

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.8
(247 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
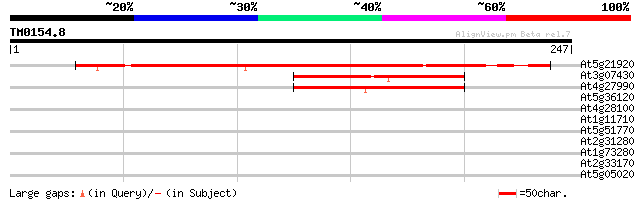
At5g21920 unknown protein 220 5e-58
At3g07430 unknown protein 80 1e-15
At4g27990 unknown protein 79 2e-15
At5g36120 unknown protein 37 0.013
At4g28100 predicted GPI-anchored protein 29 2.0
At1g11710 putative salt-inducible protein 29 2.0
At5g51770 putative protein 28 3.5
At2g31280 unknown protein 28 3.5
At1g73280 putative serine carboxypeptidase 28 3.5
At2g33170 putative receptor-like protein kinase 28 4.5
At5g05020 putative protein 28 5.9
>At5g21920 unknown protein
Length = 251
Score = 220 bits (561), Expect = 5e-58
Identities = 123/215 (57%), Positives = 150/215 (69%), Gaps = 19/215 (8%)
Query: 30 TQTPFALP-----ILKPPISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQNPFLNKIL 84
TQTP +L PP+++ + + H+S +AT+ FLHSLAS+NP + +
Sbjct: 33 TQTPLVRSNKPNLLLLPPVADSVKLIQ--DFHQSLISATEKFKGFLHSLASKNPLFQEAV 90
Query: 85 SLRSEFDSLCFQIRCSNY-RSRRWLSSHNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLV 143
L SEF LC +IR N R R +S+H FAAVLPGDSVAGLVV NG+ NFLNIYNT+LV
Sbjct: 91 RLSSEFHILCDEIRLRNTTRVRFAMSNHGFAAVLPGDSVAGLVVANGLINFLNIYNTILV 150
Query: 144 VRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASA 203
VRLVLTWFP++PPAIV PLST+CDPYLN+FRG IPPLGG LDLSPILAFLVLNAFTS++
Sbjct: 151 VRLVLTWFPSAPPAIVNPLSTLCDPYLNIFRGFIPPLGG-LDLSPILAFLVLNAFTSSAM 209
Query: 204 ALPAELPVTQQSEQGVAATLQSTDITSSQKKWMKR 238
ALP ELP S G + SS+ KW++R
Sbjct: 210 ALPCELP----SADGAVSP------ASSETKWVRR 234
>At3g07430 unknown protein
Length = 232
Score = 80.1 bits (196), Expect = 1e-15
Identities = 43/78 (55%), Positives = 56/78 (71%), Gaps = 4/78 (5%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV GI+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 149 VVAVGIKKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPIFD 207
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 208 TLDVSPLLAFAVLGTLGS 225
>At4g27990 unknown protein
Length = 218
Score = 79.0 bits (193), Expect = 2e-15
Identities = 38/77 (49%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGT 183
VV G+ +L+IY+ +L+VR++L+WFPN P + + +CDPYLNLFR +IPP+ T
Sbjct: 135 VVAAGLSKWLDIYSGVLMVRVLLSWFPNIPWDRQPLSAIRDLCDPYLNLFRNIIPPVFDT 194
Query: 184 LDLSPILAFLVLNAFTS 200
LD+SP+LAF VL S
Sbjct: 195 LDVSPLLAFAVLGTLGS 211
>At5g36120 unknown protein
Length = 174
Score = 36.6 bits (83), Expect = 0.013
Identities = 27/87 (31%), Positives = 46/87 (52%), Gaps = 8/87 (9%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP----PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPIL 190
L+ + L ++R+V++W+P P P ++ T +P L R +IPPL G +D++P++
Sbjct: 90 LSAFGFLFILRIVMSWYPKLPVDKFPYVLAYAPT--EPILVQTRKVIPPLAG-VDVTPVV 146
Query: 191 AFLVLNAFTSASAALPAELPVTQQSEQ 217
F L +F S P L V +Q
Sbjct: 147 WF-GLVSFLSEILVGPQGLLVLVSQQQ 172
>At4g28100 predicted GPI-anchored protein
Length = 304
Score = 29.3 bits (64), Expect = 2.0
Identities = 34/115 (29%), Positives = 48/115 (41%), Gaps = 20/115 (17%)
Query: 137 IYNTLLVVRLVLTWFPNSPPAIVGPLSTIC-----DPYLNLFRGLIPPLGGTLDLS---P 188
+++T+L LV PN+ PA P+ T D LF G+ G LD S P
Sbjct: 16 LFSTVLSNLLVEPVQPNTVPAF--PVETQAQSCRLDLSNELFGGVNEACGRNLDRSRCCP 73
Query: 189 ILAFLVLNAFTSASAALPAELPVTQQSE----------QGVAATLQSTDITSSQK 233
+LA + A ++ LPA P + S+ Q TLQS +T K
Sbjct: 74 VLAAWLFAAHARSALQLPAPAPTPESSDPDEPMKPDDSQKCVNTLQSALLTKQIK 128
>At1g11710 putative salt-inducible protein
Length = 657
Score = 29.3 bits (64), Expect = 2.0
Identities = 15/54 (27%), Positives = 25/54 (45%), Gaps = 5/54 (9%)
Query: 69 FLHSLASQNPFLNKILSLRSEF-----DSLCFQIRCSNYRSRRWLSSHNFAAVL 117
F H + + FL + + +F + + F C N R RRW + H F++ L
Sbjct: 2 FGHVFSRRTSFLVRCFHVAKKFSNPEPEDILFSALCLNLRQRRWNTLHQFSSSL 55
>At5g51770 putative protein
Length = 654
Score = 28.5 bits (62), Expect = 3.5
Identities = 15/49 (30%), Positives = 26/49 (52%), Gaps = 2/49 (4%)
Query: 21 KVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTATDNCFRF 69
KV+ C + ++P + P +K + LT +SPP+L F+ + F F
Sbjct: 600 KVALQCLQ--KSPVSRPSMKDVLEMLTGAISPPDLPTEFSPSPQTRFPF 646
>At2g31280 unknown protein
Length = 720
Score = 28.5 bits (62), Expect = 3.5
Identities = 18/60 (30%), Positives = 31/60 (51%), Gaps = 6/60 (10%)
Query: 194 VLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITSSQKKW------MKRLQGNRENKE 247
VLN ++++ A+ AE +T QS + +T Q+T T + + + QGN+ KE
Sbjct: 304 VLNDTSTSALAIEAERLITSQSYPRLDSTFQATSRTDKESSYHNEVFQLSENQGNKYIKE 363
>At1g73280 putative serine carboxypeptidase
Length = 441
Score = 28.5 bits (62), Expect = 3.5
Identities = 19/68 (27%), Positives = 31/68 (44%), Gaps = 6/68 (8%)
Query: 79 FLNKILSLRSEFDSLCFQIRCSNYRSR------RWLSSHNFAAVLPGDSVAGLVVGNGIQ 132
FL K L EF S F + +Y + +S N+ P ++ G V+GN +
Sbjct: 162 FLQKWLGKHQEFSSNPFYVGGDSYSGMVVPATVQEISKGNYECCNPPINLQGYVLGNPLT 221
Query: 133 NFLNIYNT 140
+F+ YN+
Sbjct: 222 DFVYDYNS 229
>At2g33170 putative receptor-like protein kinase
Length = 1124
Score = 28.1 bits (61), Expect = 4.5
Identities = 24/77 (31%), Positives = 35/77 (45%), Gaps = 5/77 (6%)
Query: 135 LNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYL--NLFRGLIPPLGGTLDLSPILAF 192
L + L ++RL F + P +G L+ + + + NLF G IPP G L I
Sbjct: 585 LGSLHQLEILRLSENRFSGNIPFTIGNLTHLTELQMGGNLFSGSIPPQLGLLSSLQIAMN 644
Query: 193 LVLNAFTSASAALPAEL 209
L N F S +P E+
Sbjct: 645 LSYNDF---SGEIPPEI 658
>At5g05020 putative protein
Length = 154
Score = 27.7 bits (60), Expect = 5.9
Identities = 26/92 (28%), Positives = 40/92 (43%), Gaps = 10/92 (10%)
Query: 119 GDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIP 178
G S+AGL +G G IY V+R ++ PN+PP P+ CD +
Sbjct: 30 GLSIAGLSLGQG-----QIYG---VLRCNISGDPNAPPVSGAPVYLKCDGSNTTIADAVT 81
Query: 179 PLGGTLDLSPILAFLVLNAFTSASAALPAELP 210
GT + +L+ + + +S L A LP
Sbjct: 82 KPDGTFRV--LLSAVQTVLISPSSCYLLANLP 111
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,656,798
Number of Sequences: 26719
Number of extensions: 233814
Number of successful extensions: 630
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 621
Number of HSP's gapped (non-prelim): 11
length of query: 247
length of database: 11,318,596
effective HSP length: 97
effective length of query: 150
effective length of database: 8,726,853
effective search space: 1309027950
effective search space used: 1309027950
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0154.8


