

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0141.1
(705 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
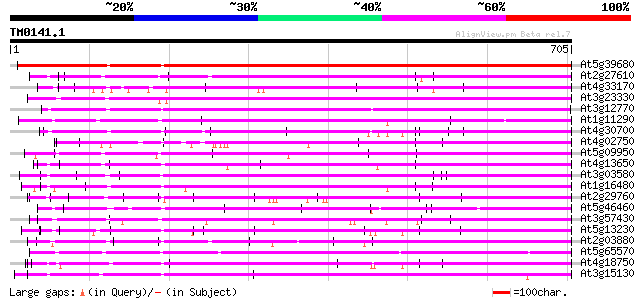
Score E
Sequences producing significant alignments: (bits) Value
At5g39680 unknown protein 688 0.0
At2g27610 putative selenium-binding protein 444 e-125
At4g33170 putative protein 439 e-123
At3g23330 hypothetical protein 436 e-122
At3g12770 unknown protein 436 e-122
At1g11290 hypothetical protein 429 e-120
At4g30700 unknown protein 428 e-120
At4g02750 hypothetical protein 427 e-119
At5g09950 selenium-binding protein-like 425 e-119
At4g13650 unknown protein 421 e-118
At3g03580 unknown protein (At3g03580) 420 e-117
At1g16480 hypothetical protein 419 e-117
At2g29760 hypothetical protein 417 e-116
At5g46460 putative protein 412 e-115
At3g57430 putative protein 412 e-115
At5g13230 putative protein 406 e-113
At2g03880 putative selenium-binding protein 404 e-113
At5g65570 putative protein 396 e-110
At4g18750 putative protein 392 e-109
At3g15130 hypothetical protein 391 e-109
>At5g39680 unknown protein
Length = 710
Score = 688 bits (1775), Expect = 0.0
Identities = 330/697 (47%), Positives = 481/697 (68%), Gaps = 7/697 (1%)
Query: 10 VVVNSRYLP-PLEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLY 68
+V S+ P P++ + +LLK+ A++ L G++IHA L+V +Q+S+ D Q+NSLI+LY
Sbjct: 20 LVPKSKKTPFPIDRLNELLKVCANSSYLRIGESIHAHLIVTNQSSRAEDAYQINSLINLY 79
Query: 69 VKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNE 128
VKC AR +FD MP R+V SW +M GY +SG EVL LFK++ S ++ PNE
Sbjct: 80 VKCRETVRARKLFDLMPERNVVSWCAMMKGYQNSGFDFEVLKLFKSMFFSGESR---PNE 136
Query: 129 YVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
+V T V SCS+SGR+ EG Q HG KYGL+S++FV++ L MY+ C A++VL+
Sbjct: 137 FVATVVFKSCSNSGRIEEGKQFHGCFLKYGLISHEFVRNTLVYMYSLCSGNGEAIRVLDD 196
Query: 189 ESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDL 248
D+ +++S L+ +E G ++E + VLR+ NE VW+N TY+ + L + + DL
Sbjct: 197 LP---YCDLSVFSSALSGYLECGAFKEGLDVLRKTANEDFVWNNLTYLSSLRLFSNLRDL 253
Query: 249 QLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAY 308
L L+VH+R++R G + LI+MYGKC VL A++VFD +N+ + T++M AY
Sbjct: 254 NLALQVHSRMVRFGFNAEVEACGALINMYGKCGKVLYAQRVFDDTHAQNIFLNTTIMDAY 313
Query: 309 LQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYV 368
Q++ FEE L+L + MD ++ PNE TF +LL + A ++LLK GDLLH + K G++N+V
Sbjct: 314 FQDKSFEEALNLFSKMDTKEVPPNEYTFAILLNSIAELSLLKQGDLLHGLVLKSGYRNHV 373
Query: 369 IVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSA 428
+V NAL+NMY+KSG I+ + FS RD+ TWN+MI G SHHGLG+EAL F M
Sbjct: 374 MVGNALVNMYAKSGSIEDARKAFSGMTFRDIVTWNTMISGCSHHGLGREALEAFDRMIFT 433
Query: 429 GESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEK 488
GE PN +TF+GVL AC+H+ V++GL Y MK ++P ++HYTC+V L +AG+ +
Sbjct: 434 GEIPNRITFIGVLQACSHIGFVEQGLHYFNQLMKKFDVQPDIQHYTCIVGLLSKAGMFKD 493
Query: 489 AENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHAT 548
AE+FM+T ++WD+VAWRTLLN+C V RNY LGK++AE ++ P D G Y LLSN+HA
Sbjct: 494 AEDFMRTAPIEWDVVAWRTLLNACYVRRNYRLGKKVAEYAIEKYPNDSGVYVLLSNIHAK 553
Query: 549 ANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAM 608
+ W+GV ++R +M R +KKEPG SW+ IRN HVF++E +HPE I KV+++++
Sbjct: 554 SREWEGVAKVRSLMNNRGVKKEPGVSWIGIRNQTHVFLAEDNQHPEITLIYAKVKEVMSK 613
Query: 609 IKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDC 668
IKPLGY P+++ HDV++EQ+E L+ HSEKLA+AYGL+K P +P+ + KN+R+CDDC
Sbjct: 614 IKPLGYSPDVAGAFHDVDEEQREDNLSYHSEKLAVAYGLIKTPEKSPLYVTKNVRICDDC 673
Query: 669 HTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
H+A+KLISK++ R+I++RD+NRFHHF DG C+C D+W
Sbjct: 674 HSAIKLISKISKRYIVIRDSNRFHHFLDGQCSCCDYW 710
>At2g27610 putative selenium-binding protein
Length = 868
Score = 444 bits (1142), Expect = e-125
Identities = 239/646 (36%), Positives = 377/646 (57%), Gaps = 12/646 (1%)
Query: 62 NSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTT 121
NSLI+LY+KCG +R AR +FDK ++ V +WN +++GY +G LE L +F ++
Sbjct: 233 NSLINLYLKCGNVRKARILFDKTEVKSVVTWNSMISGYAANGLDLEALGMFYSM----RL 288
Query: 122 QHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNL 181
+ +E F +V+ C++ +R Q H + KYG + Q +++AL Y++C +
Sbjct: 289 NYVRLSESSFASVIKLCANLKELRFTEQLHCSVVKYGFLFDQNIRTALMVAYSKCTAMLD 348
Query: 182 ALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGL 241
AL++ + G ++ + ++++ +++ EEA+ + M + + + TY ++
Sbjct: 349 ALRLF--KEIGCVGNVVSWTAMISGFLQNDGKEEAVDLFSEMKRKGVRPNEFTYSVILTA 406
Query: 242 CAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVW 301
I + VHA++++ VG+ L+D Y K V A KVF G+ ++++V W
Sbjct: 407 LPVISPSE----VHAQVVKTNYERSSTVGTALLDAYVKLGKVEEAAKVFSGIDDKDIVAW 462
Query: 302 TSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGI-ALLKHGDLLHARLE 360
+++++ Y Q E + + + + PNE TF +L CA A + G H
Sbjct: 463 SAMLAGYAQTGETEAAIKMFGELTKGGIKPNEFTFSSILNVCAATNASMGQGKQFHGFAI 522
Query: 361 KLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALR 420
K + + V++AL+ MY+K G+I+S+ VF +D+ +WNSMI GY+ HG +AL
Sbjct: 523 KSRLDSSLCVSSALLTMYAKKGNIESAEEVFKRQREKDLVSWNSMISGYAQHGQAMKALD 582
Query: 421 VFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLY 480
VF++MK + VTF+GV +AC H LV++G Y +++ KI P EH +CMV LY
Sbjct: 583 VFKEMKKRKVKMDGVTFIGVFAACTHAGLVEEGEKYFDIMVRDCKIAPTKEHNSCMVDLY 642
Query: 481 CRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYT 540
RAG LEKA ++ WRT+L +CRVH+ +LG+ AE I+ M P+D Y
Sbjct: 643 SRAGQLEKAMKVIENMPNPAGSTIWRTILAACRVHKKTELGRLAAEKIIAMKPEDSAAYV 702
Query: 541 LLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINK 600
LLSNM+A + W ++RK+M ERN+KKEPG SW+E++N + F++ HP ++I
Sbjct: 703 LLSNMYAESGDWQERAKVRKLMNERNVKKEPGYSWIEVKNKTYSFLAGDRSHPLKDQIYM 762
Query: 601 KVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIK 660
K++ L +K LGY P+ S VL D++DE KE L HSE+LAIA+GL+ P +P+ IIK
Sbjct: 763 KLEDLSTRLKDLGYEPDTSYVLQDIDDEHKEAVLAQHSERLAIAFGLIATPKGSPLLIIK 822
Query: 661 NLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHF-RDGSCTCADHW 705
NLR+C DCH +KLI+K+ R I+VRD+NRFHHF DG C+C D W
Sbjct: 823 NLRVCGDCHLVIKLIAKIEEREIVVRDSNRFHHFSSDGVCSCGDFW 868
Score = 209 bits (531), Expect = 6e-54
Identities = 140/513 (27%), Positives = 263/513 (50%), Gaps = 22/513 (4%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
+LK+SA G+ +H Q + + DV SL+ Y+K + R +FD+M
Sbjct: 99 VLKVSATLCDELFGRQLHCQCI---KFGFLDDVSVGTSLVDTYMKGSNFKDGRKVFDEMK 155
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
R+V +W L++GY + EVL LF + + T PN + F L + G
Sbjct: 156 ERNVVTWTTLISGYARNSMNDEVLTLFMRMQNEGTQ----PNSFTFAAALGVLAEEGVGG 211
Query: 146 EGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLN 205
G+Q H + K GL V ++L N+Y +C +V A + + + + +NS+++
Sbjct: 212 RGLQVHTVVVKNGLDKTIPVSNSLINLYLKCGNVRKARILFDKT---EVKSVVTWNSMIS 268
Query: 206 VLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLF 265
+G EA+G+ M + +++ V+ LCA + +L+ ++H +++ G LF
Sbjct: 269 GYAANGLDLEALGMFYSMRLNYVRLSESSFASVIKLCANLKELRFTEQLHCSVVKYGFLF 328
Query: 266 DEFVGSKLIDMYGKCCHVLNARKVFDGLQ-NRNVVVWTSLMSAYLQNRYFEETLDLLTCM 324
D+ + + L+ Y KC +L+A ++F + NVV WT+++S +LQN EE +DL + M
Sbjct: 329 DQNIRTALMVAYSKCTAMLDALRLFKEIGCVGNVVSWTAMISGFLQNDGKEEAVDLFSEM 388
Query: 325 DQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDI 384
++ PNE T+ V+L A I+ + +HA++ K ++ V AL++ Y K G +
Sbjct: 389 KRKGVRPNEFTYSVILTALPVISPSE----VHAQVVKTNYERSSTVGTALLDAYVKLGKV 444
Query: 385 DSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
+ + VFS +D+ W++M+ GY+ G + A+++F ++ G PN TF +L+ C
Sbjct: 445 EEAAKVFSGIDDKDIVAWSAMLAGYAQTGETEAAIKMFGELTKGGIKPNEFTFSSILNVC 504
Query: 445 AHL-ALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIV 503
A A + +G + +K+ +++ L + ++ +Y + G +E AE K + K D+V
Sbjct: 505 AATNASMGQGKQFHGFAIKS-RLDSSLCVSSALLTMYAKKGNIESAEEVFKRQREK-DLV 562
Query: 504 AWRTLLNSCRVH----RNYDLGKRIAESILQMD 532
+W ++++ H + D+ K + + ++MD
Sbjct: 563 SWNSMISGYAQHGQAMKALDVFKEMKKRKVKMD 595
Score = 146 bits (369), Expect = 3e-35
Identities = 95/311 (30%), Positives = 154/311 (48%), Gaps = 1/311 (0%)
Query: 200 YNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLL 259
Y SLL GR +EA + + M D + + V+ + A + D G ++H + +
Sbjct: 61 YISLLFGFSRDGRTQEAKRLFLNIHRLGMEMDCSIFSSVLKVSATLCDELFGRQLHCQCI 120
Query: 260 RGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLD 319
+ G L D VG+ L+D Y K + + RKVFD ++ RNVV WT+L+S Y +N +E L
Sbjct: 121 KFGFLDDVSVGTSLVDTYMKGSNFKDGRKVFDEMKERNVVTWTTLISGYARNSMNDEVLT 180
Query: 320 LLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYS 379
L M E T PN TF LG A + G +H + K G + V+N+LIN+Y
Sbjct: 181 LFMRMQNEGTQPNSFTFAAALGVLAEEGVGGRGLQVHTVVVKNGLDKTIPVSNSLINLYL 240
Query: 380 KSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVG 439
K G++ + +F T + V TWNSMI GY+ +GL EAL +F M+ + +F
Sbjct: 241 KCGNVRKARILFDKTEVKSVVTWNSMISGYAANGLDLEALGMFYSMRLNYVRLSESSFAS 300
Query: 440 VLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVK 499
V+ CA+L + + + L+ + + T ++V Y + + A K
Sbjct: 301 VIKLCANLKEL-RFTEQLHCSVVKYGFLFDQNIRTALMVAYSKCTAMLDALRLFKEIGCV 359
Query: 500 WDIVAWRTLLN 510
++V+W +++
Sbjct: 360 GNVVSWTAMIS 370
Score = 140 bits (353), Expect = 2e-33
Identities = 114/442 (25%), Positives = 203/442 (45%), Gaps = 14/442 (3%)
Query: 69 VKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNE 128
V RL A ++FDK P RD S+ L+ G+ G E LF N+
Sbjct: 38 VSSSRLYNAHNLFDKSPGRDRESYISLLFGFSRDGRTQEAKRLFLNIHRLGMEMDCS--- 94
Query: 129 YVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
+F++VL + G Q H K+G + V ++L + Y + + +V +
Sbjct: 95 -IFSSVLKVSATLCDELFGRQLHCQCIKFGFLDDVSVGTSLVDTYMKGSNFKDGRKVFDE 153
Query: 189 ESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDL 248
E ++ + +L++ + +E + + RM NE ++ T+ +G+ A+ G
Sbjct: 154 MK---ERNVVTWTTLISGYARNSMNDEVLTLFMRMQNEGTQPNSFTFAAALGVLAEEGVG 210
Query: 249 QLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAY 308
GL+VH +++ GL V + LI++Y KC +V AR +FD + ++VV W S++S Y
Sbjct: 211 GRGLQVHTVVVKNGLDKTIPVSNSLINLYLKCGNVRKARILFDKTEVKSVVTWNSMISGY 270
Query: 309 LQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYV 368
N E L + M +E +F ++ CA + L+ + LH + K GF
Sbjct: 271 AANGLDLEALGMFYSMRLNYVRLSESSFASVIKLCANLKELRFTEQLHCSVVKYGFLFDQ 330
Query: 369 IVNNALINMYSKSGDIDSSYNVFSDTMC-RDVTTWNSMICGYSHHGLGQEALRVFQDMKS 427
+ AL+ YSK + + +F + C +V +W +MI G+ + +EA+ +F +MK
Sbjct: 331 NIRTALMVAYSKCTAMLDALRLFKEIGCVGNVVSWTAMISGFLQNDGKEEAVDLFSEMKR 390
Query: 428 AGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLE 487
G PN T+ +L+A ++ + ++ ++ E T ++ Y + G +E
Sbjct: 391 KGVRPNEFTYSVILTALPVISPSE-----VHAQVVKTNYERSSTVGTALLDAYVKLGKVE 445
Query: 488 KAENFMKTTKVKWDIVAWRTLL 509
+A K DIVAW +L
Sbjct: 446 EAAKVFSGIDDK-DIVAWSAML 466
>At4g33170 putative protein
Length = 990
Score = 439 bits (1128), Expect = e-123
Identities = 237/648 (36%), Positives = 372/648 (56%), Gaps = 15/648 (2%)
Query: 62 NSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTT 121
NSLI++Y K + AR++FD M RD+ SWN ++ G +G +E + LF L+
Sbjct: 354 NSLINMYCKLRKFGFARTVFDNMSERDLISWNSVIAGIAQNGLEVEAVCLFMQLLRCGLK 413
Query: 122 QHACPNEYVFTTVLSSCSHSGRVREGM----QCHGYLFKYGLVSYQFVKSALFNMYARCL 177
P++Y T+VL + S + EG+ Q H + K VS FV +AL + Y+R
Sbjct: 414 ----PDQYTMTSVLKAASS---LPEGLSLSKQVHVHAIKINNVSDSFVSTALIDAYSR-- 464
Query: 178 HVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVG 237
N ++ D+ +N+++ +S + + + M + D+ T
Sbjct: 465 --NRCMKEAEILFERHNFDLVAWNAMMAGYTQSHDGHKTLKLFALMHKQGERSDDFTLAT 522
Query: 238 VMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRN 297
V C + + G +VHA ++ G D +V S ++DMY KC + A+ FD + +
Sbjct: 523 VFKTCGFLFAINQGKQVHAYAIKSGYDLDLWVSSGILDMYVKCGDMSAAQFAFDSIPVPD 582
Query: 298 VVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHA 357
V WT+++S ++N E + + M LP+E T L A + + L+ G +HA
Sbjct: 583 DVAWTTMISGCIENGEEERAFHVFSQMRLMGVLPDEFTIATLAKASSCLTALEQGRQIHA 642
Query: 358 RLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQE 417
KL N V +L++MY+K G ID +Y +F ++T WN+M+ G + HG G+E
Sbjct: 643 NALKLNCTNDPFVGTSLVDMYAKCGSIDDAYCLFKRIEMMNITAWNAMLVGLAQHGEGKE 702
Query: 418 ALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMV 477
L++F+ MKS G P+ VTF+GVLSAC+H LV + ++ + ++ I+P +EHY+C+
Sbjct: 703 TLQLFKQMKSLGIKPDKVTFIGVLSACSHSGLVSEAYKHMRSMHGDYGIKPEIEHYSCLA 762
Query: 478 VLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMG 537
RAGL+++AEN +++ ++ +RTLL +CRV + + GKR+A +L+++P D
Sbjct: 763 DALGRAGLVKQAENLIESMSMEASASMYRTLLAACRVQGDTETGKRVATKLLELEPLDSS 822
Query: 538 TYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNE 597
Y LLSNM+A A+ WD + R MMK +KK+PG SW+E++N H+FV + + ++
Sbjct: 823 AYVLLSNMYAAASKWDEMKLARTMMKGHKVKKDPGFSWIEVKNKIHIFVVDDRSNRQTEL 882
Query: 598 INKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIR 657
I +KV+ ++ IK GYVP L DVE+E+KE L HSEKLA+A+GL+ P PIR
Sbjct: 883 IYRKVKDMIRDIKQEGYVPETDFTLVDVEEEEKERALYYHSEKLAVAFGLLSTPPSTPIR 942
Query: 658 IIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
+IKNLR+C DCH A+K I+KV NR I++RDANRFH F+DG C+C D+W
Sbjct: 943 VIKNLRVCGDCHNAMKYIAKVYNREIVLRDANRFHRFKDGICSCGDYW 990
Score = 138 bits (347), Expect = 1e-32
Identities = 118/496 (23%), Positives = 217/496 (42%), Gaps = 21/496 (4%)
Query: 82 DKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHS 141
D + ++ N ++ Y+HSG Y +L F ++V S C ++ F +L++
Sbjct: 273 DASSVSEIIFRNKGLSEYLHSGQYSALLKCFADMVESDVE---C-DQVTFILMLATAVKV 328
Query: 142 GRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYN 201
+ G Q H K GL V ++L NMY + A V + S E D+ +N
Sbjct: 329 DSLALGQQVHCMALKLGLDLMLTVSNSLINMYCKLRKFGFARTVFDNMS---ERDLISWN 385
Query: 202 SLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGD-LQLGLRVHARLLR 260
S++ + ++G EA+ + ++ + D T V+ + + + L L +VH ++
Sbjct: 386 SVIAGIAQNGLEVEAVCLFMQLLRCGLKPDQYTMTSVLKAASSLPEGLSLSKQVHVHAIK 445
Query: 261 GGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDL 320
+ D FV + LID Y + + A +F+ N ++V W ++M+ Y Q+ +TL L
Sbjct: 446 INNVSDSFVSTALIDAYSRNRCMKEAEILFE-RHNFDLVAWNAMMAGYTQSHDGHKTLKL 504
Query: 321 LTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSK 380
M ++ ++ T + C + + G +HA K G+ + V++ +++MY K
Sbjct: 505 FALMHKQGERSDDFTLATVFKTCGFLFAINQGKQVHAYAIKSGYDLDLWVSSGILDMYVK 564
Query: 381 SGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGV 440
GD+ ++ F D W +MI G +G + A VF M+ G P+ T +
Sbjct: 565 CGDMSAAQFAFDSIPVPDDVAWTTMISGCIENGEEERAFHVFSQMRLMGVLPDEFTIATL 624
Query: 441 LSACAHLALVDKGLDYLYNEMK-NHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVK 499
A + L +++G N +K N +P + T +V +Y + G ++ A K ++
Sbjct: 625 AKASSCLTALEQGRQIHANALKLNCTNDPFVG--TSLVDMYAKCGSIDDAYCLFKRIEM- 681
Query: 500 WDIVAWRTLLNSCRVHRNYDLGKRIAESILQM-----DPQDMGTYTLLSNMHATANMWDG 554
+I AW +L H GK + QM P + +LS + + +
Sbjct: 682 MNITAWNAMLVGLAQHGE---GKETLQLFKQMKSLGIKPDKVTFIGVLSACSHSGLVSEA 738
Query: 555 VVEIRKMMKERNIKKE 570
+R M + IK E
Sbjct: 739 YKHMRSMHGDYGIKPE 754
Score = 135 bits (339), Expect = 1e-31
Identities = 134/552 (24%), Positives = 233/552 (41%), Gaps = 88/552 (15%)
Query: 36 LTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDL 95
L GK HA++L ++ +R +N+LI +Y KCG L AR +FDKMP RD+ SWN +
Sbjct: 55 LMLGKCTHARILTFEENPER---FLINNLISMYSKCGSLTYARRVFDKMPDRDLVSWNSI 111
Query: 96 MTGYMHSG-----NYLEVLVLFKNL--------------------------VSSSTTQHA 124
+ Y S N + +LF+ L S S +A
Sbjct: 112 LAAYAQSSECVVENIQQAFLLFRILRQDVVYTSRMTLSPMLKLCLHSGYVWASESFHGYA 171
Query: 125 CP-----NEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNM------- 172
C +E+V +++ G+V+EG + +V + + A M
Sbjct: 172 CKIGLDGDEFVAGALVNIYLKFGKVKEGKVLFEEMPYRDVVLWNLMLKAYLEMGFKEEAI 231
Query: 173 ------YARCLHVN-LALQVLNTESGGDEN-----------------DIFLYNSLLNVLV 208
++ L+ N + L++L SG D + +I N L+ +
Sbjct: 232 DLSSAFHSSGLNPNEITLRLLARISGDDSDAGQVKSFANGNDASSVSEIIFRNKGLSEYL 291
Query: 209 ESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEF 268
SG++ + M + D T++ ++ ++ L LG +VH L+ GL
Sbjct: 292 HSGQYSALLKCFADMVESDVECDQVTFILMLATAVKVDSLALGQQVHCMALKLGLDLMLT 351
Query: 269 VGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEET----LDLLTCM 324
V + LI+MY K AR VFD + R+++ W S+++ QN E + LL C
Sbjct: 352 VSNSLINMYCKLRKFGFARTVFDNMSERDLISWNSVIAGIAQNGLEVEAVCLFMQLLRCG 411
Query: 325 DQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDI 384
+ D K G++L K +H K+ + V+ ALI+ YS++ +
Sbjct: 412 LKPDQYTMTSVLKAASSLPEGLSLSKQ---VHVHAIKINNVSDSFVSTALIDAYSRNRCM 468
Query: 385 DSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
+ +F + D+ WN+M+ GY+ G + L++F M GE ++ T V C
Sbjct: 469 KEAEILF-ERHNFDLVAWNAMMAGYTQSHDGHKTLKLFALMHKQGERSDDFTLATVFKTC 527
Query: 445 AHLALVDKGLDYLYNEMKNHKIEPG--LEHYTCMVVL--YCRAGLLEKAENFMKTTKVKW 500
L +++G ++ + I+ G L+ + +L Y + G + A+ + V
Sbjct: 528 GFLFAINQG-----KQVHAYAIKSGYDLDLWVSSGILDMYVKCGDMSAAQFAFDSIPVP- 581
Query: 501 DIVAWRTLLNSC 512
D VAW T+++ C
Sbjct: 582 DDVAWTTMISGC 593
Score = 96.3 bits (238), Expect = 5e-20
Identities = 58/195 (29%), Positives = 93/195 (46%), Gaps = 5/195 (2%)
Query: 247 DLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMS 306
DL LG HAR+L + F+ + LI MY KC + AR+VFD + +R++V W S+++
Sbjct: 54 DLMLGKCTHARILTFEENPERFLINNLISMYSKCGSLTYARRVFDKMPDRDLVSWNSILA 113
Query: 307 AYLQN-----RYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEK 361
AY Q+ ++ L + Q+ + T +L C + + H K
Sbjct: 114 AYAQSSECVVENIQQAFLLFRILRQDVVYTSRMTLSPMLKLCLHSGYVWASESFHGYACK 173
Query: 362 LGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRV 421
+G V AL+N+Y K G + +F + RDV WN M+ Y G +EA+ +
Sbjct: 174 IGLDGDEFVAGALVNIYLKFGKVKEGKVLFEEMPYRDVVLWNLMLKAYLEMGFKEEAIDL 233
Query: 422 FQDMKSAGESPNNVT 436
S+G +PN +T
Sbjct: 234 SSAFHSSGLNPNEIT 248
>At3g23330 hypothetical protein
Length = 715
Score = 436 bits (1122), Expect = e-122
Identities = 237/716 (33%), Positives = 388/716 (54%), Gaps = 41/716 (5%)
Query: 23 IVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFD 82
I L+K K + K +HAQ + S S + +I +Y L A +F
Sbjct: 8 IKTLIKNPTRIKSKSQAKQLHAQFIRTQSLSHTSASI----VISIYTNLKLLHEALLLFK 63
Query: 83 KMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSG 142
+ V +W ++ + + + L F + +S CP+ VF +VL SC+
Sbjct: 64 TLKSPPVLAWKSVIRCFTDQSLFSKALASFVEMRASGR----CPDHNVFPSVLKSCTMMM 119
Query: 143 RVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLN--------TESGGDE 194
+R G HG++ + G+ + +AL NMYA+ L + + V N T + GDE
Sbjct: 120 DLRFGESVHGFIVRLGMDCDLYTGNALMNMYAKLLGMGSKISVGNVFDEMPQRTSNSGDE 179
Query: 195 N-------------------------DIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMV 229
+ D+ YN+++ +SG +E+A+ ++R MG +
Sbjct: 180 DVKAETCIMPFGIDSVRRVFEVMPRKDVVSYNTIIAGYAQSGMYEDALRMVREMGTTDLK 239
Query: 230 WDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKV 289
D+ T V+ + ++ D+ G +H ++R G+ D ++GS L+DMY K + ++ +V
Sbjct: 240 PDSFTLSSVLPIFSEYVDVIKGKEIHGYVIRKGIDSDVYIGSSLVDMYAKSARIEDSERV 299
Query: 290 FDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALL 349
F L R+ + W SL++ Y+QN + E L L M P F ++ ACA +A L
Sbjct: 300 FSRLYCRDGISWNSLVAGYVQNGRYNEALRLFRQMVTAKVKPGAVAFSSVIPACAHLATL 359
Query: 350 KHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGY 409
G LH + + GF + + + +AL++MYSK G+I ++ +F D +W ++I G+
Sbjct: 360 HLGKQLHGYVLRGGFGSNIFIASALVDMYSKCGNIKAARKIFDRMNVLDEVSWTAIIMGH 419
Query: 410 SHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPG 469
+ HG G EA+ +F++MK G PN V FV VL+AC+H+ LVD+ Y + K + +
Sbjct: 420 ALHGHGHEAVSLFEEMKRQGVKPNQVAFVAVLTACSHVGLVDEAWGYFNSMTKVYGLNQE 479
Query: 470 LEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESIL 529
LEHY + L RAG LE+A NF+ V+ W TLL+SC VH+N +L +++AE I
Sbjct: 480 LEHYAAVADLLGRAGKLEEAYNFISKMCVEPTGSVWSTLLSSCSVHKNLELAEKVAEKIF 539
Query: 530 QMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEG 589
+D ++MG Y L+ NM+A+ W + ++R M+++ ++K+P SW+E++N H FVS
Sbjct: 540 TVDSENMGAYVLMCNMYASNGRWKEMAKLRLRMRKKGLRKKPACSWIEMKNKTHGFVSGD 599
Query: 590 CKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMK 649
HP ++IN+ ++ ++ ++ GYV + S VLHDV++E K L HSE+LA+A+G++
Sbjct: 600 RSHPSMDKINEFLKAVMEQMEKEGYVADTSGVLHDVDEEHKRELLFGHSERLAVAFGIIN 659
Query: 650 MPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
IR+ KN+R+C DCH A+K ISK+T R IIVRD +RFHHF G+C+C D+W
Sbjct: 660 TEPGTTIRVTKNIRICTDCHVAIKFISKITEREIIVRDNSRFHHFNRGNCSCGDYW 715
>At3g12770 unknown protein
Length = 719
Score = 436 bits (1121), Expect = e-122
Identities = 225/665 (33%), Positives = 378/665 (56%), Gaps = 9/665 (1%)
Query: 40 KAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGY 99
K IHA+LLV + + LIH G + AR +FD +P +F WN ++ GY
Sbjct: 38 KQIHARLLV---LGLQFSGFLITKLIHASSSFGDITFARQVFDDLPRPQIFPWNAIIRGY 94
Query: 100 MHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGL 159
+ ++ + L+++ N+ + + P+ + F +L +CS ++ G H +F+ G
Sbjct: 95 SRNNHFQDALLMYSNMQLARVS----PDSFTFPHLLKACSGLSHLQMGRFVHAQVFRLGF 150
Query: 160 VSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGV 219
+ FV++ L +YA+C + A V E I + ++++ ++G EA+ +
Sbjct: 151 DADVFVQNGLIALYAKCRRLGSARTVFEGLPL-PERTIVSWTAIVSAYAQNGEPMEALEI 209
Query: 220 LRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGK 279
+M + D V V+ + DL+ G +HA +++ GL + + L MY K
Sbjct: 210 FSQMRKMDVKPDWVALVSVLNAFTCLQDLKQGRSIHASVVKMGLEIEPDLLISLNTMYAK 269
Query: 280 CCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVL 339
C V A+ +FD +++ N+++W +++S Y +N Y E +D+ M +D P+ +
Sbjct: 270 CGQVATAKILFDKMKSPNLILWNAMISGYAKNGYAREAIDMFHEMINKDVRPDTISITSA 329
Query: 340 LGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDV 399
+ ACA + L+ ++ + + +++ V +++ALI+M++K G ++ + VF T+ RDV
Sbjct: 330 ISACAQVGSLEQARSMYEYVGRSDYRDDVFISSALIDMFAKCGSVEGARLVFDRTLDRDV 389
Query: 400 TTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYN 459
W++MI GY HG +EA+ +++ M+ G PN+VTF+G+L AC H +V +G + +N
Sbjct: 390 VVWSAMIVGYGLHGRAREAISLYRAMERGGVHPNDVTFLGLLMACNHSGMVREGW-WFFN 448
Query: 460 EMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYD 519
M +HKI P +HY C++ L RAG L++A +K V+ + W LL++C+ HR+ +
Sbjct: 449 RMADHKINPQQQHYACVIDLLGRAGHLDQAYEVIKCMPVQPGVTVWGALLSACKKHRHVE 508
Query: 520 LGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIR 579
LG+ A+ + +DP + G Y LSN++A A +WD V E+R MKE+ + K+ G SW+E+R
Sbjct: 509 LGEYAAQQLFSIDPSNTGHYVQLSNLYAAARLWDRVAEVRVRMKEKGLNKDVGCSWVEVR 568
Query: 580 NVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSE 639
F HP EI ++V+ + + +K G+V N A LHD+ DE+ E L HSE
Sbjct: 569 GRLEAFRVGDKSHPRYEEIERQVEWIESRLKEGGFVANKDASLHDLNDEEAEETLCSHSE 628
Query: 640 KLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSC 699
++AIAYGL+ P P+RI KNLR C +CH A KLISK+ +R I+VRD NRFHHF+DG C
Sbjct: 629 RIAIAYGLISTPQGTPLRITKNLRACVNCHAATKLISKLVDREIVVRDTNRFHHFKDGVC 688
Query: 700 TCADH 704
+C D+
Sbjct: 689 SCGDY 693
>At1g11290 hypothetical protein
Length = 809
Score = 429 bits (1102), Expect = e-120
Identities = 245/697 (35%), Positives = 394/697 (56%), Gaps = 14/697 (2%)
Query: 12 VNSRY--LPPLEY-IVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLY 68
V RY + P+ Y LLK+ D L GK IH LLV+ S D+ + L ++Y
Sbjct: 124 VRMRYDDVEPVVYNFTYLLKVCGDEAELRVGKEIHG-LLVKSGFSL--DLFAMTGLENMY 180
Query: 69 VKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNE 128
KC ++ AR +FD+MP RD+ SWN ++ GY +G L +V S ++ P+
Sbjct: 181 AKCRQVNEARKVFDRMPERDLVSWNTIVAGYSQNGMARMAL----EMVKSMCEENLKPSF 236
Query: 129 YVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
+VL + S + G + HGY + G S + +AL +MYA+C + A Q+ +
Sbjct: 237 ITIVSVLPAVSALRLISVGKEIHGYAMRSGFDSLVNISTALVDMYAKCGSLETARQLFD- 295
Query: 189 ESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDL 248
G E ++ +NS+++ V++ +EA+ + ++M +E + + + +G + CA +GDL
Sbjct: 296 --GMLERNVVSWNSMIDAYVQNENPKEAMLIFQKMLDEGVKPTDVSVMGALHACADLGDL 353
Query: 249 QLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAY 308
+ G +H + GL + V + LI MY KC V A +F LQ+R +V W +++ +
Sbjct: 354 ERGRFIHKLSVELGLDRNVSVVNSLISMYCKCKEVDTAASMFGKLQSRTLVSWNAMILGF 413
Query: 309 LQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYV 368
QN + L+ + M P+ T+ ++ A A +++ H +H + + V
Sbjct: 414 AQNGRPIDALNYFSQMRSRTVKPDTFTYVSVITAIAELSITHHAKWIHGVVMRSCLDKNV 473
Query: 369 IVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSA 428
V AL++MY+K G I + +F R VTTWN+MI GY HG G+ AL +F++M+
Sbjct: 474 FVTTALVDMYAKCGAIMIARLIFDMMSERHVTTWNAMIDGYGTHGFGKAALELFEEMQKG 533
Query: 429 GESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEK 488
PN VTF+ V+SAC+H LV+ GL Y +N+ IE ++HY MV L RAG L +
Sbjct: 534 TIKPNGVTFLSVISACSHSGLVEAGLKCFYMMKENYSIELSMDHYGAMVDLLGRAGRLNE 593
Query: 489 AENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHAT 548
A +F+ VK + + +L +C++H+N + ++ AE + +++P D G + LL+N++
Sbjct: 594 AWDFIMQMPVKPAVNVYGAMLGACQIHKNVNFAEKAAERLFELNPDDGGYHVLLANIYRA 653
Query: 549 ANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAM 608
A+MW+ V ++R M + ++K PG S +EI+N H F S HP+S +I +++L+
Sbjct: 654 ASMWEKVGQVRVSMLRQGLRKTPGCSMVEIKNEVHSFFSGSTAHPDSKKIYAFLEKLICH 713
Query: 609 IKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDC 668
IK GYVP+ + VL VE++ KE L+ HSEKLAI++GL+ + I + KNLR+C DC
Sbjct: 714 IKEAGYVPDTNLVL-GVENDVKEQLLSTHSEKLAISFGLLNTTAGTTIHVRKNLRVCADC 772
Query: 669 HTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
H A K IS VT R I+VRD RFHHF++G+C+C D+W
Sbjct: 773 HNATKYISLVTGREIVVRDMQRFHHFKNGACSCGDYW 809
Score = 107 bits (266), Expect = 3e-23
Identities = 75/318 (23%), Positives = 153/318 (47%), Gaps = 18/318 (5%)
Query: 242 CAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVW 301
C+ + +L+ ++ + + GL + F +KL+ ++ + V A +VF+ + ++ V++
Sbjct: 47 CSSLKELR---QILPLVFKNGLYQEHFFQTKLVSLFCRYGSVDEAARVFEPIDSKLNVLY 103
Query: 302 TSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEK 361
+++ + + ++ L M +D P F LL C A L+ G +H L K
Sbjct: 104 HTMLKGFAKVSDLDKALQFFVRMRYDDVEPVVYNFTYLLKVCGDEAELRVGKEIHGLLVK 163
Query: 362 LGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRV 421
GF + L NMY+K ++ + VF RD+ +WN+++ GYS +G+ + AL +
Sbjct: 164 SGFSLDLFAMTGLENMYAKCRQVNEARKVFDRMPERDLVSWNTIVAGYSQNGMARMALEM 223
Query: 422 FQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHY----TCMV 477
+ M P+ +T V VL A + L L+ G E+ + + G + T +V
Sbjct: 224 VKSMCEENLKPSFITIVSVLPAVSALRLISVG-----KEIHGYAMRSGFDSLVNISTALV 278
Query: 478 VLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQ--MDPQD 535
+Y + G LE A ++ ++V+W +++++ + N I + +L + P D
Sbjct: 279 DMYAKCGSLETARQLF-DGMLERNVVSWNSMIDAYVQNENPKEAMLIFQKMLDEGVKPTD 337
Query: 536 MGTYTLLSNMHATANMWD 553
+ +++ +HA A++ D
Sbjct: 338 V---SVMGALHACADLGD 352
>At4g30700 unknown protein
Length = 792
Score = 428 bits (1100), Expect = e-120
Identities = 226/669 (33%), Positives = 378/669 (55%), Gaps = 14/669 (2%)
Query: 38 SGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMT 97
+G+ IH Q +V S+++ ++++ +Y K R+ AR +FD+MP +D WN +++
Sbjct: 137 AGRVIHGQAVVD---GCDSELLLGSNIVKMYFKFWRVEDARKVFDRMPEKDTILWNTMIS 193
Query: 98 GYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKY 157
GY + Y+E + +F++L++ S T+ + +L + + +R GMQ H K
Sbjct: 194 GYRKNEMYVESIQVFRDLINESCTRL---DTTTLLDILPAVAELQELRLGMQIHSLATKT 250
Query: 158 GLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAI 217
G S+ +V + ++Y++C + + + + DI YN++++ +G E ++
Sbjct: 251 GCYSHDYVLTGFISLYSKCGKIKMGSALFREFR---KPDIVAYNAMIHGYTSNGETELSL 307
Query: 218 GVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMY 277
+ + + ++T V ++ + G L L +H L+ L V + L +Y
Sbjct: 308 SLFKELMLSGARLRSSTLVSLVPVS---GHLMLIYAIHGYCLKSNFLSHASVSTALTTVY 364
Query: 278 GKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFK 337
K + +ARK+FD +++ W +++S Y QN E+ + L M + + PN T
Sbjct: 365 SKLNEIESARKLFDESPEKSLPSWNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTIT 424
Query: 338 VLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCR 397
+L ACA + L G +H + F++ + V+ ALI MY+K G I + +F +
Sbjct: 425 CILSACAQLGALSLGKWVHDLVRSTDFESSIYVSTALIGMYAKCGSIAEARRLFDLMTKK 484
Query: 398 DVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYL 457
+ TWN+MI GY HG GQEAL +F +M ++G +P VTF+ VL AC+H LV +G D +
Sbjct: 485 NEVTWNTMISGYGLHGQGQEALNIFYEMLNSGITPTPVTFLCVLYACSHAGLVKEG-DEI 543
Query: 458 YNEM-KNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHR 516
+N M + EP ++HY CMV + RAG L++A F++ ++ W TLL +CR+H+
Sbjct: 544 FNSMIHRYGFEPSVKHYACMVDILGRAGHLQRALQFIEAMSIEPGSSVWETLLGACRIHK 603
Query: 517 NYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWL 576
+ +L + ++E + ++DP ++G + LLSN+H+ + +R+ K+R + K PG + +
Sbjct: 604 DTNLARTVSEKLFELDPDNVGYHVLLSNIHSADRNYPQAATVRQTAKKRKLAKAPGYTLI 663
Query: 577 EIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNC 636
EI HVF S HP+ EI +K+++L ++ GY P LHDVE+E++E +
Sbjct: 664 EIGETPHVFTSGDQSHPQVKEIYEKLEKLEGKMREAGYQPETELALHDVEEEERELMVKV 723
Query: 637 HSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRD 696
HSE+LAIA+GL+ IRIIKNLR+C DCHT KLISK+T R I+VRDANRFHHF+D
Sbjct: 724 HSERLAIAFGLIATEPGTEIRIIKNLRVCLDCHTVTKLISKITERVIVVRDANRFHHFKD 783
Query: 697 GSCTCADHW 705
G C+C D+W
Sbjct: 784 GVCSCGDYW 792
Score = 161 bits (407), Expect = 1e-39
Identities = 117/474 (24%), Positives = 227/474 (47%), Gaps = 15/474 (3%)
Query: 43 HAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHS 102
HAQ+++ R+D+ L L G + AR +F + DVF +N LM G+ +
Sbjct: 40 HAQIILH---GFRNDISLLTKLTQRLSDLGAIYYARDIFLSVQRPDVFLFNVLMRGFSVN 96
Query: 103 GNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSY 162
+ L +F +L S+ + PN + +S+ S R G HG G S
Sbjct: 97 ESPHSSLSVFAHLRKSTDLK---PNSSTYAFAISAASGFRDDRAGRVIHGQAVVDGCDSE 153
Query: 163 QFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRR 222
+ S + MY + V A +V + E D L+N++++ ++ + E+I V R
Sbjct: 154 LLLGSNIVKMYFKFWRVEDARKVFDRM---PEKDTILWNTMISGYRKNEMYVESIQVFRD 210
Query: 223 MGNE-CMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCC 281
+ NE C D T + ++ A++ +L+LG+++H+ + G ++V + I +Y KC
Sbjct: 211 LINESCTRLDTTTLLDILPAVAELQELRLGMQIHSLATKTGCYSHDYVLTGFISLYSKCG 270
Query: 282 HVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLG 341
+ +F + ++V + +++ Y N E +L L + T L+
Sbjct: 271 KIKMGSALFREFRKPDIVAYNAMIHGYTSNGETELSLSLFKELMLSGARLRSSTLVSLVP 330
Query: 342 ACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTT 401
+ L+ +H K F ++ V+ AL +YSK +I+S+ +F ++ + + +
Sbjct: 331 VSGHLMLIY---AIHGYCLKSNFLSHASVSTALTTVYSKLNEIESARKLFDESPEKSLPS 387
Query: 402 WNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEM 461
WN+MI GY+ +GL ++A+ +F++M+ + SPN VT +LSACA L + G ++++ +
Sbjct: 388 WNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTITCILSACAQLGALSLG-KWVHDLV 446
Query: 462 KNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
++ E + T ++ +Y + G + +A K + V W T+++ +H
Sbjct: 447 RSTDFESSIYVSTALIGMYAKCGSIAEARRLF-DLMTKKNEVTWNTMISGYGLH 499
Score = 101 bits (251), Expect = 2e-21
Identities = 91/408 (22%), Positives = 176/408 (42%), Gaps = 36/408 (8%)
Query: 196 DIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNA-TYVGVMGLCAQIGDLQLGLRV 254
D+FL+N L+ + ++ V + + N+ TY + + D + G +
Sbjct: 82 DVFLFNVLMRGFSVNESPHSSLSVFAHLRKSTDLKPNSSTYAFAISAASGFRDDRAGRVI 141
Query: 255 HARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYF 314
H + + G + +GS ++ MY K V +ARKVFD + ++ ++W +++S Y +N +
Sbjct: 142 HGQAVVDGCDSELLLGSNIVKMYFKFWRVEDARKVFDRMPEKDTILWNTMISGYRKNEMY 201
Query: 315 EETLDLLTCMDQED-TLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNA 373
E++ + + E T + T +L A A + L+ G +H+ K G ++ V
Sbjct: 202 VESIQVFRDLINESCTRLDTTTLLDILPAVAELQELRLGMQIHSLATKTGCYSHDYVLTG 261
Query: 374 LINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPN 433
I++YSK G I +F + D+ +N+MI GY+ +G + +L +F+++ +G
Sbjct: 262 FISLYSKCGKIKMGSALFREFRKPDIVAYNAMIHGYTSNGETELSLSLFKELMLSGARLR 321
Query: 434 NVTFVGVLSACAHLAL------------------VDKGLDYLYNEMK---------NHKI 466
+ T V ++ HL L V L +Y+++ +
Sbjct: 322 SSTLVSLVPVSGHLMLIYAIHGYCLKSNFLSHASVSTALTTVYSKLNEIESARKLFDESP 381
Query: 467 EPGLEHYTCMVVLYCRAGLLEKAENF---MKTTKVKWDIVAWRTLLNSCRVHRNYDLGKR 523
E L + M+ Y + GL E A + M+ ++ + V +L++C LGK
Sbjct: 382 EKSLPSWNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTITCILSACAQLGALSLGKW 441
Query: 524 IAESILQMD-PQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKE 570
+ + + D + T L M+A + E R++ K E
Sbjct: 442 VHDLVRSTDFESSIYVSTALIGMYAKCG---SIAEARRLFDLMTKKNE 486
Score = 68.9 bits (167), Expect = 9e-12
Identities = 58/266 (21%), Positives = 118/266 (43%), Gaps = 16/266 (6%)
Query: 253 RVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNR 312
+ HA+++ G D + +KL + AR +F +Q +V ++ LM + N
Sbjct: 38 QTHAQIILHGFRNDISLLTKLTQRLSDLGAIYYARDIFLSVQRPDVFLFNVLMRGFSVNE 97
Query: 313 YFEETLDLLTCMDQE-DTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVN 371
+L + + + D PN T+ + A +G + G ++H + G + +++
Sbjct: 98 SPHSSLSVFAHLRKSTDLKPNSSTYAFAISAASGFRDDRAGRVIHGQAVVDGCDSELLLG 157
Query: 372 NALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDM-KSAGE 430
+ ++ MY K ++ + VF +D WN+MI GY + + E+++VF+D+ +
Sbjct: 158 SNIVKMYFKFWRVEDARKVFDRMPEKDTILWNTMISGYRKNEMYVESIQVFRDLINESCT 217
Query: 431 SPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHY------TCMVVLYCRAG 484
+ T + +L A A L + G M+ H + Y T + LY + G
Sbjct: 218 RLDTTTLLDILPAVAELQELRLG-------MQIHSLATKTGCYSHDYVLTGFISLYSKCG 270
Query: 485 LLEKAENFMKTTKVKWDIVAWRTLLN 510
++ + + K DIVA+ +++
Sbjct: 271 KIKMGSALFREFR-KPDIVAYNAMIH 295
Score = 43.9 bits (102), Expect = 3e-04
Identities = 44/204 (21%), Positives = 86/204 (41%), Gaps = 3/204 (1%)
Query: 349 LKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICG 408
+ H HA++ GF+N + + L S G I + ++F DV +N ++ G
Sbjct: 33 ISHLAQTHAQIILHGFRNDISLLTKLTQRLSDLGAIYYARDIFLSVQRPDVFLFNVLMRG 92
Query: 409 YSHHGLGQEALRVFQDM-KSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIE 467
+S + +L VF + KS PN+ T+ +SA + D+ ++ + +
Sbjct: 93 FSVNESPHSSLSVFAHLRKSTDLKPNSSTYAFAISAASGFR-DDRAGRVIHGQAVVDGCD 151
Query: 468 PGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAES 527
L + +V +Y + +E A K D + W T+++ R + Y ++
Sbjct: 152 SELLLGSNIVKMYFKFWRVEDARKVFDRMPEK-DTILWNTMISGYRKNEMYVESIQVFRD 210
Query: 528 ILQMDPQDMGTYTLLSNMHATANM 551
++ + T TLL + A A +
Sbjct: 211 LINESCTRLDTTTLLDILPAVAEL 234
>At4g02750 hypothetical protein
Length = 781
Score = 427 bits (1098), Expect = e-119
Identities = 245/702 (34%), Positives = 379/702 (53%), Gaps = 71/702 (10%)
Query: 59 VQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFK----- 113
V N +I Y++ G LAR +FD+MP RD+ SWN ++ GY+ + N + LF+
Sbjct: 96 VSYNGMISGYLRNGEFELARKLFDEMPERDLVSWNVMIKGYVRNRNLGKARELFEIMPER 155
Query: 114 -----NLVSSSTTQHAC-------------PNEYVFTTVLSSCSHSGRVREGMQCHGYLF 155
N + S Q+ C N+ + +LS+ + ++ E
Sbjct: 156 DVCSWNTMLSGYAQNGCVDDARSVFDRMPEKNDVSWNALLSAYVQNSKMEEACMLFKSRE 215
Query: 156 KYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEE 215
+ LVS+ + L + + + A Q ++ + D+ +N+++ +SG+ +E
Sbjct: 216 NWALVSW----NCLLGGFVKKKKIVEARQFFDSMN---VRDVVSWNTIITGYAQSGKIDE 268
Query: 216 AIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHAR----------------LL 259
A R++ +E V D T+ ++ G +Q + AR +L
Sbjct: 269 A----RQLFDESPVQDVFTWTAMVS-----GYIQNRMVEEARELFDKMPERNEVSWNAML 319
Query: 260 RGGL----------LFDEF------VGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTS 303
G + LFD + +I Y +C + A+ +FD + R+ V W +
Sbjct: 320 AGYVQGERMEMAKELFDVMPCRNVSTWNTMITGYAQCGKISEAKNLFDKMPKRDPVSWAA 379
Query: 304 LMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLG 363
+++ Y Q+ + E L L M++E N +F L CA + L+ G LH RL K G
Sbjct: 380 MIAGYSQSGHSFEALRLFVQMEREGGRLNRSSFSSALSTCADVVALELGKQLHGRLVKGG 439
Query: 364 FKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQ 423
++ V NAL+ MY K G I+ + ++F + +D+ +WN+MI GYS HG G+ ALR F+
Sbjct: 440 YETGCFVGNALLLMYCKCGSIEEANDLFKEMAGKDIVSWNTMIAGYSRHGFGEVALRFFE 499
Query: 424 DMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRA 483
MK G P++ T V VLSAC+H LVDKG Y Y +++ + P +HY CMV L RA
Sbjct: 500 SMKREGLKPDDATMVAVLSACSHTGLVDKGRQYFYTMTQDYGVMPNSQHYACMVDLLGRA 559
Query: 484 GLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLS 543
GLLE A N MK + D W TLL + RVH N +L + A+ I M+P++ G Y LLS
Sbjct: 560 GLLEDAHNLMKNMPFEPDAAIWGTLLGASRVHGNTELAETAADKIFAMEPENSGMYVLLS 619
Query: 544 NMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQ 603
N++A++ W V ++R M+++ +KK PG SW+EI+N H F HPE +EI ++
Sbjct: 620 NLYASSGRWGDVGKLRVRMRDKGVKKVPGYSWIEIQNKTHTFSVGDEFHPEKDEIFAFLE 679
Query: 604 QLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLR 663
+L +K GYV S VLHDVE+E+KE + HSE+LA+AYG+M++ S PIR+IKNLR
Sbjct: 680 ELDLRMKKAGYVSKTSVVLHDVEEEEKERMVRYHSERLAVAYGIMRVSSGRPIRVIKNLR 739
Query: 664 MCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
+C+DCH A+K ++++T R II+RD NRFHHF+DGSC+C D+W
Sbjct: 740 VCEDCHNAIKYMARITGRLIILRDNNRFHHFKDGSCSCGDYW 781
Score = 120 bits (300), Expect = 3e-27
Identities = 115/481 (23%), Positives = 208/481 (42%), Gaps = 50/481 (10%)
Query: 88 DVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREG 147
D+ WN ++ YM +G E L +FK + S+ + ++S +G
Sbjct: 63 DIKEWNVAISSYMRTGRCNEALRVFKRMPRWSSVS--------YNGMISGYLRNGEFELA 114
Query: 148 MQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVL 207
+ + + LVS+ + + Y R ++ A ++ E D+ +N++L+
Sbjct: 115 RKLFDEMPERDLVSW----NVMIKGYVRNRNLGKARELFEIM---PERDVCSWNTMLSGY 167
Query: 208 VESGRWEEAIGVLRRMGNE---------------------CMVW-DNATYVGVMGLCAQI 245
++G ++A V RM + CM++ + V C
Sbjct: 168 AQNGCVDDARSVFDRMPEKNDVSWNALLSAYVQNSKMEEACMLFKSRENWALVSWNCLLG 227
Query: 246 GDLQLGLRVHARLLRGGLLFDEFVG-SKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSL 304
G ++ V AR + + V + +I Y + + AR++FD ++V WT++
Sbjct: 228 GFVKKKKIVEARQFFDSMNVRDVVSWNTIITGYAQSGKIDEARQLFDESPVQDVFTWTAM 287
Query: 305 MSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGF 364
+S Y+QNR EE +L M + NE ++ +L AG + ++ + +
Sbjct: 288 VSGYIQNRMVEEARELFDKMPER----NEVSWNAML---AGYVQGERMEMAKELFDVMPC 340
Query: 365 KNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQD 424
+N V N +I Y++ G I + N+F RD +W +MI GYS G EALR+F
Sbjct: 341 RN-VSTWNTMITGYAQCGKISEAKNLFDKMPKRDPVSWAAMIAGYSQSGHSFEALRLFVQ 399
Query: 425 MKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAG 484
M+ G N +F LS CA + ++ G L+ + E G ++++YC+ G
Sbjct: 400 MEREGGRLNRSSFSSALSTCADVVALELG-KQLHGRLVKGGYETGCFVGNALLLMYCKCG 458
Query: 485 LLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQ--MDPQDMGTYTLL 542
+E+A + K K DIV+W T++ H ++ R ES+ + + P D +L
Sbjct: 459 SIEEANDLFKEMAGK-DIVSWNTMIAGYSRHGFGEVALRFFESMKREGLKPDDATMVAVL 517
Query: 543 S 543
S
Sbjct: 518 S 518
Score = 112 bits (279), Expect = 9e-25
Identities = 87/388 (22%), Positives = 178/388 (45%), Gaps = 26/388 (6%)
Query: 57 DVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLV 116
DVV N++I Y + G++ AR +FD+ P++DVF+W +++GY+ + E LF +
Sbjct: 249 DVVSWNTIITGYAQSGKIDEARQLFDESPVQDVFTWTAMVSGYIQNRMVEEARELFDKMP 308
Query: 117 SSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARC 176
NE + +L+ E M+ LF + + YA+C
Sbjct: 309 ER--------NEVSWNAMLAGYVQG----ERMEMAKELFDVMPCRNVSTWNTMITGYAQC 356
Query: 177 LHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYV 236
++ A + + + D + +++ +SG EA+ + +M E + +++
Sbjct: 357 GKISEAKNLFDKM---PKRDPVSWAAMIAGYSQSGHSFEALRLFVQMEREGGRLNRSSFS 413
Query: 237 GVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNR 296
+ CA + L+LG ++H RL++GG FVG+ L+ MY KC + A +F + +
Sbjct: 414 SALSTCADVVALELGKQLHGRLVKGGYETGCFVGNALLLMYCKCGSIEEANDLFKEMAGK 473
Query: 297 NVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLH 356
++V W ++++ Y ++ + E L M +E P++ T +L AC+ L+ G
Sbjct: 474 DIVSWNTMIAGYSRHGFGEVALRFFESMKREGLKPDDATMVAVLSACSHTGLVDKGRQYF 533
Query: 357 ARLEKLGFKNYVIVNNA-----LINMYSKSGDIDSSYNVFSDTMCR-DVTTWNSMICGYS 410
+ ++Y ++ N+ ++++ ++G ++ ++N+ + D W +++
Sbjct: 534 YTMT----QDYGVMPNSQHYACMVDLLGRAGLLEDAHNLMKNMPFEPDAAIWGTLLGASR 589
Query: 411 HHGLGQEALRVFQDMKSAGESPNNVTFV 438
HG E D A E N+ +V
Sbjct: 590 VHG-NTELAETAADKIFAMEPENSGMYV 616
Score = 82.0 bits (201), Expect = 1e-15
Identities = 77/345 (22%), Positives = 150/345 (43%), Gaps = 53/345 (15%)
Query: 194 ENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLR 253
++DI +N ++ + +GR EA+ V +RM W + +Y G++ + G+ +L +
Sbjct: 61 DSDIKEWNVAISSYMRTGRCNEALRVFKRMPR----WSSVSYNGMISGYLRNGEFELARK 116
Query: 254 V-----HARLLRGGLLFDEFVGSK----------------------LIDMYGKCCHVLNA 286
+ L+ ++ +V ++ ++ Y + V +A
Sbjct: 117 LFDEMPERDLVSWNVMIKGYVRNRNLGKARELFEIMPERDVCSWNTMLSGYAQNGCVDDA 176
Query: 287 RKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGI 346
R VFD + +N V W +L+SAY+QN EE L + + C LLG
Sbjct: 177 RSVFDRMPEKNDVSWNALLSAYVQNSKMEEACMLFKSRENWALVSWNC----LLG----- 227
Query: 347 ALLKHGDLLHAR--LEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNS 404
+K ++ AR + + ++ V+ N +I Y++SG ID + +F ++ +DV TW +
Sbjct: 228 GFVKKKKIVEARQFFDSMNVRD-VVSWNTIITGYAQSGKIDEARQLFDESPVQDVFTWTA 286
Query: 405 MICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNH 464
M+ GY + + +EA +F M E N G + + + + L++ M
Sbjct: 287 MVSGYIQNRMVEEARELFDKMPERNEVSWNAMLAGYVQG-ERMEMAKE----LFDVMPCR 341
Query: 465 KIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLL 509
+ + M+ Y + G + +A+N K D V+W ++
Sbjct: 342 NVST----WNTMITGYAQCGKISEAKNLFDKMP-KRDPVSWAAMI 381
>At5g09950 selenium-binding protein-like
Length = 995
Score = 425 bits (1092), Expect = e-119
Identities = 238/697 (34%), Positives = 394/697 (56%), Gaps = 19/697 (2%)
Query: 19 PLEYIVKLLKLS----ADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRL 74
P Y++ L A+ L G+ +H ++ + N L+++Y KCG +
Sbjct: 308 PESYVILLSSFPEYSLAEEVGLKKGREVHGHVITTGLVDFMVGIG--NGLVNMYAKCGSI 365
Query: 75 RLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTV 134
AR +F M +D SWN ++TG +G ++E + +K++ P + +
Sbjct: 366 ADARRVFYFMTDKDSVSWNSMITGLDQNGCFIEAVERYKSM----RRHDILPGSFTLISS 421
Query: 135 LSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDE 194
LSSC+ + G Q HG K G+ V +AL +YA ++N ++ S E
Sbjct: 422 LSSCASLKWAKLGQQIHGESLKLGIDLNVSVSNALMTLYAETGYLNECRKIF---SSMPE 478
Query: 195 NDIFLYNSLLNVLVESGR-WEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLR 253
+D +NS++ L S R EA+ + T+ V+ + + +LG +
Sbjct: 479 HDQVSWNSIIGALARSERSLPEAVVCFLNAQRAGQKLNRITFSSVLSAVSSLSFGELGKQ 538
Query: 254 VHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGL-QNRNVVVWTSLMSAYLQNR 312
+H L+ + + + LI YGKC + K+F + + R+ V W S++S Y+ N
Sbjct: 539 IHGLALKNNIADEATTENALIACYGKCGEMDGCEKIFSRMAERRDNVTWNSMISGYIHNE 598
Query: 313 YFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNN 372
+ LDL+ M Q + + +L A A +A L+ G +HA + ++ V+V +
Sbjct: 599 LLAKALDLVWFMLQTGQRLDSFMYATVLSAFASVATLERGMEVHACSVRACLESDVVVGS 658
Query: 373 ALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESP 432
AL++MYSK G +D + F+ R+ +WNSMI GY+ HG G+EAL++F+ MK G++P
Sbjct: 659 ALVDMYSKCGRLDYALRFFNTMPVRNSYSWNSMISGYARHGQGEEALKLFETMKLDGQTP 718
Query: 433 -NNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAEN 491
++VTFVGVLSAC+H L+++G + + ++ + P +EH++CM + RAG L+K E+
Sbjct: 719 PDHVTFVGVLSACSHAGLLEEGFKHFESMSDSYGLAPRIEHFSCMADVLGRAGELDKLED 778
Query: 492 FMKTTKVKWDIVAWRTLLNSC-RVH-RNYDLGKRIAESILQMDPQDMGTYTLLSNMHATA 549
F++ +K +++ WRT+L +C R + R +LGK+ AE + Q++P++ Y LL NM+A
Sbjct: 779 FIEKMPMKPNVLIWRTVLGACCRANGRKAELGKKAAEMLFQLEPENAVNYVLLGNMYAAG 838
Query: 550 NMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMI 609
W+ +V+ RK MK+ ++KKE G SW+ +++ H+FV+ HP+++ I KK+++L +
Sbjct: 839 GRWEDLVKARKKMKDADVKKEAGYSWVTMKDGVHMFVAGDKSHPDADVIYKKLKELNRKM 898
Query: 610 KPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGL-MKMPSPAPIRIIKNLRMCDDC 668
+ GYVP L+D+E E KE L+ HSEKLA+A+ L + S PIRI+KNLR+C DC
Sbjct: 899 RDAGYVPQTGFALYDLEQENKEEILSYHSEKLAVAFVLAAQRSSTLPIRIMKNLRVCGDC 958
Query: 669 HTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
H+A K ISK+ R II+RD+NRFHHF+DG+C+C+D W
Sbjct: 959 HSAFKYISKIEGRQIILRDSNRFHHFQDGACSCSDFW 995
Score = 171 bits (434), Expect = 1e-42
Identities = 129/472 (27%), Positives = 224/472 (47%), Gaps = 33/472 (6%)
Query: 57 DVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLV 116
DV N+LI+ Y++ G AR +FD+MPLR+ SW +++GY +G + E LV +++V
Sbjct: 35 DVYLCNNLINAYLETGDSVSARKVFDEMPLRNCVSWACIVSGYSRNGEHKEALVFLRDMV 94
Query: 117 SSSTTQHACPNEYVFTTVLSSCSHSGRV--REGMQCHGYLFKYGLVSYQFVKSALFNMYA 174
N+Y F +VL +C G V G Q HG +FK V + L +MY
Sbjct: 95 KEGIFS----NQYAFVSVLRACQEIGSVGILFGRQIHGLMFKLSYAVDAVVSNVLISMYW 150
Query: 175 RCL-HVNLAL------QVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNEC 227
+C+ V AL +V N+ S +NS+++V ++G A + M +
Sbjct: 151 KCIGSVGYALCAFGDIEVKNSVS---------WNSIISVYSQAGDQRSAFRIFSSMQYDG 201
Query: 228 MVWDNATYVGVMGLCAQI--GDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLN 285
T+ ++ + D++L ++ + + GLL D FVGS L+ + K +
Sbjct: 202 SRPTEYTFGSLVTTACSLTEPDVRLLEQIMCTIQKSGLLTDLFVGSGLVSAFAKSGSLSY 261
Query: 286 ARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAG 345
ARKVF+ ++ RN V LM ++ ++ EE L M+ + E ++ +LL +
Sbjct: 262 ARKVFNQMETRNAVTLNGLMVGLVRQKWGEEATKLFMDMNSMIDVSPE-SYVILLSSFPE 320
Query: 346 IAL-----LKHGDLLHARLEKLGFKNYVI-VNNALINMYSKSGDIDSSYNVFSDTMCRDV 399
+L LK G +H + G ++++ + N L+NMY+K G I + VF +D
Sbjct: 321 YSLAEEVGLKKGREVHGHVITTGLVDFMVGIGNGLVNMYAKCGSIADARRVFYFMTDKDS 380
Query: 400 TTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYN 459
+WNSMI G +G EA+ ++ M+ P + T + LS+CA L G ++
Sbjct: 381 VSWNSMITGLDQNGCFIEAVERYKSMRRHDILPGSFTLISSLSSCASLKWAKLG-QQIHG 439
Query: 460 EMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNS 511
E I+ + ++ LY G L + ++ + D V+W +++ +
Sbjct: 440 ESLKLGIDLNVSVSNALMTLYAETGYLNECRKIF-SSMPEHDQVSWNSIIGA 490
Score = 96.7 bits (239), Expect = 4e-20
Identities = 63/200 (31%), Positives = 99/200 (49%), Gaps = 3/200 (1%)
Query: 255 HARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYF 314
H+RL + L D ++ + LI+ Y + ++ARKVFD + RN V W ++S Y +N
Sbjct: 24 HSRLYKNRLDKDVYLCNNLINAYLETGDSVSARKVFDEMPLRNCVSWACIVSGYSRNGEH 83
Query: 315 EETLDLLTCMDQEDTLPNECTFKVLLGAC--AGIALLKHGDLLHARLEKLGFKNYVIVNN 372
+E L L M +E N+ F +L AC G + G +H + KL + +V+N
Sbjct: 84 KEALVFLRDMVKEGIFSNQYAFVSVLRACQEIGSVGILFGRQIHGLMFKLSYAVDAVVSN 143
Query: 373 ALINMYSKS-GDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGES 431
LI+MY K G + + F D ++ +WNS+I YS G + A R+F M+ G
Sbjct: 144 VLISMYWKCIGSVGYALCAFGDIEVKNSVSWNSIISVYSQAGDQRSAFRIFSSMQYDGSR 203
Query: 432 PNNVTFVGVLSACAHLALVD 451
P TF +++ L D
Sbjct: 204 PTEYTFGSLVTTACSLTEPD 223
>At4g13650 unknown protein
Length = 1024
Score = 421 bits (1083), Expect = e-118
Identities = 218/643 (33%), Positives = 362/643 (55%), Gaps = 7/643 (1%)
Query: 63 SLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQ 122
+L++LY KC + A F + + +V WN ++ Y + +F+ + +
Sbjct: 389 ALLNLYAKCADIETALDYFLETEVENVVLWNVMLVAYGLLDDLRNSFRIFRQM----QIE 444
Query: 123 HACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLA 182
PN+Y + ++L +C G + G Q H + K +V S L +MYA+ ++ A
Sbjct: 445 EIVPNQYTYPSILKTCIRLGDLELGEQIHSQIIKTNFQLNAYVCSVLIDMYAKLGKLDTA 504
Query: 183 LQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLC 242
+L +G D+ + +++ + ++A+ R+M + + D + C
Sbjct: 505 WDILIRFAG---KDVVSWTTMIAGYTQYNFDDKALTTFRQMLDRGIRSDEVGLTNAVSAC 561
Query: 243 AQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWT 302
A + L+ G ++HA+ G D + L+ +Y +C + + F+ + + + W
Sbjct: 562 AGLQALKEGQQIHAQACVSGFSSDLPFQNALVTLYSRCGKIEESYLAFEQTEAGDNIAWN 621
Query: 303 SLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKL 362
+L+S + Q+ EE L + M++E N TF + A + A +K G +HA + K
Sbjct: 622 ALVSGFQQSGNNEEALRVFVRMNREGIDNNNFTFGSAVKAASETANMKQGKQVHAVITKT 681
Query: 363 GFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVF 422
G+ + V NALI+MY+K G I + F + ++ +WN++I YS HG G EAL F
Sbjct: 682 GYDSETEVCNALISMYAKCGSISDAEKQFLEVSTKNEVSWNAIINAYSKHGFGSEALDSF 741
Query: 423 QDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCR 482
M + PN+VT VGVLSAC+H+ LVDKG+ Y + + + P EHY C+V + R
Sbjct: 742 DQMIHSNVRPNHVTLVGVLSACSHIGLVDKGIAYFESMNSEYGLSPKPEHYVCVVDMLTR 801
Query: 483 AGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLL 542
AGLL +A+ F++ +K D + WRTLL++C VH+N ++G+ A +L+++P+D TY LL
Sbjct: 802 AGLLSRAKEFIQEMPIKPDALVWRTLLSACVVHKNMEIGEFAAHHLLELEPEDSATYVLL 861
Query: 543 SNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKV 602
SN++A + WD R+ MKE+ +KKEPG SW+E++N H F HP ++EI++
Sbjct: 862 SNLYAVSKKWDARDLTRQKMKEKGVKKEPGQSWIEVKNSIHSFYVGDQNHPLADEIHEYF 921
Query: 603 QQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNL 662
Q L +GYV + ++L++++ EQK+ + HSEKLAI++GL+ +P+ PI ++KNL
Sbjct: 922 QDLTKRASEIGYVQDCFSLLNELQHEQKDPIIFIHSEKLAISFGLLSLPATVPINVMKNL 981
Query: 663 RMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
R+C+DCH +K +SKV+NR IIVRDA RFHHF G+C+C D+W
Sbjct: 982 RVCNDCHAWIKFVSKVSNREIIVRDAYRFHHFEGGACSCKDYW 1024
Score = 164 bits (415), Expect = 2e-40
Identities = 127/478 (26%), Positives = 232/478 (47%), Gaps = 19/478 (3%)
Query: 36 LTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDL 95
L G+ +H+Q+L + S+ L Y+ G L A +FD+MP R +F+WN +
Sbjct: 61 LDEGRKLHSQIL---KLGLDSNGCLSEKLFDFYLFKGDLYGAFKVFDEMPERTIFTWNKM 117
Query: 96 MTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGM--QCHGY 153
+ EV LF +VS + T PNE F+ VL +C G V + Q H
Sbjct: 118 IKELASRNLIGEVFGLFVRMVSENVT----PNEGTFSGVLEAC-RGGSVAFDVVEQIHAR 172
Query: 154 LFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRW 213
+ GL V + L ++Y+R V+LA +V + G D + ++++ L ++
Sbjct: 173 ILYQGLRDSTVVCNPLIDLYSRNGFVDLARRVFD---GLRLKDHSSWVAMISGLSKNECE 229
Query: 214 EEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKL 273
EAI + M ++ + V+ C +I L++G ++H +L+ G D +V + L
Sbjct: 230 AEAIRLFCDMYVLGIMPTPYAFSSVLSACKKIESLEIGEQLHGLVLKLGFSSDTYVCNAL 289
Query: 274 IDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNE 333
+ +Y ++++A +F + R+ V + +L++ Q Y E+ ++L M + P+
Sbjct: 290 VSLYFHLGNLISAEHIFSNMSQRDAVTYNTLINGLSQCGYGEKAMELFKRMHLDGLEPDS 349
Query: 334 CTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSD 393
T L+ AC+ L G LHA KLGF + + AL+N+Y+K DI+++ + F +
Sbjct: 350 NTLASLVVACSADGTLFRGQQLHAYTTKLGFASNNKIEGALLNLYAKCADIETALDYFLE 409
Query: 394 TMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKG 453
T +V WN M+ Y + + R+F+ M+ PN T+ +L C L ++ G
Sbjct: 410 TEVENVVLWNVMLVAYGLLDDLRNSFRIFRQMQIEEIVPNQYTYPSILKTCIRLGDLELG 469
Query: 454 LDYLYNEMKNHKIEPGLEHYTCMVV--LYCRAGLLEKAENFMKTTKVKWDIVAWRTLL 509
+ +++++ + L Y C V+ +Y + G L+ A + + K D+V+W T++
Sbjct: 470 -EQIHSQIIKTNFQ--LNAYVCSVLIDMYAKLGKLDTAWDILIRFAGK-DVVSWTTMI 523
Score = 148 bits (373), Expect = 1e-35
Identities = 113/392 (28%), Positives = 191/392 (47%), Gaps = 19/392 (4%)
Query: 126 PNEYVFTTVLSSC-SHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQ 184
PN +L C +G + EG + H + K GL S + LF+ Y + A +
Sbjct: 42 PNHQTLKWLLEGCLKTNGSLDEGRKLHSQILKLGLDSNGCLSEKLFDFYLFKGDLYGAFK 101
Query: 185 VLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQ 244
V + E IF +N ++ L E G+ RM +E + + T+ GV+ C
Sbjct: 102 VFDEMP---ERTIFTWNKMIKELASRNLIGEVFGLFVRMVSENVTPNEGTFSGVLEACRG 158
Query: 245 IGDLQLGL--RVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWT 302
G + + ++HAR+L GL V + LID+Y + V AR+VFDGL+ ++ W
Sbjct: 159 -GSVAFDVVEQIHARILYQGLRDSTVVCNPLIDLYSRNGFVDLARRVFDGLRLKDHSSWV 217
Query: 303 SLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKL 362
+++S +N E + L M +P F +L AC I L+ G+ LH + KL
Sbjct: 218 AMISGLSKNECEAEAIRLFCDMYVLGIMPTPYAFSSVLSACKKIESLEIGEQLHGLVLKL 277
Query: 363 GFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVF 422
GF + V NAL+++Y G++ S+ ++FS+ RD T+N++I G S G G++A+ +F
Sbjct: 278 GFSSDTYVCNALVSLYFHLGNLISAEHIFSNMSQRDAVTYNTLINGLSQCGYGEKAMELF 337
Query: 423 QDMKSAGESPNNVTFVGVLSACAHLALVDKGLD-YLYNE----MKNHKIEPGLEHYTCMV 477
+ M G P++ T ++ AC+ + +G + Y N+KIE L +
Sbjct: 338 KRMHLDGLEPDSNTLASLVVACSADGTLFRGQQLHAYTTKLGFASNNKIEGAL------L 391
Query: 478 VLYCRAGLLEKAENFMKTTKVKWDIVAWRTLL 509
LY + +E A ++ T+V+ ++V W +L
Sbjct: 392 NLYAKCADIETALDYFLETEVE-NVVLWNVML 422
Score = 90.1 bits (222), Expect = 4e-18
Identities = 73/298 (24%), Positives = 140/298 (46%), Gaps = 34/298 (11%)
Query: 31 ADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVF 90
A + L G+ IHAQ V + SD+ N+L+ LY +CG++ + F++ D
Sbjct: 562 AGLQALKEGQQIHAQACV---SGFSSDLPFQNALVTLYSRCGKIEESYLAFEQTEAGDNI 618
Query: 91 SWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQC 150
+WN L++G+ SGN E L +F + N + F + + + S + +++G Q
Sbjct: 619 AWNALVSGFQQSGNNEEALRVFVRMNREGIDN----NNFTFGSAVKAASETANMKQGKQV 674
Query: 151 HGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVES 210
H + K G S V +AL +MYA+C ++ A + S +E +N+++N +
Sbjct: 675 HAVITKTGYDSETEVCNALISMYAKCGSISDAEKQFLEVSTKNE---VSWNAIINAYSKH 731
Query: 211 GRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVG 270
G EA+ +M + + ++ T VGV+ C+ IG L+ G+ + E +
Sbjct: 732 GFGSEALDSFDQMIHSNVRPNHVTLVGVLSACSHIG-----------LVDKGIAYFESMN 780
Query: 271 SK------------LIDMYGKCCHVLNARKVFDGLQNR-NVVVWTSLMSAYLQNRYFE 315
S+ ++DM + + A++ + + + +VW +L+SA + ++ E
Sbjct: 781 SEYGLSPKPEHYVCVVDMLTRAGLLSRAKEFIQEMPIKPDALVWRTLLSACVVHKNME 838
>At3g03580 unknown protein (At3g03580)
Length = 882
Score = 420 bits (1079), Expect = e-117
Identities = 235/695 (33%), Positives = 390/695 (55%), Gaps = 14/695 (2%)
Query: 13 NSRYLPPLEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCG 72
NS +P + +L + + G+ +H L ++ S VV N L+ +Y+K
Sbjct: 200 NSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFAL---KSGVNSVVVVNNGLVAMYLKFR 256
Query: 73 RLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFT 132
R AR +FD+M +RD S+N ++ GY+ E + +F + P+ +
Sbjct: 257 RPTDARRVFDEMDVRDSVSYNTMICGYLKLEMVEESVRMFLENLDQFK-----PDLLTVS 311
Query: 133 TVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGG 192
+VL +C H + + Y+ K G V V++ L ++YA+C + A V N+
Sbjct: 312 SVLRACGHLRDLSLAKYIYNYMLKAGFVLESTVRNILIDVYAKCGDMITARDVFNSM--- 368
Query: 193 DENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGL 252
+ D +NS+++ ++SG EA+ + + M D+ TY+ ++ + ++ DL+ G
Sbjct: 369 ECKDTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISVSTRLADLKFGK 428
Query: 253 RVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNR 312
+H+ ++ G+ D V + LIDMY KC V ++ K+F + + V W +++SA ++
Sbjct: 429 GLHSNGIKSGICIDLSVSNALIDMYAKCGEVGDSLKIFSSMGTGDTVTWNTVISACVRFG 488
Query: 313 YFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNN 372
F L + T M + + +P+ TF V L CA +A + G +H L + G+++ + + N
Sbjct: 489 DFATGLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKEIHCCLLRFGYESELQIGN 548
Query: 373 ALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESP 432
ALI MYSK G +++S VF RDV TW MI Y +G G++AL F DM+ +G P
Sbjct: 549 ALIEMYSKCGCLENSSRVFERMSRRDVVTWTGMIYAYGMYGEGEKALETFADMEKSGIVP 608
Query: 433 NNVTFVGVLSACAHLALVDKGLDYLYNEMKNH-KIEPGLEHYTCMVVLYCRAGLLEKAEN 491
++V F+ ++ AC+H LVD+GL + +MK H KI+P +EHY C+V L R+ + KAE
Sbjct: 609 DSVVFIAIIYACSHSGLVDEGLA-CFEKMKTHYKIDPMIEHYACVVDLLSRSQKISKAEE 667
Query: 492 FMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANM 551
F++ +K D W ++L +CR + + +R++ I++++P D G L SN +A
Sbjct: 668 FIQAMPIKPDASIWASVLRACRTSGDMETAERVSRRIIELNPDDPGYSILASNAYAALRK 727
Query: 552 WDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKP 611
WD V IRK +K+++I K PG SW+E+ HVF S P+S I K ++ L +++
Sbjct: 728 WDKVSLIRKSLKDKHITKNPGYSWIEVGKNVHVFSSGDDSAPQSEAIYKSLEILYSLMAK 787
Query: 612 LGYVPNISAVLHDVEDEQKEGYLNC-HSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHT 670
GY+P+ V ++E+E+++ L C HSE+LAIA+GL+ P++++KNLR+C DCH
Sbjct: 788 EGYIPDPREVSQNLEEEEEKRRLICGHSERLAIAFGLLNTEPGTPLQVMKNLRVCGDCHE 847
Query: 671 AVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
KLISK+ R I+VRDANRFH F+DG+C+C D W
Sbjct: 848 VTKLISKIVGREILVRDANRFHLFKDGTCSCKDRW 882
Score = 190 bits (483), Expect = 2e-48
Identities = 134/494 (27%), Positives = 239/494 (48%), Gaps = 16/494 (3%)
Query: 39 GKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTG 98
G ++ Q+L SD+ N+L+ +Y + G L AR +FD+MP+RD+ SWN L++G
Sbjct: 125 GDLVYEQIL---DMGFESDLFVGNALVDMYSRMGLLTRARQVFDEMPVRDLVSWNSLISG 181
Query: 99 YMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYG 158
Y G Y E L ++ L +S P+ + ++VL + + V++G HG+ K G
Sbjct: 182 YSSHGYYEEALEIYHELKNS----WIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSG 237
Query: 159 LVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIG 218
+ S V + L MY + A +V + D D YN+++ ++ EE++
Sbjct: 238 VNSVVVVNNGLVAMYLKFRRPTDARRVFDEM---DVRDSVSYNTMICGYLKLEMVEESVR 294
Query: 219 VLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYG 278
+ ++ D T V+ C + DL L ++ +L+ G + + V + LID+Y
Sbjct: 295 MFLENLDQFKP-DLLTVSSVLRACGHLRDLSLAKYIYNYMLKAGFVLESTVRNILIDVYA 353
Query: 279 KCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKV 338
KC ++ AR VF+ ++ ++ V W S++S Y+Q+ E + L M + + T+ +
Sbjct: 354 KCGDMITARDVFNSMECKDTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLM 413
Query: 339 LLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRD 398
L+ +A LK G LH+ K G + V+NALI+MY+K G++ S +FS D
Sbjct: 414 LISVSTRLADLKFGKGLHSNGIKSGICIDLSVSNALIDMYAKCGEVGDSLKIFSSMGTGD 473
Query: 399 VTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLY 458
TWN++I G L+V M+ + P+ TF+ L CA LA G + ++
Sbjct: 474 TVTWNTVISACVRFGDFATGLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKE-IH 532
Query: 459 NEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNY 518
+ E L+ ++ +Y + G LE + + + D+V W ++ + + Y
Sbjct: 533 CCLLRFGYESELQIGNALIEMYSKCGCLENSSRVFERMS-RRDVVTWTGMIYA---YGMY 588
Query: 519 DLGKRIAESILQMD 532
G++ E+ M+
Sbjct: 589 GEGEKALETFADME 602
Score = 154 bits (389), Expect = 2e-37
Identities = 109/405 (26%), Positives = 196/405 (47%), Gaps = 6/405 (1%)
Query: 139 SHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIF 198
S S + E + H + GL S F L + Y+ +L V S +++
Sbjct: 15 SSSSNLNELRRIHALVISLGLDSSDFFSGKLIDKYSHFREPASSLSVFRRVSPA--KNVY 72
Query: 199 LYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARL 258
L+NS++ ++G + EA+ ++ + D T+ V+ CA + D ++G V+ ++
Sbjct: 73 LWNSIIRAFSKNGLFPEALEFYGKLRESKVSPDKYTFPSVIKACAGLFDAEMGDLVYEQI 132
Query: 259 LRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETL 318
L G D FVG+ L+DMY + + AR+VFD + R++V W SL+S Y + Y+EE L
Sbjct: 133 LDMGFESDLFVGNALVDMYSRMGLLTRARQVFDEMPVRDLVSWNSLISGYSSHGYYEEAL 192
Query: 319 DLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMY 378
++ + +P+ T +L A + ++K G LH K G + V+VNN L+ MY
Sbjct: 193 EIYHELKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSGVNSVVVVNNGLVAMY 252
Query: 379 SKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFV 438
K + VF + RD ++N+MICGY + +E++R+F + + P+ +T
Sbjct: 253 LKFRRPTDARRVFDEMDVRDSVSYNTMICGYLKLEMVEESVRMFLENLDQFK-PDLLTVS 311
Query: 439 GVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKV 498
VL AC HL + Y+YN M ++ +Y + G + A + + +
Sbjct: 312 SVLRACGHLRDLSLA-KYIYNYMLKAGFVLESTVRNILIDVYAKCGDMITARDVFNSMEC 370
Query: 499 KWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQ-DMGTYTLL 542
K D V+W ++++ + ++ + ++ M+ Q D TY +L
Sbjct: 371 K-DTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLML 414
Score = 152 bits (384), Expect = 6e-37
Identities = 118/466 (25%), Positives = 225/466 (47%), Gaps = 13/466 (2%)
Query: 85 PLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRV 144
P ++V+ WN ++ + +G + E L + L S + P++Y F +V+ +C+
Sbjct: 67 PAKNVYLWNSIIRAFSKNGLFPEALEFYGKLRESKVS----PDKYTFPSVIKACAGLFDA 122
Query: 145 REGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLL 204
G + + G S FV +AL +MY+R + A QV + D+ +NSL+
Sbjct: 123 EMGDLVYEQILDMGFESDLFVGNALVDMYSRMGLLTRARQVFDEM---PVRDLVSWNSLI 179
Query: 205 NVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLL 264
+ G +EEA+ + + N +V D+ T V+ + ++ G +H L+ G+
Sbjct: 180 SGYSSHGYYEEALEIYHELKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSGVN 239
Query: 265 FDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDL-LTC 323
V + L+ MY K +AR+VFD + R+ V + +++ YL+ EE++ + L
Sbjct: 240 SVVVVNNGLVAMYLKFRRPTDARRVFDEMDVRDSVSYNTMICGYLKLEMVEESVRMFLEN 299
Query: 324 MDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGD 383
+DQ P+ T +L AC + L ++ + K GF V N LI++Y+K GD
Sbjct: 300 LDQ--FKPDLLTVSSVLRACGHLRDLSLAKYIYNYMLKAGFVLESTVRNILIDVYAKCGD 357
Query: 384 IDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSA 443
+ ++ +VF+ C+D +WNS+I GY G EA+++F+ M E +++T++ ++S
Sbjct: 358 MITARDVFNSMECKDTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISV 417
Query: 444 CAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIV 503
LA + G N +K+ I L ++ +Y + G + + + D V
Sbjct: 418 STRLADLKFGKGLHSNGIKS-GICIDLSVSNALIDMYAKCGEVGDSLKIFSSMGT-GDTV 475
Query: 504 AWRTLLNSCRVHRNYDLGKRIAESILQMD-PQDMGTYTLLSNMHAT 548
W T++++C ++ G ++ + + + DM T+ + M A+
Sbjct: 476 TWNTVISACVRFGDFATGLQVTTQMRKSEVVPDMATFLVTLPMCAS 521
>At1g16480 hypothetical protein
Length = 905
Score = 419 bits (1078), Expect = e-117
Identities = 231/668 (34%), Positives = 371/668 (54%), Gaps = 11/668 (1%)
Query: 39 GKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTG 98
G+ IH ++ + S V N+L+ +Y GR A +F +MP +D+ SWN LM
Sbjct: 248 GRGIHGLVV---KMGFDSVVCVCNTLLRMYAGAGRSVEANLVFKQMPTKDLISWNSLMAS 304
Query: 99 YMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYG 158
+++ G L+ L L +++SS + N FT+ L++C +G HG + G
Sbjct: 305 FVNDGRSLDALGLLCSMISSGKSV----NYVTFTSALAACFTPDFFEKGRILHGLVVVSG 360
Query: 159 LVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIG 218
L Q + +AL +MY + ++ + +VL D+ +N+L+ E ++A+
Sbjct: 361 LFYNQIIGNALVSMYGKIGEMSESRRVLLQMP---RRDVVAWNALIGGYAEDEDPDKALA 417
Query: 219 VLRRMGNECMVWDNATYVGVMGLCAQIGDL-QLGLRVHARLLRGGLLFDEFVGSKLIDMY 277
+ M E + + T V V+ C GDL + G +HA ++ G DE V + LI MY
Sbjct: 418 AFQTMRVEGVSSNYITVVSVLSACLLPGDLLERGKPLHAYIVSAGFESDEHVKNSLITMY 477
Query: 278 GKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFK 337
KC + +++ +F+GL NRN++ W ++++A + + EE L L++ M ++ +F
Sbjct: 478 AKCGDLSSSQDLFNGLDNRNIITWNAMLAANAHHGHGEEVLKLVSKMRSFGVSLDQFSFS 537
Query: 338 VLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCR 397
L A A +A+L+ G LH KLGF++ + NA +MYSK G+I + ++ R
Sbjct: 538 EGLSAAAKLAVLEEGQQLHGLAVKLGFEHDSFIFNAAADMYSKCGEIGEVVKMLPPSVNR 597
Query: 398 DVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYL 457
+ +WN +I HG +E F +M G P +VTFV +L+AC+H LVDKGL Y
Sbjct: 598 SLPSWNILISALGRHGYFEEVCATFHEMLEMGIKPGHVTFVSLLTACSHGGLVDKGLAYY 657
Query: 458 YNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRN 517
++ +EP +EH C++ L R+G L +AE F+ +K + + WR+LL SC++H N
Sbjct: 658 DMIARDFGLEPAIEHCICVIDLLGRSGRLAEAETFISKMPMKPNDLVWRSLLASCKIHGN 717
Query: 518 YDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLE 577
D G++ AE++ +++P+D Y L SNM AT W+ V +RK M +NIKK+ SW++
Sbjct: 718 LDRGRKAAENLSKLEPEDDSVYVLSSNMFATTGRWEDVENVRKQMGFKNIKKKQACSWVK 777
Query: 578 IRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCH 637
+++ F HP++ EI K++ + +IK GYV + S L D ++EQKE L H
Sbjct: 778 LKDKVSSFGIGDRTHPQTMEIYAKLEDIKKLIKESGYVADTSQALQDTDEEQKEHNLWNH 837
Query: 638 SEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDG 697
SE+LA+AY LM P + +RI KNLR+C DCH+ K +S+V R I++RD RFHHF G
Sbjct: 838 SERLALAYALMSTPEGSTVRIFKNLRICSDCHSVYKFVSRVIGRRIVLRDQYRFHHFERG 897
Query: 698 SCTCADHW 705
C+C D+W
Sbjct: 898 LCSCKDYW 905
Score = 155 bits (392), Expect = 7e-38
Identities = 138/533 (25%), Positives = 236/533 (43%), Gaps = 35/533 (6%)
Query: 16 YLPPLEYIVKLLKL----------SADAKCLTSGKAIHAQLLVRDQASKR---SDVVQLN 62
YL +E+ K+ L S C SG + V +K SDV
Sbjct: 10 YLEGMEFFRKMCDLGIKPSSFVIASLVTACGRSGSMFREGVQVHGFVAKSGLLSDVYVST 69
Query: 63 SLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQ 122
+++HLY G + +R +F++MP R+V SW LM GY G EV+ ++K +
Sbjct: 70 AILHLYGVYGLVSCSRKVFEEMPDRNVVSWTSLMVGYSDKGEPEEVIDIYKGMRGEGV-- 127
Query: 123 HACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLA 182
C NE ++ C G Q G + K GL S V+++L +M +V+ A
Sbjct: 128 -GC-NE---NSMSLFCGLLKDESLGRQIIGQVVKSGLESKLAVENSLISMLGSMGNVDYA 182
Query: 183 LQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGV---LRRMGNECMVWDNATYVGVM 239
+ + S E D +NS+ ++G EE+ + +RR +E +T + V+
Sbjct: 183 NYIFDQMS---ERDTISWNSIAAAYAQNGHIEESFRIFSLMRRFHDEVNSTTVSTLLSVL 239
Query: 240 GLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVV 299
G + + G +H +++ G V + L+ MY + A VF + ++++
Sbjct: 240 G---HVDHQKWGRGIHGLVVKMGFDSVVCVCNTLLRMYAGAGRSVEANLVFKQMPTKDLI 296
Query: 300 VWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARL 359
W SLM++++ + + L LL M N TF L AC + G +LH +
Sbjct: 297 SWNSLMASFVNDGRSLDALGLLCSMISSGKSVNYVTFTSALAACFTPDFFEKGRILHGLV 356
Query: 360 EKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEAL 419
G I+ NAL++MY K G++ S V RDV WN++I GY+ +AL
Sbjct: 357 VVSGLFYNQIIGNALVSMYGKIGEMSESRRVLLQMPRRDVVAWNALIGGYAEDEDPDKAL 416
Query: 420 RVFQDMKSAGESPNNVTFVGVLSACAHLA-LVDKGLDYLYNEMKNHKIEPGLEHYTCMVV 478
FQ M+ G S N +T V VLSAC L+++G L+ + + E ++
Sbjct: 417 AAFQTMRVEGVSSNYITVVSVLSACLLPGDLLERGKP-LHAYIVSAGFESDEHVKNSLIT 475
Query: 479 LYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQM 531
+Y + G L +++ + +I+ W +L + H + G+ + + + +M
Sbjct: 476 MYAKCGDLSSSQDLFNGLDNR-NIITWNAML-AANAHHGH--GEEVLKLVSKM 524
Score = 151 bits (381), Expect = 1e-36
Identities = 114/419 (27%), Positives = 202/419 (48%), Gaps = 21/419 (5%)
Query: 96 MTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRV-REGMQCHGYL 154
M+G + G YLE + F+ + P+ +V +++++C SG + REG+Q HG++
Sbjct: 1 MSGIVRVGLYLEGMEFFRKMCDLGIK----PSSFVIASLVTACGRSGSMFREGVQVHGFV 56
Query: 155 FKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWE 214
K GL+S +V +A+ ++Y V+ + +V E D N + + SL+ + G E
Sbjct: 57 AKSGLLSDVYVSTAILHLYGVYGLVSCSRKVF--EEMPDRN-VVSWTSLMVGYSDKGEPE 113
Query: 215 EAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLI 274
E I + + M E V N + + C + D LG ++ ++++ GL V + LI
Sbjct: 114 EVIDIYKGMRGEG-VGCNENSMSLF--CGLLKDESLGRQIIGQVVKSGLESKLAVENSLI 170
Query: 275 DMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNEC 334
M G +V A +FD + R+ + W S+ +AY QN + EE+ + + M + N
Sbjct: 171 SMLGSMGNVDYANYIFDQMSERDTISWNSIAAAYAQNGHIEESFRIFSLMRRFHDEVNST 230
Query: 335 TFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDT 394
T LL + K G +H + K+GF + V V N L+ MY+ +G + VF
Sbjct: 231 TVSTLLSVLGHVDHQKWGRGIHGLVVKMGFDSVVCVCNTLLRMYAGAGRSVEANLVFKQM 290
Query: 395 MCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGL 454
+D+ +WNS++ + + G +AL + M S+G+S N VTF L+AC +KG
Sbjct: 291 PTKDLISWNSLMASFVNDGRSLDALGLLCSMISSGKSVNYVTFTSALAACFTPDFFEKG- 349
Query: 455 DYLYNEMKNHKIEPGLEHY----TCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLL 509
+ + GL + +V +Y + G + ++ + + D+VAW L+
Sbjct: 350 ----RILHGLVVVSGLFYNQIIGNALVSMYGKIGEMSESRRVL-LQMPRRDVVAWNALI 403
>At2g29760 hypothetical protein
Length = 738
Score = 417 bits (1072), Expect = e-116
Identities = 229/665 (34%), Positives = 368/665 (54%), Gaps = 39/665 (5%)
Query: 74 LRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTT 133
L AR +FD++P + F+WN L+ Y + + + F ++VS S PN+Y F
Sbjct: 80 LEYARKVFDEIPKPNSFAWNTLIRAYASGPDPVLSIWAFLDMVSES---QCYPNKYTFPF 136
Query: 134 VLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGD 193
++ + + + G HG K + S FV ++L + Y C ++ A +V T
Sbjct: 137 LIKAAAEVSSLSLGQSLHGMAVKSAVGSDVFVANSLIHCYFSCGDLDSACKVFTTIK--- 193
Query: 194 ENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLR 253
E D+ +NS++N V+ G ++A+ + ++M +E + + T VGV+ CA+I +L+ G +
Sbjct: 194 EKDVVSWNSMINGFVQKGSPDKALELFKKMESEDVKASHVTMVGVLSACAKIRNLEFGRQ 253
Query: 254 VHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRY 313
V + + + + + + ++DMY KC + +A+++FD ++ ++ V WT+++ Y +
Sbjct: 254 VCSYIEENRVNVNLTLANAMLDMYTKCGSIEDAKRLFDAMEEKDNVTWTTMLDGYAISED 313
Query: 314 FEETLDLLTCMDQEDTL--------------PNEC------------------TFKVLLG 341
+E ++L M Q+D + PNE T L
Sbjct: 314 YEAAREVLNSMPQKDIVAWNALISAYEQNGKPNEALIVFHELQLQKNMKLNQITLVSTLS 373
Query: 342 ACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTT 401
ACA + L+ G +H+ ++K G + V +ALI+MYSK GD++ S VF+ RDV
Sbjct: 374 ACAQVGALELGRWIHSYIKKHGIRMNFHVTSALIHMYSKCGDLEKSREVFNSVEKRDVFV 433
Query: 402 WNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEM 461
W++MI G + HG G EA+ +F M+ A PN VTF V AC+H LVD+ +
Sbjct: 434 WSAMIGGLAMHGCGNEAVDMFYKMQEANVKPNGVTFTNVFCACSHTGLVDEAESLFHQME 493
Query: 462 KNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLG 521
N+ I P +HY C+V + R+G LEKA F++ + W LL +C++H N +L
Sbjct: 494 SNYGIVPEEKHYACIVDVLGRSGYLEKAVKFIEAMPIPPSTSVWGALLGACKIHANLNLA 553
Query: 522 KRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNV 581
+ +L+++P++ G + LLSN++A W+ V E+RK M+ +KKEPG S +EI +
Sbjct: 554 EMACTRLLELEPRNDGAHVLLSNIYAKLGKWENVSELRKHMRVTGLKKEPGCSSIEIDGM 613
Query: 582 NHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQ-KEGYLNCHSEK 640
H F+S HP S ++ K+ +++ +K GY P IS VL +E+E+ KE LN HSEK
Sbjct: 614 IHEFLSGDNAHPMSEKVYGKLHEVMEKLKSNGYEPEISQVLQIIEEEEMKEQSLNLHSEK 673
Query: 641 LAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCT 700
LAI YGL+ +P IR+IKNLR+C DCH+ KLIS++ +R IIVRD RFHHFR+G C+
Sbjct: 674 LAICYGLISTEAPKVIRVIKNLRVCGDCHSVAKLISQLYDREIIVRDRYRFHHFRNGQCS 733
Query: 701 CADHW 705
C D W
Sbjct: 734 CNDFW 738
Score = 139 bits (351), Expect = 4e-33
Identities = 98/396 (24%), Positives = 186/396 (46%), Gaps = 45/396 (11%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
L+K +A+ L+ G+++H + +++ SDV NSLIH Y CG L A +F +
Sbjct: 137 LIKAAAEVSSLSLGQSLHGMAV---KSAVGSDVFVANSLIHCYFSCGDLDSACKVFTTIK 193
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
+DV SWN ++ G++ G+ + L LFK + S + VLS+C+ +
Sbjct: 194 EKDVVSWNSMINGFVQKGSPDKALELFKKMESEDVK----ASHVTMVGVLSACAKIRNLE 249
Query: 146 EGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGD------------ 193
G Q Y+ + + + +A+ +MY +C + A ++ + D
Sbjct: 250 FGRQVCSYIEENRVNVNLTLANAMLDMYTKCGSIEDAKRLFDAMEEKDNVTWTTMLDGYA 309
Query: 194 ----------------ENDIFLYNSLLNVLVESGRWEEAIGVLRRMG-NECMVWDNATYV 236
+ DI +N+L++ ++G+ EA+ V + + M + T V
Sbjct: 310 ISEDYEAAREVLNSMPQKDIVAWNALISAYEQNGKPNEALIVFHELQLQKNMKLNQITLV 369
Query: 237 GVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNR 296
+ CAQ+G L+LG +H+ + + G+ + V S LI MY KC + +R+VF+ ++ R
Sbjct: 370 STLSACAQVGALELGRWIHSYIKKHGIRMNFHVTSALIHMYSKCGDLEKSREVFNSVEKR 429
Query: 297 NVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLH 356
+V VW++++ + E +D+ M + + PN TF + AC+ L+ + L
Sbjct: 430 DVFVWSAMIGGLAMHGCGNEAVDMFYKMQEANVKPNGVTFTNVFCACSHTGLVDEAESLF 489
Query: 357 ARLEKLGFKNYVIVNN-----ALINMYSKSGDIDSS 387
++E NY IV ++++ +SG ++ +
Sbjct: 490 HQME----SNYGIVPEEKHYACIVDVLGRSGYLEKA 521
Score = 136 bits (342), Expect = 5e-32
Identities = 92/343 (26%), Positives = 165/343 (47%), Gaps = 14/343 (4%)
Query: 231 DNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDM--YGKCCHVLNARK 288
+ + ++ ++ C + L+ + H ++R G D + SKL M + ARK
Sbjct: 29 ERSRHISLIERCVSLRQLK---QTHGHMIRTGTFSDPYSASKLFAMAALSSFASLEYARK 85
Query: 289 VFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMD---QEDTLPNECTFKVLLGACAG 345
VFD + N W +L+ AY + L + +D + PN+ TF L+ A A
Sbjct: 86 VFDEIPKPNSFAWNTLIRAYASGP--DPVLSIWAFLDMVSESQCYPNKYTFPFLIKAAAE 143
Query: 346 IALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSM 405
++ L G LH K + V V N+LI+ Y GD+DS+ VF+ +DV +WNSM
Sbjct: 144 VSSLSLGQSLHGMAVKSAVGSDVFVANSLIHCYFSCGDLDSACKVFTTIKEKDVVSWNSM 203
Query: 406 ICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHK 465
I G+ G +AL +F+ M+S ++VT VGVLSACA + ++ G + + ++ ++
Sbjct: 204 INGFVQKGSPDKALELFKKMESEDVKASHVTMVGVLSACAKIRNLEFGRQ-VCSYIEENR 262
Query: 466 IEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIA 525
+ L M+ +Y + G +E A+ + K D V W T+L+ + +Y+ + +
Sbjct: 263 VNVNLTLANAMLDMYTKCGSIEDAKRLFDAMEEK-DNVTWTTMLDGYAISEDYEAAREVL 321
Query: 526 ESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIK 568
S+ Q D + L+S + ++ ++ ++N+K
Sbjct: 322 NSMPQKD--IVAWNALISAYEQNGKPNEALIVFHELQLQKNMK 362
Score = 129 bits (325), Expect = 4e-30
Identities = 98/405 (24%), Positives = 182/405 (44%), Gaps = 36/405 (8%)
Query: 144 VREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSL 203
+R+ Q HG++ + G S + S LF M A +L + N F +N+L
Sbjct: 43 LRQLKQTHGHMIRTGTFSDPYSASKLFAMAALSSFASLEYARKVFDEIPKPNS-FAWNTL 101
Query: 204 LNVLVESGRWEEAIGVLRRMGNECMVWDNA-TYVGVMGLCAQIGDLQLGLRVHARLLRGG 262
+ +I M +E + N T+ ++ A++ L LG +H ++
Sbjct: 102 IRAYASGPDPVLSIWAFLDMVSESQCYPNKYTFPFLIKAAAEVSSLSLGQSLHGMAVKSA 161
Query: 263 LLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLT 322
+ D FV + LI Y C + +A KVF ++ ++VV W S+++ ++Q ++ L+L
Sbjct: 162 VGSDVFVANSLIHCYFSCGDLDSACKVFTTIKEKDVVSWNSMINGFVQKGSPDKALELFK 221
Query: 323 CMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSG 382
M+ ED + T +L ACA I L+ G + + +E+ + + NA+++MY+K G
Sbjct: 222 KMESEDVKASHVTMVGVLSACAKIRNLEFGRQVCSYIEENRVNVNLTLANAMLDMYTKCG 281
Query: 383 DIDSSYNVFS----------DTMC---------------------RDVTTWNSMICGYSH 411
I+ + +F TM +D+ WN++I Y
Sbjct: 282 SIEDAKRLFDAMEEKDNVTWTTMLDGYAISEDYEAAREVLNSMPQKDIVAWNALISAYEQ 341
Query: 412 HGLGQEALRVFQDMK-SAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGL 470
+G EAL VF +++ N +T V LSACA + ++ G ++++ +K H I
Sbjct: 342 NGKPNEALIVFHELQLQKNMKLNQITLVSTLSACAQVGALELG-RWIHSYIKKHGIRMNF 400
Query: 471 EHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
+ ++ +Y + G LEK+ + + K D+ W ++ +H
Sbjct: 401 HVTSALIHMYSKCGDLEKSREVFNSVE-KRDVFVWSAMIGGLAMH 444
Score = 84.7 bits (208), Expect = 2e-16
Identities = 59/258 (22%), Positives = 123/258 (46%), Gaps = 8/258 (3%)
Query: 52 ASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVL 111
A + D V +++ Y AR + + MP +D+ +WN L++ Y +G E L++
Sbjct: 292 AMEEKDNVTWTTMLDGYAISEDYEAAREVLNSMPQKDIVAWNALISAYEQNGKPNEALIV 351
Query: 112 FKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFN 171
F L ++ N+ + LS+C+ G + G H Y+ K+G+ V SAL +
Sbjct: 352 FHEL---QLQKNMKLNQITLVSTLSACAQVGALELGRWIHSYIKKHGIRMNFHVTSALIH 408
Query: 172 MYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWD 231
MY++C + + +V N+ ++ D+F++++++ L G EA+ + +M + +
Sbjct: 409 MYSKCGDLEKSREVFNSV---EKRDVFVWSAMIGGLAMHGCGNEAVDMFYKMQEANVKPN 465
Query: 232 NATYVGVMGLCAQIGDL-QLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVF 290
T+ V C+ G + + H G++ +E + ++D+ G+ ++ A K
Sbjct: 466 GVTFTNVFCACSHTGLVDEAESLFHQMESNYGIVPEEKHYACIVDVLGRSGYLEKAVKFI 525
Query: 291 DGLQ-NRNVVVWTSLMSA 307
+ + + VW +L+ A
Sbjct: 526 EAMPIPPSTSVWGALLGA 543
Score = 63.2 bits (152), Expect = 5e-10
Identities = 44/165 (26%), Positives = 78/165 (46%), Gaps = 8/165 (4%)
Query: 23 IVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFD 82
+V L A L G+ IH+ + + R + ++LIH+Y KCG L +R +F+
Sbjct: 368 LVSTLSACAQVGALELGRWIHSYI---KKHGIRMNFHVTSALIHMYSKCGDLEKSREVFN 424
Query: 83 KMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSG 142
+ RDVF W+ ++ G G E + +F + ++ PN FT V +CSH+G
Sbjct: 425 SVEKRDVFVWSAMIGGLAMHGCGNEAVDMFYKMQEANVK----PNGVTFTNVFCACSHTG 480
Query: 143 RVREGMQC-HGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVL 186
V E H YG+V + + + ++ R ++ A++ +
Sbjct: 481 LVDEAESLFHQMESNYGIVPEEKHYACIVDVLGRSGYLEKAVKFI 525
>At5g46460 putative protein
Length = 697
Score = 412 bits (1060), Expect = e-115
Identities = 222/640 (34%), Positives = 365/640 (56%), Gaps = 20/640 (3%)
Query: 68 YVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPN 127
Y + RL A ++FD+MP+RDV SWN +++G + G+ + LF + S
Sbjct: 76 YTRSNRLVDALNLFDEMPVRDVVSWNSMISGCVECGDMNTAVKLFDEMPERSVVS----- 130
Query: 128 EYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLN 187
+T +++ C SG+V + + LF V +++ + Y + V+ AL++
Sbjct: 131 ---WTAMVNGCFRSGKVDQAER----LFYQMPVKDTAAWNSMVHGYLQFGKVDDALKLFK 183
Query: 188 TESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGD 247
G ++ + +++ L ++ R EA+ + + M C+ + + V+ CA
Sbjct: 184 QMPG---KNVISWTTMICGLDQNERSGEALDLFKNMLRCCIKSTSRPFTCVITACANAPA 240
Query: 248 LQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSA 307
+G++VH +++ G L++E+V + LI Y C + ++RKVFD + V VWT+L+S
Sbjct: 241 FHMGIQVHGLIIKLGFLYEEYVSASLITFYANCKRIGDSRKVFDEKVHEQVAVWTALLSG 300
Query: 308 YLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNY 367
Y N+ E+ L + + M + LPN+ TF L +C+ + L G +H KLG +
Sbjct: 301 YSLNKKHEDALSIFSGMLRNSILPNQSTFASGLNSCSALGTLDWGKEMHGVAVKLGLETD 360
Query: 368 VIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKS 427
V N+L+ MYS SG+++ + +VF + + +WNS+I G + HG G+ A +F M
Sbjct: 361 AFVGNSLVVMYSDSGNVNDAVSVFIKIFKKSIVSWNSIIVGCAQHGRGKWAFVIFGQMIR 420
Query: 428 AGESPNNVTFVGVLSACAHLALVDKG--LDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGL 485
+ P+ +TF G+LSAC+H ++KG L Y + NH I+ ++HYTCMV + R G
Sbjct: 421 LNKEPDEITFTGLLSACSHCGFLEKGRKLFYYMSSGINH-IDRKIQHYTCMVDILGRCGK 479
Query: 486 LEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNM 545
L++AE ++ VK + + W LL++CR+H + D G++ A +I +D + Y LLSN+
Sbjct: 480 LKEAEELIERMVVKPNEMVWLALLSACRMHSDVDRGEKAAAAIFNLDSKSSAAYVLLSNI 539
Query: 546 HATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQL 605
+A+A W V ++R MK+ I K+PG+SW+ IR H F S P + I +K++ L
Sbjct: 540 YASAGRWSNVSKLRVKMKKNGIMKKPGSSWVVIRGKKHEFFSG--DQPHCSRIYEKLEFL 597
Query: 606 LAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMC 665
+K LGY P+ + LHDVEDEQKE L HSE+LAIA+GL+ + + ++KNLR+C
Sbjct: 598 REKLKELGYAPDYRSALHDVEDEQKEEMLWYHSERLAIAFGLINTVEGSAVTVMKNLRVC 657
Query: 666 DDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
+DCHT +KLIS V R I++RD RFHHF++G+C+C D+W
Sbjct: 658 EDCHTVIKLISGVVGREIVLRDPIRFHHFKNGTCSCGDYW 697
Score = 130 bits (326), Expect = 3e-30
Identities = 102/419 (24%), Positives = 190/419 (45%), Gaps = 41/419 (9%)
Query: 35 CLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWND 94
C SGK A+ L K D NS++H Y++ G++ A +F +MP ++V SW
Sbjct: 138 CFRSGKVDQAERLFYQMPVK--DTAAWNSMVHGYLQFGKVDDALKLFKQMPGKNVISWTT 195
Query: 95 LMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYL 154
++ G + E L LFKN++ + P FT V+++C+++ G+Q HG +
Sbjct: 196 MICGLDQNERSGEALDLFKNMLRCCIKSTSRP----FTCVITACANAPAFHMGIQVHGLI 251
Query: 155 FKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWE 214
K G + ++V ++L YA C + + +V + + + ++ +LL+ + + E
Sbjct: 252 IKLGFLYEEYVSASLITFYANCKRIGDSRKVFDEKV---HEQVAVWTALLSGYSLNKKHE 308
Query: 215 EAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLI 274
+A+ + M ++ + +T+ + C+ +G L G +H ++ GL D FVG+ L+
Sbjct: 309 DALSIFSGMLRNSILPNQSTFASGLNSCSALGTLDWGKEMHGVAVKLGLETDAFVGNSLV 368
Query: 275 DMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNEC 334
MY +V +A VF + +++V W S++ Q+ + + M + + P+E
Sbjct: 369 VMYSDSGNVNDAVSVFIKIFKKSIVSWNSIIVGCAQHGRGKWAFVIFGQMIRLNKEPDEI 428
Query: 335 TFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDT 394
TF LL AC+ L+ G L Y SG +
Sbjct: 429 TFTGLLSACSHCGFLEKGRKLF--------------------YYMSSG---------INH 459
Query: 395 MCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKG 453
+ R + + M+ G +EA + + M PN + ++ +LSAC + VD+G
Sbjct: 460 IDRKIQHYTCMVDILGRCGKLKEAEELIERMV---VKPNEMVWLALLSACRMHSDVDRG 515
Score = 58.9 bits (141), Expect = 9e-09
Identities = 53/271 (19%), Positives = 112/271 (40%), Gaps = 32/271 (11%)
Query: 271 SKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTL 330
+K+I Y + +++A +FD + R+VV W S++S + E D+ T + D +
Sbjct: 70 TKMITGYTRSNRLVDALNLFDEMPVRDVVSWNSMISGCV------ECGDMNTAVKLFDEM 123
Query: 331 PNE--CTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSY 388
P ++ ++ C + + L ++ + N++++ Y + G +D +
Sbjct: 124 PERSVVSWTAMVNGCFRSGKVDQAERLFYQMPVKDTAAW----NSMVHGYLQFGKVDDAL 179
Query: 389 NVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLA 448
+F ++V +W +MICG + EAL +F++M + F V++ACA+
Sbjct: 180 KLFKQMPGKNVISWTTMICGLDQNERSGEALDLFKNMLRCCIKSTSRPFTCVITACANAP 239
Query: 449 LVDKG---------LDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVK 499
G L +LY E + ++ Y + + KV
Sbjct: 240 AFHMGIQVHGLIIKLGFLYEEYVS----------ASLITFYANCKRIGDSRKVF-DEKVH 288
Query: 500 WDIVAWRTLLNSCRVHRNYDLGKRIAESILQ 530
+ W LL+ +++ ++ I +L+
Sbjct: 289 EQVAVWTALLSGYSLNKKHEDALSIFSGMLR 319
Score = 45.8 bits (107), Expect = 8e-05
Identities = 37/158 (23%), Positives = 75/158 (47%), Gaps = 12/158 (7%)
Query: 367 YVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMK 426
+V + +I Y++S + + N+F + RDV +WNSMI G G A+++F +M
Sbjct: 65 HVSLYTKMITGYTRSNRLVDALNLFDEMPVRDVVSWNSMISGCVECGDMNTAVKLFDEMP 124
Query: 427 SAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLL 486
+ V++ +++ C VD+ + L+ +M + MV Y + G +
Sbjct: 125 ER----SVVSWTAMVNGCFRSGKVDQA-ERLFYQMP----VKDTAAWNSMVHGYLQFGKV 175
Query: 487 EKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRI 524
+ A K K ++++W T++ C + +N G+ +
Sbjct: 176 DDALKLFKQMPGK-NVISWTTMI--CGLDQNERSGEAL 210
Score = 38.9 bits (89), Expect = 0.010
Identities = 40/161 (24%), Positives = 74/161 (45%), Gaps = 16/161 (9%)
Query: 351 HGDLLHARLEKLGFKNY-VIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGY 409
HG + + F+N V++ N L+ S ID + VF+ V+ + MI GY
Sbjct: 22 HGKCYRSFSVTVEFQNREVLICNHLL-----SRRIDEAREVFNQVPSPHVSLYTKMITGY 76
Query: 410 SHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPG 469
+ +AL +F +M + V++ ++S C ++ + L++EM E
Sbjct: 77 TRSNRLVDALNLFDEMPVR----DVVSWNSMISGCVECGDMNTAVK-LFDEMP----ERS 127
Query: 470 LEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLN 510
+ +T MV R+G +++AE VK D AW ++++
Sbjct: 128 VVSWTAMVNGCFRSGKVDQAERLFYQMPVK-DTAAWNSMVH 167
>At3g57430 putative protein
Length = 803
Score = 412 bits (1059), Expect = e-115
Identities = 227/684 (33%), Positives = 381/684 (55%), Gaps = 25/684 (3%)
Query: 36 LTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDL 95
L GK +HA L + + + +N+L+ +Y K G+L ++ + RD+ +WN +
Sbjct: 131 LMMGKQVHAYGLRKGELNS----FIINTLVAMYGKLGKLASSKVLLGSFGGRDLVTWNTV 186
Query: 96 MTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLF 155
++ + LE L + +V P+E+ ++VL +CSH +R G + H Y
Sbjct: 187 LSSLCQNEQLLEALEYLREMVLEGVE----PDEFTISSVLPACSHLEMLRTGKELHAYAL 242
Query: 156 KYG-LVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWE 214
K G L FV SAL +MY C V +V + G + I L+N+++ ++ +
Sbjct: 243 KNGSLDENSFVGSALVDMYCNCKQVLSGRRVFD---GMFDRKIGLWNAMIAGYSQNEHDK 299
Query: 215 EAIGVLRRMGNEC-MVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKL 273
EA+ + M ++ ++ T GV+ C + G +H +++ GL D FV + L
Sbjct: 300 EALLLFIGMEESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTL 359
Query: 274 IDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTL--- 330
+DMY + + A ++F +++R++V W ++++ Y+ + + E+ L LL M +
Sbjct: 360 MDMYSRLGKIDIAMRIFGKMEDRDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSK 419
Query: 331 --------PNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSG 382
PN T +L +CA ++ L G +HA K V V +AL++MY+K G
Sbjct: 420 GASRVSLKPNSITLMTILPSCAALSALAKGKEIHAYAIKNNLATDVAVGSALVDMYAKCG 479
Query: 383 DIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLS 442
+ S VF ++V TWN +I Y HG GQEA+ + + M G PN VTF+ V +
Sbjct: 480 CLQMSRKVFDQIPQKNVITWNVIIMAYGMHGNGQEAIDLLRMMMVQGVKPNEVTFISVFA 539
Query: 443 ACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDI 502
AC+H +VD+GL Y ++ +EP +HY C+V L RAG +++A M ++
Sbjct: 540 ACSHSGMVDEGLRIFYVMKPDYGVEPSSDHYACVVDLLGRAGRIKEAYQLMNMMPRDFNK 599
Query: 503 V-AWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKM 561
AW +LL + R+H N ++G+ A++++Q++P Y LL+N++++A +WD E+R+
Sbjct: 600 AGAWSSLLGASRIHNNLEIGEIAAQNLIQLEPNVASHYVLLANIYSSAGLWDKATEVRRN 659
Query: 562 MKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAV 621
MKE+ ++KEPG SW+E + H FV+ HP+S +++ ++ L ++ GYVP+ S V
Sbjct: 660 MKEQGVRKEPGCSWIEHGDEVHKFVAGDSSHPQSEKLSGYLETLWERMRKEGYVPDTSCV 719
Query: 622 LHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNR 681
LH+VE+++KE L HSEKLAIA+G++ IR+ KNLR+C+DCH A K ISK+ +R
Sbjct: 720 LHNVEEDEKEILLCGHSEKLAIAFGILNTSPGTIIRVAKNLRVCNDCHLATKFISKIVDR 779
Query: 682 FIIVRDANRFHHFRDGSCTCADHW 705
II+RD RFH F++G+C+C D+W
Sbjct: 780 EIILRDVRRFHRFKNGTCSCGDYW 803
Score = 195 bits (495), Expect = 8e-50
Identities = 143/508 (28%), Positives = 249/508 (48%), Gaps = 28/508 (5%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
LLK AD + + GK IHA + V N+L++LY KCG +FD++
Sbjct: 16 LLKAVADLQDMELGKQIHAHVYKFGYGV--DSVTVANTLVNLYRKCGDFGAVYKVFDRIS 73
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSH---SG 142
R+ SWN L++ + L F+ ++ + P+ + +V+++CS+
Sbjct: 74 ERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVE----PSSFTLVSVVTACSNLPMPE 129
Query: 143 RVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNS 202
+ G Q H Y + G ++ F+ + L MY + L + +VL GG D+ +N+
Sbjct: 130 GLMMGKQVHAYGLRKGELN-SFIINTLVAMYGK-LGKLASSKVLLGSFGG--RDLVTWNT 185
Query: 203 LLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGG 262
+L+ L ++ + EA+ LR M E + D T V+ C+ + L+ G +HA L+ G
Sbjct: 186 VLSSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNG 245
Query: 263 LLFD-EFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLL 321
L + FVGS L+DMY C VL+ R+VFDG+ +R + +W ++++ Y QN + +E L L
Sbjct: 246 SLDENSFVGSALVDMYCNCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLF 305
Query: 322 TCMDQE-DTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSK 380
M++ L N T ++ AC + +H + K G V N L++MYS+
Sbjct: 306 IGMEESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSR 365
Query: 381 SGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMK------SAGES--- 431
G ID + +F RD+ TWN+MI GY ++AL + M+ S G S
Sbjct: 366 LGKIDIAMRIFGKMEDRDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVS 425
Query: 432 --PNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKA 489
PN++T + +L +CA L+ + KG + +KN+ + + + +V +Y + G L+ +
Sbjct: 426 LKPNSITLMTILPSCAALSALAKGKEIHAYAIKNN-LATDVAVGSALVDMYAKCGCLQMS 484
Query: 490 ENFMKTTKVKWDIVAWRTLLNSCRVHRN 517
K +++ W ++ + +H N
Sbjct: 485 RKVFDQIPQK-NVITWNVIIMAYGMHGN 511
Score = 147 bits (372), Expect = 2e-35
Identities = 121/452 (26%), Positives = 212/452 (46%), Gaps = 33/452 (7%)
Query: 126 PNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGL-VSYQFVKSALFNMYARCLHVNLALQ 184
P+ Y F +L + + + G Q H +++K+G V V + L N+Y +C +
Sbjct: 8 PDNYAFPALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDFGAVYK 67
Query: 185 VLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQ 244
V + S E + +NSL++ L +WE A+ R M +E + + T V V+ C+
Sbjct: 68 VFDRIS---ERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSN 124
Query: 245 I---GDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVW 301
+ L +G +VHA LR G L + F+ + L+ MYGK + +++ + R++V W
Sbjct: 125 LPMPEGLMMGKQVHAYGLRKGEL-NSFIINTLVAMYGKLGKLASSKVLLGSFGGRDLVTW 183
Query: 302 TSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEK 361
+++S+ QN E L+ L M E P+E T +L AC+ + +L+ G LHA K
Sbjct: 184 NTVLSSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALK 243
Query: 362 LG-FKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALR 420
G V +AL++MY + S VF R + WN+MI GYS + +EAL
Sbjct: 244 NGSLDENSFVGSALVDMYCNCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALL 303
Query: 421 VFQDM-KSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEH----YTC 475
+F M +SAG N+ T GV+ AC V G + ++ GL+
Sbjct: 304 LFIGMEESAGLLANSTTMAGVVPAC-----VRSGAFSRKEAIHGFVVKRGLDRDRFVQNT 358
Query: 476 MVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNS-----------CRVHRNYDLGKRI 524
++ +Y R G ++ A + + D+V W T++ +H+ +L +++
Sbjct: 359 LMDMYSRLGKIDIAMRIFGKMEDR-DLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKV 417
Query: 525 AE--SILQMDPQDMGTYTLLSNMHATANMWDG 554
++ S + + P + T+L + A + + G
Sbjct: 418 SKGASRVSLKPNSITLMTILPSCAALSALAKG 449
>At5g13230 putative protein
Length = 822
Score = 406 bits (1043), Expect = e-113
Identities = 230/644 (35%), Positives = 358/644 (54%), Gaps = 8/644 (1%)
Query: 63 SLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQ 122
+LI+ Y CG + AR++F+ + +D+ W +++ Y+ +G + + L L+S
Sbjct: 186 ALINAYSVCGSVDSARTVFEGILCKDIVVWAGIVSCYVENGYFEDSL----KLLSCMRMA 241
Query: 123 HACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLA 182
PN Y F T L + G HG + K V V L +Y + ++ A
Sbjct: 242 GFMPNNYTFDTALKASIGLGAFDFAKGVHGQILKTCYVLDPRVGVGLLQLYTQLGDMSDA 301
Query: 183 LQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLC 242
+V N +ND+ ++ ++ ++G EA+ + RM +V + T ++ C
Sbjct: 302 FKVFNEMP---KNDVVPWSFMIARFCQNGFCNEAVDLFIRMREAFVVPNEFTLSSILNGC 358
Query: 243 AQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWT 302
A LG ++H +++ G D +V + LID+Y KC + A K+F L ++N V W
Sbjct: 359 AIGKCSGLGEQLHGLVVKVGFDLDIYVSNALIDVYAKCEKMDTAVKLFAELSSKNEVSWN 418
Query: 303 SLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKL 362
+++ Y + + + E TF LGACA +A + G +H K
Sbjct: 419 TVIVGYENLGEGGKAFSMFREALRNQVSVTEVTFSSALGACASLASMDLGVQVHGLAIKT 478
Query: 363 GFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVF 422
V V+N+LI+MY+K GDI + +VF++ DV +WN++I GYS HGLG++ALR+
Sbjct: 479 NNAKKVAVSNSLIDMYAKCGDIKFAQSVFNEMETIDVASWNALISGYSTHGLGRQALRIL 538
Query: 423 QDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCR 482
MK PN +TF+GVLS C++ L+D+G + + +++H IEP LEHYTCMV L R
Sbjct: 539 DIMKDRDCKPNGLTFLGVLSGCSNAGLIDQGQECFESMIRDHGIEPCLEHYTCMVRLLGR 598
Query: 483 AGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLL 542
+G L+KA ++ + ++ WR +L++ N + +R AE IL+++P+D TY L+
Sbjct: 599 SGQLDKAMKLIEGIPYEPSVMIWRAMLSASMNQNNEEFARRSAEEILKINPKDEATYVLV 658
Query: 543 SNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKV 602
SNM+A A W V IRK MKE +KKEPG SW+E + H F HP+ IN +
Sbjct: 659 SNMYAGAKQWANVASIRKSMKEMGVKKEPGLSWIEHQGDVHYFSVGLSDHPDMKLINGML 718
Query: 603 QQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPA-PIRIIKN 661
+ L GYVP+ +AVL D++DE+K+ L HSE+LA+AYGL++MPS I I+KN
Sbjct: 719 EWLNMKATRAGYVPDRNAVLLDMDDEEKDKRLWVHSERLALAYGLVRMPSSRNRILIMKN 778
Query: 662 LRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
LR+C DCH+A+K+IS + R +++RD NRFHHF G C+C DHW
Sbjct: 779 LRICSDCHSAMKVISSIVQRDLVIRDMNRFHHFHAGVCSCGDHW 822
Score = 171 bits (434), Expect = 1e-42
Identities = 128/478 (26%), Positives = 230/478 (47%), Gaps = 16/478 (3%)
Query: 38 SGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMT 97
S KAIH +L + D+ N L++ YVK G + A ++FD+MP R+ S+ L
Sbjct: 67 SAKAIHCDILKKGSCL---DLFATNILLNAYVKAGFDKDALNLFDEMPERNNVSFVTLAQ 123
Query: 98 GYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKY 157
GY + + L+ L N +VFT+ L + H + K
Sbjct: 124 GYACQ----DPIGLYSRLHREGHEL----NPHVFTSFLKLFVSLDKAEICPWLHSPIVKL 175
Query: 158 GLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAI 217
G S FV +AL N Y+ C V+ A V G DI ++ +++ VE+G +E+++
Sbjct: 176 GYDSNAFVGAALINAYSVCGSVDSARTVF---EGILCKDIVVWAGIVSCYVENGYFEDSL 232
Query: 218 GVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMY 277
+L M + +N T+ + +G VH ++L+ + D VG L+ +Y
Sbjct: 233 KLLSCMRMAGFMPNNYTFDTALKASIGLGAFDFAKGVHGQILKTCYVLDPRVGVGLLQLY 292
Query: 278 GKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFK 337
+ + +A KVF+ + +VV W+ +++ + QN + E +DL M + +PNE T
Sbjct: 293 TQLGDMSDAFKVFNEMPKNDVVPWSFMIARFCQNGFCNEAVDLFIRMREAFVVPNEFTLS 352
Query: 338 VLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCR 397
+L CA G+ LH + K+GF + V+NALI++Y+K +D++ +F++ +
Sbjct: 353 SILNGCAIGKCSGLGEQLHGLVVKVGFDLDIYVSNALIDVYAKCEKMDTAVKLFAELSSK 412
Query: 398 DVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYL 457
+ +WN++I GY + G G +A +F++ S VTF L ACA LA +D G+
Sbjct: 413 NEVSWNTVIVGYENLGEGGKAFSMFREALRNQVSVTEVTFSSALGACASLASMDLGVQVH 472
Query: 458 YNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
+K + + + ++ +Y + G ++ A++ + D+ +W L++ H
Sbjct: 473 GLAIKTNNAKK-VAVSNSLIDMYAKCGDIKFAQSVFNEMET-IDVASWNALISGYSTH 528
Score = 125 bits (314), Expect = 8e-29
Identities = 82/319 (25%), Positives = 147/319 (45%), Gaps = 7/319 (2%)
Query: 127 NEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVL 186
+ + + +L C H + K G F + L N Y + AL +
Sbjct: 48 DSHAYGAMLRRCIQKNDPISAKAIHCDILKKGSCLDLFATNILLNAYVKAGFDKDALNLF 107
Query: 187 NTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIG 246
+ E N F+ L + ++ IG+ R+ E + + + L +
Sbjct: 108 D-EMPERNNVSFV------TLAQGYACQDPIGLYSRLHREGHELNPHVFTSFLKLFVSLD 160
Query: 247 DLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMS 306
++ +H+ +++ G + FVG+ LI+ Y C V +AR VF+G+ +++VVW ++S
Sbjct: 161 KAEICPWLHSPIVKLGYDSNAFVGAALINAYSVCGSVDSARTVFEGILCKDIVVWAGIVS 220
Query: 307 AYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKN 366
Y++N YFE++L LL+CM +PN TF L A G+ +H ++ K +
Sbjct: 221 CYVENGYFEDSLKLLSCMRMAGFMPNNYTFDTALKASIGLGAFDFAKGVHGQILKTCYVL 280
Query: 367 YVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMK 426
V L+ +Y++ GD+ ++ VF++ DV W+ MI + +G EA+ +F M+
Sbjct: 281 DPRVGVGLLQLYTQLGDMSDAFKVFNEMPKNDVVPWSFMIARFCQNGFCNEAVDLFIRMR 340
Query: 427 SAGESPNNVTFVGVLSACA 445
A PN T +L+ CA
Sbjct: 341 EAFVVPNEFTLSSILNGCA 359
Score = 99.4 bits (246), Expect = 6e-21
Identities = 86/374 (22%), Positives = 156/374 (40%), Gaps = 46/374 (12%)
Query: 231 DNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVF 290
D+ Y ++ C Q D +H +L+ G D F + L++ Y K +A +F
Sbjct: 48 DSHAYGAMLRRCIQKNDPISAKAIHCDILKKGSCLDLFATNILLNAYVKAGFDKDALNLF 107
Query: 291 DGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLK 350
D + RN V + +L Y ++ + L + + +E N F L + +
Sbjct: 108 DEMPERNNVSFVTLAQGYA----CQDPIGLYSRLHREGHELNPHVFTSFLKLFVSLDKAE 163
Query: 351 HGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYS 410
LH+ + KLG+ + V ALIN YS G +DS+ VF +C+D+ W ++ Y
Sbjct: 164 ICPWLHSPIVKLGYDSNAFVGAALINAYSVCGSVDSARTVFEGILCKDIVVWAGIVSCYV 223
Query: 411 HHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVD------------------- 451
+G +++L++ M+ AG PNN TF L A L D
Sbjct: 224 ENGYFEDSLKLLSCMRMAGFMPNNYTFDTALKASIGLGAFDFAKGVHGQILKTCYVLDPR 283
Query: 452 --KGLDYLY-------------NEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENF---M 493
GL LY NEM + + P ++ M+ +C+ G +A + M
Sbjct: 284 VGVGLLQLYTQLGDMSDAFKVFNEMPKNDVVP----WSFMIARFCQNGFCNEAVDLFIRM 339
Query: 494 KTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMD-PQDMGTYTLLSNMHATANMW 552
+ V + ++LN C + + LG+++ ++++ D+ L +++A
Sbjct: 340 REAFVVPNEFTLSSILNGCAIGKCSGLGEQLHGLVVKVGFDLDIYVSNALIDVYAKCEKM 399
Query: 553 DGVVEIRKMMKERN 566
D V++ + +N
Sbjct: 400 DTAVKLFAELSSKN 413
Score = 85.1 bits (209), Expect = 1e-16
Identities = 65/295 (22%), Positives = 139/295 (47%), Gaps = 13/295 (4%)
Query: 16 YLPPLEYIVK-LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRL 74
++ P E+ + +L A KC G+ +H ++ + D+ N+LI +Y KC ++
Sbjct: 343 FVVPNEFTLSSILNGCAIGKCSGLGEQLHGLVV---KVGFDLDIYVSNALIDVYAKCEKM 399
Query: 75 RLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTV 134
A +F ++ ++ SWN ++ GY + G + +F+ + + + E F++
Sbjct: 400 DTAVKLFAELSSKNEVSWNTVIVGYENLGEGGKAFSMFREALRNQVS----VTEVTFSSA 455
Query: 135 LSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDE 194
L +C+ + G+Q HG K V ++L +MYA+C + A V N +
Sbjct: 456 LGACASLASMDLGVQVHGLAIKTNNAKKVAVSNSLIDMYAKCGDIKFAQSVFNEM---ET 512
Query: 195 NDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRV 254
D+ +N+L++ G +A+ +L M + + T++GV+ C+ G + G
Sbjct: 513 IDVASWNALISGYSTHGLGRQALRILDIMKDRDCKPNGLTFLGVLSGCSNAGLIDQGQEC 572
Query: 255 HARLLRG-GLLFDEFVGSKLIDMYGKCCHVLNARKVFDGL-QNRNVVVWTSLMSA 307
++R G+ + ++ + G+ + A K+ +G+ +V++W +++SA
Sbjct: 573 FESMIRDHGIEPCLEHYTCMVRLLGRSGQLDKAMKLIEGIPYEPSVMIWRAMLSA 627
Score = 70.5 bits (171), Expect = 3e-12
Identities = 58/212 (27%), Positives = 101/212 (47%), Gaps = 23/212 (10%)
Query: 39 GKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTG 98
G +H + + A K V NSLI +Y KCG ++ A+S+F++M DV SWN L++G
Sbjct: 468 GVQVHGLAIKTNNAKK---VAVSNSLIDMYAKCGDIKFAQSVFNEMETIDVASWNALISG 524
Query: 99 YMHSG---NYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLF 155
Y G L +L + K+ + PN F VLS CS++G + +G +C +
Sbjct: 525 YSTHGLGRQALRILDIMKD-------RDCKPNGLTFLGVLSGCSNAGLIDQGQECFESMI 577
Query: 156 K-YGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWE 214
+ +G+ + + + R ++ A++++ E E + ++ ++L+ + E
Sbjct: 578 RDHGIEPCLEHYTCMVRLLGRSGQLDKAMKLI--EGIPYEPSVMIWRAMLSASMNQNNEE 635
Query: 215 EAIGVLRRMGNECM---VWDNATYVGVMGLCA 243
A RR E + D ATYV V + A
Sbjct: 636 FA----RRSAEEILKINPKDEATYVLVSNMYA 663
>At2g03880 putative selenium-binding protein
Length = 630
Score = 404 bits (1039), Expect = e-113
Identities = 215/575 (37%), Positives = 336/575 (58%), Gaps = 8/575 (1%)
Query: 131 FTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTES 190
++ ++ C + V EG +L+ G F+ + L NMY + +N A Q+ +
Sbjct: 64 YSELIKCCISNRAVHEGNLICRHLYFNGHRPMMFLVNVLINMYVKFNLLNDAHQLFDQMP 123
Query: 191 GGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQL 250
+ ++ + ++++ + ++A+ +L M + + + TY V+ C + D+++
Sbjct: 124 ---QRNVISWTTMISAYSKCKIHQKALELLVLMLRDNVRPNVYTYSSVLRSCNGMSDVRM 180
Query: 251 GLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQ 310
+H +++ GL D FV S LID++ K +A VFD + + +VW S++ + Q
Sbjct: 181 ---LHCGIIKEGLESDVFVRSALIDVFAKLGEPEDALSVFDEMVTGDAIVWNSIIGGFAQ 237
Query: 311 NRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIV 370
N + L+L M + + + T +L AC G+ALL+ G H + K + +I+
Sbjct: 238 NSRSDVALELFKRMKRAGFIAEQATLTSVLRACTGLALLELGMQAHVHIVK--YDQDLIL 295
Query: 371 NNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGE 430
NNAL++MY K G ++ + VF+ RDV TW++MI G + +G QEAL++F+ MKS+G
Sbjct: 296 NNALVDMYCKCGSLEDALRVFNQMKERDVITWSTMISGLAQNGYSQEALKLFERMKSSGT 355
Query: 431 SPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAE 490
PN +T VGVL AC+H L++ G Y + K + I+P EHY CM+ L +AG L+ A
Sbjct: 356 KPNYITIVGVLFACSHAGLLEDGWYYFRSMKKLYGIDPVREHYGCMIDLLGKAGKLDDAV 415
Query: 491 NFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATAN 550
+ + + D V WRTLL +CRV RN L + A+ ++ +DP+D GTYTLLSN++A +
Sbjct: 416 KLLNEMECEPDAVTWRTLLGACRVQRNMVLAEYAAKKVIALDPEDAGTYTLLSNIYANSQ 475
Query: 551 MWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIK 610
WD V EIR M++R IKKEPG SW+E+ H F+ HP+ E++KK+ QL+ +
Sbjct: 476 KWDSVEEIRTRMRDRGIKKEPGCSWIEVNKQIHAFIIGDNSHPQIVEVSKKLNQLIHRLT 535
Query: 611 PLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHT 670
+GYVP + VL D+E EQ E L HSEKLA+A+GLM +P IRI KNLR+C DCH
Sbjct: 536 GIGYVPETNFVLQDLEGEQMEDSLRHHSEKLALAFGLMTLPIEKVIRIRKNLRICGDCHV 595
Query: 671 AVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
KL SK+ R I++RD R+HHF+DG C+C D+W
Sbjct: 596 FCKLASKLEIRSIVIRDPIRYHHFQDGKCSCGDYW 630
Score = 159 bits (403), Expect = 4e-39
Identities = 107/378 (28%), Positives = 189/378 (49%), Gaps = 17/378 (4%)
Query: 34 KCLTSGKAIHAQLLVRDQA---SKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVF 90
KC S +A+H L+ R + +N LI++YVK L A +FD+MP R+V
Sbjct: 69 KCCISNRAVHEGNLICRHLYFNGHRPMMFLVNVLINMYVKFNLLNDAHQLFDQMPQRNVI 128
Query: 91 SWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQC 150
SW +++ Y + + L L ++ + PN Y +++VL SC+ VR
Sbjct: 129 SWTTMISAYSKCKIHQKALELLVLMLRDNVR----PNVYTYSSVLRSCNGMSDVR---ML 181
Query: 151 HGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVES 210
H + K GL S FV+SAL +++A+ AL V + G D ++NS++ ++
Sbjct: 182 HCGIIKEGLESDVFVRSALIDVFAKLGEPEDALSVFDEMVTG---DAIVWNSIIGGFAQN 238
Query: 211 GRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVG 270
R + A+ + +RM + + AT V+ C + L+LG++ H +++ D +
Sbjct: 239 SRSDVALELFKRMKRAGFIAEQATLTSVLRACTGLALLELGMQAHVHIVKYDQ--DLILN 296
Query: 271 SKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTL 330
+ L+DMY KC + +A +VF+ ++ R+V+ W++++S QN Y +E L L M T
Sbjct: 297 NALVDMYCKCGSLEDALRVFNQMKERDVITWSTMISGLAQNGYSQEALKLFERMKSSGTK 356
Query: 331 PNECTFKVLLGACAGIALLKHGDLLHARLEKL-GFKNYVIVNNALINMYSKSGDIDSSYN 389
PN T +L AC+ LL+ G ++KL G +I++ K+G +D +
Sbjct: 357 PNYITIVGVLFACSHAGLLEDGWYYFRSMKKLYGIDPVREHYGCMIDLLGKAGKLDDAVK 416
Query: 390 VFSDTMCR-DVTTWNSMI 406
+ ++ C D TW +++
Sbjct: 417 LLNEMECEPDAVTWRTLL 434
Score = 69.3 bits (168), Expect = 7e-12
Identities = 45/166 (27%), Positives = 82/166 (49%), Gaps = 10/166 (6%)
Query: 23 IVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFD 82
+ +L+ L G H ++ DQ D++ N+L+ +Y KCG L A +F+
Sbjct: 263 LTSVLRACTGLALLELGMQAHVHIVKYDQ-----DLILNNALVDMYCKCGSLEDALRVFN 317
Query: 83 KMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSG 142
+M RDV +W+ +++G +G E L LF+ + SS T PN VL +CSH+G
Sbjct: 318 QMKERDVITWSTMISGLAQNGYSQEALKLFERMKSSGTK----PNYITIVGVLFACSHAG 373
Query: 143 RVREGMQCHGYLFK-YGLVSYQFVKSALFNMYARCLHVNLALQVLN 187
+ +G + K YG+ + + ++ + ++ A+++LN
Sbjct: 374 LLEDGWYYFRSMKKLYGIDPVREHYGCMIDLLGKAGKLDDAVKLLN 419
Score = 57.0 bits (136), Expect = 4e-08
Identities = 41/165 (24%), Positives = 73/165 (43%), Gaps = 11/165 (6%)
Query: 302 TSLMSAYLQNRYFEETLDLLTCMDQEDT---LPNECTFKVLLGACAGIALLKHGDLLHAR 358
T L+S + + Y + + MD + + T+ L+ C + G+L+
Sbjct: 27 TLLLSEFTRLCYQRDLPRAMKAMDSLQSHGLWADSATYSELIKCCISNRAVHEGNLICRH 86
Query: 359 LEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEA 418
L G + + + N LINMY K ++ ++ +F R+V +W +MI YS + Q+A
Sbjct: 87 LYFNGHRPMMFLVNVLINMYVKFNLLNDAHQLFDQMPQRNVISWTTMISAYSKCKIHQKA 146
Query: 419 LRVFQDMKSAGESPNNVTFVGVLSAC--------AHLALVDKGLD 455
L + M PN T+ VL +C H ++ +GL+
Sbjct: 147 LELLVLMLRDNVRPNVYTYSSVLRSCNGMSDVRMLHCGIIKEGLE 191
>At5g65570 putative protein
Length = 738
Score = 396 bits (1017), Expect = e-110
Identities = 238/685 (34%), Positives = 379/685 (54%), Gaps = 20/685 (2%)
Query: 25 KLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKM 84
+LL+ D + ++ K I A +L ++ S + L+ +KCG + AR +FD M
Sbjct: 70 QLLRQCIDERSISGIKTIQAHMLKSGFPAEISG----SKLVDASLKCGDIDYARQVFDGM 125
Query: 85 PLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRV 144
R + +WN L+ + E + +++ ++ T + P+EY ++V + S
Sbjct: 126 SERHIVTWNSLIAYLIKHRRSKEAVEMYRLMI----TNNVLPDEYTLSSVFKAFSDLSLE 181
Query: 145 REGMQCHGYLFKYGL-VSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSL 203
+E + HG GL VS FV SAL +MY + A VL+ +E D+ L +L
Sbjct: 182 KEAQRSHGLAVILGLEVSNVFVGSALVDMYVKFGKTREAKLVLDRV---EEKDVVLITAL 238
Query: 204 LNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGL 263
+ + G EA+ + M E + + TY V+ C + D+ G +H +++ G
Sbjct: 239 IVGYSQKGEDTEAVKAFQSMLVEKVQPNEYTYASVLISCGNLKDIGNGKLIHGLMVKSG- 297
Query: 264 LFDEFVGSK--LIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLL 321
F+ + S+ L+ MY +C V ++ +VF ++ N V WTSL+S +QN E L
Sbjct: 298 -FESALASQTSLLTMYLRCSLVDDSLRVFKCIEYPNQVSWTSLISGLVQNGREEMALIEF 356
Query: 322 TCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKS 381
M ++ PN T L C+ +A+ + G +H + K GF + LI++Y K
Sbjct: 357 RKMMRDSIKPNSFTLSSALRGCSNLAMFEEGRQIHGIVTKYGFDRDKYAGSGLIDLYGKC 416
Query: 382 GDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVL 441
G D + VF DV + N+MI Y+ +G G+EAL +F+ M + G PN+VT + VL
Sbjct: 417 GCSDMARLVFDTLSEVDVISLNTMIYSYAQNGFGREALDLFERMINLGLQPNDVTVLSVL 476
Query: 442 SACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWD 501
AC + LV++G + L++ + KI +HY CMV L RAG LE+AE + T + D
Sbjct: 477 LACNNSRLVEEGCE-LFDSFRKDKIMLTNDHYACMVDLLGRAGRLEEAE-MLTTEVINPD 534
Query: 502 IVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKM 561
+V WRTLL++C+VHR ++ +RI IL+++P D GT L+SN++A+ W+ V+E++
Sbjct: 535 LVLWRTLLSACKVHRKVEMAERITRKILEIEPGDEGTLILMSNLYASTGKWNRVIEMKSK 594
Query: 562 MKERNIKKEPGASWLEIRNVNHVFVS-EGCKHPESNEINKKVQQLLAMIKPLGYVPNISA 620
MK+ +KK P SW+EI H F++ + HP S +I + +++L+ K LGYV + S
Sbjct: 595 MKDMKLKKNPAMSWVEINKETHTFMAGDLFSHPNSEQILENLEELIKKSKDLGYVEDKSC 654
Query: 621 VLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTN 680
V D+E+ KE L+ HSEKLAIA+ + + IRI+KNLR+C DCH+ +K++S+V
Sbjct: 655 VFQDMEETAKERSLHQHSEKLAIAFAVWRNVG-GSIRILKNLRVCVDCHSWIKIVSRVMK 713
Query: 681 RFIIVRDANRFHHFRDGSCTCADHW 705
R II RD+ RFHHFRDGSC+C D+W
Sbjct: 714 REIICRDSKRFHHFRDGSCSCGDYW 738
>At4g18750 putative protein
Length = 871
Score = 392 bits (1008), Expect = e-109
Identities = 222/680 (32%), Positives = 366/680 (53%), Gaps = 13/680 (1%)
Query: 28 KLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLR 87
K + + + G+ +H +L + +R+ V NSL+ Y+K R+ AR +FD+M R
Sbjct: 203 KSFSSLRSVHGGEQLHGFIL-KSGFGERNSVG--NSLVAFYLKNQRVDSARKVFDEMTER 259
Query: 88 DVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREG 147
DV SWN ++ GY+ +G + L +F ++ S +V + C+ S + G
Sbjct: 260 DVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLA----TIVSVFAGCADSRLISLG 315
Query: 148 MQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVL 207
H K + L +MY++C ++ A V S + + Y S++
Sbjct: 316 RAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAVFREMS---DRSVVSYTSMIAGY 372
Query: 208 VESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDE 267
G EA+ + M E + D T V+ CA+ L G RVH + L FD
Sbjct: 373 AREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFDI 432
Query: 268 FVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQE 327
FV + L+DMY KC + A VF ++ ++++ W +++ Y +N Y E L L + +E
Sbjct: 433 FVSNALMDMYAKCGSMQEAELVFSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLEE 492
Query: 328 DTL-PNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDS 386
P+E T +L ACA ++ G +H + + G+ + V N+L++MY+K G +
Sbjct: 493 KRFSPDERTVACVLPACASLSAFDKGREIHGYIMRNGYFSDRHVANSLVDMYAKCGALLL 552
Query: 387 SYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAH 446
++ +F D +D+ +W MI GY HG G+EA+ +F M+ AG + ++FV +L AC+H
Sbjct: 553 AHMLFDDIASKDLVSWTVMIAGYGMHGFGKEAIALFNQMRQAGIEADEISFVSLLYACSH 612
Query: 447 LALVDKGLDYLYNEMKNH-KIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAW 505
LVD+G + +N M++ KIEP +EHY C+V + R G L KA F++ + D W
Sbjct: 613 SGLVDEGWRF-FNIMRHECKIEPTVEHYACIVDMLARTGDLIKAYRFIENMPIPPDATIW 671
Query: 506 RTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKER 565
LL CR+H + L +++AE + +++P++ G Y L++N++A A W+ V +RK + +R
Sbjct: 672 GALLCGCRIHHDVKLAEKVAEKVFELEPENTGYYVLMANIYAEAEKWEQVKRLRKRIGQR 731
Query: 566 NIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDV 625
++K PG SW+EI+ ++FV+ +PE+ I ++++ A + GY P L D
Sbjct: 732 GLRKNPGCSWIEIKGRVNIFVAGDSSNPETENIEAFLRKVRARMIEEGYSPLTKYALIDA 791
Query: 626 EDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIV 685
E+ +KE L HSEKLA+A G++ IR+ KNLR+C DCH K +SK+T R I++
Sbjct: 792 EEMEKEEALCGHSEKLAMALGIISSGHGKIIRVTKNLRVCGDCHEMAKFMSKLTRREIVL 851
Query: 686 RDANRFHHFRDGSCTCADHW 705
RD+NRFH F+DG C+C W
Sbjct: 852 RDSNRFHQFKDGHCSCRGFW 871
Score = 191 bits (484), Expect = 2e-48
Identities = 147/500 (29%), Positives = 242/500 (48%), Gaps = 25/500 (5%)
Query: 23 IVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLN---SLIHLYVKCGRLRLARS 79
+ +L+L AD+K L GK + +R V+ N L +Y CG L+ A
Sbjct: 97 LCSVLQLCADSKSLKDGKEVDN--FIRGNGF----VIDSNLGSKLSLMYTNCGDLKEASR 150
Query: 80 MFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCS 139
+FD++ + WN LM SG++ + LFK ++SS + Y F+ V S S
Sbjct: 151 VFDEVKIEKALFWNILMNELAKSGDFSGSIGLFKKMMSSGVEM----DSYTFSCVSKSFS 206
Query: 140 HSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFL 199
V G Q HG++ K G V ++L Y + V+ A +V + + E D+
Sbjct: 207 SLRSVHGGEQLHGFILKSGFGERNSVGNSLVAFYLKNQRVDSARKVFDEMT---ERDVIS 263
Query: 200 YNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLL 259
+NS++N V +G E+ + V +M + D AT V V CA + LG VH+ +
Sbjct: 264 WNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLATIVSVFAGCADSRLISLGRAVHSIGV 323
Query: 260 RGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLD 319
+ ++ + L+DMY KC + +A+ VF + +R+VV +TS+++ Y + E +
Sbjct: 324 KACFSREDRFCNTLLDMYSKCGDLDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVK 383
Query: 320 LLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEK--LGFKNYVIVNNALINM 377
L M++E P+ T +L CA LL G +H +++ LGF + V+NAL++M
Sbjct: 384 LFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFD--IFVSNALMDM 441
Query: 378 YSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQ-DMKSAGESPNNVT 436
Y+K G + + VFS+ +D+ +WN++I GYS + EAL +F ++ SP+ T
Sbjct: 442 YAKCGSMQEAELVFSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERT 501
Query: 437 FVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHY-TCMVVLYCRAGLLEKAENFMKT 495
VL ACA L+ DKG + M+N H +V +Y + G L A
Sbjct: 502 VACVLPACASLSAFDKGREIHGYIMRNGYFSD--RHVANSLVDMYAKCGALLLAHMLFDD 559
Query: 496 TKVKWDIVAWRTLLNSCRVH 515
K D+V+W ++ +H
Sbjct: 560 IASK-DLVSWTVMIAGYGMH 578
Score = 143 bits (360), Expect = 4e-34
Identities = 103/396 (26%), Positives = 190/396 (47%), Gaps = 13/396 (3%)
Query: 20 LEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARS 79
L IV + AD++ ++ G+A+H+ + +A + N+L+ +Y KCG L A++
Sbjct: 296 LATIVSVFAGCADSRLISLGRAVHS---IGVKACFSREDRFCNTLLDMYSKCGDLDSAKA 352
Query: 80 MFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCS 139
+F +M R V S+ ++ GY G E + LF+ + + P+ Y T VL+ C+
Sbjct: 353 VFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGIS----PDVYTVTAVLNCCA 408
Query: 140 HSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFL 199
+ EG + H ++ + L FV +AL +MYA+C + A V S DI
Sbjct: 409 RYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQEAELVF---SEMRVKDIIS 465
Query: 200 YNSLLNVLVESGRWEEAIGVLRRMGNECMVW-DNATYVGVMGLCAQIGDLQLGLRVHARL 258
+N+++ ++ EA+ + + E D T V+ CA + G +H +
Sbjct: 466 WNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGYI 525
Query: 259 LRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETL 318
+R G D V + L+DMY KC +L A +FD + ++++V WT +++ Y + + +E +
Sbjct: 526 MRNGYFSDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEAI 585
Query: 319 DLLTCMDQEDTLPNECTFKVLLGACAGIALLKHG-DLLHARLEKLGFKNYVIVNNALINM 377
L M Q +E +F LL AC+ L+ G + + + V +++M
Sbjct: 586 ALFNQMRQAGIEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVDM 645
Query: 378 YSKSGDIDSSYNVFSD-TMCRDVTTWNSMICGYSHH 412
+++GD+ +Y + + D T W +++CG H
Sbjct: 646 LARTGDLIKAYRFIENMPIPPDATIWGALLCGCRIH 681
Score = 124 bits (310), Expect = 2e-28
Identities = 106/383 (27%), Positives = 162/383 (41%), Gaps = 50/383 (13%)
Query: 201 NSLLNVLVESGRWEEAIGVLRRMGNECMVWD--NATYVGVMGLCAQIGDLQLGLRVHARL 258
N+ L ESG E A+ +L G WD T V+ LCA L+ G V +
Sbjct: 65 NTQLRRFCESGNLENAVKLLCVSGK----WDIDPRTLCSVLQLCADSKSLKDGKEVDNFI 120
Query: 259 LRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETL 318
G + D +GSKL MY C + A +VFD ++ + W LM+ ++ F ++
Sbjct: 121 RGNGFVIDSNLGSKLSLMYTNCGDLKEASRVFDEVKIEKALFWNILMNELAKSGDFSGSI 180
Query: 319 DLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMY 378
L M + TF + + + + + G+ LH + K GF V N+L+ Y
Sbjct: 181 GLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGFGERNSVGNSLVAFY 240
Query: 379 SKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFV 438
K+ +DS+ VF + RDV +WNS+I GY +GL ++ L VF M +G + T V
Sbjct: 241 LKNQRVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLATIV 300
Query: 439 GVLSACAHLALVDKG----------------------LDY------------LYNEMKNH 464
V + CA L+ G LD ++ EM +
Sbjct: 301 SVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAVFREMSDR 360
Query: 465 KIEPGLEHYTCMVVLYCRAGLLEKAENF---MKTTKVKWDIVAWRTLLNSCRVHRNYDLG 521
+ YT M+ Y R GL +A M+ + D+ +LN C +R D G
Sbjct: 361 SV----VSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEG 416
Query: 522 KRIAESILQMDPQDMGTYTLLSN 544
KR+ E I + D+G +SN
Sbjct: 417 KRVHEWIKE---NDLGFDIFVSN 436
>At3g15130 hypothetical protein
Length = 689
Score = 391 bits (1004), Expect = e-109
Identities = 220/691 (31%), Positives = 370/691 (52%), Gaps = 18/691 (2%)
Query: 23 IVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFD 82
+V +L++ G +H LL ++ +++ N LI +Y KC +A +FD
Sbjct: 9 LVSILRVCTRKGLSDQGGQVHCYLL---KSGSGLNLITSNYLIDMYCKCREPLMAYKVFD 65
Query: 83 KMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSG 142
MP R+V SW+ LM+G++ +G+ L LF S Q PNE+ F+T L +C
Sbjct: 66 SMPERNVVSWSALMSGHVLNGDLKGSLSLF----SEMGRQGIYPNEFTFSTNLKACGLLN 121
Query: 143 RVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNS 202
+ +G+Q HG+ K G V ++L +MY++C +N A +V + + +N+
Sbjct: 122 ALEKGLQIHGFCLKIGFEMMVEVGNSLVDMYSKCGRINEAEKVFRRIV---DRSLISWNA 178
Query: 203 LLNVLVESGRWEEAIGVLRRM--GNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLR 260
++ V +G +A+ M N D T ++ C+ G + G ++H L+R
Sbjct: 179 MIAGFVHAGYGSKALDTFGMMQEANIKERPDEFTLTSLLKACSSTGMIYAGKQIHGFLVR 238
Query: 261 GGLLFDEF--VGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETL 318
G + L+D+Y KC ++ +ARK FD ++ + ++ W+SL+ Y Q F E +
Sbjct: 239 SGFHCPSSATITGSLVDLYVKCGYLFSARKAFDQIKEKTMISWSSLILGYAQEGEFVEAM 298
Query: 319 DLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMY 378
L + + ++ + ++G A ALL+ G + A KL V N++++MY
Sbjct: 299 GLFKRLQELNSQIDSFALSSIIGVFADFALLRQGKQMQALAVKLPSGLETSVLNSVVDMY 358
Query: 379 SKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFV 438
K G +D + F++ +DV +W +I GY HGLG++++R+F +M P+ V ++
Sbjct: 359 LKCGLVDEAEKCFAEMQLKDVISWTVVITGYGKHGLGKKSVRIFYEMLRHNIEPDEVCYL 418
Query: 439 GVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKV 498
VLSAC+H ++ +G + ++ H I+P +EHY C+V L RAG L++A++ + T +
Sbjct: 419 AVLSACSHSGMIKEGEELFSKLLETHGIKPRVEHYACVVDLLGRAGRLKEAKHLIDTMPI 478
Query: 499 KWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEI 558
K ++ W+TLL+ CRVH + +LGK + + +L++D ++ Y ++SN++ A W+
Sbjct: 479 KPNVGIWQTLLSLCRVHGDIELGKEVGKILLRIDAKNPANYVMMSNLYGQAGYWNEQGNA 538
Query: 559 RKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIK-PLGYVPN 617
R++ + +KKE G SW+EI H F S HP + I + +++ ++ LGYV
Sbjct: 539 RELGNIKGLKKEAGMSWVEIEREVHFFRSGEDSHPLTPVIQETLKEAERRLREELGYVYG 598
Query: 618 ISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMK---MPSPAPIRIIKNLRMCDDCHTAVKL 674
+ LHD++DE KE L HSEKLAI L IR+ KNLR+C DCH +K
Sbjct: 599 LKHELHDIDDESKEENLRAHSEKLAIGLALATGGLNQKGKTIRVFKNLRVCVDCHEFIKG 658
Query: 675 ISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
+SK+T +VRDA RFH F DG C+C D+W
Sbjct: 659 LSKITKIAYVVRDAVRFHSFEDGCCSCGDYW 689
Score = 84.7 bits (208), Expect = 2e-16
Identities = 70/302 (23%), Positives = 137/302 (45%), Gaps = 10/302 (3%)
Query: 7 GPMVVVNSRYLPPLEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIH 66
G M N + P + LLK + + +GK IH LVR S SL+
Sbjct: 197 GMMQEANIKERPDEFTLTSLLKACSSTGMIYAGKQIHG-FLVRSGFHCPSSATITGSLVD 255
Query: 67 LYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACP 126
LYVKCG L AR FD++ + + SW+ L+ GY G ++E + LFK L ++
Sbjct: 256 LYVKCGYLFSARKAFDQIKEKTMISWSSLILGYAQEGEFVEAMGLFKRLQELNSQ----I 311
Query: 127 NEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVL 186
+ + ++++ + +R+G Q K V +++ +MY +C V+ A +
Sbjct: 312 DSFALSSIIGVFADFALLRQGKQMQALAVKLPSGLETSVLNSVVDMYLKCGLVDEAEKCF 371
Query: 187 NTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIG 246
+ D+ + ++ + G ++++ + M + D Y+ V+ C+ G
Sbjct: 372 ---AEMQLKDVISWTVVITGYGKHGLGKKSVRIFYEMLRHNIEPDEVCYLAVLSACSHSG 428
Query: 247 DLQLGLRVHARLLR-GGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNR-NVVVWTSL 304
++ G + ++LL G+ + ++D+ G+ + A+ + D + + NV +W +L
Sbjct: 429 MIKEGEELFSKLLETHGIKPRVEHYACVVDLLGRAGRLKEAKHLIDTMPIKPNVGIWQTL 488
Query: 305 MS 306
+S
Sbjct: 489 LS 490
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,747,442
Number of Sequences: 26719
Number of extensions: 675506
Number of successful extensions: 9375
Number of sequences better than 10.0: 444
Number of HSP's better than 10.0 without gapping: 395
Number of HSP's successfully gapped in prelim test: 50
Number of HSP's that attempted gapping in prelim test: 1522
Number of HSP's gapped (non-prelim): 2145
length of query: 705
length of database: 11,318,596
effective HSP length: 106
effective length of query: 599
effective length of database: 8,486,382
effective search space: 5083342818
effective search space used: 5083342818
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0141.1


