

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0157a.4
(594 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
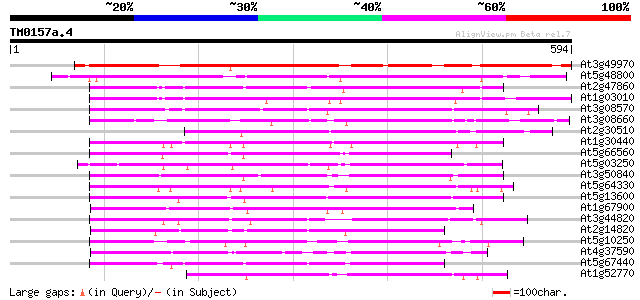
Score E
Sequences producing significant alignments: (bits) Value
At3g49970 putative protein 437 e-123
At5g48800 non-phototropic hypocotyl-like protein 327 2e-89
At2g47860 unknown protein 297 1e-80
At1g03010 hypothetical protein 293 2e-79
At3g08570 hypothetical protein 269 4e-72
At3g08660 putative non-phototropic hypocotyl 247 1e-65
At2g30510 unknown protein 245 4e-65
At1g30440 non-phototropic hypocotyl, putative 244 1e-64
At5g66560 unknown protein 241 8e-64
At5g03250 photoreceptor-interacting protein - like 229 4e-60
At3g50840 putative protein 228 6e-60
At5g64330 non-phototropic hypocotyl 3 (NPH3) 227 1e-59
At5g13600 photoreceptor-interacting protein-like; non-phototropi... 226 3e-59
At1g67900 unknown protein 224 8e-59
At3g44820 non-phototropic hypocotyl 3-like protein 215 5e-56
At2g14820 hypothetical protein 212 5e-55
At5g10250 non-phototropic hypocotyl 3-like protein 209 3e-54
At4g37590 unknown protein 196 4e-50
At5g67440 photoreceptor-interacting protein-like 195 5e-50
At1g52770 putative non-phototropic hypocotyl 189 5e-48
>At3g49970 putative protein
Length = 526
Score = 437 bits (1125), Expect = e-123
Identities = 260/540 (48%), Positives = 345/540 (63%), Gaps = 83/540 (15%)
Query: 69 DVTVQIDNASFSLHKMLRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHM 128
D++ +I+ + + ML C AEYLEMTE++S+ NLV + YLNEV L+++S +V VL
Sbjct: 56 DISFEINTENIA---MLLCAAEYLEMTEEHSVENLVETIEVYLNEVILKSLSKSVKVLQK 112
Query: 129 PETLLQVAEKAKLVNRCIDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVHWW 188
+ LL +AE+ +LV+RCID+IAY ++SQ +V WW
Sbjct: 113 SQDLLPIAERVRLVDRCIDSIAYAICQESQ---------------------SNEDIVDWW 151
Query: 189 GEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGL-----------EIFGK 237
+DL VL+ID+F+RVL+AM+ RGFK++ LGP++ LYA+K+L GL +IFGK
Sbjct: 152 ADDLAVLKIDMFRRVLVAMIARGFKRYSLGPVLKLYAEKALRGLVRFLNFLTEQCDIFGK 211
Query: 238 GRKELEPQHEHEKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRV 297
K++E + EHEKR+++ET+VSLLPRE+N++SVS LS+LLRAAIYLETTVACRLDLEKR+
Sbjct: 212 EAKKMEAEQEHEKRLILETIVSLLPRERNSVSVSFLSILLRAAIYLETTVACRLDLEKRM 271
Query: 298 AMQLGQAVLDDLLIPSYSFPG-NTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQ 356
+QL QAV+DDLLIP YSF G NT+ DVDTVQRI+++YLEFE+ +S A D
Sbjct: 272 GLQLRQAVIDDLLIPYYSFNGDNTMLDVDTVQRILMNYLEFEVEGNSADFASD------- 324
Query: 357 RGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFL 416
+G+LME YLAEIA+DRN+ +KFI AE IP+QS YRAIDI+LK HP +
Sbjct: 325 -----IGELMETYLAEIASDRNINFAKFIGFAECIPKQS-----RMYRAIDIFLKTHPNI 374
Query: 417 SEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDG 476
SE+EKK VCS+M C+KLSRD AHAAQNDR Q++L + + +
Sbjct: 375 SEVEKKKVCSLMDCKKLSRDVYAHAAQNDRF------------QENLSNSDSPAPATAEK 422
Query: 477 TSIPDKLNVYSAELNPVSKVISNLRRENEELKREIVKLKMKLQEIEK-PAHESSAPSSPL 535
T P +L+ Y EL S L REN+ LK E++K+KMK +E+EK A E + S
Sbjct: 423 TLSPPELSSYKNEL-------SKLNRENQYLKLELLKVKMKFKELEKEKAFEVMSGSDCS 475
Query: 536 ISAYSPSVNKPPLPRKSFMKSVSRKLGRLY-PFSRAAADTVTTPFKDRLKPEKKIRHSIS 594
S + SV KP LPRKSF+ SVS+KLG+L PF T K K RHSIS
Sbjct: 476 SSVSTASVAKPRLPRKSFINSVSQKLGKLINPFGLKQGQT---------KQPKSRRHSIS 526
>At5g48800 non-phototropic hypocotyl-like protein
Length = 614
Score = 327 bits (837), Expect = 2e-89
Identities = 216/613 (35%), Positives = 329/613 (53%), Gaps = 97/613 (15%)
Query: 45 RKKELKSSAMKKTSEWNFPQEIPSDVTVQIDNASFSLHK---------MLRCVAE----- 90
+K L S+ + SEW F +++PSD+T++++ +F+LHK + R VAE
Sbjct: 21 QKLSLAKSSRQSCSEWIF-RDVPSDITIEVNGGNFALHKFPLVSRSGRIRRIVAEHRDSD 79
Query: 91 -------------------------------------------YLEMTEDNSIGNLVGRS 107
YLEMTE+ S NL R+
Sbjct: 80 ISKVELLNLPGGAETFELAAKFCYGINFEITSSNVAQLFCVSDYLEMTEEYSKDNLASRT 139
Query: 108 DFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNRCIDAIAYMASKQSQLCSPARSQC 167
+ YL + + + V VL E LL +A++ ++ RCIDAIA A + S +R +
Sbjct: 140 EEYLESIVCKNLEMCVQVLKQSEILLPLADELNIIGRCIDAIASKACAEQIASSFSRLEY 199
Query: 168 SSDG-VMSSSMASHRRPVVHWWGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQ 226
SS G + S WW EDL+VLRID++QRV+ AM RG + +G ++ YA+
Sbjct: 200 SSSGRLHMSRQVKSSGDGGDWWIEDLSVLRIDLYQRVMNAMKCRGVRPESIGASLVSYAE 259
Query: 227 KSLEGLEIFGKGRKELEPQHEHEKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETT 286
+EL + EHE + +VET+V+LLP E + +S L LLR A+ L+T+
Sbjct: 260 -------------RELTKRSEHE-QTIVETIVTLLPVENLVVPISFLFGLLRRAVILDTS 305
Query: 287 VACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSS 346
V+CRLDLE+R+ QL A LDDLLIPS+ G+TLFD+DTV RI+V++ + + G+ S
Sbjct: 306 VSCRLDLERRLGSQLDMATLDDLLIPSFRHAGDTLFDIDTVHRILVNFSQ-QGGDDSEDE 364
Query: 347 AD----DEYFSPSQRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGK 402
D SPSQ + +V KL+++YLAEIA D NL SKF+ IAE +P + DG
Sbjct: 365 ESVFECDSPHSPSQTAMFKVAKLVDSYLAEIAPDANLDLSKFLLIAEALPPHARTLHDGL 424
Query: 403 YRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKH 462
YRAID+YLKAH LS+ +KK + ++ QKLS++A AHAAQN+RLP+Q+++QVL F+Q
Sbjct: 425 YRAIDLYLKAHQGLSDSDKKKLSKLIDFQKLSQEAGAHAAQNERLPLQSIVQVLYFEQLK 484
Query: 463 LRETMNDGGVNWDGTSIPDKLNVYSAELNPVSKVIS------NLRRENEELKREIVKLKM 516
LR ++ + + + + +S +S +LRREN ELK E+ +L+M
Sbjct: 485 LRSSLCSSYSDEEPKPKQQQQQSWRINSGALSATMSPKDNYASLRRENRELKLELARLRM 544
Query: 517 KLQEIEKPAHESSAPSSPLISAYSPSVNKPPLPRKSFMKSVSRKLGRLYPFSRAAADTVT 576
+L ++EK E + ++S + FM S S+K+G+L F +++ +
Sbjct: 545 RLNDLEK---EHICMKRDMQRSHS----------RKFMSSFSKKMGKLSFFGHSSSRGSS 591
Query: 577 TPFKDRLKPEKKI 589
+P K + + K+
Sbjct: 592 SPSKQSFRTDSKV 604
>At2g47860 unknown protein
Length = 635
Score = 297 bits (760), Expect = 1e-80
Identities = 182/463 (39%), Positives = 269/463 (57%), Gaps = 35/463 (7%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRCVA YLEMTED S NL R++ YL + IS +++VLH E LL VAE+ LV R
Sbjct: 110 LRCVAHYLEMTEDLSEKNLEARTEAYLKDSIFNDISNSITVLHSCERLLPVAEEINLVGR 169
Query: 145 CIDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVHWWGEDLTVLRIDIFQRVL 204
++AIA A K+ QL S D S + +P WWG L +L++D FQRVL
Sbjct: 170 LVNAIAVNACKE-QLAS---GLLKLDQSFSCGVPETAKPC-DWWGRSLPILKLDFFQRVL 224
Query: 205 IAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVSLLPRE 264
AM +G + I+M YA+KSL+ + + + + Q + +R+V+E +V LLP +
Sbjct: 225 SAMKSKGLNHDIISDILMSYARKSLQIIREPNLVKSDSDLQRK--QRIVLEAVVGLLPTQ 282
Query: 265 KNAMSV--SSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLF 322
N S+ S LS LL+ AI T+V+CR DLE+R++ L QA+L+D+LIP+ ++
Sbjct: 283 ANKSSIPISFLSSLLKTAIGSGTSVSCRSDLERRISHLLDQAILEDILIPANI---GAMY 339
Query: 323 DVDTVQRIVVSYLEFEIGNHSVSSADDEYF---------------SPSQRGIVRVGKLME 367
D D+VQRI +L + + D+E SP Q I +V KLM+
Sbjct: 340 DTDSVQRIFSMFLNLDECEYRDDDDDEEDAVDESEMAMYDFEGAESPKQSSIFKVSKLMD 399
Query: 368 NYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSV 427
+YLAE+A D +LP SKFI++AEL+P+ + DG YRA+DI+LK HP + + E+ +C
Sbjct: 400 SYLAEVALDSSLPPSKFIALAELLPDHARVVCDGLYRAVDIFLKVHPHMKDSERYRLCKT 459
Query: 428 MHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTS-----IPDK 482
+ C+KLS+DA +HAAQN+RLPVQ +QVL ++Q L+ M GG P++
Sbjct: 460 VSCKKLSQDASSHAAQNERLPVQIAVQVLFYEQTRLKNAMTSGGGTGGSNQSQFFLFPNR 519
Query: 483 --LNVYSAELNPVSKVISNLRRENEELKREIVKLKMKLQEIEK 523
+ S ++P S +RREN EL+ E+ +++M+L ++EK
Sbjct: 520 SGSGMASGAISPRDNYAS-VRRENRELRLEVARMRMRLTDLEK 561
>At1g03010 hypothetical protein
Length = 634
Score = 293 bits (749), Expect = 2e-79
Identities = 205/543 (37%), Positives = 300/543 (54%), Gaps = 50/543 (9%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC + YLEMTED S NL +++ +L E +I ++ VLH ETL+ V+E LVNR
Sbjct: 109 LRCASHYLEMTEDFSEENLASKTEHFLKETIFPSILNSIIVLHHCETLIPVSEDLNLVNR 168
Query: 145 CIDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVHWWGEDLTVLRIDIFQRVL 204
I A+A A K+ QL S D S + + P+ WWG+ L VL +D FQRV+
Sbjct: 169 LIIAVANNACKE-QLTS---GLLKLDYSFSGTNIEPQTPL-DWWGKSLAVLNLDFFQRVI 223
Query: 205 IAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFG-KGRKE--LEPQHEHEKRVVVETLVSLL 261
A+ +G Q + I++ Y KSL+GL + K KE L+ + + ++R++VET+V LL
Sbjct: 224 SAVKSKGLIQDVISKILISYTNKSLQGLIVRDPKLEKERVLDSEGKKKQRLIVETIVRLL 283
Query: 262 PREKNAMSV-----SSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIP-SYS 315
P + SV SSL ++ A +T +CR DLE+R+ +QL QA+L+D+LIP + +
Sbjct: 284 PTQGRRSSVPMAFLSSLLKMVIATSSSASTGSCRSDLERRIGLQLDQAILEDVLIPINLN 343
Query: 316 FPGNTLFDVDTVQRIVVSYLEF-----EIGNHSVSSAD--------DEYFSPSQRGIVRV 362
NT++D+D++ RI +L E +H + D D SP Q I++V
Sbjct: 344 GTNNTMYDIDSILRIFSIFLNLDEDDEEEEHHHLQFRDETEMIYDFDSPGSPKQSSILKV 403
Query: 363 GKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKK 422
KLM+NYLAEIA D NL SKFI++AEL+P+ + DG YRA+DIYLK HP + + E+
Sbjct: 404 SKLMDNYLAEIAMDPNLTTSKFIALAELLPDHARIISDGLYRAVDIYLKVHPNIKDSERY 463
Query: 423 NVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDG------GVNWDG 476
+C + QKLS++A +HAAQN+RLPVQ +QVL F+Q LR M+ N +
Sbjct: 464 RLCKTIDSQKLSQEACSHAAQNERLPVQMAVQVLYFEQIRLRNAMSSSIGPTQFLFNSNC 523
Query: 477 TSIPDK--LNVYSAELNPVSKVISNLRRENEELKREIVKLKMKLQEIEKPAHESSAPSSP 534
P + S ++P S +RREN ELK E+ +++M+L ++EK
Sbjct: 524 HQFPQRSGSGAGSGAISPRDNYAS-VRRENRELKLEVARMRMRLTDLEKDH--------- 573
Query: 535 LISAYSPSVNKPPLPR--KSFMKSVSRKLGRLYPFSRAAAD-TVTTPFKDRLKPEKKIRH 591
IS V P + KSF K +S KL L+ FS + + R ++K RH
Sbjct: 574 -ISIKQELVKSNPGTKLFKSFAKKIS-KLNSLFSFSSLKPSLSGKASSESRFLFQRKRRH 631
Query: 592 SIS 594
S+S
Sbjct: 632 SVS 634
>At3g08570 hypothetical protein
Length = 612
Score = 269 bits (687), Expect = 4e-72
Identities = 177/498 (35%), Positives = 281/498 (55%), Gaps = 32/498 (6%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLL--QVAEKAKLV 142
+RC A YLEMTED NL+ R++ YL +VA ++ +V VL ETL +AE A +
Sbjct: 107 IRCAAGYLEMTEDFKEENLIARTETYLEQVAFRSLEKSVEVLCSCETLYPQDIAETAHIP 166
Query: 143 NRCIDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVHWWGEDLTVLRIDIFQR 202
+RC++AIA A ++ + +R + G S + P WW EDL+ LRID + R
Sbjct: 167 DRCVEAIAVNACREQLVLGLSRL---NRGTESGELKRGDSP--EWWIEDLSALRIDYYAR 221
Query: 203 VLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVSLLP 262
V+ AM G + + +M YAQ+SL+G+ + K E+E+R V+E +VSL P
Sbjct: 222 VVSAMARTGLRSESIITSLMHYAQESLKGIRNCKERTKLDSGTFENEQRNVLEAIVSLFP 281
Query: 263 REKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLF 322
+ + +S L +LR I + ++CRL+LE+R+A QL LDDLLIP G++++
Sbjct: 282 NDN--VPLSFLFGMLRVGITINVAISCRLELERRIAQQLETVSLDDLLIPVVR-DGDSMY 338
Query: 323 DVDTVQRIVVSYL-----EFEIGNHSVSSADDEYFSPSQ--RGIVRVGKLMENYLAEIAA 375
DVDTV RI+V +L E E + E + S +++VG++M+ YLAEIA
Sbjct: 339 DVDTVHRILVCFLKKIEEEEEYDEDCCYENETENLTGSMCHSSLLKVGRIMDAYLAEIAP 398
Query: 376 DRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSR 435
D L KF+++ E++P+ + +DG YRAID++LK HP L+E E K++C + QKLS+
Sbjct: 399 DPCLSLHKFMALIEILPDYARVMDDGLYRAIDMFLKGHPSLNEQECKSLCKFIDTQKLSQ 458
Query: 436 DARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTSIPDKLNVYSAELNPVS- 494
+A H AQNDRLP+Q V++VL +Q ++ M+ G + +G + + + VS
Sbjct: 459 EACNHVAQNDRLPMQMVVRVLYSEQLRMKNVMS--GESGEGLLLSSQKHSSENPSRAVSP 516
Query: 495 -KVISNLRRENEELKREIVKLKMKLQEIEKP-------AHESSAPSSPLISAYSPSVNKP 546
++LRREN ELK EI +++++L E+EK E S L+++ S + +
Sbjct: 517 RDTYASLRRENRELKLEISRVRVRLTELEKEQILMKQGMMEKSGHGGTLLTSLSKGIGRI 576
Query: 547 PL----PRKSFMKSVSRK 560
+ P + +++ +RK
Sbjct: 577 SIFGGGPTEGKLRNANRK 594
>At3g08660 putative non-phototropic hypocotyl
Length = 582
Score = 247 bits (631), Expect = 1e-65
Identities = 178/519 (34%), Positives = 280/519 (53%), Gaps = 58/519 (11%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC A YLEMTED NL+ R++ YL+++ + +V VL ET ++AE ++ +R
Sbjct: 109 LRCAAGYLEMTEDYKEQNLIFRAENYLDQIVFRSFHESVLVLCSCETQ-EIAETYEIPDR 167
Query: 145 CIDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVHWWGEDLTVLRIDIFQRVL 204
C++AIA A C V S R + W E+L+ L ID + +V+
Sbjct: 168 CVEAIAMNA-------------CRKQLVSGLSEELKGRDCLEMWTEELSALGIDYYVQVV 214
Query: 205 IAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVSLLPR- 263
AM + + ++ YA+ SL+G+ ++ E+R +VE +V+LLP
Sbjct: 215 SAMARLSVRSESIVASLVHYAKTSLKGII----------DRNCQEQRKIVEAMVNLLPND 264
Query: 264 EKNAMSVSSLSM-----LLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPG 318
EK + S+S + + +L+ ++ ++CRL+LE+R+ QL A LDDLLIPS
Sbjct: 265 EKGSYSLSIIPLGFLFGMLKVGTIIDIEISCRLELERRIGHQLETASLDDLLIPSVQNE- 323
Query: 319 NTLFDVDTVQRIVVSYLEFEIGNHSVSSADDE--YFSPS---QRGIVRVGKLMENYLAEI 373
++++DVDTV RI+ +LE + DDE Y S S +++VG++M+ YL EI
Sbjct: 324 DSMYDVDTVHRILTFFLE------RIEEEDDECGYDSDSTGQHSSLLKVGRIMDAYLVEI 377
Query: 374 AADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKL 433
A D L KF +I E +PE S +DG YRAID+YLKAHP L+E E+K +C+ + C+KL
Sbjct: 378 APDPYLSLHKFTAIIETLPEHSRIVDDGIYRAIDMYLKAHPLLTEEERKKLCNFIDCKKL 437
Query: 434 SRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTSIPDKLNVYSAELNPV 493
S++A H AQNDRLPVQ V++VL +Q L++ ++ G + +G+ + V S ++P
Sbjct: 438 SQEASNHVAQNDRLPVQMVVRVLYTEQLRLKKALS--GDSEEGSWVLPS-GVQSRAVSP- 493
Query: 494 SKVISNLRRENEELKREIVKLKMKLQEIEKPAHESSAPSSPLISAYSPSVNKPPLPRKSF 553
+ LRREN ELK EI ++++++ E+EK + + K +F
Sbjct: 494 RDTYAALRRENRELKLEISRMRVRVSELEKEHNLMK----------HEMMEKSGNNGGTF 543
Query: 554 MKSVSRKLGRLYPFSRAAADTVTTPFKDRLKPEKKIRHS 592
+ S+S+ +GR+ F V K R E+K S
Sbjct: 544 LTSLSKGIGRIATFGGETRQKVNR--KSRSVSERKSSRS 580
>At2g30510 unknown protein
Length = 415
Score = 245 bits (626), Expect = 4e-65
Identities = 151/396 (38%), Positives = 231/396 (58%), Gaps = 19/396 (4%)
Query: 186 HWWGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGL--EIFGKGRKELE 243
+WW E+L +L +D F V+ +M RG K L I+ Y +KSL L + G+G K +
Sbjct: 9 NWWTEELCILDVDFFSDVVSSMKQRGVKPSSLASAIITYTEKSLRDLVRDHSGRGVKYSD 68
Query: 244 P-----QHEHEKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVA 298
P ++R +V+++VSLLP +K V+ L LLR A++L+T++ C+ +LEKR++
Sbjct: 69 PGDNESDERSQQRDLVQSIVSLLPSDKGLFPVNFLCSLLRCAVFLDTSLTCKNELEKRIS 128
Query: 299 MQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRG 358
+ L +DDLLIPS+++ G L D+D+V+RI+ +++E E N V + D
Sbjct: 129 VVLEHVSVDDLLIPSFTYDGERLLDLDSVRRIISAFVEKE-KNVGVFNGGDFNRGVCSVS 187
Query: 359 IVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSE 418
+ RV K +++YLAEIA +L SKF +IA L+P+ + ++D YRAIDI+LKAHP L E
Sbjct: 188 LQRVAKTVDSYLAEIATYGDLTISKFNAIANLVPKSARKSDDDLYRAIDIFLKAHPNLDE 247
Query: 419 MEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTS 478
+E++ VCS M KLS DAR HA+QN RLPV VL L + Q LR + +
Sbjct: 248 IEREKVCSSMDPLKLSYDARLHASQNKRLPVNIVLHALYYDQLKLRSGVAEQEER-AVVV 306
Query: 479 IPDKLNVYSAELNPVSKVISNLRRENEELKREIVKLKMKLQEIEKPAHESSAPSSPLISA 538
+P+ L S + + L +ENE L+ E++K+KM + +++K + + A SS S
Sbjct: 307 LPEALKTRSQ-----LQADTTLAKENEALRSELMKMKMYVSDMQKNKNGAGASSSNSSSL 361
Query: 539 YSPSVNKPPLPRKSFMKSVSRKLGRLYPFSRAAADT 574
S +K +F SVS+KLG+L PF + DT
Sbjct: 362 VSSKKSK-----HTFFSSVSKKLGKLNPFKNGSKDT 392
>At1g30440 non-phototropic hypocotyl, putative
Length = 665
Score = 244 bits (622), Expect = 1e-64
Identities = 163/475 (34%), Positives = 261/475 (54%), Gaps = 39/475 (8%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC AE+LEMTE++ GNL+ +++ + N+V L++ ++ LH + +L+ A++ + +
Sbjct: 104 LRCAAEHLEMTEEHGEGNLISQTETFFNQVVLKSWKDSIKALHSCDEVLEYADELNITKK 163
Query: 145 CIDAIAYMASKQSQLCS--------PARSQCSS---DGVMSSSMASHRRPVVHWWGEDLT 193
CI+++A AS L P +S S +G+ + + H WW ED +
Sbjct: 164 CIESLAMRASTDPNLFGWPVVEHGGPMQSPGGSVLWNGISTGARPKHTSS--DWWYEDAS 221
Query: 194 VLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGL-------EIFGKGRKELEPQH 246
+L +F+R++ M RG ++ + + Y +K L GL E G+ L +
Sbjct: 222 MLSFPLFKRLITVMESRGIREDIIAGSLTYYTRKHLPGLKRRRGGPESSGRFSTPLGSGN 281
Query: 247 ---EHEKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQ 303
E E++ ++E + LL +K + +LR A L+ + C +LEKR+ MQL Q
Sbjct: 282 VLSEEEQKNLLEEIQELLRMQKGLVPTKFFVDMLRIAKILKASPDCIANLEKRIGMQLDQ 341
Query: 304 AVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFE------IGNHSVSSADDEYFSPSQR 357
A L+DL++PS+S TL+DVD+VQRI+ +L + +G+ SS DD S +
Sbjct: 342 AALEDLVMPSFSHTMETLYDVDSVQRILDHFLGTDQIMPGGVGS-PCSSVDDGNLIGSPQ 400
Query: 358 GIV---RVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHP 414
I V KL++ YLAE+A D NL KF ++A IPE + +DG YRAIDIYLK HP
Sbjct: 401 SITPMTAVAKLIDGYLAEVAPDVNLKLPKFQALAASIPEYARLLDDGLYRAIDIYLKHHP 460
Query: 415 FLSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGV-- 472
+L+E E++N+C ++ CQKLS +A HAAQN+RLP++ ++QVL F+Q LR ++ +
Sbjct: 461 WLAETERENLCRLLDCQKLSLEACTHAAQNERLPLRIIVQVLFFEQLQLRTSVAGCFLVS 520
Query: 473 -NWDGTSIPDKLNVYSAELNP---VSKVISNLRRENEELKREIVKLKMKLQEIEK 523
N DG S + Y N + REN+ LK + ++M++ E+EK
Sbjct: 521 DNLDGGSRQLRSGGYVGGPNEGGGGGGGWATAVRENQVLKVGMDSMRMRVCELEK 575
>At5g66560 unknown protein
Length = 668
Score = 241 bits (615), Expect = 8e-64
Identities = 149/404 (36%), Positives = 232/404 (56%), Gaps = 28/404 (6%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC AE+LEMTE+ S NL+ +++ +L+ +++ ++ L E++ +A + +
Sbjct: 139 LRCAAEHLEMTEEYSPDNLISKTERFLSHSVYKSLRESIKALKACESVSPLAGSLGITEQ 198
Query: 145 CIDAIAYMASKQSQLC--------------SPARSQCSSDGVMSSSMASHRRPVVHWWGE 190
CID+I AS S Q G S R + W E
Sbjct: 199 CIDSIVSRASSADPSLFGWPVNDGGGRGNISATDLQLIPGGAAKSRKKPSRDSNMELWFE 258
Query: 191 DLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQH---- 246
DLT L + IF+ V+++M + ++ YA+K + G I RK
Sbjct: 259 DLTQLSLPIFKTVILSMRSGDLSSDIIESCLICYAKKHIPG--ILRSNRKPPSSSSTAVS 316
Query: 247 EHEKRVVVETLVSLLPREKNAMSVSS--LSMLLRAAIYLETTVACRLDLEKRVAMQLGQA 304
E+E+R ++ET+ S LP +K+++S ++ L LLR AI L CR LE+++ QL +A
Sbjct: 317 ENEQRELLETITSNLPLDKSSISSTTRFLFGLLRTAIILNAAEICRDLLERKIGSQLERA 376
Query: 305 VLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLE-FEIGNHSVSSADDEYFSPSQRGIVRVG 363
LDDLL+PSYS+ TL+DVD V+RI+ +L+ E N ++ D + SPS ++ VG
Sbjct: 377 TLDDLLVPSYSYLNETLYDVDLVERILGHFLDTLEQSNTAIVEVDGK--SPS---LMLVG 431
Query: 364 KLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKN 423
KL++ +LAEIA+D NL + KF ++A +P+Q+ +DG YRA+D+YLKAHP++SE E++
Sbjct: 432 KLIDGFLAEIASDANLKSDKFYNLAISLPDQARLYDDGLYRAVDVYLKAHPWVSEAEREK 491
Query: 424 VCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETM 467
+C VM CQKL+ +A HAAQN+RLP++ V+QVL F+Q LR +
Sbjct: 492 ICGVMDCQKLTLEACTHAAQNERLPLRAVVQVLFFEQLQLRHAI 535
>At5g03250 photoreceptor-interacting protein - like
Length = 592
Score = 229 bits (583), Expect = 4e-60
Identities = 148/479 (30%), Positives = 258/479 (52%), Gaps = 40/479 (8%)
Query: 72 VQIDNASFSLHKMLRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPET 131
V+I+ +F++ LRC AEYLEMT++ GNLVG ++ +LNEV + ++ L E
Sbjct: 92 VKIELTAFNVVS-LRCAAEYLEMTDNYGEGNLVGMTETFLNEV-FGNWTDSIKALQTCEE 149
Query: 132 LLQVAEKAKLVNRCIDAIAYMASKQSQLCS---------PARSQCSSDGVMSSSMASHRR 182
++ AE +++RC+D++A A L + + + + + +++ +
Sbjct: 150 VIDYAEDLHIISRCVDSLAVKACADPSLFNWPVGGGKNATSGQNTEDESHLWNGISASGK 209
Query: 183 PVVH----WWGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEI---F 235
+ H WW +D + L + +F+R++ A+ RG K ++ +M Y +K + +
Sbjct: 210 MLQHTGEDWWFDDASFLSLPLFKRLITAIEARGMKLENIAMAVMYYTRKHVPLMNRQVNM 269
Query: 236 GKGRKELEPQHEHEKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEK 295
+ E E +++ +E +V LLP +K L LL+ A+ L + + R +LE+
Sbjct: 270 DEQVIETPNPSEEDQKTCLEEIVGLLPSKKGVNPTKFLLRLLQTAMVLHASQSSRENLER 329
Query: 296 RVAMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLE-------------FEIGNH 342
R+ QL QA L DLLIP+ + TL+DV+ V R++ ++ E G+
Sbjct: 330 RIGNQLDQAALVDLLIPNMGY-SETLYDVECVLRMIEQFVSSTEQAGIVPSPCIIEEGHL 388
Query: 343 SVSSADDEYFSPSQRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGK 402
AD +P+ V L++ YLAE+A D NL +KF +IA IP+ + P +DG
Sbjct: 389 VKDGAD--LLTPT----TLVATLVDGYLAEVAPDVNLKLAKFEAIAAAIPDYARPLDDGV 442
Query: 403 YRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKH 462
Y AID+YLKAHP++++ E++++C +M+CQKLS +A HAAQN+RLP++ ++QVL F+Q
Sbjct: 443 YHAIDVYLKAHPWITDSEREHICRLMNCQKLSLEASTHAAQNERLPLRVIVQVLFFEQLR 502
Query: 463 LRETMNDGGVNWDGTSIPDKLNVYSAELNPVSKVISNLRRENEELKREIVKLKMKLQEI 521
LR +++ + PD N + A + N+R EL++E + +K +L ++
Sbjct: 503 LRTSVSGWFFVSENLDNPD--NQHGANGGLLKPRGENVRERVSELEKECMNMKQELHKL 559
>At3g50840 putative protein
Length = 567
Score = 228 bits (582), Expect = 6e-60
Identities = 153/453 (33%), Positives = 244/453 (53%), Gaps = 35/453 (7%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC AEYLEMTE+ S NL+ +++ +L+E + ++ L E++ +AE + +
Sbjct: 91 LRCAAEYLEMTEEYSPENLISKTEKFLSEFVFTNVQESIKALKACESVSSLAESLCITEQ 150
Query: 145 CIDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPV-VHWWGEDLTVLRIDIFQRV 203
CID+I + AS S ++ G+ + + W EDLT L IF+RV
Sbjct: 151 CIDSIVFQASSTDP-SSFYGWPINNGGIFTVDRKKQSKDSKTELWFEDLTELSFPIFRRV 209
Query: 204 LIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQH-----EHEKRVVVETLV 258
+++M + ++ YA+K + G+ E+++R ++ET+
Sbjct: 210 ILSMKSSVLSPEIVERSLLTYAKKHIPGISRSSSASSSSSSSSTTIASENQQRELLETIT 269
Query: 259 SLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPG 318
S LP A + SL LLRAAI L + CR LEK++ L +A LDDLLIPSYS+
Sbjct: 270 SDLPL--TATTTRSLFGLLRAAIILNASENCRKFLEKKIGSNLEKATLDDLLIPSYSYLN 327
Query: 319 NTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKLMENYLAEIAADRN 378
TL+D+D V+R++ +LE N +VSS+ + VG+L++ L EIA+D N
Sbjct: 328 ETLYDIDLVERLLRRFLE----NVAVSSSS----------LTVVGRLIDGVLGEIASDAN 373
Query: 379 LPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDAR 438
L +F ++A L+P Q+ +DG YRA+DIY K H ++ E EK+ +CSVM C+KL+ +
Sbjct: 374 LKPEQFYNLAVLLPVQARVYDDGLYRAVDIYFKTHSWILEEEKEKICSVMDCRKLTVEGC 433
Query: 439 AHAAQNDRLPVQTVLQVLSFQQKHLRE-------TMNDGGVNWDGTSIPDKLNVYSAELN 491
HAAQN+RLP++ V+QVL +Q LR+ T DG D T + + + N
Sbjct: 434 THAAQNERLPLRAVVQVLFLEQLQLRQVITGTLLTEEDG----DKTVVDLGRWKEAVKEN 489
Query: 492 PVSKV-ISNLRRENEELKREIVKLKMKLQEIEK 523
V ++ + +R +L++E + LK + +I+K
Sbjct: 490 QVLRLDMDTMRTRVNQLEKECLYLKKVIAKIDK 522
>At5g64330 non-phototropic hypocotyl 3 (NPH3)
Length = 746
Score = 227 bits (579), Expect = 1e-59
Identities = 181/575 (31%), Positives = 269/575 (46%), Gaps = 131/575 (22%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC AEYLEMTED GNL+ +++ +L+ V L + ++ VL E L AE ++V R
Sbjct: 128 LRCAAEYLEMTEDLEEGNLIFKTEAFLSYVVLSSWRDSILVLKSCEKLSPWAENLQIVRR 187
Query: 145 CIDAIAYMASKQ---------SQLCSPARSQCS-----------SDGVMSSSMASHRRPV 184
C ++IA+ A + SP+ + + S S S ++ +PV
Sbjct: 188 CSESIAWKACSNPKGIRWAYTGKAPSPSTTNFAGSSPRWNESKDSSFYCSPSRNTNSQPV 247
Query: 185 -VHWWGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGL----------- 232
WW ED+++LRID F RV+ A+ +G + LG +IM YA K L GL
Sbjct: 248 PPDWWFEDVSILRIDHFVRVITAIKVKGMRFELLGAVIMHYAGKWLPGLIKEGGVAIAPA 307
Query: 233 ------EIFGKGRKELE---------------------------PQHEHEKRVVVETLVS 259
G G E+ + H+ V +
Sbjct: 308 MSSAIGGGLGLGGDEMSISCGSNSSGGSSGPDWKGGLHMVLSAGKTNGHQDSVACLAGLG 367
Query: 260 LLPREKNAMSVSSLSML---------------LRAAIYLETTVACRLDLEKRVAMQLGQA 304
+ P+++ + S +S++ LRAA L+ A +LEKRV MQ QA
Sbjct: 368 ISPKDQRMIVESLISIIPPQKDSVTCSFLLRLLRAANMLKVAPALITELEKRVGMQFEQA 427
Query: 305 VLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPS--------- 355
L DLLIP Y+ G T++DVD VQR++ +L E + SPS
Sbjct: 428 TLQDLLIPGYNNKGETMYDVDLVQRLLEHFLVQE----QTEGSSPSRMSPSPSQSMYADI 483
Query: 356 ---------------QRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTED 400
Q +RV +L+++YL E+A DRNLP +KF +AE +PE + +D
Sbjct: 484 PRGNNNNGGGGGGNNQNAKMRVARLVDSYLTEVARDRNLPLTKFQVLAEALPESARTCDD 543
Query: 401 GKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQ 460
G YRAID YLKAHP LSE E+K +C VM CQKLS DA HAAQN+RLP++ V+QVL +Q
Sbjct: 544 GLYRAIDSYLKAHPTLSEHERKRLCRVMDCQKLSMDACMHAAQNERLPLRVVVQVLFSEQ 603
Query: 461 KHLRETMNDGGVNWDGTSIPDKLNVYS---------AELNPVS---------KVISNLRR 502
+ + + + + T++ + + Y E P S K I+ L+
Sbjct: 604 VKISNALANTSLK-ESTTLGEAMGTYQPMIPNRKTLIEATPQSFQEGWAAAKKDINTLKF 662
Query: 503 ENEELKREIVKLKMKLQ----EIEKPAHESSAPSS 533
E E +K + V+L+ +++ + EK + PSS
Sbjct: 663 ELETVKTKYVELQNEMEVMQRQFEKTGKVKNTPSS 697
>At5g13600 photoreceptor-interacting protein-like; non-phototropic
hypocotyl-like protein
Length = 591
Score = 226 bits (576), Expect = 3e-59
Identities = 147/459 (32%), Positives = 242/459 (52%), Gaps = 26/459 (5%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMP--ETLLQVAEKAKLV 142
LRC AEYL+M+E+ NL+ ++ +LN+ ++ L +L +AE+ +V
Sbjct: 101 LRCAAEYLQMSENYGDANLIYLTESFLNDHVFVNWEDSIKALEKSCEPKVLPLAEELHIV 160
Query: 143 NRCIDAIAYMASKQSQLCSPARSQCSSDGVMSSSM----ASHRRPVVHWWGEDLT-VLRI 197
+RCI ++A A + +G ++++ + +WW D++ L +
Sbjct: 161 SRCIGSLAMKACAEDNTSFFNWPISLPEGTTTTTIYWNGIQTKATSENWWFNDVSSFLDL 220
Query: 198 DIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQHE---------- 247
+++R + + RG + + YA+++L + G RK P E
Sbjct: 221 PMYKRFIKTVESRGVNAGIIAASVTHYAKRNLP---LLGCSRKSGSPSEEGTNYGDDMYY 277
Query: 248 --HEKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAV 305
E+R ++E +V LLP +K S L LLR ++ L + + LEKR+ MQL +A
Sbjct: 278 SHEEQRSLLEEIVELLPGKKCVTSTKFLLRLLRTSMVLHASQVTQETLEKRIGMQLDEAA 337
Query: 306 LDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKL 365
L+DLLIP+ + G TL+D D+VQRI+ ++ + V S + I +V L
Sbjct: 338 LEDLLIPNMKYSGETLYDTDSVQRILDHFM-LTFDSSIVEEKQMMGDSHPLKSITKVASL 396
Query: 366 MENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVC 425
++ YLAE+A+D NL SKF ++ LIPE P +DG YRAIDIY+KAHP+L+E E++ +C
Sbjct: 397 IDGYLAEVASDENLKLSKFQALGALIPEDVRPMDDGIYRAIDIYIKAHPWLTESEREQLC 456
Query: 426 SVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTSIPDKLNV 485
+M+CQKLS +A HAAQN+RLP++ ++QVL F+Q LR ++ G + D
Sbjct: 457 LLMNCQKLSLEACTHAAQNERLPLRVIVQVLFFEQMRLRTSI--AGWLFGSEENNDTSGA 514
Query: 486 YSAELNP-VSKVISNLRRENEELKREIVKLKMKLQEIEK 523
N + V+ +R EL++E + +K L ++ K
Sbjct: 515 LEGNKNTNANMVMHGMRERVFELEKECMSMKQDLDKLVK 553
>At1g67900 unknown protein
Length = 631
Score = 224 bits (572), Expect = 8e-59
Identities = 151/444 (34%), Positives = 231/444 (52%), Gaps = 45/444 (10%)
Query: 86 RCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNRC 145
RC AEYL+M+E+ GNLV + + + N L ++ L + +E + +RC
Sbjct: 100 RCAAEYLQMSEEVEKGNLVYKLEVFFNSCILNGWRDSIVTLQTTKAFPLWSEDLAITSRC 159
Query: 146 IDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVH--WWGEDLTVLRIDIFQRV 203
I+AIA S + S S V M+S+R WW ED+ L ID++ R
Sbjct: 160 IEAIASKVLSHPSKVSLSHSH--SRRVRDDDMSSNRAAASSRGWWAEDIAELGIDLYWRT 217
Query: 204 LIAMMGRGFKQFDL-GPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEK----------RV 252
+IA+ G L G + +YA K L L+ + RK ++ + + + R+
Sbjct: 218 MIAIKSGGKVPASLIGDALRVYASKWLPTLQ---RNRKVVKKKEDSDSDSDTDTSSKHRL 274
Query: 253 VVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIP 312
++E+++SLLP EK A+S S L LL+AA L + + +++L +RVA+QL +A + DLLIP
Sbjct: 275 LLESIISLLPAEKGAVSCSFLLKLLKAANILNASTSSKMELARRVALQLEEATVSDLLIP 334
Query: 313 SYSFPGNTLFDVDTVQRIVVSYL-------------------EFEIGNHSVSSADDEY-- 351
S+ L+DVD V I+ ++ + + S + D E+
Sbjct: 335 PMSYKSELLYDVDIVATILEQFMVQGQTSPPTSPLRGKKGMMDRRRRSRSAENIDLEFQE 394
Query: 352 ----FSPSQRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAID 407
S S ++V KL++ YL +IA D NLP SKF+++AE +PE S D YRAID
Sbjct: 395 SRRSSSASHSSKLKVAKLVDGYLQQIARDVNLPLSKFVTLAESVPEFSRLDHDDLYRAID 454
Query: 408 IYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETM 467
IYLKAH L++ E+K VC V+ C+KLS +A HAAQN+ LP++ V+QVL ++Q
Sbjct: 455 IYLKAHKNLNKSERKRVCRVLDCKKLSMEACMHAAQNEMLPLRVVVQVLFYEQARAAAAT 514
Query: 468 NDGGVNWDGTSIPDKLNVYSAELN 491
N+G N T +P + A N
Sbjct: 515 NNGEKN--TTELPSNIKALLAAHN 536
>At3g44820 non-phototropic hypocotyl 3-like protein
Length = 661
Score = 215 bits (548), Expect = 5e-56
Identities = 152/498 (30%), Positives = 243/498 (48%), Gaps = 69/498 (13%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
+ C AEYLEMT + NL+ + + +L++ L + L +L+ AEK +++ +
Sbjct: 145 IHCAAEYLEMTNEYGEDNLISQVETFLHKHVLRNWKDCILALQSSSPVLKSAEKLQMIPK 204
Query: 145 CIDAIAYMASKQSQLCS-----PARSQCSSDGVMSSSM---ASHRRPVVHWWGEDLTVLR 196
++A++ M L Q ++ + + A R WW ED++ L
Sbjct: 205 LMNAVSTMVCTDPSLFGWPMMMYGTLQSPGGSILWNGINTGARMRSSGSDWWYEDISYLS 264
Query: 197 IDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVV--- 253
+D+F+R++ M +G + L +M YA+K L GL G+ + + +RVV
Sbjct: 265 VDLFKRLIKTMETKGIRAESLAGAMMYYARKYLPGL---GRWQSGTSDSSKSRRRVVSFN 321
Query: 254 ------------------VETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEK 295
+ET++SLLP ++ L LLR A L C LEK
Sbjct: 322 LAKASSPSSMPPLDQIALLETILSLLPEKRGRSFCKFLLGLLRVAFILGVDGNCVKKLEK 381
Query: 296 RVAMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPS 355
R+ MQL A LD+LLI +YS TL++VD V+RIV +
Sbjct: 382 RIGMQLELATLDNLLILNYS-DSETLYNVDCVERIVRHF--------------------- 419
Query: 356 QRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPF 415
+L+++Y+AE+A+D NL K S+A +PE S P DG YRA DIY K HP+
Sbjct: 420 -------WRLVDSYMAEVASDVNLKPDKMRSLAAALPESSRPLYDGLYRAFDIYFKEHPW 472
Query: 416 LSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWD 475
LS+ +K+ +C++M Q+LS DA AHA+ NDRLP++ VLQVL F+Q HLR T GG+N
Sbjct: 473 LSDRDKEQLCNIMDYQRLSIDACAHASHNDRLPLRVVLQVLFFEQMHLR-TALAGGLNVA 531
Query: 476 GTSIPDKLNVYSAELNPVSKVIS-----NLRRENEELKREIVKLKMKLQEIEKPAHESSA 530
T + + +++ + R+N+ LK ++ K++ ++ E+E+
Sbjct: 532 NTETAHAVTIPGGRTG--QEIVQRDGWVTVVRQNQVLKVDMQKMRSRVGELEEEFQSIKQ 589
Query: 531 PSSPLISAYSPSVNKPPL 548
+S S S++ P L
Sbjct: 590 EMKKRVSKSSSSMSSPRL 607
>At2g14820 hypothetical protein
Length = 634
Score = 212 bits (539), Expect = 5e-55
Identities = 140/391 (35%), Positives = 215/391 (54%), Gaps = 24/391 (6%)
Query: 86 RCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNRC 145
RC AEYL M E GNL+ + D +L+ + ++ VL + L ++E KLV+ C
Sbjct: 104 RCAAEYLGMHETVEKGNLIYKIDVFLSSSLFRSWKDSIIVLQTTKPFLPLSEDLKLVSLC 163
Query: 146 IDAIAYMA----SKQSQLCSPARSQCSSDGVMSSSMASHRRPVVH-WWGEDLTVLRIDIF 200
IDAIA A S + + + + + + S+ + R V H WW EDL L ID +
Sbjct: 164 IDAIATKACVDVSHVEWSYTYNKKKLAEENNGADSIKA--RDVPHDWWVEDLCELEIDYY 221
Query: 201 QRVLIAMMGRGFKQFD-LGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVS 259
+RV++ + + + +G + Y + L G F KG E +H + ++ETLV
Sbjct: 222 KRVIMNIKTKCILGGEVIGEALKAYGYRRLSG---FNKGVMEQGDLVKH--KTIIETLVW 276
Query: 260 LLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGN 319
LLP EKN++S L LL+A + + + L +R+ QL +A + +LLI S+
Sbjct: 277 LLPAEKNSVSCGFLLKLLKAVTMVNSGEVVKEQLVRRIGQQLEEASMAELLIKSHQ-GSE 335
Query: 320 TLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSP----------SQRGIVRVGKLMENY 369
TL+DVD VQ+IV+ ++ + + D++ F S+ + V K++++Y
Sbjct: 336 TLYDVDLVQKIVMEFMRRDKNSEIEVQDDEDGFEVQEVRKLPGILSEASKLMVAKVIDSY 395
Query: 370 LAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMH 429
L EIA D NLPASKFI +AE + P D YRAID++LK HP +++ EKK +C +M
Sbjct: 396 LTEIAKDPNLPASKFIDVAESVTSIPRPAHDALYRAIDMFLKEHPGITKGEKKRMCKLMD 455
Query: 430 CQKLSRDARAHAAQNDRLPVQTVLQVLSFQQ 460
C+KLS +A HA QNDRLP++ V+QVL F+Q
Sbjct: 456 CRKLSVEACMHAVQNDRLPLRVVVQVLFFEQ 486
>At5g10250 non-phototropic hypocotyl 3-like protein
Length = 607
Score = 209 bits (532), Expect = 3e-54
Identities = 156/486 (32%), Positives = 236/486 (48%), Gaps = 75/486 (15%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC +EYL MTE+ GNL+ +++ ++ V L + ++VL L AE ++V R
Sbjct: 127 LRCASEYLYMTEEFEAGNLISKTEAFITFVVLASWRDTLTVLRSCTNLSPWAENLQIVRR 186
Query: 145 CIDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVHWWGEDLTVLRIDIFQRVL 204
C D +A+ A C+ + + + + R + + D+ L ID F RV+
Sbjct: 187 CCDLLAWKA-------------CNDNNIPEDVVDRNERCLYN----DIATLDIDHFMRVI 229
Query: 205 IAMMGRGFKQFDLGPIIMLYAQK-------SLEGLEIFGKGRKELEPQHEH--------- 248
M R K G IIM YA LEG++ +G G+ EL+
Sbjct: 230 TTMKARRAKPQITGKIIMKYADNFLPVINDDLEGIKGYGLGKNELQFSVNRGRMEESNSL 289
Query: 249 ---EKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAV 305
E + +E+LVS+LP + A+S L +L+ +I + A DLEKRV M L A
Sbjct: 290 GCQEHKETIESLVSVLPPQSGAVSCHFLLRMLKTSIVYSASPALISDLEKRVGMALEDAN 349
Query: 306 LDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKL 365
+ DLLIP++ + Q+ V EF + + G + KL
Sbjct: 350 VCDLLIPNFK---------NEEQQERVRIFEFFLMHEQQQVL----------GKPSISKL 390
Query: 366 MENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVC 425
++NYLAEIA D LP +KF +AE++PE + DG YRAID++LK HP LS+ +++ +C
Sbjct: 391 LDNYLAEIAKDPYLPITKFQVLAEMLPENAWKCHDGLYRAIDMFLKTHPSLSDHDRRRLC 450
Query: 426 SVMHCQKLSRDARAHAAQNDRLPVQTVL----QVLSFQQKHLRETMNDGGVNWDGTSIPD 481
M+C+KLS DA HAAQNDRLP++T++ QVL +Q +R M D +P+
Sbjct: 451 KTMNCEKLSLDACLHAAQNDRLPLRTIVQINTQVLFSEQVKMRMMMQD--------KLPE 502
Query: 482 KLNVYSAELNPVSKVISNLRRENE---ELKREIVKLKMKLQEIEKPAHESSAPSSPLISA 538
K E N + + R+NE LK E+ +K K+ E++ +E L S
Sbjct: 503 K-----EEENSGGREDKRMSRDNEIIKTLKEELENVKKKMSELQSDYNELQQEYERLSSK 557
Query: 539 YSPSVN 544
S N
Sbjct: 558 QKSSHN 563
>At4g37590 unknown protein
Length = 580
Score = 196 bits (497), Expect = 4e-50
Identities = 139/423 (32%), Positives = 222/423 (51%), Gaps = 42/423 (9%)
Query: 86 RCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNRC 145
RC AE+LEM E GNLV + + +LN L++ ++ VL L +E+ KL RC
Sbjct: 115 RCAAEFLEMYETVEKGNLVYKIEVFLNSSILQSWKDSIIVLQTTRALSPYSEELKLTGRC 174
Query: 146 IDAIAYMASKQSQLCSPARSQCSSDGVMSSSMASHRRPVVHWWGEDLTVLRIDIFQRVLI 205
+D+IA AS + + + + + + + WW EDL L ID+++R L
Sbjct: 175 LDSIASRASIDTSKVEWSYTYSKKKN-LDNGLRKPQAVPRDWWVEDLCDLHIDLYKRALA 233
Query: 206 AMMGRGFKQFD-LGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVSLLPRE 264
+ RG D +G + YA K + G F K ++ + R + ++++ L+P E
Sbjct: 234 TIEARGNVSADVIGEALHAYAIKRIPG---FSKS-SSVQVTDFAKYRALADSIIELIPDE 289
Query: 265 KNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLFDV 324
K ++S S L+ LLRA+I+L L+ RV +L +A L D+L L+DV
Sbjct: 290 KRSVSSSFLTKLLRASIFLGCDEVA--GLKNRVGERLDEANLGDVL----------LYDV 337
Query: 325 DTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKLMENYLAEIAADR-NLPASK 383
+ +Q +V +L+ S +D+ + + V KL++ YLAE + D NLP K
Sbjct: 338 ELMQSLVEVFLK------SRDPREDDVTAKAS-----VAKLVDGYLAEKSRDSDNLPLQK 386
Query: 384 FISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAAQ 443
F+S+AE++ + DG YRAID++LK HP +++ EKK +C +M C+KLS +A AHA Q
Sbjct: 387 FLSLAEMVSSFPRQSHDGVYRAIDMFLKEHPEMNKSEKKRICRLMDCRKLSAEACAHAVQ 446
Query: 444 NDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTSIPDKLNVYSAELNPVSKVISNLRRE 503
N+RLP++ V+QVL F+Q N+ G + G S P E+ P S+ + +E
Sbjct: 447 NERLPMRVVVQVLFFEQVR----ANNNGSSSTGNSTP--------EVIPASRSTNTTDQE 494
Query: 504 NEE 506
+ E
Sbjct: 495 DTE 497
>At5g67440 photoreceptor-interacting protein-like
Length = 579
Score = 195 bits (496), Expect = 5e-50
Identities = 133/382 (34%), Positives = 211/382 (54%), Gaps = 19/382 (4%)
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
+RC AEYLEM E GNLV + + +LN L + ++ VL + +E KL R
Sbjct: 106 VRCAAEYLEMYESIENGNLVYKMEVFLNSSVLRSWKDSIIVLQTTRSFYPWSEDVKLDVR 165
Query: 145 CIDAIAYMASKQSQLCSPARSQCSSD----GVMSSSMASHRRPVVHWWGEDLTVLRIDIF 200
C+++IA A+ PAR S ++ M ++ P WW EDL L ID+F
Sbjct: 166 CLESIALKAAMD-----PARVDWSYTYNRRKLLPPEMNNNSVPR-DWWVEDLAELSIDLF 219
Query: 201 QRVLIAMMGRGFKQFD-LGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVS 259
+RV+ + +G + +G + +YA K + G I + E E +R ++ETLVS
Sbjct: 220 KRVVSTIRRKGGVLPEVIGEALEVYAAKRIPGFMIQNDDNDDEEDVME--QRSLLETLVS 277
Query: 260 LLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGN 319
+LP EK ++S L LL++++ E R +L +R+ +L +A + DLLI + G
Sbjct: 278 MLPSEKQSVSCGFLIKLLKSSVSFECGEEERKELSRRIGEKLEEANVGDLLIRAPE-GGE 336
Query: 320 TLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKLMENYLAEIAA-DRN 378
T++D+D V+ ++ ++ + +DD S V KL++ YLAEI+ + N
Sbjct: 337 TVYDIDIVETLIDEFVTQTEKRDELDCSDDINDSSK----ANVAKLIDGYLAEISRIETN 392
Query: 379 LPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDAR 438
L +KFI+IAE + + DG YRAID++LK HP +++ EKK+ +M C+KLS +A
Sbjct: 393 LSTTKFITIAEKVSTFPRQSHDGVYRAIDMFLKQHPGITKSEKKSSSKLMDCRKLSPEAC 452
Query: 439 AHAAQNDRLPVQTVLQVLSFQQ 460
AHA QN+RLP++ V+Q+L F+Q
Sbjct: 453 AHAVQNERLPLRVVVQILFFEQ 474
>At1g52770 putative non-phototropic hypocotyl
Length = 454
Score = 189 bits (479), Expect = 5e-48
Identities = 118/362 (32%), Positives = 195/362 (53%), Gaps = 34/362 (9%)
Query: 188 WGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLE--IFGKGRKELEPQ 245
W +D +L ID F + + + +G + +G II+ YA + L L + ++ +PQ
Sbjct: 28 WFDDGCILGIDYFVKTIAGIKSKGVRPDLIGSIIVHYASQWLPDLSDIVLNSDDQQPQPQ 87
Query: 246 HEHE----------KRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEK 295
+ E KR VETL+ ++P E++++ L LLR A + + +LE
Sbjct: 88 QQSESFSVTAFVMKKRSFVETLIGIIPPERDSVPCDFLLRLLRTANMVGADANYKAELEA 147
Query: 296 RVAMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPS 355
R++ QL QA L +L+IPS+S TL DV+ + R+V + + N V S
Sbjct: 148 RISWQLDQASLKELMIPSFSHTCGTLLDVELMTRLVKKFAGLD--NEGVKSG-------- 197
Query: 356 QRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPF 415
+++V KL+++YLAE A D +L S+FIS+ E +P + TEDG YRAID YLKAHP
Sbjct: 198 -ASLIKVAKLVDSYLAEAALDGDLTLSEFISLVEALPNHARVTEDGLYRAIDTYLKAHPN 256
Query: 416 LSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWD 475
+++ E+K +C ++ KLS +A HAAQNDRLPV+T++QVL +Q L ++ + W
Sbjct: 257 VTKQERKRLCGLIDSNKLSMEASLHAAQNDRLPVRTIIQVLFSEQAKLSHRSHN-NIEWS 315
Query: 476 GTSI------PDKLNVYSAELNPV----SKVISNLRRENEELKREIVKLKMKLQEIEKPA 525
G+S P+ + ++ P + I+ + E L+ ++ KLK + + ++
Sbjct: 316 GSSFSGVRSSPNPSGSHYSDSGPARCTSKREINVQQAEIRRLREDMAKLKCECEAMQTQL 375
Query: 526 HE 527
H+
Sbjct: 376 HK 377
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,570,405
Number of Sequences: 26719
Number of extensions: 522258
Number of successful extensions: 2199
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 23
Number of HSP's that attempted gapping in prelim test: 1993
Number of HSP's gapped (non-prelim): 119
length of query: 594
length of database: 11,318,596
effective HSP length: 105
effective length of query: 489
effective length of database: 8,513,101
effective search space: 4162906389
effective search space used: 4162906389
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0157a.4


