

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.6
(156 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
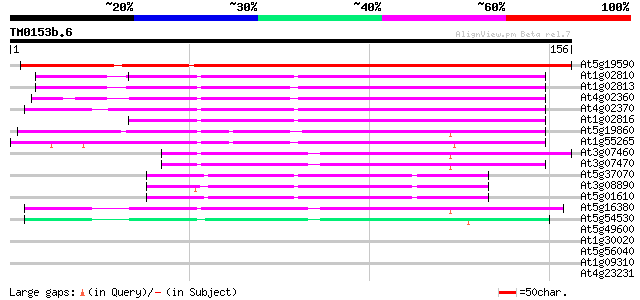
Score E
Sequences producing significant alignments: (bits) Value
At5g19590 unknown protein 196 4e-51
At1g02810 hypothetical protein 80 4e-16
At1g02813 unknown protein 80 4e-16
At4g02360 putative protein 77 3e-15
At4g02370 unknown protein 76 8e-15
At1g02816 Unknown protein 75 1e-14
At5g19860 putative protein 73 5e-14
At1g55265 unknown protein 70 4e-13
At3g07460 unknown protein 56 7e-09
At3g07470 unknown protein 53 8e-08
At5g37070 putative protein 51 2e-07
At3g08890 unknown protein 47 3e-06
At5g01610 unknown protein 46 9e-06
At5g16380 unknown protein 43 8e-05
At5g54530 unknown protein 40 4e-04
At5g49600 putative protein 37 0.006
At1g30020 unknown protein 30 0.69
At5g56040 receptor protein kinase-like protein 28 2.0
At1g09310 unknown protein 28 2.6
At4g23231 unknown protein 27 3.4
>At5g19590 unknown protein
Length = 151
Score = 196 bits (498), Expect = 4e-51
Identities = 100/153 (65%), Positives = 120/153 (78%), Gaps = 3/153 (1%)
Query: 4 KILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTS 63
KI + L L S Q+P P+ K P+ AH ELTN+GFP+GLLP +V + +NQTS
Sbjct: 2 KISLFLLSILILLPISLQDPTPEVKK--PTRAHAELTNHGFPIGLLP-LSVKDYFLNQTS 58
Query: 64 GDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSS 123
GDFS+ L GACKITLPPDNY+ATYSN +TG+I +G+IAEL GIRVRAFF+ WSITGIRSS
Sbjct: 59 GDFSLFLNGACKITLPPDNYIATYSNKVTGRISQGKIAELQGIRVRAFFKSWSITGIRSS 118
Query: 124 GDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
GDN+VFEV +TAKYPSKNFD+S CEG+RSSS
Sbjct: 119 GDNLVFEVAGITAKYPSKNFDESLDCEGKRSSS 151
>At1g02810 hypothetical protein
Length = 887
Score = 80.1 bits (196), Expect = 4e-16
Identities = 48/142 (33%), Positives = 74/142 (51%), Gaps = 8/142 (5%)
Query: 8 LFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFS 67
+F +FL L S S K S + L NY P G+LP+ V + +N+ +G F
Sbjct: 577 IFIIFLLLLSTSVSVSGQKK-----RSVYQVLENYTLPRGILPEG-VHDYDLNRRTGVFK 630
Query: 68 VRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNI 127
VR C+ ++ D+Y Y I+G I +GR+ L G+ V+ F W +I+ + GD++
Sbjct: 631 VRFNTTCQFSI--DSYKVKYKPVISGIITRGRVIRLIGVSVKVLFFWINISEVSRDGDDV 688
Query: 128 VFEVGMVTAKYPSKNFDDSPAC 149
F VG + ++ SK F DSP C
Sbjct: 689 EFFVGAASEEFSSKYFVDSPKC 710
Score = 75.1 bits (183), Expect = 1e-14
Identities = 37/116 (31%), Positives = 66/116 (56%), Gaps = 2/116 (1%)
Query: 34 SAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITG 93
+A+ L +Y FPVG+LP V+ + +++++G F +C L +Y Y +TI+G
Sbjct: 752 TAYTLLQSYNFPVGILPKG-VVSYDLDKSTGQFHAYFNKSCSFALQ-GSYQLDYKSTISG 809
Query: 94 KIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I + +I +L G++V+ F W +I + +GD + F VG+ +A + F +SP C
Sbjct: 810 YISENKITKLTGVKVKVLFLWLNIVEVIRNGDELEFSVGITSANFEIDEFYESPQC 865
>At1g02813 unknown protein
Length = 149
Score = 80.1 bits (196), Expect = 4e-16
Identities = 48/142 (33%), Positives = 74/142 (51%), Gaps = 8/142 (5%)
Query: 8 LFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFS 67
+F +FL L S S K S + L NY P G+LP+ V + +N+ +G F
Sbjct: 3 IFIIFLLLLSTSVSVSGQKK-----RSVYQVLENYTLPRGILPEG-VHDYDLNRRTGVFK 56
Query: 68 VRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNI 127
VR C+ ++ D+Y Y I+G I +GR+ L G+ V+ F W +I+ + GD++
Sbjct: 57 VRFNTTCQFSI--DSYKVKYKPVISGIITRGRVIRLIGVSVKVLFFWINISEVSRDGDDV 114
Query: 128 VFEVGMVTAKYPSKNFDDSPAC 149
F VG + ++ SK F DSP C
Sbjct: 115 EFFVGAASEEFSSKYFVDSPKC 136
>At4g02360 putative protein
Length = 154
Score = 77.4 bits (189), Expect = 3e-15
Identities = 45/143 (31%), Positives = 74/143 (51%), Gaps = 12/143 (8%)
Query: 7 ILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDF 66
I+F LF + SF KP +A+ + Y P G+LP V+ + +N +G+F
Sbjct: 10 IVFFLFFTV---SFAVSGQKP------TAYDAVKLYNLPPGILPKG-VVDYELNPKTGNF 59
Query: 67 SVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDN 126
V C+ T+ +Y Y +TI+G I G + L G+ V+ F W +I + G +
Sbjct: 60 KVYFNDTCEFTI--QSYQLKYKSTISGVISPGHVKNLKGVSVKVLFFWVNIAEVSLDGAD 117
Query: 127 IVFEVGMVTAKYPSKNFDDSPAC 149
+ F VG+ +A +P+ NF++SP C
Sbjct: 118 LDFSVGIASASFPAANFEESPQC 140
>At4g02370 unknown protein
Length = 167
Score = 75.9 bits (185), Expect = 8e-15
Identities = 44/145 (30%), Positives = 77/145 (52%), Gaps = 6/145 (4%)
Query: 5 ILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSG 64
ILI LFL+ + + +S +A+ L +Y FPVG+LP V+ + ++ T+G
Sbjct: 6 ILIASCLFLSSLTAAVVTA----AESDTPTAYSLLQSYNFPVGILPKG-VVAYDLDTTTG 60
Query: 65 DFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSG 124
F +C L +Y Y +TI+G I + ++ +L G++V+ F W +I + +G
Sbjct: 61 KFHAYFNDSCSFNLV-GSYQLNYKSTISGYISENKLKKLTGVKVKVLFLWLNIVEVIRNG 119
Query: 125 DNIVFEVGMVTAKYPSKNFDDSPAC 149
D + F VG+ +A + + F +SP C
Sbjct: 120 DEMEFSVGITSANFAIQEFLESPQC 144
>At1g02816 Unknown protein
Length = 166
Score = 75.1 bits (183), Expect = 1e-14
Identities = 37/116 (31%), Positives = 66/116 (56%), Gaps = 2/116 (1%)
Query: 34 SAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITG 93
+A+ L +Y FPVG+LP V+ + +++++G F +C L +Y Y +TI+G
Sbjct: 31 TAYTLLQSYNFPVGILPKG-VVSYDLDKSTGQFHAYFNKSCSFALQ-GSYQLDYKSTISG 88
Query: 94 KIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I + +I +L G++V+ F W +I + +GD + F VG+ +A + F +SP C
Sbjct: 89 YISENKITKLTGVKVKVLFLWLNIVEVIRNGDELEFSVGITSANFEIDEFYESPQC 144
>At5g19860 putative protein
Length = 181
Score = 73.2 bits (178), Expect = 5e-14
Identities = 51/150 (34%), Positives = 74/150 (49%), Gaps = 9/150 (6%)
Query: 3 PKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQT 62
P + +F L L + + + N P S S+ + L YG P GLLPDT V F ++
Sbjct: 5 PFFISIFIFSLTLFTTTTHSLNEPDPDSI-STVYELLPKYGLPSGLLPDT-VTDFTLSD- 61
Query: 63 SGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIR- 121
G F V L +C+I +Y+ Y TI+G+I G I EL GI+V+ FF W + I+
Sbjct: 62 DGRFVVHLPNSCEIEF---DYLVHYDKTISGRIGYGSITELKGIQVKKFFIWLDVDEIKV 118
Query: 122 --SSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
D+I F+VG + K F +C
Sbjct: 119 DLPPSDSIYFKVGFINKKLDIDQFKTIHSC 148
>At1g55265 unknown protein
Length = 175
Score = 70.5 bits (171), Expect = 4e-13
Identities = 52/170 (30%), Positives = 78/170 (45%), Gaps = 25/170 (14%)
Query: 1 MNPKILILFA-LFLNLTSKS------------------FQNPNPKPPKSTPSSAHIELTN 41
M +L LF+ LFL+L S S F + N P H L
Sbjct: 1 MASSLLALFSCLFLSLLSLSSSLNLRRPIFSQSNDLDLFSSLNLDRPSLAADDIHDLLPR 60
Query: 42 YGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIA 101
YGFP GLLP+ V + ++ GDF+V L +C + + + Y I GK+ G +
Sbjct: 61 YGFPKGLLPNN-VKSYTISD-DGDFTVDLISSCYVKF--SDQLVFYGKNIAGKLSYGSVK 116
Query: 102 ELNGIRVRAFFQWWSITGIRS--SGDNIVFEVGMVTAKYPSKNFDDSPAC 149
++ GI+ + F W IT + S S +VF VG V+ P+ F++ P+C
Sbjct: 117 DVRGIQAKEAFLWLPITAMESDPSSATVVFSVGFVSKTLPASMFENVPSC 166
>At3g07460 unknown protein
Length = 177
Score = 56.2 bits (134), Expect = 7e-09
Identities = 37/117 (31%), Positives = 56/117 (47%), Gaps = 7/117 (5%)
Query: 43 GFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAE 102
G P+GL P V GF VN +G FSV L +C+ + + Y ++G I +I +
Sbjct: 38 GLPLGLFPKG-VKGFTVNGETGRFSVYLNQSCQAKYETELH---YDEIVSGTIGYAQIRD 93
Query: 103 LNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
L+GI + F W + GIR S I F+VG++ +Y F+ C R +
Sbjct: 94 LSGISAQELFLWLQVKGIRVDVPSSGLIFFDVGVLRKQYSLSLFETPRDCVAVRGDA 150
>At3g07470 unknown protein
Length = 169
Score = 52.8 bits (125), Expect = 8e-08
Identities = 36/110 (32%), Positives = 53/110 (47%), Gaps = 7/110 (6%)
Query: 43 GFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAE 102
G P G+ P V F + +G FSV L AC+ + + Y ITG I +I++
Sbjct: 39 GLPSGIFPKG-VREFTFDVETGRFSVYLNQACEAKYETEIH---YDANITGTIGSAQISD 94
Query: 103 LNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
L+GI + F W+ + GIR S I F+VG+V +Y F+ C
Sbjct: 95 LSGISAQELFLWFPVKGIRVDVPSSGLIYFDVGVVRKQYSLSLFETPRDC 144
>At5g37070 putative protein
Length = 170
Score = 51.2 bits (121), Expect = 2e-07
Identities = 26/95 (27%), Positives = 49/95 (51%), Gaps = 3/95 (3%)
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L +G PVG+ P + N+ +G +V + C++ D+ V +S T+TG + KG
Sbjct: 58 LKEFGLPVGIFPQDAT-NYEFNEETGKLTVFIPETCEVGYR-DSSVLRFSTTVTGYLEKG 115
Query: 99 RIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGM 133
++AE+ G++ + W +T I + + F G+
Sbjct: 116 KLAEVEGMKTKVMI-WVKVTCISADSSKVYFTAGI 149
>At3g08890 unknown protein
Length = 170
Score = 47.4 bits (111), Expect = 3e-06
Identities = 26/96 (27%), Positives = 49/96 (50%), Gaps = 5/96 (5%)
Query: 39 LTNYGFPVGLLP-DTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVK 97
L +G PVG+ P D T + N+ + +V + C++ D V ++ T+TG + K
Sbjct: 58 LKEFGLPVGIFPRDAT--NYEFNEQTRKLTVFIPSICEVGYK-DTSVLRFTTTVTGFLEK 114
Query: 98 GRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGM 133
G++A++ G++ + W +T I + + F GM
Sbjct: 115 GKLADVEGMKTKVMI-WVKVTSISADSSKVHFTAGM 149
>At5g01610 unknown protein
Length = 170
Score = 45.8 bits (107), Expect = 9e-06
Identities = 22/95 (23%), Positives = 46/95 (48%), Gaps = 3/95 (3%)
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L Y P+G+ P + ++ + +V + C++ D+ V ++ T+TG + KG
Sbjct: 58 LKEYDLPIGIFPGDAT-NYEFDEETKKLTVLIPSICEVGYK-DSSVLKFTTTVTGHLEKG 115
Query: 99 RIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGM 133
++ ++ GI+ + W +T I + + F GM
Sbjct: 116 KLTDVEGIKTKVMI-WVKVTSISTDASKVYFTAGM 149
>At5g16380 unknown protein
Length = 195
Score = 42.7 bits (99), Expect = 8e-05
Identities = 37/153 (24%), Positives = 63/153 (40%), Gaps = 17/153 (11%)
Query: 5 ILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSG 64
I++LF+ L S +P S + L P G++P V F+++ +G
Sbjct: 11 IIVLFSSILFPQLSSLPDP----------SFYDYLRESNLPAGIVPKG-VTNFSIDIKTG 59
Query: 65 DFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIR--- 121
F+V L C + + I+G + GRI L+G+ + F W+++ GI
Sbjct: 60 RFTVALPVPCDAKFENQFH---FDYNISGVLSDGRIGNLSGVTQKELFLWFAVKGIHVDP 116
Query: 122 SSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRS 154
S I F+VG+ + F+ C S
Sbjct: 117 QSSGLIHFDVGVADKQLSLSLFESPRDCTAAES 149
>At5g54530 unknown protein
Length = 161
Score = 40.4 bits (93), Expect = 4e-04
Identities = 35/149 (23%), Positives = 58/149 (38%), Gaps = 18/149 (12%)
Query: 5 ILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSG 64
IL+L L L+L+ S P + H L + G P GLLP + + G
Sbjct: 9 ILLLTTLRLSLSLSSPSYP----------TVHDVLRSEGLPAGLLPQE--VDSYILHNDG 56
Query: 65 DFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSG 124
V L C + + + + G + G + + G+ + F W + I
Sbjct: 57 RLEVFLAAPCYAKFETNVH---FEAVVRGNLSYGSLVGVEGLSQKELFLWLQVKDIVVEN 113
Query: 125 DN---IVFEVGMVTAKYPSKNFDDSPACE 150
N IVF++G+ + F+D P C+
Sbjct: 114 PNSGVIVFDIGVAFKQLSLSLFEDPPKCK 142
>At5g49600 putative protein
Length = 171
Score = 36.6 bits (83), Expect = 0.006
Identities = 24/70 (34%), Positives = 36/70 (51%), Gaps = 6/70 (8%)
Query: 81 DNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGI---RSSGDNIVF--EVGMVT 135
DN V + + +T RI +L G++ + F W S+ I RSSG I F EVG+++
Sbjct: 77 DNVVVCFEDEVTAYFEPNRIKKLTGVKAKEFMVWISLGEIQVNRSSG-LITFKTEVGLLS 135
Query: 136 AKYPSKNFDD 145
P F+D
Sbjct: 136 KSLPLSVFED 145
>At1g30020 unknown protein
Length = 157
Score = 29.6 bits (65), Expect = 0.69
Identities = 21/78 (26%), Positives = 33/78 (41%), Gaps = 4/78 (5%)
Query: 45 PVGLLP--DTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAE 102
P GLLP D T +G+ N+T G +R+ + T Y IT + R+
Sbjct: 37 PTGLLPLKDMTEVGY--NKTKGFVWMRMRSKIEHTFREIGRRVLYDTEITAFVEDRRMRR 94
Query: 103 LNGIRVRAFFQWWSITGI 120
L G++ + W + I
Sbjct: 95 LTGVKSKELMIWVPVNDI 112
>At5g56040 receptor protein kinase-like protein
Length = 1090
Score = 28.1 bits (61), Expect = 2.0
Identities = 24/77 (31%), Positives = 37/77 (47%), Gaps = 13/77 (16%)
Query: 47 GLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKI--VKGRIAELN 104
G LP+ L +VNQ SG L K+T ++ +N I+G+I + G++ L
Sbjct: 334 GNLPNLQELQLSVNQLSGTIPEELANCTKLT-----HLEIDNNQISGEIPPLIGKLTSLT 388
Query: 105 GIRVRAFFQWWS-ITGI 120
FF W + +TGI
Sbjct: 389 -----MFFAWQNQLTGI 400
>At1g09310 unknown protein
Length = 179
Score = 27.7 bits (60), Expect = 2.6
Identities = 17/82 (20%), Positives = 34/82 (40%)
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L P GLLP + ++ SG ++ + + + +Y +T + G
Sbjct: 29 LKEISMPNGLLPLKDIEEVGYDRESGVVWLKQKKSITHKFTEIDKLVSYGTEVTAIVETG 88
Query: 99 RIAELNGIRVRAFFQWWSITGI 120
+I +L G++ + W +I I
Sbjct: 89 KIKKLTGVKAKELLIWVTINEI 110
>At4g23231 unknown protein
Length = 627
Score = 27.3 bits (59), Expect = 3.4
Identities = 12/25 (48%), Positives = 15/25 (60%)
Query: 8 LFALFLNLTSKSFQNPNPKPPKSTP 32
LFA + T ++ Q P P PP STP
Sbjct: 237 LFAFYNETTVRTQQAPPPLPPSSTP 261
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,762,555
Number of Sequences: 26719
Number of extensions: 154508
Number of successful extensions: 507
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 473
Number of HSP's gapped (non-prelim): 38
length of query: 156
length of database: 11,318,596
effective HSP length: 91
effective length of query: 65
effective length of database: 8,887,167
effective search space: 577665855
effective search space used: 577665855
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0153b.6


