

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0142.11
(270 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
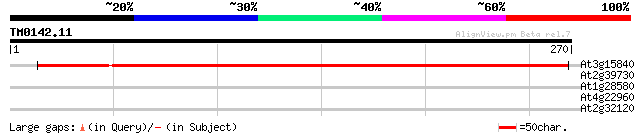
At3g15840 unknown protein 370 e-103
At2g39730 Rubisco activase 40 0.001
At1g28580 lipase like protein 29 2.3
At4g22960 putative protein 28 6.8
At2g32120 70kD heat shock protein 28 6.8
>At3g15840 unknown protein
Length = 268
Score = 370 bits (951), Expect = e-103
Identities = 174/256 (67%), Positives = 208/256 (80%), Gaps = 1/256 (0%)
Query: 14 PSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFEL 73
PS P++S+ +FL S L + K+ + A VA P +EI+EYK+PSWA FE+
Sbjct: 14 PSQPLSSKRSFLYSSRIGPILRRFPRKKLDLQVKA-VATTLAPLEEIKEYKLPSWAMFEM 72
Query: 74 GKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAML 133
G A VYWKTMNGLPPTSGEKLKLFYNP +++L NE++G+AFNGGFNQPIMCGGEPRAML
Sbjct: 73 GTAPVYWKTMNGLPPTSGEKLKLFYNPAASKLTLNEDYGVAFNGGFNQPIMCGGEPRAML 132
Query: 134 RKDRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEG 193
+KDRGKAD+PIY++QIC+PKHA+NLIFSFTNGVDWDGPYRLQFQVPK QNKPIEFFNEG
Sbjct: 133 KKDRGKADSPIYTMQICIPKHAVNLIFSFTNGVDWDGPYRLQFQVPKRWQNKPIEFFNEG 192
Query: 194 LAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLA 253
LA EL ++GACE+AIFPDSN V T+C M+ NLTVEGGDRC+L+LV GC D +S +NP A
Sbjct: 193 LANELSQDGACERAIFPDSNVVPTRCTMIANLTVEGGDRCNLDLVPGCMDTNSEHFNPYA 252
Query: 254 NVDDGTCPLDLDSDSE 269
NVDDG+CPL+L E
Sbjct: 253 NVDDGSCPLELSDSDE 268
>At2g39730 Rubisco activase
Length = 474
Score = 40.4 bits (93), Expect = 0.001
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>At1g28580 lipase like protein
Length = 390
Score = 29.3 bits (64), Expect = 2.3
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 5/47 (10%)
Query: 205 EQAIFPDSNKVITKCAMLG---NLTVEGGDRCDLNLVEGCTDPSSHL 248
E A F N+ ++ C G N TV G +C ++VE C DPS ++
Sbjct: 300 EPAKFGFMNRPLSACCGAGGPYNYTV--GRKCGTDIVESCDDPSKYV 344
>At4g22960 putative protein
Length = 487
Score = 27.7 bits (60), Expect = 6.8
Identities = 25/100 (25%), Positives = 45/100 (45%), Gaps = 2/100 (2%)
Query: 6 DLLFFSVPPSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKI 65
D F +V N +S+ T + +S+ + L +N ++ I A + +Q+ + +
Sbjct: 292 DETFGNVETEN--SSKMTDVLISVEKTNLESTKNDIISEAITLVNADLSEKSQDDDVSQS 349
Query: 66 PSWANFELGKASVYWKTMNGLPPTSGEKLKLFYNPTSTQL 105
LG++S K GL GE +K F+ + TQL
Sbjct: 350 VYEVQSLLGQSSPEGKGTKGLTQEEGELIKGFFEESVTQL 389
>At2g32120 70kD heat shock protein
Length = 563
Score = 27.7 bits (60), Expect = 6.8
Identities = 13/37 (35%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Query: 206 QAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCT 242
Q +F + +++ +C L + V GGD DL +V GC+
Sbjct: 329 QKVFEECERLVVQC--LRDARVNGGDIDDLIMVGGCS 363
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,761,885
Number of Sequences: 26719
Number of extensions: 299134
Number of successful extensions: 525
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 523
Number of HSP's gapped (non-prelim): 5
length of query: 270
length of database: 11,318,596
effective HSP length: 98
effective length of query: 172
effective length of database: 8,700,134
effective search space: 1496423048
effective search space used: 1496423048
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0142.11


