

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.2
(402 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
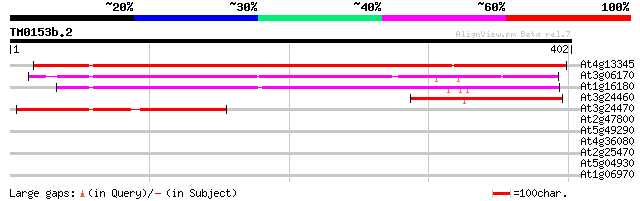
At4g13345 unknown protein 566 e-162
At3g06170 unknown protein 209 2e-54
At1g16180 193 1e-49
At3g24460 unknown protein 190 1e-48
At3g24470 hypothetical protein 169 2e-42
At2g47800 MRP-like ABC transporter 30 1.8
At5g49290 disease resistance protein-like 29 4.0
At4g36080 ATM-like protein 29 4.0
At2g25470 putative disease resistance protein 29 4.0
At5g04930 ATPase 29 5.2
At1g06970 hypothetical protein 28 8.9
>At4g13345 unknown protein
Length = 394
Score = 566 bits (1459), Expect = e-162
Identities = 263/382 (68%), Positives = 311/382 (80%), Gaps = 3/382 (0%)
Query: 18 LKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTCKDPK 77
+K SW QFRN NP MARYVY LIFL+ NLLAWA RD GR LT+++ K CK+
Sbjct: 15 IKNGSWFIQFRNGCNPWMARYVYGLIFLLANLLAWALRDY--GRGALTEMRKFKNCKEGG 72
Query: 78 DCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFL 137
DCLGT GVLRVS GCFLF+ +MF ST SK + +RD WHSGWW K+ + + +TIFPFL
Sbjct: 73 DCLGTEGVLRVSFGCFLFYFIMFLSTVGTSKTHSSRDKWHSGWWFAKLFMLLGLTIFPFL 132
Query: 138 LPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASY 197
LPS +I YGE+AHFGAGVFL IQLISIISFITWLN+ F ++K AERC +HVML AT +Y
Sbjct: 133 LPSSIIQFYGEIAHFGAGVFLLIQLISIISFITWLNECFQAQKDAERCHVHVMLLATTAY 192
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYV 257
+C++GVILMYIWY P+PSCLLNI FIT+TL L+Q+MTS+SLHPK+NAG L+P LMGLYV
Sbjct: 193 TVCILGVILMYIWYVPEPSCLLNIFFITWTLFLIQLMTSISLHPKINAGFLTPALMGLYV 252
Query: 258 VFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCFQYRK 317
VF+CWCAIRSEP G C RK++ ++TDW IIS VV +LA+VIATFSTG+DS+CFQ+RK
Sbjct: 253 VFICWCAIRSEPVGETCNRKAEGSSRTDWLTIISFVVALLAMVIATFSTGVDSQCFQFRK 312
Query: 318 GDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNE 377
D+ EED +PYGYGFFHFVFATGAMYFAMLLVGWN HHSMKKWTIDVGWTS WVRIVNE
Sbjct: 313 -DENHEEDAIPYGYGFFHFVFATGAMYFAMLLVGWNIHHSMKKWTIDVGWTSTWVRIVNE 371
Query: 378 WLAVCVYLWMLIAPIIWKNIHT 399
WLAV VY+WML+AP++ K+ T
Sbjct: 372 WLAVGVYIWMLVAPMVLKSRQT 393
>At3g06170 unknown protein
Length = 409
Score = 209 bits (532), Expect = 2e-54
Identities = 127/404 (31%), Positives = 200/404 (49%), Gaps = 39/404 (9%)
Query: 14 CGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTC 73
CG+ V+S +S+ K AR Y +F +++W R+ G +L KL + T
Sbjct: 14 CGLCSSVASGISR-------KSARIAYCGLFGASLVVSWILRET--GAPLLEKLPWINTS 64
Query: 74 KD-PKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVT 132
K+ VLRVS G FLFF + N+ RD+WH G W +K+++W +
Sbjct: 65 DSYTKEWYQQQAVLRVSFGNFLFFAIYALIMIGVKDQNDRRDSWHHGGWGLKMIVWFLLV 124
Query: 133 IFPFLLPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLF 192
+ F +P+ ++ LYG ++ FGAG FL +Q++ ++ ND + EK ++ I +++
Sbjct: 125 VLMFFVPNVIVSLYGTLSKFGAGAFLLVQVVLLLDATHNWNDSWV-EKDEKKWYIALLVI 183
Query: 193 ATASYFICMVGVILMYIWYAPQ-PSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPG 251
+ Y +++IW+ P C LN+ FI ++L + ++LHP VN +L
Sbjct: 184 SIVCYIATYTFSGILFIWFNPSGQDCGLNVFFIVMPMILAFVFAIIALHPAVNGSLLPAS 243
Query: 252 LMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATF------- 304
++ +Y ++C+ + SEP Y C + NK+ N +L++G+L V++
Sbjct: 244 VISVYCAYVCYTGLSSEPHDYVC----NGLNKSKAVNASTLILGMLTTVLSVLYSALRAG 299
Query: 305 -STGIDSKCFQYRKGDK--------------PAEEDDVPYGYGFFHFVFATGAMYFAMLL 349
ST S R G K AE V Y Y FFH +FA +MY AMLL
Sbjct: 300 SSTTFLSPPSSPRSGVKDALLGDPEDGKKSGEAEARPVSYSYSFFHIIFALASMYAAMLL 359
Query: 350 VGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPII 393
GW + S IDVGWTS WV+I W+ +Y+W LIAP+I
Sbjct: 360 SGW-TDSSESATLIDVGWTSVWVKICTGWVTAGLYIWTLIAPLI 402
>At1g16180
Length = 412
Score = 193 bits (491), Expect = 1e-49
Identities = 119/383 (31%), Positives = 189/383 (49%), Gaps = 26/383 (6%)
Query: 34 KMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTC-KDP-KDCLGTNGVLRVSMG 91
+ AR Y +F + +++W R+ ++ KL + K P ++ T+ VLRVS+G
Sbjct: 29 RSARIAYCGLFALSLIVSWILREV--AAPLMEKLPWINHFHKTPDREWFETDAVLRVSLG 86
Query: 92 CFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAH 151
FLFF ++ + RD H G W +KI+ W + IF F LP+E+I Y ++
Sbjct: 87 NFLFFSILSVMMIGVKNQKDPRDGIHHGGWMMKIICWCILVIFMFFLPNEIISFYESMSK 146
Query: 152 FGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFAT-ASYFICMVGVILMYIW 210
FGAG FL +Q++ ++ F+ ND + Y E+ +L + Y V ++ W
Sbjct: 147 FGAGFFLLVQVVLLLDFVHGWNDTWVG--YDEQFWYAALLVVSLVCYLATFVFSGFLFHW 204
Query: 211 YAPQ-PSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEP 269
+ P C LN FI TL+ + + V LHP V IL ++ LY ++LC+ + SEP
Sbjct: 205 FTPSGHDCGLNTFFIIMTLIFVFVFAIVVLHPTVGGSILPASVISLYCMYLCYSGLASEP 264
Query: 270 EGYECI-RKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCF---QYRKGDKP---- 321
YEC + S + I L+ +L++V + G + + +KP
Sbjct: 265 RDYECNGLHNHSKAVSTGTMTIGLLTTVLSVVYSAVRAGSSTTLLSPPDSPRAEKPLLPI 324
Query: 322 ---AEEDD-------VPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAW 371
AEE + V Y Y FFH +F+ +MY AMLL GW++ +DVGW S W
Sbjct: 325 DGKAEEKEEKENKKPVSYSYAFFHIIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVW 384
Query: 372 VRIVNEWLAVCVYLWMLIAPIIW 394
VR+V W +++W L+API++
Sbjct: 385 VRVVTSWATAGLFIWSLVAPILF 407
>At3g24460 unknown protein
Length = 126
Score = 190 bits (482), Expect = 1e-48
Identities = 86/113 (76%), Positives = 100/113 (88%), Gaps = 4/113 (3%)
Query: 288 NII-SLVVGILALVIATFSTGIDSKCFQYRKGDKPAEE---DDVPYGYGFFHFVFATGAM 343
NI+ S VV +LA+VIATFSTGIDS+CFQ++K + EE DDVPYGYGFFHFVFATGAM
Sbjct: 6 NIVQSFVVALLAMVIATFSTGIDSQCFQFKKDENDQEEEAEDDVPYGYGFFHFVFATGAM 65
Query: 344 YFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKN 396
YFAMLL+GWN+HH MKKWTIDVGWTS WVR+VNEWLAVCVY+WML+AP+I K+
Sbjct: 66 YFAMLLIGWNTHHPMKKWTIDVGWTSTWVRVVNEWLAVCVYIWMLVAPLILKS 118
>At3g24470 hypothetical protein
Length = 151
Score = 169 bits (429), Expect = 2e-42
Identities = 82/150 (54%), Positives = 103/150 (68%), Gaps = 8/150 (5%)
Query: 6 DNSSNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLT 65
+N S + +K SW +QFRN NP MARYVY LIFL+ NLLAWA+RD GR L
Sbjct: 10 NNQSIRNDSYEAIKNGSWFNQFRNGCNPWMARYVYGLIFLIANLLAWAARDY--GRGALR 67
Query: 66 KLKGLKTCKDPKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKI 125
K+ K CK ++CLGT+GVLR LF+ +MF ST SK + +RD WHSGWW +K+
Sbjct: 68 KVTRFKNCKGGENCLGTDGVLR------LFYFVMFLSTLGTSKTHSSRDRWHSGWWFVKL 121
Query: 126 VLWVAVTIFPFLLPSELIDLYGEVAHFGAG 155
++W A+TI PFLLPS +I LYGE+AHFGAG
Sbjct: 122 IMWPALTIIPFLLPSSIIHLYGEIAHFGAG 151
>At2g47800 MRP-like ABC transporter
Length = 1516
Score = 30.4 bits (67), Expect = 1.8
Identities = 31/143 (21%), Positives = 58/143 (39%), Gaps = 24/143 (16%)
Query: 119 GWWSIKIVLWVAVTIFPFLLPSELIDLYGEVA----HFGAGVFLFIQLISIISFITWLND 174
GWW I +VL+ ++T L+ S+ Y A F A VF+ +I +
Sbjct: 953 GWWGIVLVLFFSLTWQGSLMASDYWLAYETSAKNAISFDASVFILGYVIIAL-------- 1004
Query: 175 FFASEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIM 234
+ ++L + SY++ +G+ I++ + +L+ F +
Sbjct: 1005 ------------VSIVLVSIRSYYVTHLGLKTAQIFFRQILNSILHAPMSFFDTTPSGRI 1052
Query: 235 TSVSLHPKVNAGILSPGLMGLYV 257
S + + N IL P ++GL V
Sbjct: 1053 LSRASTDQTNVDILIPFMLGLVV 1075
>At5g49290 disease resistance protein-like
Length = 888
Score = 29.3 bits (64), Expect = 4.0
Identities = 18/75 (24%), Positives = 32/75 (42%), Gaps = 9/75 (12%)
Query: 309 DSKCFQYRKGDKPA---EEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDV 365
D+ C + ++ A EEDD F ++T Y L+ + +D
Sbjct: 813 DTSCETKKNSEENANGGEEDDKEVAIDMLVFYWSTAGTYVTALI------GILVLMCVDC 866
Query: 366 GWTSAWVRIVNEWLA 380
W AW+R+V+ ++A
Sbjct: 867 SWRRAWLRLVDAFIA 881
>At4g36080 ATM-like protein
Length = 3738
Score = 29.3 bits (64), Expect = 4.0
Identities = 9/20 (45%), Positives = 14/20 (70%)
Query: 381 VCVYLWMLIAPIIWKNIHTE 400
V +LW+L+ PI+W +H E
Sbjct: 2464 VAYHLWVLVFPIVWATLHKE 2483
>At2g25470 putative disease resistance protein
Length = 910
Score = 29.3 bits (64), Expect = 4.0
Identities = 17/64 (26%), Positives = 28/64 (43%), Gaps = 6/64 (9%)
Query: 317 KGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVN 376
+ D EE+D F F+T ++Y L+ + D W AW+RIV+
Sbjct: 846 EADNGQEEEDDKAAIDMMVFYFSTASIYVTALI------GVLVLMCFDCPWRRAWLRIVD 899
Query: 377 EWLA 380
++A
Sbjct: 900 AFIA 903
>At5g04930 ATPase
Length = 1158
Score = 28.9 bits (63), Expect = 5.2
Identities = 29/127 (22%), Positives = 49/127 (37%), Gaps = 20/127 (15%)
Query: 284 TDWQNIISLVV--GILALVIATFSTGIDSKCFQ-----YRKGDKPAEEDDVPYGYGFFHF 336
T+W +++ V+ I ++I + + Y G + + Y
Sbjct: 947 TEWSSVLYSVIYTAIPTIIIGILDKDLGRQTLLDHPQLYGVGQRAEGYSTTLFWYTMIDT 1006
Query: 337 VFATGAMYFAMLLVGWNSHHSMKKWTIDVG-----WTSAWVRIVNEWLAVCVYLWMLIA- 390
++ + A++F + W S TID WT A V +VN LA+ V W I
Sbjct: 1007 IWQSAAIFFIPMFAYWGS-------TIDTSSLGDLWTIAAVVVVNLHLAMDVIRWNWITH 1059
Query: 391 PIIWKNI 397
IW +I
Sbjct: 1060 AAIWGSI 1066
>At1g06970 hypothetical protein
Length = 829
Score = 28.1 bits (61), Expect = 8.9
Identities = 16/79 (20%), Positives = 35/79 (44%)
Query: 160 IQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQPSCLL 219
+++I + IT+ F + + C I + + + +C GVI +Y + +L
Sbjct: 359 VKIIEAVILITYGCKFLGTAAASAYCNIQIGDAFSLALLMCCQGVIEIYTCVMWKDEKVL 418
Query: 220 NISFITFTLVLLQIMTSVS 238
N ++ L ++T +S
Sbjct: 419 NTECFNLLIITLLLVTGIS 437
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.140 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,510,576
Number of Sequences: 26719
Number of extensions: 398364
Number of successful extensions: 977
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 957
Number of HSP's gapped (non-prelim): 12
length of query: 402
length of database: 11,318,596
effective HSP length: 102
effective length of query: 300
effective length of database: 8,593,258
effective search space: 2577977400
effective search space used: 2577977400
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0153b.2


