSA 71
Query: 77 GSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWG 136
G+LLASKK LDPVAP+YKYCL+SV+NILTTTCQYEALKYVSFPVQTLAKCAKMIPVM+WG
Sbjct: 72 GALLASKKVLDPVAPVYKYCLISVTNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMVWG 131
Query: 137 TIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDG 196
T+IMQK+Y+G DY++AFLVTLGCSVFIL+P
Sbjct: 132 TLIMQKKYKGFDYLVAFLVTLGCSVFILFP------------------------------ 161
Query: 197 FTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIA 256
NQIFYTT ++ GLILQGHL+PA++FV H DC
Sbjct: 162 ------------------NQIFYTTHYVNLMVDVGLILQGHLLPAVDFVSLHRDCLLPET 203
Query: 257 LLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAV 313
VATASQFFISYTIRTFGALTF + + L SIMLSC+WFSHPLSWEQ IG+V
Sbjct: 204 NNVQVATASQFFISYTIRTFGALTFVICLLIK-LASIMLSCIWFSHPLSWEQCIGSV 259
>At2g02810 unknown protein
Length = 332
Score = 142 bits (358), Expect = 3e-34
Identities = 101/296 (34%), Positives = 146/296 (49%), Gaps = 6/296 (2%)
Query: 28 VSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSAGSLLASKKALD 87
+SGI + GVLQE + +G ++ F+H FL + S + A +
Sbjct: 18 ISGIWSAYIYQGVLQETLSTKRFGPDEKRFEHLAFLNLAQSVVCLIWSYIMIKLWSNAGN 77
Query: 88 PVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMQKRYQGP 147
AP + Y ++N + EALKY+S+P Q LAK +KMIPVM+ GT++ RY P
Sbjct: 78 GGAPWWTYWSAGITNTIGPAMGIEALKYISYPAQVLAKSSKMIPVMLMGTLVYGIRYTFP 137
Query: 148 DYMLAFLVTLGCSVF-ILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDKLF 206
+YM FLV G S+F +L + + IS + N G L + L DGFT+ QD +
Sbjct: 138 EYMCTFLVAGGVSIFALLKTSSKTISKLA--HPNAPLGYALCSLNLAFDGFTNATQDSIA 195
Query: 207 RGY-DMEIHNQIFYTTLCSCILSLTGL--ILQGHLIPAIEFVYRHHDCFFDIALLSTVAT 263
Y E + + L I ++ + + QG AI+F H + +DI
Sbjct: 196 SRYPKTEAWDIMLGMNLWGTIYNMIYMFGLPQGIGFKAIQFCKLHPEAAWDILKYCICGA 255
Query: 264 ASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSL 319
Q FI TI FG+L TI TTR+ VSI++S V +PLS +QW +VFG L
Sbjct: 256 VGQNFIFMTISNFGSLANTTITTTRKFVSIVVSSVMSGNPLSLKQWGCVSMVFGGL 311
>At1g14360 putative UDP-galactose transporter MSS4
Length = 331
Score = 135 bits (341), Expect = 3e-32
Identities = 93/298 (31%), Positives = 144/298 (48%), Gaps = 6/298 (2%)
Query: 26 FAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSAGSLLASKKA 85
F V+GI + G+LQE + +G + F+H FL + S +
Sbjct: 16 FCVAGIWAAYIYQGILQETLSTKKFGEDGKRFEHLAFLNLAQNVICLVWSYIMIKLWSNG 75
Query: 86 LDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMQKRYQ 145
AP + Y ++N + EALKY+S+P Q LAK +KMIPVM+ G+++ RY
Sbjct: 76 GSGGAPWWTYWSAGITNTIGPAMGIEALKYISYPAQVLAKSSKMIPVMLMGSLVYGIRYT 135
Query: 146 GPDYMLAFLVTLGCSVF-ILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDK 204
P+Y+ FLV G S+F +L + + IS + +G+ + L DGFT+ QD
Sbjct: 136 LPEYLCTFLVAGGVSMFALLKTSSKTISKLAHPNAPLGYGLCFLN--LAFDGFTNATQDS 193
Query: 205 LFRGY-DMEIHNQIFYTTLCSCILSLTGL--ILQGHLIPAIEFVYRHHDCFFDIALLSTV 261
+ Y + + L I ++ + + G A++F +H + +DI +
Sbjct: 194 ITARYPKTNAWDIMLGMNLWGTIYNMVYMFGLPHGSGFEAVQFCKQHPEAAWDILMYCLC 253
Query: 262 ATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSL 319
Q FI TI FG+L TI TTR+ VSI++S V +PLS +QW +VFG L
Sbjct: 254 GAVGQNFIFLTISRFGSLANTTITTTRKFVSIVVSSVLSGNPLSSKQWGCVSMVFGGL 311
>At4g23010 unknown protein
Length = 345
Score = 82.0 bits (201), Expect = 4e-16
Identities = 81/345 (23%), Positives = 148/345 (42%), Gaps = 33/345 (9%)
Query: 18 DNKLWKG-IFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHS-----LFLVFCNRITT 71
D W+ + SG + GV +E + + YF LFL++ TT
Sbjct: 16 DKPTWQQFLICTSGFFFGYLVNGVCEEYVYNRLQFSFGWYFTFIQGFVYLFLIYLQGFTT 75
Query: 72 SAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIP 131
+ V P+ Y +S + + +L Y+++P Q + K K++P
Sbjct: 76 KHI--------------VNPMRTYVKLSAVLMGSHGLTKGSLAYLNYPAQIMFKSTKVLP 121
Query: 132 VMIWGTII--MQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMT 189
VMI G I ++++Y +Y+ AFL+ LG +F L A +SP ++ G++++T
Sbjct: 122 VMIMGAFIPGLRRKYPVHEYISAFLLVLGLILFTL--ADAQMSP-----NFSMIGIMMIT 174
Query: 190 GYLGCDGFTSTFQDKLFR-GYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRH 248
G L D F Q+ +F + +F +T+ ++L G + A +H
Sbjct: 175 GALIMDAFLGNLQEAIFTMNPETTQMEMLFCSTVVGLPFLFVPMVLTGEVFRAWTACAQH 234
Query: 249 HDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQ 308
+ + + Q + I FGA T A I T R+ V+++LS + F+ PL+ +
Sbjct: 235 PYVYGVLVFEAMATFIGQVSVLSLIALFGAATTALITTARKGVTLLLSYLIFTKPLTEQH 294
Query: 309 WIGAVIVFGSLYAK--SFTRKAPQKTTSSDSIPLVQSGDSNNLKD 351
G +++ + K KAP K + ++ + GD + +D
Sbjct: 295 GSGLLLIAMGIVLKMVPMDSKAPAKIPARPAV-RIAGGDGDREED 338
>At1g12600 hypothetical protein
Length = 155
Score = 50.1 bits (118), Expect = 2e-06
Identities = 29/97 (29%), Positives = 54/97 (54%), Gaps = 9/97 (9%)
Query: 112 ALKYVSFPVQTLAKCAKMIPVMIWGTII--MQKRYQGPDYMLAFLVTLGCSVFILYPAGE 169
+L Y+++P Q + K K++PVM+ G I ++++Y +Y+ A L+ +G +F L A
Sbjct: 33 SLAYLNYPAQIMFKSTKVLPVMVMGAFIPGLRRKYPVHEYISAMLLVIGLILFTL--ADA 90
Query: 170 DISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDKLF 206
SP ++ GV++++G L D F Q+ +F
Sbjct: 91 HTSP-----NFSIIGVMMISGALIMDAFLGNLQEAIF 122
>At1g77610 unknown protein
Length = 336
Score = 37.4 bits (85), Expect = 0.012
Identities = 28/114 (24%), Positives = 50/114 (43%), Gaps = 2/114 (1%)
Query: 203 DKLFRGYDMEIHNQIFYTT-LCSCILSLTGLILQGH-LIPAIEFVYRHHDCFFDIALLST 260
+ L GY + N ++Y + IL + L+L+G ++ E I
Sbjct: 175 ESLLHGYKFDSINTVYYMAPFATMILGIPALLLEGSGILSWFEAHPAPWSALIIILSSGV 234
Query: 261 VATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVI 314
+A F I Y I + A+TF + V++M+S + F +P+S+ +G I
Sbjct: 235 LAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVMVSWLIFRNPISYMNAVGCGI 288
>At3g29560 hypothetical protein
Length = 120
Score = 32.3 bits (72), Expect = 0.40
Identities = 26/79 (32%), Positives = 37/79 (45%), Gaps = 8/79 (10%)
Query: 20 KLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNR--ITTSAVSAG 77
K WK + + I T Y V + + G Q SL+ F +R +T+SAV AG
Sbjct: 3 KFWK-VQPILPIPETKTQYRVASDNL-----GFQASVLSASLYSGFAHRSSLTSSAVLAG 56
Query: 78 SLLASKKALDPVAPIYKYC 96
SLL+S A P+ +C
Sbjct: 57 SLLSSPLARTPLLSYLDHC 75
>At1g21870 glucose 6 phosphate/phosphate translocator, putative
Length = 341
Score = 32.3 bits (72), Expect = 0.40
Identities = 27/116 (23%), Positives = 53/116 (45%), Gaps = 6/116 (5%)
Query: 203 DKLFRGYDMEIHNQIFYTT-LCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALL--- 258
+ L GY + N ++Y + IL L +L+ + I +++ H + + +L
Sbjct: 181 ESLLHGYKFDSINTVYYMAPFATMILGLPAFLLERNGI--LDWFEAHPSPWSALIILFNS 238
Query: 259 STVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVI 314
+A F I Y I++ A+TF + V++ +S + F +P+S +G I
Sbjct: 239 GVLAFCLNFSIFYVIQSTTAVTFNVAGNLKVAVAVFVSWMIFRNPISPMNAVGCGI 294
>At3g17430 unknown protein
Length = 375
Score = 28.9 bits (63), Expect = 4.4
Identities = 50/215 (23%), Positives = 79/215 (36%), Gaps = 22/215 (10%)
Query: 92 IYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYML 151
IY C+V +S ++ + Y+ V + ++PV T IM ++
Sbjct: 78 IYATCVVPISAFFASSLWFGNTAYLHISVAFIQMLKALMPV---ATFIMA--------VV 126
Query: 152 AFLVTLGCSVF---ILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDKLF-- 206
C VF +L G IS Y N V V +TG + + L
Sbjct: 127 CGTDKPRCDVFSNMLLVSVGVVISSYGEIHFNIVGTVYQVTG-IFAEALRLVLTQVLLQK 185
Query: 207 RGYDMEIHNQIFYTTLCSCI-LSLTGLILQGHLIPAIEFVYRHHDCFFDIALLSTVATAS 265
+G + ++Y CS + L+L +L+ + + + FF AL A A
Sbjct: 186 KGLTLNPITSLYYIAPCSFVFLALPWYVLEKPTMEVSQIQFNFW-IFFSNAL---CALAL 241
Query: 266 QFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWF 300
F I I GA+T + + I LS V F
Sbjct: 242 NFSIFLVIGRTGAVTIRVAGVLKDWILIALSTVIF 276
>At5g59470 unknown protein
Length = 148
Score = 28.5 bits (62), Expect = 5.8
Identities = 15/42 (35%), Positives = 23/42 (54%), Gaps = 6/42 (14%)
Query: 276 FGALTFATIMTTRQLVSIMLSCVW-FSHPLSWEQWIGAVIVF 316
FG L F I I+++C++ FS PLS W+ A++ F
Sbjct: 91 FGELAFLLIQAL-----ILVACIYYFSQPLSVTTWVKAILYF 127
>At1g32810 hypothetical protein
Length = 654
Score = 28.5 bits (62), Expect = 5.8
Identities = 15/38 (39%), Positives = 23/38 (60%), Gaps = 2/38 (5%)
Query: 313 VIVFGSLYAKSFTRKAPQKTTSSDSIPLV--QSGDSNN 348
V+ G + S T K+P+ +TS +SIP + Q GD +N
Sbjct: 148 VVCIGKTSSSSATEKSPKPSTSRNSIPGLKQQPGDDDN 185
>At5g53500 unknown protein
Length = 654
Score = 27.7 bits (60), Expect = 9.9
Identities = 13/32 (40%), Positives = 19/32 (58%), Gaps = 2/32 (6%)
Query: 172 SPYSRGRENTVWGVLLMTGYLGCDGFTSTFQD 203
S Y R + WG++++TG G DG TFQ+
Sbjct: 618 SSYQRASSSLSWGMVIVTG--GWDGQIRTFQN 647
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.137 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,593,705
Number of Sequences: 26719
Number of extensions: 291174
Number of successful extensions: 949
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 931
Number of HSP's gapped (non-prelim): 14
length of query: 353
length of database: 11,318,596
effective HSP length: 100
effective length of query: 253
effective length of database: 8,646,696
effective search space: 2187614088
effective search space used: 2187614088
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0159.10


