

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.14
(906 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
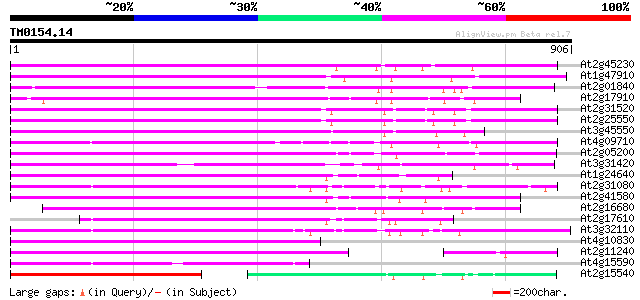
Score E
Sequences producing significant alignments: (bits) Value
At2g45230 putative non-LTR retroelement reverse transcriptase 526 e-149
At1g47910 reverse transcriptase, putative 517 e-146
At2g01840 putative non-LTR retroelement reverse transcriptase 507 e-143
At2g17910 putative non-LTR retroelement reverse transcriptase 481 e-136
At2g31520 putative non-LTR retroelement reverse transcriptase 476 e-134
At2g25550 putative non-LTR retroelement reverse transcriptase 473 e-133
At3g45550 putative protein 462 e-130
At4g09710 RNA-directed DNA polymerase -like protein 461 e-129
At2g05200 putative non-LTR retroelement reverse transcriptase 451 e-127
At3g31420 hypothetical protein 432 e-121
At1g24640 hypothetical protein 430 e-120
At2g31080 putative non-LTR retroelement reverse transcriptase 424 e-119
At2g41580 putative non-LTR retroelement reverse transcriptase 414 e-116
At2g16680 putative non-LTR retroelement reverse transcriptase 414 e-116
At2g17610 putative non-LTR retroelement reverse transcriptase 405 e-113
At3g32110 non-LTR reverse transcriptase, putative 405 e-113
At4g10830 putative protein 399 e-111
At2g11240 pseudogene 394 e-109
At4g15590 reverse transcriptase like protein 339 5e-93
At2g15540 putative non-LTR retroelement reverse transcriptase 266 4e-71
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 526 bits (1356), Expect = e-149
Identities = 322/930 (34%), Positives = 464/930 (49%), Gaps = 56/930 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V+ A F ++P K PG DG+ YQ+FW +GD ++ +NKT + L
Sbjct: 432 EVQRATFSINPHKCPGPDGMNGFLYQQFWETMGDQITEMVQAFFRSGSIEEGMNKTNICL 491
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ K FR ISLCNVI+K+I K +ANRLK ILP LI E+Q AFV GRLI+DN L
Sbjct: 492 IPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKKILPSLISETQAAFVKGRLISDNIL 551
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H + +A+K D+SKAYDRVEWPFL ++ +GF W+ LIM CV
Sbjct: 552 IAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFADHWIRLIMECVK 611
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V + +++NG P P RGLRQGDPLSPYLF++C E+ ++ A + + G+K+AR
Sbjct: 612 SVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQKNQITGLKVAR 671
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP ISHLLFADDS+ + + + I I+ Y ASGQ +N KS + +++ ++
Sbjct: 672 GAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKSSIYFGKHISEER 731
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+++ L ++ YLGLP SK +++K+R+ KK+ GW+ L G+E+
Sbjct: 732 RCLVKRKLGIEREGGEGVYLGLPESFQGSKVATLSYLKDRLGKKVLGWQSNFLSPGGKEI 791
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAVA A+P+Y MSCF +P +C QIE +++ F+W RG+H +W L + K G
Sbjct: 792 LLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWKAWCHLSRPKAVG 851
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGF++ + FN AL+ K WR+ + +SLM KV+++ YF + A G RPS+AW SI
Sbjct: 852 GLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSDPLNAPLGSRPSFAWKSI 911
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHF-------GVSLVSDL 533
+ + G IGNG+++ W+D W+ H + +V DL
Sbjct: 912 YEAQVLIKQGIRAVIGNGETINVWTDPWIGAKPAKAAQAVKRSHLVSQYAANSIHVVKDL 971
Query: 534 MLPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFL-- 591
+LP WN L++ + T I+A+ D W +S GHYS KSGY +
Sbjct: 972 LLPDGRDWNWNLVSLLFPDNTQENILALRPGGKETRDRFTWEYSRSGHYSVKSGYWVMTE 1031
Query: 592 ----RHGAEAVVASSSQVPSLPRADWRKLWKADA--------------LLPVRAYLHARG 633
R+ + V+ PSL ++++WK D L V + L R
Sbjct: 1032 IINQRNNPQEVLQ-----PSLDPI-FQQIWKLDVPPKIHHFLWRCVNNCLSVASNLAYRH 1085
Query: 634 LVTDPTCPRCLVALETVQHALLFCPMVQPIW------------FASSLGFRLAHECRVHE 681
L + +C RC ETV H L CP + W +A SL + H VH+
Sbjct: 1086 LAREKSCVRCPSHGETVNHLLFKCPFARLTWAISPLPAPPGGEWAESLFRNMHHVLSVHK 1145
Query: 682 FVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPM 741
Q + +L+ +W RN+L FK + T Q++ A
Sbjct: 1146 -----SQPEESDHHALIPWILWRLWKNRNDLVFKGREFTAPQVILKATEDMDAWNNRKEP 1200
Query: 742 VPQV-----HRPNTWSRPHPGTFKINFDAAMA-PTGDAGFGLVARNSRGEVLAAACSSHG 795
PQV R W P G K N D A + G+ G G V RN G +L +
Sbjct: 1201 QPQVTSSTRDRCVKWQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLLWLGLRALP 1260
Query: 796 PAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRL 855
S L E + RWA+ +RRV+FE D L + L I D R L
Sbjct: 1261 SQQSVLETEVEALRWAVLSLSRFNYRRVIFESDSQYLVSLIQNEMDIPSLAPRIQDIRNL 1320
Query: 856 SSGFDAFDFSFIRRTGNSVADALAKRSFTL 885
F+ F F RR GN+VAD A+ S +L
Sbjct: 1321 LRHFEEVKFQFTRREGNNVADRTARESLSL 1350
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 517 bits (1331), Expect = e-146
Identities = 317/936 (33%), Positives = 479/936 (50%), Gaps = 49/936 (5%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V ALF +HP KAPG DG+ ALF+QK W II D+ L + +N T + L
Sbjct: 216 EVRAALFMIHPEKAPGPDGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDKRLNTTNICL 275
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK ++P + R ISLCNV +K+I+K L RLK +LP+LI E+Q+AFV GRLI+DN L
Sbjct: 276 IPKTERPTRMTELRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGRLISDNIL 335
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E FH ++ S + MA+K DMSKAYD+VEW F+ ++L++MGF W+S IM C++
Sbjct: 336 IAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWISWIMWCIT 395
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
TV + +++NG P+ P RGLRQGDPLSPYLFILC EV A I KA + + GIK+A
Sbjct: 396 TVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLITGIKVAT 455
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
+P +SHLLFADDS+ F +A ++ I IL YE SGQ IN KS + V D
Sbjct: 456 PSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFGHKVEDSI 515
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+++ +L + + YLGLP +G SKT++F+FV++R+ ++ GW + L + G+EV
Sbjct: 516 KADIKLILGIHNLGGMGSYLGLPESLGGSKTKVFSFVRDRLQSRINGWSAKFLSKGGKEV 575
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA +P YVMSCF LP + +++ +++F+W + RGMH ++W LC +K +G
Sbjct: 576 MIKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDG 635
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFR+ FNSAL+AK WR+ + P+SL KV++ YF + + + Y PSY W S+
Sbjct: 636 GLGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSM 695
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVS-DLMLPGLL 539
+ + + G + +G+G S+ W+D W+P +G S+V L + L+
Sbjct: 696 ISARSLVYKGLIKRVGSGASISVWNDPWIPA------QFPRPAKYGGSIVDPSLKVKSLI 749
Query: 540 -----AWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHG 594
WN +L+ + P I A+P+ +D L W F+ G+Y+ KSGY R
Sbjct: 750 DSRSNFWNIDLLKELFDPEDVPLISALPIGNPNMEDTLGWHFTKAGNYTVKSGYHTARLD 809
Query: 595 A-EAVVASSSQVPSLPRADWRK---------LWK-ADALLPVRAYLHARGLVTDPTCPRC 643
E + +L W+ LW+ +PV L RG++ D C C
Sbjct: 810 LNEGTTLIGPDLTTLKAYIWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSC 869
Query: 644 LVALETVQHALLFCPMVQPIWFASSLGFR---LAHECRVHEFVSDFMQVADMSSLGAFLT 700
+ E++ H L C + IW S + F ++ +
Sbjct: 870 GASEESINHTLFQCHPARQIWALSQIPTAPGIFPSNSIFTNLDHLFWRIPSGVDSAPYPW 929
Query: 701 LLYAIWMARNE---------------LCFKEKASTLE-QILTHAASLTPLPPLEGPMVPQ 744
+++ IW ARNE L KE S E Q+ H+ L V
Sbjct: 930 IIWYIWKARNEKVFENVDKDPMEILLLAVKEAQSWQEAQVELHSERHGSLSIDSRIRVRD 989
Query: 745 VHRPNTWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLA 803
V + T+S F+ D + + +G G +S GE ++ + S L
Sbjct: 990 VSQDTTFS-----GFRCFIDGSWKASDQFSGTGWFCLSSLGESPTMGAANVRRSLSPLHT 1044
Query: 804 EGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFD 863
E + WA+ + + V F DC L + + + + F F
Sbjct: 1045 EMEALLWAMKCMIGADNQNVAFFTDCSDLVKMVSSPTEWPAFSVYLEELQSDREEFTNFS 1104
Query: 864 FSFIRRTGNSVADALAKRSFTLGVWV-WIEEIPQDF 898
S I R+ N AD LA++ T+ V ++ IP+D+
Sbjct: 1105 LSLISRSANVKADKLARKIRTVPHHVTYVNNIPRDW 1140
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 507 bits (1305), Expect = e-143
Identities = 314/922 (34%), Positives = 462/922 (50%), Gaps = 71/922 (7%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLH---GDIS*SIINKTL 57
+V +A+F + +APG DG A FY W +IG+DV CL V H D+ + IN+T
Sbjct: 794 EVRDAVFAIGADRAPGFDGFTAAFYHHLWDLIGNDV---CLMVRHFFESDVMDNQINQTQ 850
Query: 58 LVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITD 117
+ LIPK+ P H + +R ISLC +KII+K L RLK L D+I +SQ AFVPG+ I+D
Sbjct: 851 ICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCLGDVISDSQAAFVPGQNISD 910
Query: 118 NALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMT 177
N LVA E H +K + +G +A+K D+SKAYDRVEW FL V+ Q+GF WV IMT
Sbjct: 911 NVLVAHELLHSLKSRRECQSGYVAVKTDISKAYDRVEWNFLEKVMIQLGFAPRWVKWIMT 970
Query: 178 CVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIK 237
CV++V + +++NG+P P RG+RQGDPLSPYLF+ C EV S ++ KA + +HG+K
Sbjct: 971 CVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCAEVLSNMLRKAEVNKQIHGMK 1030
Query: 238 IARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVP 297
I + ISHLLFADDS+ F RA+ Q + I YE+ASGQ IN KS + + +P
Sbjct: 1031 ITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSIIFGQKIP 1090
Query: 298 DDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAG 357
L +LL + V KYLGLP +G+ K ++F ++ +V ++ +GW L AG
Sbjct: 1091 TMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVTKVKERTEGWAYNYLSPAG 1150
Query: 358 REVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAK 417
+E+++KA+A A+P Y M+CFLLP +CN+I +I+ F+WG +
Sbjct: 1151 KEIVIKAIAMALPVYSMNCFLLPTLICNEINSLITAFWWG------------------KE 1192
Query: 418 DNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAW 477
+ G LGF+D FN AL+AK WRI +NP+SL+ ++Y+ +Y+P T A +G SY W
Sbjct: 1193 NEGDLGFKDLHQFNRALLAKQAWRILTNPQSLLARLYKGLYYPNTTYLRANKGGHASYGW 1252
Query: 478 SSIMRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSL-VSDLMLP 536
+SI + G +G+G++ K W D W+P TL + V+DL
Sbjct: 1253 NSIQEGKLLLQQGLRVRLGDGQTTKIWEDPWLP---TLPPRPARGPILDEDMKVADLWRE 1309
Query: 537 GLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH--- 593
W+ + + P + ++ L D W ++ + Y+ +SGY H
Sbjct: 1310 NKREWDPVIFEGVLNPEDQQLAKSLYLSNYAARDSYKWAYTRNTQYTVRSGYWVATHVNL 1369
Query: 594 -GAEAVVASSSQVPSLPRADWRK---------LWKA-DALLPVRAYLHARGLVTDPTCPR 642
E + VP L + WR +W+ L L R + DPTC R
Sbjct: 1370 TEEEIINPLEGDVP-LKQEIWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQR 1428
Query: 643 CLVALETVQHALLFCPMVQPIWFASSL--GFRLAHECRVHEFVSDFMQVADMSSLGAF-- 698
C A ET+ H + C Q +W +++ RL + E + +Q +L
Sbjct: 1429 CCNADETINHIIFTCSYAQVVWRSANFSGSNRLCFTDNLEENIRLILQGKKNQNLPILNG 1488
Query: 699 ---LTLLYAIWMARNELCFKE-------KASTLEQILTH---------AASLTPLPPLEG 739
+++ +W +RNE F++ A EQ T A S +
Sbjct: 1489 LMPFWIMWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATEWVETMVNDTAISHNTAQSNDR 1548
Query: 740 PMVPQVHRPNTWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAA 798
P+ R WS P G K NFD+ D G + R+ G VL + C+ +
Sbjct: 1549 PL----SRSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSY 1604
Query: 799 SSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSG 858
S+L AE L F AL + G+ V FE D L L + +L++L+ D R +
Sbjct: 1605 SALQAEALGFLHALQMVWIRGYCYVWFEGDNLELTNLINKTEDHHLLETLLYDIRFWMTK 1664
Query: 859 FDAFDFSFIRRTGNSVADALAK 880
++ R N AD L K
Sbjct: 1665 LPFSSIGYVNRERNLAADKLTK 1686
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 481 bits (1238), Expect = e-136
Identities = 306/862 (35%), Positives = 439/862 (50%), Gaps = 51/862 (5%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SII----NKT 56
+V A+F ++ APG DG ALF+Q+ W D V H L + G ++ N T
Sbjct: 410 EVYNAVFSINKESAPGPDGFTALFFQQHW----DLVKHQILTEIFGFFETGVLPQDWNHT 465
Query: 57 LLVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLIT 116
+ LIPK+ P + R ISLC+V++KII+K L RLK LP ++ +Q+AFVP RLI+
Sbjct: 466 HICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLIS 525
Query: 117 DNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIM 176
DN LVA E H ++ MA K DMSKAYDRVEWPFL +++ +GF W+S IM
Sbjct: 526 DNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIM 585
Query: 177 TCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGI 236
CV++V +S+++NG P P RG+RQGDPLSP LF+LC E ++NKA A + GI
Sbjct: 586 NCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKITGI 645
Query: 237 KIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNV 296
+ ++HLLFADD+++ +AT QE + L+ Y Q SGQ+INL+KS ++ +NV
Sbjct: 646 QFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNV 705
Query: 297 PDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRA 356
++ + KYLGLP + SK +F F+KE++ +L GW ++L +
Sbjct: 706 DIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQG 765
Query: 357 GREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKA 416
G+EVL+K++A A+P YVMSCF LP LC ++ ++ F+W +R +H LSW+ L
Sbjct: 766 GKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLP 825
Query: 417 KDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYA 476
KD G GF+D + FN AL+AK WR+ SL +V+++ YF A RG RPSYA
Sbjct: 826 KDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYA 885
Query: 477 WSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWV-PGTDTLVYSVGLAQHFGVSL-VSDLM 534
W SI+ E+ G IGNG+ W+DKW+ G++ + + V L VS L+
Sbjct: 886 WRSILFGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNR--RPLNRRRFINVDLKVSQLI 943
Query: 535 LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFL--- 591
P WN ++ + P I+ PL +D W S +G YS K+GY+FL
Sbjct: 944 DPTSRNWNLNMLRDL-FPWKDVEIILKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQ 1002
Query: 592 -RHGAEAVVASSSQVPSLPRADWRK---------LWKA-DALLPVRAYLHARGLVTDPTC 640
H V SL W LWKA +PV L RG+ +D C
Sbjct: 1003 VHHRLYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGC 1062
Query: 641 PRCLVALETVQHALLFCPMVQPIW---FASSLGFRLAHECRVHEFVSDFMQVADMSSLGA 697
C ET+ H L CP+ + +W SS G ++ V+ +S + + + L
Sbjct: 1063 LMCDTENETINHILFECPLARQVWAITHLSSAGSEFSNS--VYTNMSRLIDLTQQNDLPH 1120
Query: 698 FLT-----LLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPN--- 749
L +L+ +W RN L F+ K S ++ A Q H N
Sbjct: 1121 HLRFVSPWILWFLWKNRNALLFEGKGSITTTLVDKA-----YEAYHEWFSAQTHMQNDEK 1175
Query: 750 -----TWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLA 803
W P PG K N A + +G V R+S+G+VL + S S A
Sbjct: 1176 HLKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSA 1235
Query: 804 EGLSFRWALGLAMDLGFRRVVF 825
+ S+ WAL F RV+F
Sbjct: 1236 KIRSWEWALESMTHHHFDRVIF 1257
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 476 bits (1224), Expect = e-134
Identities = 306/929 (32%), Positives = 459/929 (48%), Gaps = 61/929 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +A+ Q+ KAPG DGL A FY+ W I+G DV + IN T + +
Sbjct: 588 EIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICM 647
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV++K+I+K L NRLK L ++ +SQ AF+PGR+I DN +
Sbjct: 648 IPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVM 707
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF W+ IM V
Sbjct: 708 IAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVK 767
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++NG+P P RG+RQGDPLSPYLFILCG++ S LIN + L G++I
Sbjct: 768 SVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLRGVRIGN 827
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP I+HL FADDS+ F +A V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 828 GAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGST 887
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+L+Q+L + KYLGLP G+ K ++F ++ +RV K+ W R L AG+E+
Sbjct: 888 QSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEI 947
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA A+P Y MSCF LP G+ ++IE ++ F+W +RG+ ++WK L +K G
Sbjct: 948 MLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEG 1007
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WR+ P SL +V +A YF +I AK + SY W+S+
Sbjct: 1008 GLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASL 1067
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYS-----VGLAQHFGVSLVSDLM- 534
+ + G IG+G++++ G D +V S + + + +++L
Sbjct: 1068 LDGIALLKKGTRHLIGDGQNIRI-------GLDNIVDSHPPRPLNTEETYKEMTINNLFE 1120
Query: 535 -LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH 593
W+ I+ I I L D + W +++ G Y+ +SGY L H
Sbjct: 1121 RKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTH 1180
Query: 594 GAEAVVASSSQ----------------VPSLPRADWRKLWKADALLPVRAYLHARGLVTD 637
+ + + +P L WR L +A L L RG+ D
Sbjct: 1181 DPSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQA---LATTERLTTRGMRID 1237
Query: 638 PTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEF------VSDFMQVAD 691
P CPRC E++ HAL CP W+ S + ++ ++F + +F+Q
Sbjct: 1238 PICPRCHRENESINHALFTCPFATMAWWLSDSSL-IRNQLMSNDFEENISNILNFVQDTT 1296
Query: 692 MSSLGAFLT--LLYAIWMARNELCFKE------------KASTLEQI-LTHAASLTPLPP 736
MS L L++ IW ARN + F + KA T + + T + TP P
Sbjct: 1297 MSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPT 1356
Query: 737 LEGPMVPQVHRPNTWSRPHPGTFKINFDAAM-APTGDAGFGLVARNSRGEVLAAACSSHG 795
+ W P K NFDA +A G + RN G ++
Sbjct: 1357 RQ-----IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLA 1411
Query: 796 PAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRL 855
++ L AE + AL G+ +V E DC L S L + + D
Sbjct: 1412 HTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLINGISFHSSLANHLEDISFW 1471
Query: 856 SSGFDAFDFSFIRRTGNSVADALAKRSFT 884
++ F + F FIRR GN +A LAK T
Sbjct: 1472 ANKFASIQFGFIRRKGNKLAHVLAKYGCT 1500
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 473 bits (1218), Expect = e-133
Identities = 305/929 (32%), Positives = 458/929 (48%), Gaps = 61/929 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +A+ Q+ KAPG DGL A FY+ W I+G DV + IN T + +
Sbjct: 814 EIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICM 873
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV++K+I+K L NRLK L ++ +SQ AF+PGR+I DN +
Sbjct: 874 IPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVM 933
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF W+ IM V
Sbjct: 934 IAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVK 993
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++NG+P P RG+RQGDPLSPYLFILCG++ S LIN + L G++I
Sbjct: 994 SVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLRGVRIGN 1053
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP I+HL FADDS+ F +A V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 1054 GAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGST 1113
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
L+Q+L + KYLGLP G+ K ++F ++ +RV K+ W R L AG+E+
Sbjct: 1114 QSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEI 1173
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA A+P Y MSCF LP G+ ++IE ++ F+W +RG+ ++WK L +K G
Sbjct: 1174 MLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEG 1233
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WR+ P SL +V +A YF +I AK + SY W+S+
Sbjct: 1234 GLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASL 1293
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYS-----VGLAQHFGVSLVSDLM- 534
+ + G IG+G++++ G D +V S + + + +++L
Sbjct: 1294 LDGIALLKKGTRHLIGDGQNIRI-------GLDNIVDSHPPRPLNTEETYKEMTINNLFE 1346
Query: 535 -LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH 593
W+ I+ I I L D + W +++ G Y+ +SGY L H
Sbjct: 1347 RKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTH 1406
Query: 594 GAEAVVASSSQ----------------VPSLPRADWRKLWKADALLPVRAYLHARGLVTD 637
+ + + +P L WR L +A L L RG+ D
Sbjct: 1407 DPSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQA---LATTERLTTRGMRID 1463
Query: 638 PTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEF------VSDFMQVAD 691
P+CPRC E++ HAL CP W S + ++ ++F + +F+Q
Sbjct: 1464 PSCPRCHRENESINHALFTCPFATMAWRLSDSSL-IRNQLMSNDFEENISNILNFVQDTT 1522
Query: 692 MSSLGAFLT--LLYAIWMARNELCFKE------------KASTLEQI-LTHAASLTPLPP 736
MS L L++ IW ARN + F + KA T + + T + TP P
Sbjct: 1523 MSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPT 1582
Query: 737 LEGPMVPQVHRPNTWSRPHPGTFKINFDAAM-APTGDAGFGLVARNSRGEVLAAACSSHG 795
+ W P K NFDA +A G + RN G ++
Sbjct: 1583 RQ-----IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLA 1637
Query: 796 PAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRL 855
++ L AE + AL G+ +V E DC L S L + + D
Sbjct: 1638 HTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLINGISFHSSLANHLEDISFW 1697
Query: 856 SSGFDAFDFSFIRRTGNSVADALAKRSFT 884
++ F + F FIR+ GN +A LAK T
Sbjct: 1698 ANKFASIQFGFIRKKGNKLAHVLAKYGCT 1726
>At3g45550 putative protein
Length = 851
Score = 462 bits (1189), Expect = e-130
Identities = 276/799 (34%), Positives = 406/799 (50%), Gaps = 38/799 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ EA+ Q+ KAPG DGL A FY++ W I+G+DV + +N T + +
Sbjct: 53 EIFEAICQIGDDKAPGPDGLTARFYKQCWDIVGNDVIKEVKLFFESSHMKTSVNHTNICM 112
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK++ P + +R I+LCNV++K+I+K + NRLK L ++ +SQ AF+PGR+I DN +
Sbjct: 113 IPKIQNPQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGRIINDNVM 172
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF W+ IM V
Sbjct: 173 IAHEIMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGWIMAAVK 232
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++NG+P P RG+RQGDPLSPYLFILCG++ S LI + + G++I
Sbjct: 233 SVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIRGVRIGN 292
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP I+HL FADDS+ F +A V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 293 GAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGST 352
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
L+ LL + KYLGLP G+ K ++FN++ +RV ++ W + L AG+E+
Sbjct: 353 QTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEI 412
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+K+VA A+P Y MSCF LP G+ ++IE ++ F+W +RG+ ++WK L +K G
Sbjct: 413 LLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRLQYSKKEG 472
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WRI P SL +V +A YF +I AK + SY WSS+
Sbjct: 473 GLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKDNSIIDAKTRSQQSYGWSSL 532
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPG-LL 539
+ + G + IG+GK+++ D V QH G+SL + G
Sbjct: 533 LSGIALLRKGTRYVIGDGKTIRLGIDNVVDSHPPRPLLTD-EQHNGLSLDNLFQHRGHSR 591
Query: 540 AWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVV 599
W+ + I I L D L W ++S G Y+ +SGY H +
Sbjct: 592 CWDNAKLQTFVDQSDHDYIKRIYLSTRSKTDRLIWSYNSTGDYTVRSGYWLSTHDPSNTI 651
Query: 600 ASSSQ----------------VPSLPRADWRKLWKADALLPVRAYLHARGLVTDPTCPRC 643
+ ++ +P L WR L KA LP L RG+ DP CPRC
Sbjct: 652 PTMAKPHGSVDLKTKIWNLPIMPKLKHFLWRILSKA---LPTTDRLTTRGMRIDPGCPRC 708
Query: 644 LVALETVQHALLFCPMVQPIWFASSLGFRLAH--ECRVHEFVSDFM------QVADMSSL 695
E++ HAL CP W S + + + +S+ + + D L
Sbjct: 709 RRENESINHALFTCPFATMAWRLSDTPLYRSSILSNNIEDNISNILLLLQNTTITDSQKL 768
Query: 696 GAFLTLLYAIWMARNELCFKEKASTLEQILTHAASLT--------PLPPLEGPMVPQVHR 747
F LL+ IW ARN + F + + A + T P P
Sbjct: 769 IPF-WLLWRIWKARNNVVFNNLRESPSITVVRAKAETNEWLNATQTQGPRRLPKRTTAAG 827
Query: 748 PNTWSRPHPGTFKINFDAA 766
TW +P K NFDA+
Sbjct: 828 NTTWVKPQMPYIKCNFDAS 846
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 461 bits (1185), Expect = e-129
Identities = 315/918 (34%), Positives = 448/918 (48%), Gaps = 58/918 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+++EALF + KAPG DG A F+ +W II DVS +N+T + L
Sbjct: 373 EIKEALFSISADKAPGPDGFSASFFHAYWDIIEADVSRDIRSFFVDSCLSPRLNETHVTL 432
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV +KI+ K L RL+ L +LI Q+AFVPGR I DN L
Sbjct: 433 IPKISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLSELISLHQSAFVPGRAIADNVL 492
Query: 121 VAFECFHYMKKKISGCNGV--MALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTC 178
+ E H+++ +SG MA+K DMSKAYDR++W FL VL ++GF W+ +M C
Sbjct: 493 ITHEILHFLR--VSGAKKYCSMAIKTDMSKAYDRIKWNFLQEVLMRLGFHDKWIRWVMQC 550
Query: 179 VSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKI 238
V TV +S ++NG+PQ P RGLRQGDPLSPYLFILC EV S L KA + GI++
Sbjct: 551 VCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEVLSGLCRKAQEKGVMVGIRV 610
Query: 239 ARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPD 298
AR +P ++HLLFADD++ F + ++ IL YE ASGQ INL KS ++ S P
Sbjct: 611 ARGSPQVNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQSINLAKSAITFSSKTPQ 670
Query: 299 DCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGR 358
D ++ L + KYLGLP G+ K IF+ + +R+ ++ W R L AG+
Sbjct: 671 DIKRRVKLSLRIDNEGGIGKYLGLPEHFGRRKRDIFSSIVDRIRQRSHSWSIRFLSSAGK 730
Query: 359 EVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKD 418
++L+KAV ++PSY M CF LP LC QI+ +++RF+W +R M +SW L +
Sbjct: 731 QILLKAVLSSMPSYAMMCFKLPASLCKQIQSVLTRFWWDSKPDKRKMAWVSWDKLTLPIN 790
Query: 419 NGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYA-- 476
G LGFR+ + AK WRI P SL+ +V Y + PS+A
Sbjct: 791 EGGLGFREIE-------AKLSWRILKEPHSLLSRVLLGKYCNTSSFMDCSAS--PSFASH 841
Query: 477 -WSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSL-VSDLM 534
W I+ ++ G W+IG G S+ W++ W+ + +G L V DL+
Sbjct: 842 GWRGILAGRDLLRKGLGWSIGQGDSINVWTEAWLSPSSPQT-PIGPPTETNKDLSVHDLI 900
Query: 535 LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHG 594
+ +WN E I P I I + +P D L W G Y+TK+GY
Sbjct: 901 CHDVKSWNVEAIR-KHLPQYEDQIRKITINALPLQDSLVWLPVKSGEYTTKTGY------ 953
Query: 595 AEAVVASSSQVPSLPRADWRK--------------LWKA-DALLPVRAYLHARGLVTDPT 639
A+ +S S +W+K LWKA LPV L R + + T
Sbjct: 954 --ALAKLNSFPASQLDFNWQKNIWKIHTSPKVKHFLWKAMKGALPVGEALSRRNIEAEVT 1011
Query: 640 CPRCLVALETVQHALLFCPMVQPIWFASSLGF---RLAHECRVHEFVSDFMQVA----DM 692
C RC E+ H +L CP + +W + + F H V VA +
Sbjct: 1012 CKRC-GQTESSLHLMLLCPYAKKVWELAPVLFNPSEATHSSVALLLVDAKRMVALPPTGL 1070
Query: 693 SSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPNTWS 752
S + LL+ +W ARN L F + S E+ L A L +E ++ +H P+ S
Sbjct: 1071 GSAPLYPWLLWHLWKARNRLIF-DNHSCSEEGLVLKAILDARAWMEAQLL--IHHPSPIS 1127
Query: 753 -----RPHPGTFKINFDAAMAPTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLS 807
P+ DAA +G G G ++ + SS S+L+AE L+
Sbjct: 1128 DYPSPTPNLKVTSCFVDAAWTTSGYCGMGWFLQDPYKVKIKENQSSSSFVGSALMAETLA 1187
Query: 808 FRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFI 867
AL A+ G R++ DC L L L+ D R LS F F FI
Sbjct: 1188 VHLALVDALSTGVRQLNVFSDCKELISLLNSGKSIVELRGLLHDIRELSVSFTHLCFFFI 1247
Query: 868 RRTGNSVADALAKRSFTL 885
R N VAD+LAK + ++
Sbjct: 1248 PRLSNVVADSLAKSALSV 1265
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 451 bits (1161), Expect = e-127
Identities = 296/904 (32%), Positives = 455/904 (49%), Gaps = 39/904 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V++A+F ++ +KAPG DG A FY +WHII DV +N+T + L
Sbjct: 327 EVKDAVFSINASKAPGPDGFTAGFYHSYWHIISTDVGREIRLFFTSKNFPRRMNETHIRL 386
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK P +R I+LCN+ +KI+ K + R+++ILP LI E+Q+AFVPGR+I+DN L
Sbjct: 387 IPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLISENQSAFVPGRVISDNVL 446
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+ E H+++ + + MA+K DMSKAYDRVEW FL VLQ+ GF + W+ ++ CV+
Sbjct: 447 ITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVLQRFGFHSIWIDWVLECVT 506
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S ++NG PQ P RGLRQGDPLSP LFILC EV S L +A L G++++
Sbjct: 507 SVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSGLCTRAQRLRQLPGVRVSI 566
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P ++HLLFADD++ F+++ + + IL+ Y +ASGQ IN KS ++ S P
Sbjct: 567 NGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFSSKTPRSV 626
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+++++L ++ KYLGLP G+ K IF + +++ +K W R L +AG++V
Sbjct: 627 KGQVKRILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQV 686
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++KAV ++P Y MSCF LP LC +I+ +++RF+W R ++W L K+ G
Sbjct: 687 MLKAVLASMPLYSMSCFKLPSALCRKIQSLLTRFWWDTKPDVRKTSWVAWSKLTNPKNAG 746
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD + N +L+AK WR+ ++PESL+ ++ Y + K +PS+ W SI
Sbjct: 747 GLGFRDIERCNDSLLAKLGWRLLNSPESLLSRILLGKYCHSSSFMECKLPSQPSHGWRSI 806
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLV-YSVGLAQHFGVSLVSDLMLPGLL 539
+ EI G W I NG+ V W+D W+ + LV L +H + VS L+ L
Sbjct: 807 IAGREILKEGLGWLITNGEKVSIWNDPWLSISKPLVPIGPALREHQDLR-VSALINQNTL 865
Query: 540 AWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVV 599
W+ I I P I +P P D L W G Y+++SGY +
Sbjct: 866 QWDWNKIAVI-LPNYENLIKQLPAPSSRGVDKLAWLPVKSGQYTSRSGYG---------I 915
Query: 600 ASSSQVPSLPRA--DWR-KLWKADAL--------------LPVRAYLHARGLVTDPTCPR 642
AS + +P +P+ +W+ LWK L LPV L R + C R
Sbjct: 916 ASVASIP-IPQTQFNWQSNLWKLQTLPKIKHLMWKAAMEALPVGIQLVRRHISPSAACHR 974
Query: 643 CLVALETVQHALLFCPMVQPIWFASSL-GFRLAHECRVHEFVSDFMQVADMSSLGAFLTL 701
C A E+ H C +W + L + + + +S + + G
Sbjct: 975 C-GAPESTTHLFFHCEFAAQVWELAPLQETTVPPGSSMLDALSLLKKAIILPPTGVTSAA 1033
Query: 702 LYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPNTWSRPHPGTFKI 761
L+ W+ + L ++ LP + P P S G F
Sbjct: 1034 LFP-WICGIYGKLGTMTRAILDALAWQSAQRCLPKTRNVVHPISQLPVLRS----GYFCF 1088
Query: 762 NFDAAMAPTGDAGFGLVARNSRGEVLAAACSSHG--PAASSLLAEGLSFRWALGLAMDLG 819
A +A + AG G V +++ A S G S+L AE + + AL A+ LG
Sbjct: 1089 VDAAWIAQSSLAGSGWVFQSATALEKETATYSAGCRRLPSALSAEAWAIKSALLHALQLG 1148
Query: 820 FRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGNSVADALA 879
++ D + +A + + L+++ R L F + F+FI R+ N++ADA A
Sbjct: 1149 RTDLMVLSDSKSVVDALTSNISINEIYGLLMEIRALRVSFHSLCFNFISRSANAIADATA 1208
Query: 880 KRSF 883
K SF
Sbjct: 1209 KLSF 1212
>At3g31420 hypothetical protein
Length = 1491
Score = 432 bits (1112), Expect = e-121
Identities = 291/914 (31%), Positives = 428/914 (45%), Gaps = 103/914 (11%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++EE++F + P++ P DG A FYQ+FW I V + N T L L
Sbjct: 617 EIEESVFSVAPSRTPDPDGFTADFYQQFWPDIKQKVIDEVTRFFERSELDERHNHTNLCL 676
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPKV+ P +FR I+LCNV +KII+K L NRLK L I E+Q AFVPGRLIT+NA+
Sbjct: 677 IPKVETPTTIAKFRPIALCNVSYKIISKILVNRLKKHLGGAITENQAAFVPGRLITNNAI 736
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E ++ +K + N MALK D++KAYDR+EW FL ++QMGF W+ IM CV+
Sbjct: 737 IAHEVYYALKARKRQANSYMALKTDITKAYDRLEWDFLEETMRQMGFNTKWIERIMICVT 796
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
V FS+++NG+P +P RG+R GDPLSPYLFILC EV S +I +A + L GI+++
Sbjct: 797 MVRFSVLINGSPHGTIKPERGIRHGDPLSPYLFILCAEVLSHMIKQAEINKKLKGIRLST 856
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P ISHLLFADDS+ F A + +IK D
Sbjct: 857 QGPFISHLLFADDSIFFTLANQRSCTAIKE---------------------------PDT 889
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+R LL + KYLGLP K K ++FN++ E+V K +GW ++ L + G+EV
Sbjct: 890 KRRMRHLLGIHNEGGEGKYLGLPEQFNKKKKELFNYIIEKVKDKTQGWSKKFLSQGGKEV 949
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAVA A+P Y M+ F L +C +I+ +++RF+W +GMH +WK + K G
Sbjct: 950 LLKAVALAMPVYSMNIFKLTKEVCEEIDSLLARFWWSSGNETKGMHWFTWKRMSIPKKEG 1009
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGF++ + FN AL+ K W + +P LM +V R YFP + A +G R S+ W SI
Sbjct: 1010 GLGFKELENFNLALLGKQTWHLLQHPNCLMARVLRGRYFPETNVMNAVQGRRASFVWKSI 1069
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
+ + G +G+G + AW D W+P L P
Sbjct: 1070 LHGRNLLKKGLRCCVGDGSLINAWLDPWLP----------------------LHSPRAPY 1107
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH------- 593
+ + TAR DD++ W ++ DG Y+ KS Y H
Sbjct: 1108 KQEDAPEQLLVCSTAR------------DDLIGWHYTKDGMYTVKSAYWLATHLPGTTGT 1155
Query: 594 -------GAEAVVASSSQVPSLPRADWRKLWKADALLPVRAYLHARGLVTDPTCPRCLVA 646
+ ++ + P + W+ L A A + Y H + C RC
Sbjct: 1156 HPPPGDIKLKQLLWKTKTAPKIKHFCWKILSGAIATGEMLRYRH---INKQSICKRCCRD 1212
Query: 647 LETVQHALLFCPMVQPIWFASSLGFRLAHECRV---HEFVSDFMQVADMSSLGAFLT--L 701
ET QH C + W + L + + V +F + F ++ L +
Sbjct: 1213 EETSQHLFFECDYAKATWRGAGLPNLIFQDSIVTLEEKFRAMFTFNPSTTNYWRQLPFWI 1272
Query: 702 LYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRP-----------NT 750
L+ +W +RN L F++K E + A L + QV P N
Sbjct: 1273 LWRLWRSRNILTFQQKHIPWE-VTVQLAKQDALEWQDIEDRVQVINPLSRRHSNRYAANR 1331
Query: 751 WSRPHPGTFKINFDAAMAPTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRW 810
W+RP G K N+D + + ++ G V R+ G+ + + G ++ L L
Sbjct: 1332 WTRPVCGWKKCNYDGSYSTIINSKAGWVIRDEHGQFIGGG-QAKGKHTNNALESALQ--- 1387
Query: 811 ALGLAMDL----GFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSF 866
AL +AM G +V FE D + +Y+ + + I D + F F +
Sbjct: 1388 ALIIAMQSCWSHGHTKVCFEGDNIEVYQILNEGKARFDVYNWIRDIQAWKRRFQECRFLW 1447
Query: 867 IRRTGNSVADALAK 880
I R N AD LAK
Sbjct: 1448 INRRNNKPADTLAK 1461
>At1g24640 hypothetical protein
Length = 1270
Score = 430 bits (1106), Expect = e-120
Identities = 251/729 (34%), Positives = 382/729 (51%), Gaps = 32/729 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+++EA+F + P APG DG+ ALF+Q +W +G+ V+ + I + N T L L
Sbjct: 395 EIKEAVFSIKPASAPGPDGMSALFFQHYWSTVGNQVTSEVKKFFADGIMPAEWNYTHLCL 454
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK + P R ISLC+V++KII+K +A RL+ LP+++ ++Q+AFV RLITDN L
Sbjct: 455 IPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIVSDTQSAFVSERLITDNIL 514
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
VA E H +K + MA+K DMSKAYDRVEW +L S+L +GF WV+ IM CVS
Sbjct: 515 VAHELVHSLKVHPRISSEFMAVKSDMSKAYDRVEWSYLRSLLLSLGFHLKWVNWIMVCVS 574
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++N P RGLRQGDPLSP+LF+LC E + L+NKA +L GI+ +
Sbjct: 575 SVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSE 634
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P++ HLLFADDS+ +A+ +++L ++ IL Y A+GQ INL+KS ++ V +
Sbjct: 635 NGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKSSITFGEKVDEQL 694
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+R L + YLGLP SK + +++K+R+ +KL W R L + G+EV
Sbjct: 695 KGTIRTCLGIFTEGGAGTYLGLPECFSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEV 754
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+K+VA A+P + MSCF LP C +E ++ F+W R +H SW+ LC KD+G
Sbjct: 755 LLKSVALAMPVFAMSCFKLPITTCENLESAMASFWWDSCDHSRKIHWQSWERLCLPKDSG 814
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD + FN AL+AK WR+ P+ L+ ++ ++ YF A RPS+ W SI
Sbjct: 815 GLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDATDFLDAALSQRPSFGWRSI 874
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWV-------PGTDTLVYSVGLAQHFGVSLVSDL 533
+ E+ + G +G+G S+ W D W+ P L+Y V L V L
Sbjct: 875 LFGRELLSKGLQKRVGDGASLFVWIDPWIDDNGFRAPWRKNLIYDVTLK-------VKAL 927
Query: 534 MLPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH 593
+ P W+ E+++ + P I AI P++ D W + G +S KS Y
Sbjct: 928 LNPRTGFWDEEVLHDLFLPEDILRIKAIK-PVISQADFFVWKLNKSGDFSVKSAYWLAYQ 986
Query: 594 GAEAVVASSSQVPSLPRADWRKLWKADALLPVRAYLHARGLVTDPTCPRCLVALETVQHA 653
+ S + ++W ++ +L C E+ H
Sbjct: 987 TKSQNLRSEVSMQPSTLGLKTQVWNLQTDPKIKIFL----------WKVCGELGESTNHT 1036
Query: 654 LLFCPMVQPIWFASSLGFRL--AHECRVHEFVSDFMQVAD-----MSSLGAFLTLLYAIW 706
L CP+ + IW S F ++ ++ ++ D ++ F +L+ IW
Sbjct: 1037 LFLCPLSRQIWALSDYPFPPDGFSNGSIYSNINHLLENKDNKEWPINLRKIFPWILWRIW 1096
Query: 707 MARNELCFK 715
RN F+
Sbjct: 1097 KNRNSFIFE 1105
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 424 bits (1091), Expect = e-119
Identities = 313/938 (33%), Positives = 446/938 (47%), Gaps = 88/938 (9%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V A+ M KAPG DG +FYQ+ W +G V+ F L+ + + N LLVL
Sbjct: 296 EVVSAVKSMGRFKAPGPDGYQPVFYQQCWETVGPSVTRFVLEFFETGVLPASTNDALLVL 355
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
I KV KP QFR +SLCNV+FKIITK + RLK ++ LI +Q +F+PGRL DN +
Sbjct: 356 IAKVAKPERIQQFRPVSLCNVLFKIITKMMVTRLKNVISKLIGPAQASFIPGRLSIDNIV 415
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+ E H M++K G G M LKLD+ KAYDRV W FL L+ G W S IM V+
Sbjct: 416 LVQEAVHSMRRK-KGRKGWMLLKLDLEKAYDRVRWDFLQETLEAAGLSEGWTSRIMAGVT 474
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
S++ NG + F P RGLRQGDPLSPYLF+LC E LI +V I ++
Sbjct: 475 DPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGKREWKPIAVSC 534
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
+SH+ FADD ++FA A+V + I+ +L + +ASGQ ++L+KSK+ S NV +
Sbjct: 535 GGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHNVSREM 594
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ + + + KYLG+P + + + F V ERV +L GWK RSL AGR
Sbjct: 595 EQLISEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLAGWKGRSLSLAGRIT 654
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L KAV +IP +VMS LLP + ++ F WG + ++ H LSW+ +CK K G
Sbjct: 655 LTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEG 714
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
+G R + N ALVAK WR+ + ESL +V R Y G +P WSS
Sbjct: 715 GIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKKYKVGGV--QDTSWLKPQPRWSST 772
Query: 481 MRS-----GEIFATGGMWNIGNGKSVKAWSDKWV-------PGTDTLVYSVGLAQHFGVS 528
RS E+ G W G+G +++ W D+W+ GTD + + +
Sbjct: 773 WRSVAVGLREVVVKGVGWVPGDGCTIRFWLDRWLLQEPLVELGTDM------IPEGERIK 826
Query: 529 LVSDLMLPGLLAWNRELINFICCPPTARAIMAIPLPL-VPHDDVLYWPFSSDGHYSTKSG 587
+ +D LPG WN E++ R ++++ + + + + D + W + DG ++ +S
Sbjct: 827 VAADYWLPG-SGWNLEILGLYLPETVKRRLLSVVVQVFLGNGDEISWKGTQDGAFTVRSA 885
Query: 588 YQFLRHGAEAVVASSSQVPSLPRADWRKLWKADALLPVRAYLH--------------ARG 633
Y L + V + S + ++WK VR ++ R
Sbjct: 886 YSLL----QGDVGDRPNMGSF----FNRIWKLITPERVRVFIWLVSQNVIMTNVERVRRH 937
Query: 634 LVTDPTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEFVSDFMQVADMS 693
L + C C A ET+ H L CP ++PIW RL R HEF S + +
Sbjct: 938 LSENAICSVCNGAEETILHVLRDCPAMEPIW------RRLLPLRRHHEFFSQSLLEWLFT 991
Query: 694 SL----GAFLTLL-YAIWMA---------------RNELCF-KEKASTLEQILTHAASLT 732
++ G + TL IW A R+ L F K+ A + ++ A
Sbjct: 992 NMDPVKGIWPTLFGMGIWWAWKWRCCDVFGERKICRDRLKFIKDMAEEVRRVHVGAVG-- 1049
Query: 733 PLPPLEGPMVPQVHRPNTWSRPHPGTFKINFD-AAMAPTGDAGFGLVARNSRGEVLAAAC 791
P +V R W P G KI D A+ G A G RN +GE L
Sbjct: 1050 -----NRPNGVRVERMIRWQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFA 1104
Query: 792 SSHGPAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILD 851
+ G A+ LAE + L +A D GFRRV ++DC L+ G S L
Sbjct: 1105 LNIGSCAAP-LAELWGAYYGLLIAWDKGFRRVELDLDCKLVVGFLST--GVSNAHPLSF- 1160
Query: 852 CRRLSSGFDAFDF----SFIRRTGNSVADALAKRSFTL 885
RL GF D+ S + R N +AD LA +FTL
Sbjct: 1161 LVRLCQGFFTRDWLVRVSHVYREANRLADGLANYAFTL 1198
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 414 bits (1065), Expect = e-116
Identities = 260/857 (30%), Positives = 428/857 (49%), Gaps = 42/857 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +A+F ++ APG DG ALF+Q+ W ++ + + I N T L L
Sbjct: 161 EIYKAVFSINAESAPGPDGFTALFFQRQWPLVKNQIISDIELFFQTGILPEDWNHTHLCL 220
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ KP R ISLC+V++KII+K L+ RLK LP ++ +Q+AFV RL++DN +
Sbjct: 221 IPKITKPARMADIRPISLCSVMYKIISKILSARLKKYLPVIVSPTQSAFVAERLVSDNII 280
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H ++ M K DMSKAYDRVEWPFL +L +GF ++W++ +M CVS
Sbjct: 281 LAHEIVHNLRTNEKISKDFMVFKTDMSKAYDRVEWPFLKGILLALGFNSTWINWMMACVS 340
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++NG P PHRGLRQGDPLSP+LF+LC E ++N+A + GI+
Sbjct: 341 SVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEALIHILNQAEKIGKISGIQFNG 400
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
T P ++HLLFADD+++ +A+ E I L+ Y SGQ+IN +KS ++ V ++
Sbjct: 401 TGPSVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISGQMINSEKSAITFGAKVNEET 460
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ ++ KYLGLP SK +F F+KE++ +L GW ++L + G+++
Sbjct: 461 KQWIMNRSGIQTEGGTGKYLGLPECFQGSKQVLFGFIKEKLQSRLSGWYAKTLSQGGKDI 520
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+K++A A P Y M+CF L LC ++ ++ F+W ++ +H + + L K G
Sbjct: 521 LLKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFWWNSVQDKKKIHWIGAQKLMLPKFLG 580
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
GF+D + FN AL+AK R+ ++ +SL+ ++ ++ Y+ A +G RPSYAW SI
Sbjct: 581 GFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRYYMNSDFLSATKGTRPSYAWQSI 640
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWV-------PGTDTLVYSVGLAQHFGVSLVSDL 533
+ E+ +G IGNG++ W D W+ P + ++ + L VS L
Sbjct: 641 LYGRELLVSGLKKIIGNGENTYVWMDNWIFDDKPRRPESLQIMVDIQLK-------VSQL 693
Query: 534 MLPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFL-R 592
+ P WN ++ + P I+ P+ D W ++ G Y+ KS Y R
Sbjct: 694 IDPFSRNWNLNMLRDL-FPWKEIQIICQQRPMASRQDSFCWFGTNHGLYTVKSEYDLCSR 752
Query: 593 HGAEAVVASSSQVPSLPRADWRKLWKADALLPVRAY--------------LHARGLVTDP 638
+ + + + PSL + K+W ++ ++ + L RG++ +
Sbjct: 753 QVHKQMFKEAEEQPSL-NPLFGKIWNLNSAPKIKVFLWKVLKGAVAVEDRLRTRGVLIED 811
Query: 639 TCPRCLVALETVQHALLFCPMVQPIWFASSL-----GFRLAHECRVHEFVSDFMQVADMS 693
C C ET+ H L CP+ + +W + + GF + V+ + + ++S
Sbjct: 812 GCSMCPEKNETLNHILFQCPLARQVWALTPMQSPNHGFGDSIFTNVNHVIGN-CHNTELS 870
Query: 694 SLGAFLT--LLYAIWMARNELCFKEKASTLEQILTHAASLTP--LPPLEGPMVPQVHRPN 749
+++ +++ +W RN+ F+ S I+ A L E + +
Sbjct: 871 PHLRYVSPWIIWILWKNRNKRLFEGIGSVSLSIVGKALEDCKEWLKAHELICSKEPTKDL 930
Query: 750 TWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSF 808
TW P K N A + AG V RN +G VL + S +S A+ +
Sbjct: 931 TWIPPLMNELKCNIGIAWSKKHQMAGVSWVVRNWKGRVLLHSRCSFSQISSHFDAKIKGW 990
Query: 809 RWALGLAMDLGFRRVVF 825
+A+ F RV F
Sbjct: 991 NYAVESMDQFKFDRVTF 1007
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 414 bits (1065), Expect = e-116
Identities = 263/801 (32%), Positives = 407/801 (49%), Gaps = 36/801 (4%)
Query: 54 NKTLLVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGR 113
N T L LIPK P + R ISLC+V++KII+K L+ +LK LP ++ SQ+AF R
Sbjct: 437 NHTHLYLIPKFTNPQRMSDIRPISLCSVLYKIISKILSFKLKKHLPSIVSPSQSAFFAER 496
Query: 114 LITDNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVS 173
LI+DN L+A E H ++ M K DMSKAYDRVEW FL +L +GF W+S
Sbjct: 497 LISDNILIAHEIVHSLRTNDKISKEFMVFKTDMSKAYDRVEWSFLQEILVALGFNDKWIS 556
Query: 174 LIMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSL 233
IM CV++V +S+++NG P RG+RQGDP+SP+LF+LC E ++ +A + +
Sbjct: 557 WIMGCVTSVTYSVLINGQHFGHITPERGIRQGDPISPFLFVLCTEALIHILQQAENSKKV 616
Query: 234 HGIKIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCS 293
GI+ + P ++HLLF DD+ + RAT + + L+ Y SGQ+IN++KS ++
Sbjct: 617 SGIQFNGSGPSVNHLLFVDDTQLVCRATKSDCEQMMLCLSQYGHISGQLINVEKSSITFG 676
Query: 294 RNVPDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSL 353
V +D ++ + KYLGLP + SK +F ++KE++ L GW +++L
Sbjct: 677 VKVDEDTKRWIKNRSGIHLEGGTGKYLGLPENLSGSKQDLFGYIKEKLQSHLSGWYDKTL 736
Query: 354 YRAGREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDL 413
+ G+E+L+K++A A+P Y+M+CF LP GLC ++ ++ F+W +H + K L
Sbjct: 737 SQGGKEILLKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFWWNSMEFSNKIHWIGGKKL 796
Query: 414 CKAKDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRP 473
K G GF+D + FN AL+AK WR+ S+ +S++ +++++ YF A++G RP
Sbjct: 797 TLPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNTDFLNARQGTRP 856
Query: 474 SYAWSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWV-PGTDTLVYSVGLAQHFGVSLVSD 532
SY W SI+ E+ G IGNG+ W DKW+ G ++ + + VS
Sbjct: 857 SYTWRSILYGRELLNGGLKRLIGNGEQTNVWIDKWLFDGHSRRPMNLHSLMNIHMK-VSH 915
Query: 533 LMLPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQ--- 589
L+ P WN + + + + IM PL+ +D W +++G Y+ KSGY+
Sbjct: 916 LIDPLTRNWNLKKLTELFHEKDVQLIMH-QRPLISSEDSYCWAGTNNGLYTVKSGYERSS 974
Query: 590 ---FLRHGAEAVVASS-----SQVPSLPRADWRK--LWKA-DALLPVRAYLHARGLVTDP 638
F EA V S +V SL K +WKA L V L +RG+ T
Sbjct: 975 RETFKNLFKEADVYPSVNPLFDKVWSLETVPKIKVFMWKALKGALAVEDRLRSRGIRTAD 1034
Query: 639 TCPRCLVALETVQHALLFCPMVQPIWF-----ASSLGFRLAHECRVHEFV---SDFMQVA 690
C C +ET+ H L CP + +W A + GF + ++ + +F
Sbjct: 1035 GCLFCKEEIETINHLLFQCPFARQVWALSLIQAPATGFGTSIFSNINHVIQNSQNFGIPR 1094
Query: 691 DMSSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAAS-----LTPLPPLEGPMVPQV 745
M ++ + LL+ IW RN+ F+ T +I+ A + G + P
Sbjct: 1095 HMRTVSPW--LLWEIWKNRNKTLFQGTGLTSSEIVAKAYEECNLWINAQEKSSGGVSPSE 1152
Query: 746 HRPNTWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLAE 804
H+ W+ P G K N A + AG V R+S G+VL + S+ S A+
Sbjct: 1153 HK---WNPPPAGELKCNIGVAWSRQKQLAGVSWVLRDSMGQVLLHSRRSYSQVYSLFDAK 1209
Query: 805 GLSFRWALGLAMDLGFRRVVF 825
S+ WAL F +V F
Sbjct: 1210 IKSWDWALESMDHFHFDKVTF 1230
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 405 bits (1041), Expect = e-113
Identities = 228/638 (35%), Positives = 341/638 (52%), Gaps = 51/638 (7%)
Query: 114 LITDNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVS 173
+ITDN L+A E H + K +A KLD++KA+D++EW F+ ++++QMGF W +
Sbjct: 1 MITDNILIAHELIHSLHTK-KLVQPFVATKLDITKAFDKIEWGFIEAIMKQMGFSEKWCN 59
Query: 174 LIMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSL 233
IMTC++T +SI++NG P P RG+RQGDP+SPYL++LC E SALI ++ A L
Sbjct: 60 WIMTCITTTTYSILINGQPVRRIIPKRGIRQGDPISPYLYLLCTEGLSALIQASIKAKQL 119
Query: 234 HGIKIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCS 293
HG K +R P ISHLLFA DS++F +AT++E +++ +L YE+ASGQ +N KS +
Sbjct: 120 HGFKASRNGPAISHLLFAHDSLVFCKATLEECMTLVNVLKLYEKASGQAVNFQKSAILFG 179
Query: 294 RNVPDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSL 353
+ + +L QLL + E + +YLGLP +G++KT F+F+ + + +K+ W + L
Sbjct: 180 KGLDFRTSEQLSQLLGIYKTEGFGRYLGLPEFVGRNKTNAFSFIAQTMDQKMDNWYNKLL 239
Query: 354 YRAGREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDL 413
AG+EVL+K++ AIP+Y MSCFLLP L +QI + F+W + + ++W L
Sbjct: 240 SPAGKEVLIKSIVTAIPTYSMSCFLLPMRLIHQITSAMRWFWWSNTKVKHKIPWVAWSKL 299
Query: 414 CKAKDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRP 473
K G L RD K FN AL+AK WRI P SLM +V++A YFP+ + AK +
Sbjct: 300 NDPKKMGGLAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKYFPKERLLDAKATSQS 359
Query: 474 SYAWSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWVP--------GTDTLVYSVGLAQHF 525
SYAW SI+ ++ + G + GNG +++ W D W+P GT +YS
Sbjct: 360 SYAWKSILHGTKLISRGLKYIAGNGNNIQLWKDNWLPLNPPRPPVGTCDSIYS------- 412
Query: 526 GVSLVSDLMLPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTK 585
VSDL++ G WN +L+ + I AI + +D + W ++ DG+YS K
Sbjct: 413 -QLKVSDLLIEG--RWNEDLLCKLIHQNDIPHIRAIRPSITGANDAITWIYTHDGNYSVK 469
Query: 586 SGYQFLRHGAEAVVASSSQVPSLPRAD-------WRKLWKADA--------------LLP 624
SGY LR S Q SLP + + +WK +A LP
Sbjct: 470 SGYHLLRK------LSQQQHASLPSPNEVSAQTVFTNIWKQNAPPKIKHFWWRSAHNALP 523
Query: 625 VRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEFVS 684
L R L+TD TC RC A E V H L C + + IW + + + F
Sbjct: 524 TAGNLKRRRLITDDTCQRCGEASEDVNHLLFQCRVSKEIWEQAHIKLCPGDSLMSNSFNQ 583
Query: 685 DFMQVADMS-----SLGAFLTLLYAIWMARNELCFKEK 717
+ + ++ + F + + IW RN+L F K
Sbjct: 584 NLESIQKLNQSARKDVSLFPFIGWRIWKMRNDLIFNNK 621
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 405 bits (1040), Expect = e-113
Identities = 300/953 (31%), Positives = 448/953 (46%), Gaps = 73/953 (7%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+VE A+ M KAPG DG +FYQ+ W ++G+ V+ F + N L+VL
Sbjct: 972 EVEGAIRSMGKYKAPGPDGFQPVFYQQGWEVVGESVTKFVMDFFSSGSFPQETNDVLVVL 1031
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
I KV KP QFR ISLCNV+FK ITK + RLK ++ LI +QT+F+PGRL TDN +
Sbjct: 1032 IAKVLKPEKITQFRPISLCNVLFKTITKVMVGRLKGVINKLIGPAQTSFIPGRLSTDNIV 1091
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
V E H M++K G G M LKLD+ KAYDR+ W L L+ G P +WV IM CV
Sbjct: 1092 VVQEVVHSMRRK-KGVKGWMLLKLDLEKAYDRIRWDLLEDTLKAAGLPGTWVQWIMKCVE 1150
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
++ NG + F+P RGLRQGDPLSPYLF+LC E LI ++ A IKI++
Sbjct: 1151 GPSMRLLWNGEKTDAFKPLRGLRQGDPLSPYLFVLCIERLCHLIESSIAAKKWKPIKISQ 1210
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
+ P +SH+ FADD ++FA A++ + ++ +L + ASGQ ++L+KSK+ S+NV D
Sbjct: 1211 SGPRLSHICFADDLILFAEASIDQIRVLRGVLEKFCGASGQKVSLEKSKIYFSKNVLRDL 1270
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ + +KA + KYLG+P + + + F V +R +L GWK R L AGR
Sbjct: 1271 GKRISEESGMKATKDLGKYLGVPILQKRINKETFGEVIKRFSSRLAGWKGRMLSFAGRLT 1330
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L KAV +I + MS LP + ++ + F WG + ++ H ++W +C + G
Sbjct: 1331 LTKAVLTSILVHTMSTIKLPQSTLDGLDKVSRAFLWGSTLEKKKQHLVAWTRVCLPRREG 1390
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LG R N AL+AK WR+ ++ SL +V R+ Y G + S WSS
Sbjct: 1391 GLGIRSATAMNKALIAKVGWRVLNDGSSLWAQVVRSKY-KVGDVHDRNWTVAKS-NWSST 1448
Query: 481 MRS-----GEIFATGGMWNIGNGKSVKAWSDKWV---PGTDTLVYSVGLAQHFGVSLVSD 532
RS ++ W IG+G+ ++ W+D+W+ P D + + LAQ + D
Sbjct: 1449 WRSVGVGLRDVIWREQHWVIGDGRQIRFWTDRWLSETPIADDSIVPLSLAQM--LCTARD 1506
Query: 533 LMLPGLLAWN-RELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFL 591
L G W+ ++ F+ + I + D L W +SDG ++ KS + L
Sbjct: 1507 LWRDG-TGWDMSQIAPFVTDNKRLDLLAVIVDSVTGAHDRLAWGMTSDGRFTVKSAFAML 1565
Query: 592 RHGAEAVVASSSQVPSLPRAD----WRKLWKADALLPVRAYL--------------HARG 633
+ PR D + ++WK A VR +L R
Sbjct: 1566 TN------------DDSPRQDMSSLYGRVWKVQAPERVRVFLWLVVNQAIMTNSERKRRH 1613
Query: 634 LVTDPTCPRCLVALETVQHALLFCPMVQPIW------------FASSLGFRLAHECRVHE 681
L C C +E++ H L CP + IW F SL L R
Sbjct: 1614 LCDSDVCQVCRGGIESILHVLRDCPAMSGIWDRIVPRRLQQSFFTMSLLEWLYSNLR--- 1670
Query: 682 FVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQI-----LTHAASLTPLPP 736
+ +D S++ F ++ W R F E + +++ L S+
Sbjct: 1671 -QGLMTEGSDWSTM--FAMAVWWGWKWRCSNIFGENKTCRDRVRFIKDLAVEVSIAYSRE 1727
Query: 737 LEGPMVP-QVHRPNTWSRPHPGTFKINFD-AAMAPTGDAGFGLVARNSRGEVLAAACSSH 794
+E + +V++P W+ P G +KIN D A+ G A G V RNS G +
Sbjct: 1728 VELRLSGLRVNKPIRWTPPMEGWYKINTDGASRGNPGLASAGGVLRNSAGAWCGGFAVNI 1787
Query: 795 GPAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRR-GGCSILDSLILDCR 853
G S+ LAE + L +A + E+D ++ K G L L+ C
Sbjct: 1788 G-RCSAPLAELWGVYYGLYMAWAKQLTHLELEVDSEVVVGFLKTGIGETHPLSFLVRLCH 1846
Query: 854 RLSSGFDAFDFSFIRRTGNSVADALAKRSFTLGVWVWI-EEIPQDFYGLVQSD 905
S S + R NS+AD LA +F+L + + + +EIP L+ D
Sbjct: 1847 NFLSKDWTVRISHVYREANSLADGLANHAFSLSLGLHVFDEIPISLVMLLSED 1899
>At4g10830 putative protein
Length = 1294
Score = 399 bits (1025), Expect = e-111
Identities = 198/501 (39%), Positives = 300/501 (59%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ A+ + KAPG DGL A FY+ W I+G DV IN T + +
Sbjct: 794 EIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYMKQSINHTNICM 853
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV++KII+K L RLK L ++ +SQ AF+PGRL+ DN +
Sbjct: 854 IPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVNDNVM 913
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF +W+ IM V
Sbjct: 914 IAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIMGAVK 973
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V++S+++NG P QP RG+RQGDPLSPYLFILC ++ + LI V + GI+I
Sbjct: 974 SVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDIRGIRIGN 1033
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P ++HL FADDS+ F ++ V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 1034 GVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGSRVHGTT 1093
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ L+ +L +++ KYLGLP G+ K +FN++ ERV K+ W + L AG+E+
Sbjct: 1094 QNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGKEI 1153
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA ++P Y MSCF LP + ++IE ++ F+W + +R + ++WK L +K G
Sbjct: 1154 MLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQYSKKEG 1213
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WR+ +NP SL ++ +A YF +I AKR SY W+S+
Sbjct: 1214 GLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFREDSILDAKRQRYQSYGWTSM 1273
Query: 481 MRSGEIFATGGMWNIGNGKSV 501
+ ++ G + +G+GK+V
Sbjct: 1274 LAGLDVIKKGSRFIVGDGKTV 1294
>At2g11240 pseudogene
Length = 1044
Score = 394 bits (1011), Expect = e-109
Identities = 200/546 (36%), Positives = 310/546 (56%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+++ A F +H KAPG DG A F+Q W +G ++ INKT + L
Sbjct: 258 EIKSACFSIHADKAPGPDGFSASFFQSNWMTVGPNIVLEIQSFFSSSTLQPTINKTHITL 317
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK++ +R I+LC V +KII+K L+ RL+ IL ++I E+Q+AFVP R DN L
Sbjct: 318 IPKIQSLKRMVDYRPIALCTVFYKIISKLLSRRLQPILQEIISENQSAFVPKRASNDNVL 377
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+ E HY+K + MA+K +MSKAYDR+EW F+ V+Q+MGF +W+S I+ C++
Sbjct: 378 ITHEALHYLKSLGAEKRCFMAVKTNMSKAYDRIEWDFIKLVMQEMGFHQTWISWILQCIT 437
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
TV +S +LNG+ Q P RGLRQGDPLSP+LFI+C EV S L KA L SL G+++++
Sbjct: 438 TVSYSFLLNGSAQGAVTPERGLRQGDPLSPFLFIICSEVLSGLCRKAQLDGSLLGLRVSK 497
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P ++HLLFADD++ F R+ ++ + IL YE+ASGQ+IN KS ++ SR PD
Sbjct: 498 GNPRVNHLLFADDTIFFCRSDLKSCKTFLCILKKYEEASGQMINKSKSAITFSRKTPDHI 557
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
E +Q+L ++ V KYLGLP + G+ K +FN + +R+ ++ W R L AG+
Sbjct: 558 KTEAQQILGIQLVGGLGKYLGLPKMFGRKKRDLFNQIVDRIRQRSLSWSSRFLSTAGKTT 617
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+V ++P+Y MSCF L LC +I+ ++ F+W ++ M ++W + K K G
Sbjct: 618 MLKSVLASMPTYTMSCFKLLVSLCKRIQSALTHFWWDSSADKKKMCWIAWSKMAKNKKEG 677
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGF+D FN AL+AK WRI +P ++ ++ Y + S+ W I
Sbjct: 678 GLGFKDITNFNDALLAKLSWRIVQSPSCVLVRILLGKYCRTSSFLDCSVTAASSHGWRGI 737
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
++ + IG+G W++ W+ + + + F V+ L+ +
Sbjct: 738 CTGKDLIKSQLGKVIGSGLDTLVWNEPWLSLSTSSTPMGPALEQFKSMTVAQLICQTTKS 797
Query: 541 WNRELI 546
W+RE +
Sbjct: 798 WDREKV 803
Score = 54.7 bits (130), Expect = 2e-07
Identities = 54/194 (27%), Positives = 87/194 (44%), Gaps = 14/194 (7%)
Query: 701 LLYAIWMARNELCFKEKASTLEQILTHAASLTPL---PPLEGPMVPQVHRPNTWSRPHPG 757
+L+ +W RN+L F++K + +++ + S + ++ V + S
Sbjct: 848 ILWNLWNCRNKLIFEQKHISSMDLISQSISQSTEWLGAQIQASKSKIVIPGISPSEIDLD 907
Query: 758 TFKINFDAAMAP-TGDAGFGLVARNSRGEVLAAACSSHGPAA-----SSLLAEGLSFRWA 811
T + + DA+ T AGFG V + + SH AA S LLA+ + A
Sbjct: 908 TIQCSTDASWREETLQAGFGWVFVDHSNHL-----ESHHKAAAMNIRSPLLAKASALSLA 962
Query: 812 LGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTG 871
+ A DLGF+++V D L + L ++ D LS F+ FSF++R
Sbjct: 963 IQHAADLGFKKLVVASDSQQLVKVLNGEPHPMELHGIVFDISVLSLNFEENSFSFVKREN 1022
Query: 872 NSVADALAKRSFTL 885
NS ADALAK + L
Sbjct: 1023 NSKADALAKAALLL 1036
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 339 bits (869), Expect = 5e-93
Identities = 196/483 (40%), Positives = 260/483 (53%), Gaps = 20/483 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V A+ M KAPG DG +FYQ+ W +G+ VS F ++ + N LLVL
Sbjct: 248 EVVVAVRSMGRFKAPGPDGYQPVFYQQCWETVGESVSKFVMEFFESGVLPKSTNDVLLVL 307
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
+ KV KP QFR +SLCNV+FKIITK + RLK ++ LI +Q +F+PGRL DN +
Sbjct: 308 LAKVAKPERITQFRPVSLCNVLFKIITKMMVIRLKNVISKLIGPAQASFIPGRLSFDNIV 367
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
V E H M++K G G M LKLD+ KAYDR+ W FLA L+ G W+ IM CV+
Sbjct: 368 VVQEAVHSMRRK-KGRKGWMLLKLDLEKAYDRIRWDFLAETLEAAGLSEGWIKRIMECVA 426
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+ S++ NG + F P RGLRQGDP+SPYLF+LC E I AV I I++
Sbjct: 427 GPEMSLLWNGEKTDSFTPERGLRQGDPISPYLFVLCIERLCHQIETAVGRGDWKSISISQ 486
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P +SH+ FADD ++FA A+V Q ++L+KSK+ S NV D
Sbjct: 487 GGPKVSHVCFADDLILFAEASV-----------------AQKVSLEKSKIFFSNNVSRDL 529
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ + + KYLG+P + + F V ERV +L GWK RSL AGR
Sbjct: 530 EGLITAETGIGSTRELGKYLGMPVLQKRINKDTFGEVLERVSSRLSGWKSRSLSLAGRIT 589
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L KAV +IP + MS LLP L Q++ + F WG V +R H LSWK +C+ K G
Sbjct: 590 LTKAVLMSIPIHTMSSILLPASLLEQLDKVSRNFLWGSTVEKRKQHLLSWKKVCRPKAAG 649
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LG R K N AL+AK WR+ ++ SL +V R Y + T P WSS
Sbjct: 650 GLGLRASKDMNRALLAKVGWRLLNDKVSLWARVLRRKY--KVTDVHDSSWLVPKATWSST 707
Query: 481 MRS 483
RS
Sbjct: 708 WRS 710
>At2g15540 putative non-LTR retroelement reverse transcriptase
Length = 1225
Score = 266 bits (680), Expect = 4e-71
Identities = 137/310 (44%), Positives = 197/310 (63%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V+ ALF ++P KAPG DG+ A FYQ +W + G D+ +N+T + L
Sbjct: 379 EVKRALFSLNPDKAPGPDGMTAFFYQHYWDLTGPDLIKLVQNFHSTGFFDERLNETNICL 438
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK ++P +FR ISLCNV +K+I+K L++RLK +LP+LI E+Q+AFV RLITDN L
Sbjct: 439 IPKTERPRKMAEFRPISLCNVSYKVISKVLSSRLKRLLPELISETQSAFVAERLITDNIL 498
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E FH ++ + MA+K DMSKAYDRVEW FL +++ +MGF WV I+ C+S
Sbjct: 499 IAQENFHALRTNPACKKKYMAIKTDMSKAYDRVEWSFLRALMLKMGFAQKWVDWIIFCIS 558
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V + I+LNG+P+ +P RG+RQGDP+SP+LFILC E A + A + G++I+R
Sbjct: 559 SVSYKILLNGSPKGFIKPSRGIRQGDPISPFLFILCTEALVAKLKDAEWHGRIQGLQISR 618
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
+P SHLLFADDS+ F +A + I IL Y +ASGQ +N DKS + V +
Sbjct: 619 ASPSTSHLLFADDSLFFCKADPLQGKEIIDILRLYGEASGQQLNPDKSSVMFGHEVDNSI 678
Query: 301 FHELRQLLTV 310
+ ++ L +
Sbjct: 679 RNTIKVSLGI 688
Score = 137 bits (346), Expect = 2e-32
Identities = 134/541 (24%), Positives = 202/541 (36%), Gaps = 63/541 (11%)
Query: 385 NQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNGRLGFRDFKMFNSALVAKNWWRIKS 444
+++ +++ F+W G+H ++W LC +G LGFR + FN L+AK WR+
Sbjct: 696 SKLSSVVANFWWKTREESNGIHWIAWDKLCTPFSDGGLGFRTLEEFNLVLLAKQLWRLIR 755
Query: 445 NPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSIMRSGEIFATGGMWNIGNGKSVKAW 504
P SL+ +V R YF + RPS+ W SIM + + +G IG+G + W
Sbjct: 756 FPNSLLSRVLRGRYFRYSDPIQIGKANRPSFGWRSIMAAKPLLLSGLRRTIGSGMLTRVW 815
Query: 505 SDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLAWNRELINFICCPPTARAIMAIPLP 564
D W+P L V+DL+ P W + + P I+ I
Sbjct: 816 EDPWIPSFPPRPAKSILNIRDTHLYVNDLIDPVTKQWKLGRLQELVDPSDIPLILGIRPS 875
Query: 565 LVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVASSSQVPSLPRADWRKLWK------ 618
D W F+ G+Y+ KSGY R + Q PS+ A ++WK
Sbjct: 876 RTYKSDDFSWSFTKSGNYTVKSGYWAARDLSRPTCDLPFQGPSV-SALQAQVWKIKTTRK 934
Query: 619 --------ADALLPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPIWFAS--- 667
L L +R + T+ CPRC E++ H L CP + IW S
Sbjct: 935 FKHFEWQCLSGCLATNQRLFSRHIGTEKVCPRCGAEEESINHLLFLCPPSRQIWALSPIP 994
Query: 668 ---------SLGFR---LAHECRVHEFVSDFMQVADMSSLGAFLTLLYAIWMARNELCFK 715
SL + L + + D M++ F +L+ IW +RN F+
Sbjct: 995 SSEYIFPRNSLFYNFDFLLSRGKEFDIAEDIMEI--------FPWILWYIWKSRNRFIFE 1046
Query: 716 EKASTLEQILTHAAS-------------LTPLPPLEGPMVPQVHRPNTWSRPHPGTFKIN 762
+ + IL A T PP P V + P P
Sbjct: 1047 NVIESPQVILDFAIQEANVWKQANSKEVATEYPP---PQVVPANLP-------PTRNVCQ 1096
Query: 763 FDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFR 821
FDA+ +G G V + + VL S + S L AE S WA+ + LG
Sbjct: 1097 FDASWHLKDTLSGHGWVLVD-QDIVLLLGLKSARKSLSPLHAEVDSLLWAMECMISLGVS 1155
Query: 822 RVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGNSVADALAKR 881
F D + + + L F +F F R N AD L+K+
Sbjct: 1156 DCSFASDSADFISLLENPSEWPTFVAELATFSSLVCFFPSFSIKFFSRIYNVRADCLSKK 1215
Query: 882 S 882
+
Sbjct: 1216 A 1216
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.139 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,792,075
Number of Sequences: 26719
Number of extensions: 888411
Number of successful extensions: 2878
Number of sequences better than 10.0: 121
Number of HSP's better than 10.0 without gapping: 92
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 2438
Number of HSP's gapped (non-prelim): 208
length of query: 906
length of database: 11,318,596
effective HSP length: 108
effective length of query: 798
effective length of database: 8,432,944
effective search space: 6729489312
effective search space used: 6729489312
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0154.14


