

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0150.12
(266 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
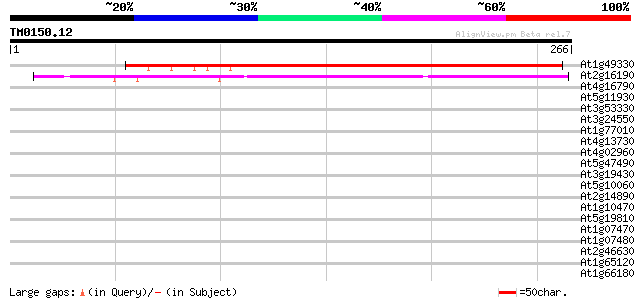
Score E
Sequences producing significant alignments: (bits) Value
At1g49330 hypothetical protein 202 2e-52
At2g16190 hypothetical protein 174 6e-44
At4g16790 glycoprotein homolog 33 0.16
At5g11930 glutaredoxin - like protein 33 0.21
At3g53330 putative protein 32 0.27
At3g24550 protein kinase, putative 32 0.35
At1g77010 32 0.35
At4g13730 unknown protein 32 0.46
At4g02960 putative polyprotein of LTR transposon 32 0.46
At5g47490 putative protein 31 0.60
At3g19430 putative late embryogenesis abundant protein 31 0.60
At5g10060 unknown protein 31 0.78
At2g14890 arabinogalactan-protein AGP9 31 0.78
At1g10470 putative response regulator 3 31 0.78
At5g19810 proline-rich protein 30 1.0
At1g07470 putative transcription factor IIA large subunit protein 30 1.0
At1g07480 transcription factor IIA large subunit 30 1.0
At2g46630 putative extensin 30 1.3
At1g65120 hypothetical protein 30 1.3
At1g66180 unknown protein 30 1.7
>At1g49330 hypothetical protein
Length = 331
Score = 202 bits (513), Expect = 2e-52
Identities = 107/221 (48%), Positives = 142/221 (63%), Gaps = 14/221 (6%)
Query: 56 PVHTTPTYM-NPFHVSLSPPS------PPSTFEPPLAS-TNDSID---SSSSRRVRRRV- 103
P H P +M N F + +PP PPS PP ++ T + + S R R R
Sbjct: 100 PSHQIPLWMSNYFQQTPNPPQLVTHFFPPSGLAPPSSNLTPPPVKRPVTGSVRIYRSRST 159
Query: 104 --EKSEIIPPPFPWATDRRARVYTLSHLLQNQIFTITGEVQCKRCERKFEMGFDLKKNFS 161
+KS+ I PPFPWAT+RR + +L +L NQI TITGEVQC+ CE+ +++ ++L++ F+
Sbjct: 160 VSKKSDTISPPFPWATNRRGEIQSLEYLESNQITTITGEVQCRHCEKVYQVSYNLRERFA 219
Query: 162 EVSSFIAKYECTMHDRAPTIWLNPKLPTCKHCGQENSVKPVMAEKKRSVNWLFLLLGQML 221
EV F + M DRA W P+ C+ CG+E +VKPV+AE+K +NWLFLLLGQ L
Sbjct: 220 EVVKFYLTEKRKMRDRAHKDWAYPEQRRCELCGREKAVKPVIAERKSQINWLFLLLGQTL 279
Query: 222 GCCTLEQLRYFCKHTKNHRTGAKDRVLYRAYVELCKQLCPE 262
G CTLEQL+ FCKH+KNHRTGAKDRVLY Y+ LCK L P+
Sbjct: 280 GFCTLEQLKNFCKHSKNHRTGAKDRVLYLTYMGLCKMLQPK 320
>At2g16190 hypothetical protein
Length = 303
Score = 174 bits (440), Expect = 6e-44
Identities = 105/267 (39%), Positives = 141/267 (52%), Gaps = 18/267 (6%)
Query: 12 QLICSSISSSTKSHPFFLSQMEMKKAENVKNNINDPL-ALSLSVEPVHT---TPTYMNPF 67
QL+ S +T+ P +M N N++ P AL +V P + TP P
Sbjct: 40 QLLTSDPPQNTQPSP--PQPNDMTSFANGTNHVIVPTQALEQAVPPPNVSVRTPLPYQPS 97
Query: 68 HVSLSPPSPPSTFEPPLASTNDSIDSSSSRR---------VRRRVEKSEIIPPPFPWATD 118
L PP LA+ R V R V EI+PP +PWAT
Sbjct: 98 EEVLPPPQLNQVATVALATPRRGRPPGGQARRNSKRPVAGVERNVGDREIVPP-YPWATK 156
Query: 119 RRARVYTLSHLLQNQIFTITGEVQCKRCERKFEMGFDLKKNFSEVSSFIAKYECTMHDRA 178
+ ++ + L N I I+G+V CK C+R + ++L++ FSE+ +I + M RA
Sbjct: 157 KPGKIQSFRDLSSNNINVISGQVHCKTCDRTDTVEYNLEEKFSELYGYIKVNKEEMRHRA 216
Query: 179 PTIWLNPKLPTCKHCGQENSVKPVMAEKKRSVNWLFLLLGQMLGCCTLEQLRYFCKHTKN 238
P W PKL C+ C E +KPVM+E+K +NWLFLLLGQMLGCCTL+QLRYFC+
Sbjct: 217 PGSWSTPKLIPCRTCKSE--MKPVMSERKEEINWLFLLLGQMLGCCTLDQLRYFCQLNSK 274
Query: 239 HRTGAKDRVLYRAYVELCKQLCPEWHF 265
HRTG+KDRV+Y Y+ LCKQL PE F
Sbjct: 275 HRTGSKDRVVYITYLSLCKQLDPEGPF 301
>At4g16790 glycoprotein homolog
Length = 473
Score = 33.1 bits (74), Expect = 0.16
Identities = 36/116 (31%), Positives = 49/116 (42%), Gaps = 22/116 (18%)
Query: 17 SISSSTKSHPFFLSQMEMKKAENVKNNI---NDPLALSLSVEPVHTTPTYMNP------- 66
S SSS+ S S E++ +VKN + PL +L+ ++P NP
Sbjct: 222 SRSSSSSSS----SSKEVESLPSVKNLTTVESQPLIKNLTPPSSFSSPRKSNPIPNLASE 277
Query: 67 FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRR---VEKS-----EIIPPPFP 114
FH S PP PP P +++ D RV RR V K+ E PPP P
Sbjct: 278 FHPSPPPPPPPPPPLPAFYNSSSRKDHPGIYRVERRESSVHKTKFAGGEFHPPPPP 333
>At5g11930 glutaredoxin - like protein
Length = 148
Score = 32.7 bits (73), Expect = 0.21
Identities = 23/78 (29%), Positives = 34/78 (43%), Gaps = 12/78 (15%)
Query: 34 MKKAENVKNNINDPLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPPLASTNDSID- 92
MK ++N ND + L L+V P P PP PST AST+ S D
Sbjct: 1 MKTMRGLRNCSNDAVTLDLTVHPPPPPPL----------PPPAPSTVSSSTASTSLSFDE 50
Query: 93 -SSSSRRVRRRVEKSEII 109
+S ++ R + + +I
Sbjct: 51 EETSESKIGRLISEHPVI 68
>At3g53330 putative protein
Length = 310
Score = 32.3 bits (72), Expect = 0.27
Identities = 31/94 (32%), Positives = 40/94 (41%), Gaps = 11/94 (11%)
Query: 20 SSTKSHPFFLSQMEMKKAENVKNNINDPLALSLSVEPVHTTPTYMNPFHVSLSPPSPPST 79
SS KSH L E + N +DPLA++ S P T P ++ SPP P T
Sbjct: 94 SSMKSHVRSLHHHEARPM-----NGHDPLAITPSPPPPSKTHERSRP--ITPSPPPPSKT 146
Query: 80 FEPPLAST-NDSIDSSSSRRVRRRVEKSEIIPPP 112
EP +T S + RR+ S PPP
Sbjct: 147 HEPSRPNTPPPPPPPSKTHEPSRRITPS---PPP 177
>At3g24550 protein kinase, putative
Length = 652
Score = 32.0 bits (71), Expect = 0.35
Identities = 14/32 (43%), Positives = 18/32 (55%)
Query: 59 TTPTYMNPFHVSLSPPSPPSTFEPPLASTNDS 90
T+P +P SL PPSPP + PPL + S
Sbjct: 52 TSPPPSSPLPPSLPPPSPPGSLTPPLPQPSPS 83
>At1g77010
Length = 695
Score = 32.0 bits (71), Expect = 0.35
Identities = 28/104 (26%), Positives = 40/104 (37%), Gaps = 16/104 (15%)
Query: 158 KNFSEVSSFIAKYECTMHDRAPTIWLNPKLPTCKHCGQENSVKPVMA--EKKRSVNWLFL 215
K FSEV S+ TI LN + CG+ + K V E K ++W +
Sbjct: 374 KLFSEVESY------------DTILLNSMIKVYFSCGRIDDAKRVFERIENKSLISWNSM 421
Query: 216 LLGQMLGCCTLEQLRYFCKHTKNHRTGAKDRVLYRAYVELCKQL 259
G CT+E L YF H + D V + + C +
Sbjct: 422 TNGFSQNGCTVETLEYF--HQMHKLDLPTDEVSLSSVISACASI 463
>At4g13730 unknown protein
Length = 449
Score = 31.6 bits (70), Expect = 0.46
Identities = 17/45 (37%), Positives = 26/45 (57%), Gaps = 1/45 (2%)
Query: 65 NPFHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVR-RRVEKSEI 108
NP S++PPSPP PP +S+ +I ++ + + R KSEI
Sbjct: 28 NPAATSITPPSPPIVERPPFSSSPSAISTAPAPPLPVRPPSKSEI 72
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 31.6 bits (70), Expect = 0.46
Identities = 32/112 (28%), Positives = 47/112 (41%), Gaps = 8/112 (7%)
Query: 5 TAAIYSSQLICSSISSSTKSHPFFLS----QMEMKKAENVKNNINDPLALSLSVEPVHTT 60
T + SS L SSISS + S P S Q + + +N N P+ + P +
Sbjct: 789 TTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSNSNSPILNN----PNPNS 844
Query: 61 PTYMNPFHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
P+ +P S P SP S+ P ST+ S +S S ++P P
Sbjct: 845 PSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSSSSTSTPPLPPVLPAP 896
>At5g47490 putative protein
Length = 1361
Score = 31.2 bits (69), Expect = 0.60
Identities = 19/65 (29%), Positives = 36/65 (55%), Gaps = 7/65 (10%)
Query: 32 MEMKKAENVKNNINDPLALSLSVEPVHTTPT------YMNPFHVSLSPPSPPSTFEPPLA 85
M M++ ++ N + L +S++ + H PT + F+ S +PP+ PSTF+P L
Sbjct: 1287 MMMQRYPSMDNIQRNGLGISVNGDN-HQPPTSRRTASWSGNFNTSFTPPTSPSTFKPVLL 1345
Query: 86 STNDS 90
+++ S
Sbjct: 1346 NSSSS 1350
>At3g19430 putative late embryogenesis abundant protein
Length = 550
Score = 31.2 bits (69), Expect = 0.60
Identities = 17/58 (29%), Positives = 32/58 (54%), Gaps = 5/58 (8%)
Query: 54 VEPVHTTPTYMNPFHVSLSPPSPPST-----FEPPLASTNDSIDSSSSRRVRRRVEKS 106
V P TP+ +P V+ +PP+PPS P + +D +++ ++RVR + ++S
Sbjct: 213 VTPTPPTPSVPSPPDVTPTPPTPPSVPTPSGSPPYVPPPSDEEEAAGAKRVRCKKQRS 270
>At5g10060 unknown protein
Length = 469
Score = 30.8 bits (68), Expect = 0.78
Identities = 25/85 (29%), Positives = 37/85 (43%), Gaps = 18/85 (21%)
Query: 14 ICSSISSSTKSHPFFLSQMEMKKAENVKNNINDPLALSLSVEPV--------------HT 59
I + +++ST SH S + AE K + L+ S S PV +
Sbjct: 326 IAAMLTASTSSHMIMQSVLSSFAAEATKTS---GLSKSESTVPVSDTNASFPSYNNSQNQ 382
Query: 60 TPTYMNPFHVSLSPPSPPSTFEPPL 84
TPT +HV +PP PP +PP+
Sbjct: 383 TPTTQGQYHVIPNPP-PPQFLKPPV 406
>At2g14890 arabinogalactan-protein AGP9
Length = 191
Score = 30.8 bits (68), Expect = 0.78
Identities = 24/76 (31%), Positives = 31/76 (40%), Gaps = 7/76 (9%)
Query: 44 INDPLALSLSVEPVHTTPTYMNPFHVSLSPP--SPPSTFEPPLASTNDSIDSSSSRRVRR 101
++ P ++ S PV T P NP SPP SPP PP+AS + S
Sbjct: 48 VSAPPPVTTSPPPVTTAPPPANPPPPVSSPPPASPPPATPPPVASPPPPVASPPP----- 102
Query: 102 RVEKSEIIPPPFPWAT 117
PPP P A+
Sbjct: 103 ATPPPVATPPPAPLAS 118
>At1g10470 putative response regulator 3
Length = 259
Score = 30.8 bits (68), Expect = 0.78
Identities = 24/93 (25%), Positives = 44/93 (46%), Gaps = 8/93 (8%)
Query: 8 IYSSQLICSSISSSTK--SHPFFLSQMEMKKAENVKNNINDPLALSLSVEPVHTTPTYMN 65
I SS+ + + I + + F L +++ + +++++ + LS + P +
Sbjct: 123 IMSSENVLTRIDRCLEEGAQDFLLKPVKLADVKRLRSHLTKDVKLSNGNK--RKLPEDSS 180
Query: 66 PFHVSLSPPSPPSTF----EPPLASTNDSIDSS 94
+ SL PPSPP T PPL + +S DSS
Sbjct: 181 SVNSSLPPPSPPLTISPESSPPLTVSTESSDSS 213
>At5g19810 proline-rich protein
Length = 249
Score = 30.4 bits (67), Expect = 1.0
Identities = 17/37 (45%), Positives = 20/37 (53%), Gaps = 3/37 (8%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPP 83
P+ LS PV+ +P P V LSPP PP F PP
Sbjct: 129 PVLLSPPPPPVNLSPP---PPPVLLSPPPPPVLFSPP 162
Score = 29.3 bits (64), Expect = 2.3
Identities = 16/42 (38%), Positives = 21/42 (49%), Gaps = 3/42 (7%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPPLASTN 88
P+ +S PV+ +P P V+LSPP PP PP N
Sbjct: 48 PVNISSPPPPVNLSPP---PPPVNLSPPPPPVNLSPPPPPVN 86
Score = 29.3 bits (64), Expect = 2.3
Identities = 16/37 (43%), Positives = 20/37 (53%), Gaps = 3/37 (8%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPP 83
P+ LS PV+ +P P V+LSPP PP PP
Sbjct: 93 PVLLSPPPPPVNLSPP---PPPVNLSPPPPPVLLSPP 126
Score = 29.3 bits (64), Expect = 2.3
Identities = 17/42 (40%), Positives = 21/42 (49%), Gaps = 3/42 (7%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPPLASTN 88
P+ LS PV+ +P P V+LSPP PP PP N
Sbjct: 66 PVNLSPPPPPVNLSPP---PPPVNLSPPPPPVLLSPPPPPVN 104
Score = 28.5 bits (62), Expect = 3.9
Identities = 17/42 (40%), Positives = 20/42 (47%), Gaps = 3/42 (7%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPPLASTN 88
P+ LS PV+ +P P V LSPP PP PP N
Sbjct: 75 PVNLSPPPPPVNLSPP---PPPVLLSPPPPPVNLSPPPPPVN 113
Score = 28.1 bits (61), Expect = 5.1
Identities = 17/42 (40%), Positives = 20/42 (47%), Gaps = 3/42 (7%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPPLASTN 88
P+ LS PV+ +P P V LSPP PP PP N
Sbjct: 102 PVNLSPPPPPVNLSPP---PPPVLLSPPPPPVLLSPPPPPVN 140
Score = 27.7 bits (60), Expect = 6.6
Identities = 16/37 (43%), Positives = 19/37 (51%), Gaps = 3/37 (8%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPP 83
P+ LS PV +P P V+LSPP PP PP
Sbjct: 84 PVNLSPPPPPVLLSPP---PPPVNLSPPPPPVNLSPP 117
>At1g07470 putative transcription factor IIA large subunit protein
Length = 375
Score = 30.4 bits (67), Expect = 1.0
Identities = 28/89 (31%), Positives = 35/89 (38%), Gaps = 17/89 (19%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEP-PLASTNDS-----IDSSSSR--- 97
PL L+V P T Y P L PP+P T P PL T D+ I + SS
Sbjct: 71 PLTHDLNV-PYEGTEEYETPTAEMLFPPTPLQTPLPTPLPGTADNSSMYNIPTGSSDYPT 129
Query: 98 -------RVRRRVEKSEIIPPPFPWATDR 119
+ + S +PPP PW R
Sbjct: 130 PGTENGVNIDVKARPSPYMPPPSPWTNPR 158
>At1g07480 transcription factor IIA large subunit
Length = 375
Score = 30.4 bits (67), Expect = 1.0
Identities = 28/89 (31%), Positives = 35/89 (38%), Gaps = 17/89 (19%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEP-PLASTNDS-----IDSSSSR--- 97
PL L+V P T Y P L PP+P T P PL T D+ I + SS
Sbjct: 71 PLTHDLNV-PYEGTEEYETPTAEMLFPPTPLQTPLPTPLPGTADNSSMYNIPTGSSDYPT 129
Query: 98 -------RVRRRVEKSEIIPPPFPWATDR 119
+ + S +PPP PW R
Sbjct: 130 PGTENGVNIDVKARPSPYMPPPSPWTNPR 158
>At2g46630 putative extensin
Length = 394
Score = 30.0 bits (66), Expect = 1.3
Identities = 21/72 (29%), Positives = 28/72 (38%), Gaps = 9/72 (12%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVS----LSPPSPPSTFEPPLASTNDSIDSSSSRRVRRR 102
PL P + P +P+H +SPP+PP PP S S S +
Sbjct: 73 PLTPPRQKAPPTSPPQERSPYHSPPSRHMSPPTPPKAATPPPPPPRSSYTSPPSPK---- 128
Query: 103 VEKSEIIPPPFP 114
E E +PP P
Sbjct: 129 -EVQEALPPRKP 139
>At1g65120 hypothetical protein
Length = 1191
Score = 30.0 bits (66), Expect = 1.3
Identities = 20/66 (30%), Positives = 32/66 (48%), Gaps = 7/66 (10%)
Query: 41 KNNINDPLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVR 100
KN N + S+S + VH + + P ++PPSP ST E + + D+ SS R R
Sbjct: 758 KNRSNKRTSASMSKDDVHESSVNLEP---KVTPPSPKSTEEDSM----EPEDTLSSERGR 810
Query: 101 RRVEKS 106
+ +
Sbjct: 811 LEISSN 816
>At1g66180 unknown protein
Length = 430
Score = 29.6 bits (65), Expect = 1.7
Identities = 27/112 (24%), Positives = 45/112 (40%), Gaps = 19/112 (16%)
Query: 1 MSFHTAAIYSSQLICSSISSSTKSHPFFLSQMEMKKAE------NVKNNINDPLALSLSV 54
+S T+ L IS++T SH F S + K N ++ +AL +S+
Sbjct: 17 VSLSTSLSLHLPLTSLPISTTTNSHRFTTSLLSRKNPSPSSPPYNFRSRFKYSMALIISL 76
Query: 55 EPVHTTPTYMNPF------------HVSLSPPSPPSTFEPPLASTNDSIDSS 94
P+ T P H PP P ++F+P L+S+ ++ S
Sbjct: 77 -PIGTPPQAQQMVLDTGSQLSWIQCHRKKLPPKPKTSFDPSLSSSFSTLPCS 127
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.133 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,289,341
Number of Sequences: 26719
Number of extensions: 267380
Number of successful extensions: 1483
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 1379
Number of HSP's gapped (non-prelim): 127
length of query: 266
length of database: 11,318,596
effective HSP length: 98
effective length of query: 168
effective length of database: 8,700,134
effective search space: 1461622512
effective search space used: 1461622512
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0150.12


