

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.6
(227 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
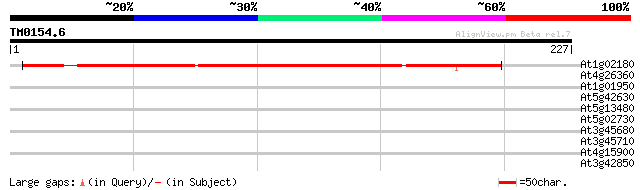
Score E
Sequences producing significant alignments: (bits) Value
At1g02180 unknown protein 141 2e-34
At4g26360 putative protein 32 0.36
At1g01950 unknown protein 29 2.3
At5g42630 unknown protein 28 3.0
At5g13480 putative protein 28 5.2
At5g02730 pathogenesis related protein - like 28 5.2
At3g45680 putative transporter protein 28 5.2
At3g45710 putative transporter protein 27 6.8
At4g15900 PRL1 protein 27 8.8
At3g42850 arabinose kinase - like protein 27 8.8
>At1g02180 unknown protein
Length = 226
Score = 141 bits (356), Expect = 2e-34
Identities = 75/195 (38%), Positives = 119/195 (60%), Gaps = 8/195 (4%)
Query: 6 LVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQ 65
L +IL FL +S +++ K+ N A ++V+ +N+NRTA K+ +L ++ GL C+ALQ
Sbjct: 8 LELILLFLSLSSVLASS-----KLHGNSAHEMVSILNQNRTARKLGKLNESPGLGCMALQ 62
Query: 66 YIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEA 125
Y++ +G+C V + + PE F +VFAPNCGV+ + ITG LGC +KY A
Sbjct: 63 YVELCEGNCN-VNNTLSCEHPEDDFTQVFAPNCGVELPTFGTITGHILGCSSKYAAPEVA 121
Query: 126 FSEVLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFE-GGVAK 184
FS++L ++ +L +L +++HT+VG + G+ +FWC+LFS G NS+F E G
Sbjct: 122 FSDILFRDSSALSVLRNRSHTEVGVGMARLHKGT-FFWCLLFSDGVKNSSFVLEDNGRGI 180
Query: 185 LTKPGCFSGANDECS 199
+ GC+SG+ CS
Sbjct: 181 KQRTGCYSGSAFPCS 195
>At4g26360 putative protein
Length = 1141
Score = 31.6 bits (70), Expect = 0.36
Identities = 14/39 (35%), Positives = 24/39 (60%)
Query: 129 VLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLF 167
++++ QRSL +L + +VG A+ + G+ F CVLF
Sbjct: 824 MVVEEQRSLGLLFASQKMKVGLALEDSPSGTKRFVCVLF 862
>At1g01950 unknown protein
Length = 893
Score = 28.9 bits (63), Expect = 2.3
Identities = 15/44 (34%), Positives = 25/44 (56%), Gaps = 2/44 (4%)
Query: 113 LGCQTKYVHAPEAFSEVLIQNQRSLEILHSKNHTQVGAAVTGTD 156
L CQ +Y+ + + E LI NQR+ E + K + +V VT ++
Sbjct: 466 LKCQMEYMESVKKLEEKLISNQRNHE--NGKRNGEVNGVVTASE 507
>At5g42630 unknown protein
Length = 276
Score = 28.5 bits (62), Expect = 3.0
Identities = 21/71 (29%), Positives = 30/71 (41%), Gaps = 9/71 (12%)
Query: 99 GVKASSLAPITGRFLGCQTKYVHAPEAF---------SEVLIQNQRSLEILHSKNHTQVG 149
G+K S AP +VHA + S + + N + L + H K+H Q+
Sbjct: 98 GLKRSIRAPRMRWTSTLHAHFVHAVQLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMY 157
Query: 150 AAVTGTDGGSP 160
V TD GSP
Sbjct: 158 RTVKCTDKGSP 168
>At5g13480 putative protein
Length = 679
Score = 27.7 bits (60), Expect = 5.2
Identities = 26/98 (26%), Positives = 39/98 (39%), Gaps = 17/98 (17%)
Query: 135 RSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNST----------FAFEGGVAK 184
RS+ H++N+ V+G DGG+ +W + K N T F G + K
Sbjct: 169 RSMVWSHNENYM-----VSGDDGGTLKYWQNNMNNVKANKTAHKESIRDLRIGFLGYIGK 223
Query: 185 LTKPGCFSGANDECSGAHDWSPLSVMWLFAASVLIALG 222
F C+ H W SV W S+L++ G
Sbjct: 224 GVIEIRFGVM--PCNAGHGWDVKSVDWHPTKSLLVSGG 259
>At5g02730 pathogenesis related protein - like
Length = 205
Score = 27.7 bits (60), Expect = 5.2
Identities = 26/103 (25%), Positives = 46/103 (44%), Gaps = 10/103 (9%)
Query: 4 LYLVVILSFLHSASTGSAATNTQIKVT--------NNPADKLVAAINENRTAYKVSELYD 55
L L++I S ++ G+++ +T+ N + + + A N R A S L
Sbjct: 19 LLLLLIFSGEFPSTAGTSSPDTKAAAARATNRGRRNKQSAEFLLAHNAARVASGASNLRW 78
Query: 56 NAGLACIALQYIKAYQGDCGAV--GGPDAKKPPESQFAEVFAP 96
+ GLA A ++ K + DC GGP + Q +E ++P
Sbjct: 79 DQGLARFASKWAKQRKSDCKMTHSGGPYGENIFRYQRSENWSP 121
>At3g45680 putative transporter protein
Length = 558
Score = 27.7 bits (60), Expect = 5.2
Identities = 10/20 (50%), Positives = 15/20 (75%)
Query: 206 PLSVMWLFAASVLIALGFAF 225
P+SV+WLF V++ +G AF
Sbjct: 436 PMSVLWLFPPLVIVGIGEAF 455
>At3g45710 putative transporter protein
Length = 556
Score = 27.3 bits (59), Expect = 6.8
Identities = 12/27 (44%), Positives = 18/27 (66%), Gaps = 3/27 (11%)
Query: 199 SGAHDWSPLSVMWLFAASVLIALGFAF 225
+G H P+SV+WL A V++ +G AF
Sbjct: 430 NGGH---PMSVLWLVPALVMVGIGEAF 453
>At4g15900 PRL1 protein
Length = 486
Score = 26.9 bits (58), Expect = 8.8
Identities = 20/68 (29%), Positives = 32/68 (46%), Gaps = 6/68 (8%)
Query: 100 VKASSLAPITGRFLGCQ---TKYVHAPEA-FSEVLIQNQRSLEILHSKNHTQVGAAVTGT 155
V+A +L P F TK P+ F ++ Q++ I+++ + G VTG
Sbjct: 347 VRAMTLHPKENAFASASADNTKKFSLPKGEFCHNMLSQQKT--IINAMAVNEDGVMVTGG 404
Query: 156 DGGSPYFW 163
D GS +FW
Sbjct: 405 DNGSIWFW 412
>At3g42850 arabinose kinase - like protein
Length = 964
Score = 26.9 bits (58), Expect = 8.8
Identities = 15/37 (40%), Positives = 22/37 (58%), Gaps = 2/37 (5%)
Query: 130 LIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVL 166
L+QN +L+ ++N T GA +TG GGS CV+
Sbjct: 880 LVQNMENLKSSKTENGTLYGAKITG--GGSGGTVCVI 914
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,181,799
Number of Sequences: 26719
Number of extensions: 217021
Number of successful extensions: 413
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 405
Number of HSP's gapped (non-prelim): 10
length of query: 227
length of database: 11,318,596
effective HSP length: 96
effective length of query: 131
effective length of database: 8,753,572
effective search space: 1146717932
effective search space used: 1146717932
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0154.6


