

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0148.11
(156 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
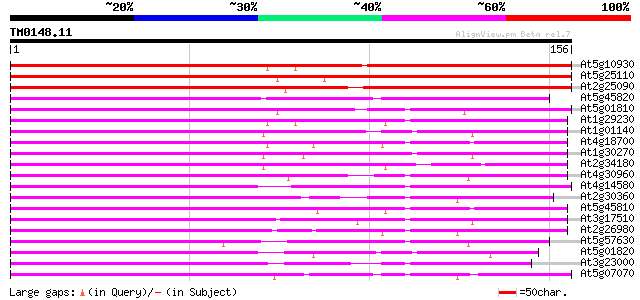
Score E
Sequences producing significant alignments: (bits) Value
At5g10930 serine/threonine protein kinase -like protein 164 2e-41
At5g25110 serine/threonine protein kinase-like protein 163 4e-41
At2g25090 CBL-interacting protein kinase 16 (CIPK16) 142 7e-35
At5g45820 CBL-interacting protein kinase 20 (CIPK20) 130 3e-31
At5g01810 serine/threonine protein kinase ATPK10 109 5e-25
At1g29230 CBL-interacting protein kinase 18 (CIPK18) 109 5e-25
At1g01140 SOS2-like protein kinase PKS6/CBL-interacting protein ... 107 3e-24
At4g18700 putative protein kinase 104 2e-23
At1g30270 serine/threonine kinase, putative 104 2e-23
At2g34180 putative protein kinase 101 2e-22
At4g30960 CBL-interacting protein kinase 6 (CIPK6) 96 1e-20
At4g14580 CBL-interacting protein kinase 4 (CIPK4) 94 2e-20
At2g30360 putative protein kinase 94 3e-20
At5g45810 CBL-interacting protein kinase 19 (CIPK19) 94 4e-20
At3g17510 CBL-interacting protein kinase 1 (CIPK1) 93 5e-20
At2g26980 putative protein kinase (At2g26980) 93 5e-20
At5g57630 CBL-interacting protein kinase 21 (CIPK21) 92 9e-20
At5g01820 unknown protein 91 2e-19
At3g23000 SNF1 related protein kinase (ATSRPK1) 91 3e-19
At5g07070 serine/threonine protein kinase-like protein 89 1e-18
>At5g10930 serine/threonine protein kinase -like protein
Length = 445
Score = 164 bits (414), Expect = 2e-41
Identities = 84/163 (51%), Positives = 111/163 (67%), Gaps = 8/163 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAGCLPFQ ENLM MY K+ RA+F+FPPWFSPE+++LISK+LV DP+ RI+I +IM
Sbjct: 203 LYVLLAGCLPFQDENLMNMYRKIFRADFEFPPWFSPEARRLISKLLVVDPDRRISIPAIM 262
Query: 61 RVSWFQKGFS--ASIPIPDP-----DESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSM 113
R W +K F+ + I +P ++N + + E T+ + + KF NAFEFISSM
Sbjct: 263 RTPWLRKNFTPPLAFKIDEPICSQSSKNNEEEEEDGDCENQTEPI-SPKFFNAFEFISSM 321
Query: 114 SSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
SSGFDLS FE KR+ SVFTS+ S +E+ KIE K + +
Sbjct: 322 SSGFDLSSLFESKRKVQSVFTSRSSATEVMEKIETVTKEMNMK 364
>At5g25110 serine/threonine protein kinase-like protein
Length = 488
Score = 163 bits (412), Expect = 4e-41
Identities = 84/161 (52%), Positives = 110/161 (68%), Gaps = 5/161 (3%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPFQ ENLM MY K+ ++EF++PPWFSPESK+LISK+LV DPN RI+I +IM
Sbjct: 233 LYVLLAGFLPFQDENLMKMYRKIFKSEFEYPPWFSPESKRLISKLLVVDPNKRISIPAIM 292
Query: 61 RVSWFQKGFSASI--PIPDPDESNFNSD---LNSSSEQSTKVVAAAKFINAFEFISSMSS 115
R WF+K ++ I I + + N + +++ +T + KF NAFEFISSMSS
Sbjct: 293 RTPWFRKNINSPIEFKIDELEIQNVEDETPTTTATTATTTTTPVSPKFFNAFEFISSMSS 352
Query: 116 GFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
GFDLS FE KR+ S+FTS+ S SEI K+EG K + +
Sbjct: 353 GFDLSSLFESKRKLRSMFTSRWSASEIMGKLEGIGKEMNMK 393
>At2g25090 CBL-interacting protein kinase 16 (CIPK16)
Length = 469
Score = 142 bits (358), Expect = 7e-35
Identities = 74/165 (44%), Positives = 102/165 (60%), Gaps = 13/165 (7%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALLAG LPF EN+MT+Y K+ +AE +FPPWFS ESK+L+S++LV DP RI++S I
Sbjct: 214 LYALLAGFLPFIDENVMTLYTKIFKAECEFPPWFSLESKELLSRLLVPDPEQRISMSEIK 273
Query: 61 RVSWFQKGFSASIPI---------PDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFIS 111
+ WF+K F+ S+ P+P DLN + A+ + NAF+FI+
Sbjct: 274 MIPWFRKNFTPSVAFSIDETIPSPPEPPTKKKKKDLNEKEDDG----ASPRSFNAFQFIT 329
Query: 112 SMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
SMSSGFDLS FE KR+ +FTSK + ++E AA+ + R
Sbjct: 330 SMSSGFDLSNLFEIKRKPKRMFTSKFPAKSVKERLETAAREMDMR 374
>At5g45820 CBL-interacting protein kinase 20 (CIPK20)
Length = 439
Score = 130 bits (327), Expect = 3e-31
Identities = 74/150 (49%), Positives = 90/150 (59%), Gaps = 3/150 (2%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPF +NL+ MY K+ + EF+ P WF PE KKL+S+IL +PN RI I IM
Sbjct: 202 LYVLLAGFLPFHEQNLVEMYRKITKGEFKCPNWFPPEVKKLLSRILDPNPNSRIKIEKIM 261
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
SWFQKGF I P ES+ L S + V + NAF+ ISS+S GFDLS
Sbjct: 262 ENSWFQKGFK-KIETPKSPESHQIDSLISDVHAAFSVKPMS--YNAFDLISSLSQGFDLS 318
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASKIEGAA 150
G FE++ R S FT+K EI SK E A
Sbjct: 319 GLFEKEERSESKFTTKKDAKEIVSKFEEIA 348
>At5g01810 serine/threonine protein kinase ATPK10
Length = 421
Score = 109 bits (273), Expect = 5e-25
Identities = 62/159 (38%), Positives = 96/159 (59%), Gaps = 7/159 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF+ NLM +Y K+ +AE +FP W +P +K+L+ +IL +PN R++ IM
Sbjct: 202 LFVLLAGYLPFRDSNLMELYKKIGKAEVKFPNWLAPGAKRLLKRILDPNPNTRVSTEKIM 261
Query: 61 RVSWFQKGFSASI--PIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFD 118
+ SWF+KG + + + E + ++ N+S+E+ K +NAFE I S+S+GFD
Sbjct: 262 KSSWFRKGLQEEVKESVEEETEVDAEAEGNASAEKEKK---RCINLNAFEII-SLSTGFD 317
Query: 119 LSGFFEE-KRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
LSG FE+ + + FTS SEI K+ K L+ +
Sbjct: 318 LSGLFEKGEEKEEMRFTSNREASEITEKLVEIGKDLKMK 356
>At1g29230 CBL-interacting protein kinase 18 (CIPK18)
Length = 520
Score = 109 bits (273), Expect = 5e-25
Identities = 67/175 (38%), Positives = 99/175 (56%), Gaps = 21/175 (12%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG +PF +N+M MY K+ + EF+ P WFS + +L++++L +P+ RITI IM
Sbjct: 264 LFVLMAGHIPFYDKNIMVMYKKIYKGEFRCPRWFSSDLVRLLTRLLDTNPDTRITIPEIM 323
Query: 61 RVSWFQKGFS-ASIPIPDP----DESNFNSDLNSSSEQSTKVVAAAKF------------ 103
+ WF+KGF I D ++ + + +SS ST + A+F
Sbjct: 324 KNRWFKKGFKHVKFYIEDDKLCREDEDEEEEASSSGRSSTVSESDAEFDVKRMGIGSMPR 383
Query: 104 ---INAFEFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
+NAF+ I S SSGFDLSG FEE+ G+ F S VS+I SK+E AK + F
Sbjct: 384 PSSLNAFDII-SFSSGFDLSGLFEEEGGEGTRFVSGAPVSKIISKLEEIAKIVSF 437
>At1g01140 SOS2-like protein kinase PKS6/CBL-interacting protein
kinase 9 (CIPK9)
Length = 447
Score = 107 bits (267), Expect = 3e-24
Identities = 68/166 (40%), Positives = 96/166 (56%), Gaps = 16/166 (9%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF NLMT+Y ++ +AEF PPWFS +K++I +IL +P RI+I+ ++
Sbjct: 210 LFVLMAGYLPFDEPNLMTLYKRICKAEFSCPPWFSQGAKRVIKRILEPNPITRISIAELL 269
Query: 61 RVSWFQKGF--------SASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISS 112
WF+KG+ I I D D + NS +E+ K V+ +NAFE ISS
Sbjct: 270 EDEWFKKGYKPPSFDQDDEDITIDDVDAAFSNSKECLVTEKKEKPVS----MNAFELISS 325
Query: 113 MSSGFDLSGFFEEKR---RGGSVFTSKCSVSEIASKIEGAAKGLRF 155
SS F L FE++ + + FTS+ S SEI SK+E AK L F
Sbjct: 326 -SSEFSLENLFEKQAQLVKKETRFTSQRSASEIMSKMEETAKPLGF 370
>At4g18700 putative protein kinase
Length = 489
Score = 104 bits (260), Expect = 2e-23
Identities = 69/177 (38%), Positives = 93/177 (51%), Gaps = 24/177 (13%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF N+M MY K+ R EF+ P WFS E +L+SK+L +P R T IM
Sbjct: 216 LFVLMAGYLPFHDRNVMAMYKKIYRGEFRCPRWFSTELTRLLSKLLETNPEKRFTFPEIM 275
Query: 61 RVSWFQKGFS-ASIPIPDPDESNF--NSDLNSSSEQSTKVVAAAK--------------- 102
SWF+KGF + D N + +L S S +S + AA++
Sbjct: 276 ENSWFKKGFKHIKFYVEDDKLCNVVDDDELESDSVESDRDSAASESEIEYLEPRRRVGGL 335
Query: 103 ----FINAFEFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
+NAF+ I S S GFDLSG F++ GS F S VS+I SK+E AK + F
Sbjct: 336 PRPASLNAFDII-SFSQGFDLSGLFDDDGE-GSRFVSGAPVSKIISKLEEIAKVVSF 390
>At1g30270 serine/threonine kinase, putative
Length = 482
Score = 104 bits (260), Expect = 2e-23
Identities = 62/166 (37%), Positives = 94/166 (56%), Gaps = 12/166 (7%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF+ NL ++Y K+ +AEF PPWFS +KKLI +IL +P RIT + ++
Sbjct: 222 LFVLMAGYLPFEDSNLTSLYKKIFKAEFTCPPWFSASAKKLIKRILDPNPATRITFAEVI 281
Query: 61 RVSWFQKGF------SASIPIPDPDE--SNFNSDLNSSSEQSTKVVAAAKFINAFEFISS 112
WF+KG+ +A + + D D + N E+ + + +NAFE IS+
Sbjct: 282 ENEWFKKGYKAPKFENADVSLDDVDAIFDDSGESKNLVVERREEGLKTPVTMNAFELIST 341
Query: 113 MSSGFDLSGFFEEKR---RGGSVFTSKCSVSEIASKIEGAAKGLRF 155
S G +L FE++ + + FTSK S +EI +KIE AA + F
Sbjct: 342 -SQGLNLGSLFEKQMGLVKRKTRFTSKSSANEIVTKIEAAAAPMGF 386
>At2g34180 putative protein kinase
Length = 502
Score = 101 bits (251), Expect = 2e-22
Identities = 62/176 (35%), Positives = 95/176 (53%), Gaps = 26/176 (14%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF +N++ MY K+ + +F+ P WFSPE +L++++L +P+ RITI IM
Sbjct: 247 LFVLMAGYLPFDDKNILVMYTKIYKGQFKCPKWFSPELARLVTRMLDTNPDTRITIPEIM 306
Query: 61 RVSWFQKGF--------SASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKF--------- 103
+ WF+KGF + + D D + +S SS ST A+F
Sbjct: 307 KHRWFKKGFKHVKFYIENDKLCREDDDNDDDDSSSLSSGRSSTASEGDAEFDIKRVDSMP 366
Query: 104 ----INAFEFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
+NAF+ +S DLSG FEE +G F S +++I SK+E AK ++F
Sbjct: 367 RPASLNAFDILSFS----DLSGLFEEGGQGAR-FVSAAPMTKIISKLEEIAKEVKF 417
>At4g30960 CBL-interacting protein kinase 6 (CIPK6)
Length = 441
Score = 95.5 bits (236), Expect = 1e-20
Identities = 61/164 (37%), Positives = 93/164 (56%), Gaps = 17/164 (10%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPFQ +NL+ MY K+ R +F+ P W S ++++L++K+L +PN RITI +M
Sbjct: 214 LFVLLAGYLPFQDDNLVNMYRKIYRGDFKCPGWLSSDARRLVTKLLDPNPNTRITIEKVM 273
Query: 61 RVSWFQKGFSASIPIP-----DPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSS 115
WF+K + S P E + + ++ S E++ + +NAF I ++S
Sbjct: 274 DSPWFKKQATRSRNEPVAATITTTEEDVDFLVHKSKEET-------ETLNAFHII-ALSE 325
Query: 116 GFDLSGFFEEK----RRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
GFDLS FEEK +R TS+ + S I+S E A G +F
Sbjct: 326 GFDLSPLFEEKKKEEKREMRFATSRPASSVISSLEEAARVGNKF 369
>At4g14580 CBL-interacting protein kinase 4 (CIPK4)
Length = 426
Score = 94.4 bits (233), Expect = 2e-20
Identities = 52/156 (33%), Positives = 82/156 (52%), Gaps = 10/156 (6%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG +PF N++ MY K+ + +++FP W S ++ +I K+L +P R++I ++M
Sbjct: 213 LFVLLAGYVPFDDANIVAMYRKIHKRDYRFPSWISKPARSIIYKLLDPNPETRMSIEAVM 272
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
WFQK + S F S + K ++ I AF+ I S+SSG DLS
Sbjct: 273 GTVWFQKSL---------EISEFQSSVFELDRFLEKEAKSSNAITAFDLI-SLSSGLDLS 322
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
G FE ++R FT++ S + K + L FR
Sbjct: 323 GLFERRKRKEKRFTARVSAERVVEKAGMIGEKLGFR 358
>At2g30360 putative protein kinase
Length = 435
Score = 94.0 bits (232), Expect = 3e-20
Identities = 57/154 (37%), Positives = 83/154 (53%), Gaps = 13/154 (8%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF N+M MY K+ + E++FP W SP+ K+ +S++L +P RITI I+
Sbjct: 213 LFVLVAGYLPFNDPNVMNMYKKIYKGEYRFPRWMSPDLKRFVSRLLDINPETRITIDEIL 272
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
+ WF +G I D + + + SS E A K +NAF+ I S SSG DLS
Sbjct: 273 KDPWFVRGGFKQIKFHDDEIE--DQKVESSLE-------AVKSLNAFDLI-SYSSGLDLS 322
Query: 121 GFF---EEKRRGGSVFTSKCSVSEIASKIEGAAK 151
G F F S+ S +A ++EG A+
Sbjct: 323 GLFAGCSNSSGESERFLSEKSPEMLAEEVEGFAR 356
>At5g45810 CBL-interacting protein kinase 19 (CIPK19)
Length = 483
Score = 93.6 bits (231), Expect = 4e-20
Identities = 63/179 (35%), Positives = 86/179 (47%), Gaps = 26/179 (14%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF N+M MY K+ R +F+ P WF E +L+ ++L P R T+ IM
Sbjct: 218 LFVLMAGFLPFHDRNVMAMYKKIYRGDFRCPRWFPVEINRLLIRMLETKPERRFTMPDIM 277
Query: 61 RVSWFQKGFSASIPIPDPDESNFN-------SDLNSSSEQSTKVVAAAKF---------- 103
SWF+KGF + D N + S S +S+ V F
Sbjct: 278 ETSWFKKGFKHIKFYVEDDHQLCNVADDDEIESIESVSGRSSTVSEPEDFESFDGRRRGG 337
Query: 104 -------INAFEFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
+NAF+ I S S GFDLSG FE+ GS F S V +I SK+E A+ + F
Sbjct: 338 SMPRPASLNAFDLI-SFSPGFDLSGLFEDDGE-GSRFVSGAPVGQIISKLEEIARIVSF 394
>At3g17510 CBL-interacting protein kinase 1 (CIPK1)
Length = 444
Score = 93.2 bits (230), Expect = 5e-20
Identities = 59/162 (36%), Positives = 84/162 (51%), Gaps = 9/162 (5%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY +L GCLPF NL +Y K+ + + P W SP ++ +I ++L +P RIT+ I
Sbjct: 211 LYVILTGCLPFDDRNLAVLYQKICKGDPPIPRWLSPGARTMIKRMLDPNPVTRITVVGIK 270
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQST-----KVVAAAKFINAFEFISSMSS 115
WF+ + SIP D DE ++D ++ S Q K + INAF+ I MSS
Sbjct: 271 ASEWFKLEYIPSIP-DDDDEEEVDTDDDAFSIQELGSEEGKGSDSPTIINAFQLI-GMSS 328
Query: 116 GFDLSGFFEEKR--RGGSVFTSKCSVSEIASKIEGAAKGLRF 155
DLSGFFE++ FTS S ++ KIE A + F
Sbjct: 329 FLDLSGFFEQENVSERRIRFTSNSSAKDLLEKIETAVTEMGF 370
>At2g26980 putative protein kinase (At2g26980)
Length = 441
Score = 93.2 bits (230), Expect = 5e-20
Identities = 61/163 (37%), Positives = 88/163 (53%), Gaps = 11/163 (6%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPF NLM +Y K+ EF PPW S + KLI++IL +P R+T +
Sbjct: 205 LYVLLAGYLPFDDSNLMNLYKKISSGEFNCPPWLSLGAMKLITRILDPNPMTRVTPQEVF 264
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAK------FINAFEFISSMS 114
WF+K + + + D+SN + D+++ + S + + K INAFE I SMS
Sbjct: 265 EDEWFKKDYKPPV-FEERDDSNMD-DIDAVFKDSEEHLVTEKREEQPAAINAFEII-SMS 321
Query: 115 SGFDLSGFF--EEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
G +L F E++ + + T + +EI KIE AAK L F
Sbjct: 322 RGLNLENLFDPEQEFKRETRITLRGGANEIIEKIEEAAKPLGF 364
>At5g57630 CBL-interacting protein kinase 21 (CIPK21)
Length = 416
Score = 92.4 bits (228), Expect = 9e-20
Identities = 59/153 (38%), Positives = 85/153 (54%), Gaps = 11/153 (7%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISS-I 59
L+ LLAG PF L +Y K+LRA++ FPP F+ E K+LI IL +P RIT++ I
Sbjct: 199 LFELLAGYPPFDDHTLPVLYKKILRADYTFPPGFTGEQKRLIFNILDPNPLSRITLAEII 258
Query: 60 MRVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDL 119
++ SWF+ G++ P + + + + A++ FINAF+ I +MSS DL
Sbjct: 259 IKDSWFKIGYT-------PVYHQLSDSIKDNVAEINAATASSNFINAFQII-AMSSDLDL 310
Query: 120 SGFFEEK--RRGGSVFTSKCSVSEIASKIEGAA 150
SG FEE +R + SK + E KIE AA
Sbjct: 311 SGLFEENDDKRYKTRIGSKNTAQETIKKIEAAA 343
>At5g01820 unknown protein
Length = 442
Score = 91.3 bits (225), Expect = 2e-19
Identities = 53/152 (34%), Positives = 86/152 (55%), Gaps = 14/152 (9%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L AG LPF NLM MY K+ + EF+ P W SP+ ++L++++L +P RITI I+
Sbjct: 212 LFVLNAGYLPFNDHNLMVMYRKIYKGEFRIPKWTSPDLRRLLTRLLDTNPQTRITIEEII 271
Query: 61 RVSWFQKGFSASIPIPDPDESNF---NSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGF 117
WF++G+ D S F +SD+ ++++ + A + +NAF+ IS S GF
Sbjct: 272 HDPWFKQGY-------DDRMSKFHLEDSDMKLPADETDSEMGARR-MNAFDIISG-SPGF 322
Query: 118 DLSGFFEEKRRGGSV--FTSKCSVSEIASKIE 147
+LSG F + R+ V F S + + ++E
Sbjct: 323 NLSGLFGDARKYDRVERFVSAWTAERVVERLE 354
>At3g23000 SNF1 related protein kinase (ATSRPK1)
Length = 429
Score = 90.5 bits (223), Expect = 3e-19
Identities = 50/145 (34%), Positives = 83/145 (56%), Gaps = 13/145 (8%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LL G +PF N+ MY K+ R +++FP W S ++K +I ++L +P R++I ++M
Sbjct: 216 LFVLLVGDVPFDDSNIAAMYRKIHRRDYRFPSWISKQAKSIIYQMLDPNPVTRMSIETVM 275
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
+ +WF+K S + + F+S++ S ++ I AF+ I S+SSG DLS
Sbjct: 276 KTNWFKKSLETS----EFHRNVFDSEVEMKSSVNS--------ITAFDLI-SLSSGLDLS 322
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASK 145
G FE K++ FT+K S E+ K
Sbjct: 323 GLFEAKKKKERRFTAKVSGVEVEEK 347
>At5g07070 serine/threonine protein kinase-like protein
Length = 456
Score = 89.0 bits (219), Expect = 1e-18
Identities = 64/171 (37%), Positives = 90/171 (52%), Gaps = 21/171 (12%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF NLM MY K+ +A+F+ P WF+PE K+L+ K+L + RITI+ I
Sbjct: 202 LFVLLAGYLPFHDTNLMEMYRKIGKADFKCPSWFAPEVKRLLCKMLDPNHETRITIAKIK 261
Query: 61 RVSWFQKGFSAS------------IPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFE 108
SWF+KG +P E+ S N + E A +NAF+
Sbjct: 262 ESSWFRKGLHLKQKKMEKMEKQQVREATNPMEAG-GSGQNENGENHEPPRLAT--LNAFD 318
Query: 109 FISSMSSGFDLSGFF---EEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
I ++S+GF L+G F +KR S F S+ SEI SK+ AK L+ +
Sbjct: 319 II-ALSTGFGLAGLFGDVYDKRE--SRFASQKPASEIISKLVEVAKCLKLK 366
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,286,378
Number of Sequences: 26719
Number of extensions: 122471
Number of successful extensions: 651
Number of sequences better than 10.0: 147
Number of HSP's better than 10.0 without gapping: 105
Number of HSP's successfully gapped in prelim test: 42
Number of HSP's that attempted gapping in prelim test: 488
Number of HSP's gapped (non-prelim): 155
length of query: 156
length of database: 11,318,596
effective HSP length: 91
effective length of query: 65
effective length of database: 8,887,167
effective search space: 577665855
effective search space used: 577665855
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0148.11


