

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.7
(571 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
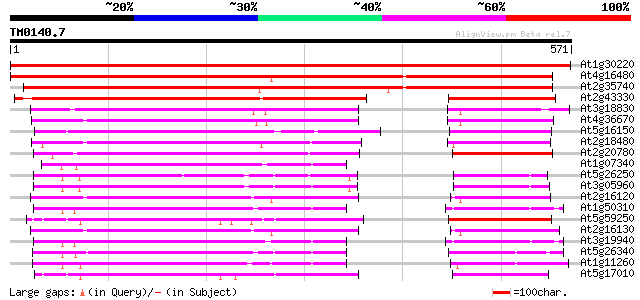
Score E
Sequences producing significant alignments: (bits) Value
At1g30220 unknown protein 858 0.0
At4g16480 membrane transporter like protein 685 0.0
At2g35740 putative sugar transporter 636 0.0
At2g43330 membrane transporter like protein 397 e-111
At3g18830 sugar transporter like protein 207 1e-53
At4g36670 sugar transporter like protein 202 5e-52
At5g16150 sugar transporter like protein 196 3e-50
At2g18480 putative sugar transporter 196 4e-50
At2g20780 putative sugar transporter 183 3e-46
At1g07340 hexose transporter like protein 182 6e-46
At5g26250 hexose transporter - like protein 179 5e-45
At3g05960 putative hexose transporter 178 6e-45
At2g16120 putative sugar transporter 178 6e-45
At1g50310 hexose transporter, putative 177 1e-44
At5g59250 D-xylose-H+ symporter - like protein 175 5e-44
At2g16130 putative sugar transporter 174 2e-43
At3g19940 putative monosaccharide transport protein, STP4 173 3e-43
At5g26340 hexose transporter - like protein 167 1e-41
At1g11260 glucose transporter 167 1e-41
At5g17010 sugar transporter - like protein 166 2e-41
>At1g30220 unknown protein
Length = 580
Score = 858 bits (2218), Expect = 0.0
Identities = 412/575 (71%), Positives = 497/575 (85%), Gaps = 5/575 (0%)
Query: 1 MEGGVPE--ADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDF 58
MEGG+ AD SAF+EC SL+WKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDF
Sbjct: 1 MEGGIIHGGADESAFKECFSLTWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDF 60
Query: 59 KAVDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPN 118
K+VDR TWLQE IVS A+AGAI+GAA+GGW ND+ GRR AI++AD LFL+G++IMAAAPN
Sbjct: 61 KSVDRNTWLQEMIVSMAVAGAIVGAAIGGWANDKLGRRSAILMADFLFLLGAIIMAAAPN 120
Query: 119 PAILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAF 178
P++L+ GRVFVG GVGMASM +PLYISEASP ++RGALVS N FLITGGQFLSYLINLAF
Sbjct: 121 PSLLVVGRVFVGLGVGMASMTAPLYISEASPAKIRGALVSTNGFLITGGQFLSYLINLAF 180
Query: 179 TNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAE 238
T+ GTWRWMLG+A PA++Q VLM +LPESPRWLYRKGREEE+K+IL++IY+ EDV+ E
Sbjct: 181 TDVTGTWRWMLGIAGIPALLQFVLMFTLPESPRWLYRKGREEEAKAILRRIYSAEDVEQE 240
Query: 239 IEALKESVESEI-EESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPT 297
I ALK+SVE+EI EE + KI+M+ L K TVRRGL AG+GLQ FQQFVGINTVMYYSPT
Sbjct: 241 IRALKDSVETEILEEGSSEKINMIKLCKAKTVRRGLIAGVGLQVFQQFVGINTVMYYSPT 300
Query: 298 IVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTV 357
IVQLAGFASNRTALLLSL+T+GLNAFGSI+SIYFID+ GRKKL +ISL GV++SL +LT
Sbjct: 301 IVQLAGFASNRTALLLSLVTAGLNAFGSIISIYFIDRIGRKKLLIISLFGVIISLGILTG 360
Query: 358 AFRESELHAPTVSAIETSHYNN-TCPDFRAAVNPGRWTCMTCLKA-SPSCGFCAASPNKL 415
F E+ HAP +S++ET +NN +CPD+++A+N W CMTCLKA SPSCG+C++ K
Sbjct: 361 VFYEAATHAPAISSLETQRFNNISCPDYKSAMNTNAWDCMTCLKASSPSCGYCSSPIGKE 420
Query: 416 LPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINS 475
PGAC ISDD+ KD+C N++R WYTRGCPS FGW AL+GL LYII FSPGMGTVPW++NS
Sbjct: 421 HPGACWISDDSVKDLCHNENRLWYTRGCPSNFGWFALLGLGLYIIFFSPGMGTVPWIVNS 480
Query: 476 EIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVF 535
EIYPLR+RG+CGGIA+T WISNLIV+QSFLSLT+ IGT+WTF++FG+++V+A+ FV+V
Sbjct: 481 EIYPLRFRGICGGIAATANWISNLIVAQSFLSLTEAIGTSWTFLIFGVISVIALLFVMVC 540
Query: 536 VPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEK 570
VPETKG+PMEE+EKMLE+RS++FKFW+K+ EK
Sbjct: 541 VPETKGMPMEEIEKMLERRSMEFKFWKKKSKLVEK 575
>At4g16480 membrane transporter like protein
Length = 582
Score = 685 bits (1767), Expect = 0.0
Identities = 337/562 (59%), Positives = 429/562 (75%), Gaps = 12/562 (2%)
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
+EGG+ +AD + F EC +WK PY++RLA SAGIGGLLFGYDTGVISGALL+I++DF
Sbjct: 2 VEGGIAKADKTEFTECWRTTWKTPYIMRLALSAGIGGLLFGYDTGVISGALLFIKEDFDE 61
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
VD+KTWLQ IVS A+AGAI+GAAVGGWIND+FGRR++I+IAD LFLIG+++MA AP P
Sbjct: 62 VDKKTWLQSTIVSMAVAGAIVGAAVGGWINDKFGRRMSILIADVLFLIGAIVMAFAPAPW 121
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
+++ GR+FVGFGVGMASM SPLYISEASP R+RGALVS N LITGGQF SYLINLAF +
Sbjct: 122 VIIVGRIFVGFGVGMASMTSPLYISEASPARIRGALVSTNGLLITGGQFFSYLINLAFVH 181
Query: 181 APGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIE 240
PGTWRWMLGVA PAI+Q VLMLSLPESPRWLYRK R ES++IL++IY ++V+AE+E
Sbjct: 182 TPGTWRWMLGVAGVPAIVQFVLMLSLPESPRWLYRKDRIAESRAILERIYPADEVEAEME 241
Query: 241 ALKESVESEIEESKTSKISMMTLLK----TTTVRRGLYAGMGLQFFQQFVGINTVMYYSP 296
ALK SVE+E + S LK VRRGL AG+ +Q QQFVGINTVMYYSP
Sbjct: 242 ALKLSVEAEKADEAIIGDSFSAKLKGAFGNPVVRRGLAAGITVQVAQQFVGINTVMYYSP 301
Query: 297 TIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLT 356
+IVQ AG+ASN+TA+ LSLITSGLNA GSI+S+ F+D+ GR+KL +IS+ G++ L +L
Sbjct: 302 SIVQFAGYASNKTAMALSLITSGLNALGSIVSMMFVDRYGRRKLMIISMFGIIACLIILA 361
Query: 357 VAFRESELHAPTVSAIETSHY--NNTCPDFR--AAVN--PGRWTCMTCLKASPSCGFCAA 410
F ++ +HAP + A E+ + N TC + AA N P RW CM CL++ CGFCA+
Sbjct: 362 TVFSQAAIHAPKIDAFESRTFAPNATCSAYAPLAAENAPPSRWNCMKCLRS--ECGFCAS 419
Query: 411 SPNKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVP 470
PGAC++ D K C + R ++ GCPSKFG+ A+V L LYI+ ++PGMGTVP
Sbjct: 420 GVQPYAPGACVVLSDDMKATCSSRGRTFFKDGCPSKFGFLAIVFLGLYIVVYAPGMGTVP 479
Query: 471 WVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIF 530
W++NSEIYPLRYRG+ GGIA+ + W+SNLIVS+SFLSLT +G++ TF++F + + +F
Sbjct: 480 WIVNSEIYPLRYRGLGGGIAAVSNWVSNLIVSESFLSLTHALGSSGTFLLFAGFSTIGLF 539
Query: 531 FVIVFVPETKGVPMEEVEKMLE 552
F+ + VPETKG+ EEVEK+LE
Sbjct: 540 FIWLLVPETKGLQFEEVEKLLE 561
>At2g35740 putative sugar transporter
Length = 580
Score = 636 bits (1640), Expect = 0.0
Identities = 309/548 (56%), Positives = 406/548 (73%), Gaps = 12/548 (2%)
Query: 15 ECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVST 74
E + +W+ PY++RLA SAGIGGLLFGY+TGVI+GALLYI+++F VD KTWLQE IVS
Sbjct: 15 EVWTTTWETPYIMRLALSAGIGGLLFGYNTGVIAGALLYIKEEFGEVDNKTWLQEIIVSM 74
Query: 75 AIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVG 134
+AGAI+GAA+GGW ND+FGRR++++IAD LFL+G+++M A P +++ GR+ VGFGVG
Sbjct: 75 TVAGAIVGAAIGGWYNDKFGRRMSVLIADVLFLLGALVMVIAHAPWVIILGRLLVGFGVG 134
Query: 135 MASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAA 194
MASM SPLYISE SP R+RGALVS N LITGGQFLSYLINLAF + PGTWRWMLGV+A
Sbjct: 135 MASMTSPLYISEMSPARIRGALVSTNGLLITGGQFLSYLINLAFVHTPGTWRWMLGVSAI 194
Query: 195 PAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEE-- 252
PAIIQ LML+LPESPRWLYR R+ ES+ IL++IY E V+AEI ALKESV +E +
Sbjct: 195 PAIIQFCLMLTLPESPRWLYRNDRKAESRDILERIYPAEMVEAEIAALKESVRAETADED 254
Query: 253 --SKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTA 310
T + L VR GL AG+ +Q QQFVGINTVMYYSPTI+Q AG+ASN+TA
Sbjct: 255 IIGHTFSDKLRGALSNPVVRHGLAAGITVQVAQQFVGINTVMYYSPTILQFAGYASNKTA 314
Query: 311 LLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVS 370
+ L+LITSGLNA GS++S+ F+D+ GR+KL +IS+ G++ L +L F E+ HAP +
Sbjct: 315 MALALITSGLNAVGSVVSMMFVDRYGRRKLMIISMFGIITCLVILAAVFNEASNHAPKID 374
Query: 371 AIETSHY--NNTCPDF----RAAVNPGRWTCMTCLKASPSCGFCAASPNKLLPGACLISD 424
++ ++ N TCP F + P W CM CL+ CGFC+ + PGAC++
Sbjct: 375 KRDSRNFAKNATCPAFAPFTASRSPPSNWNCMKCLQY--DCGFCSNGAQEYAPGACIVQS 432
Query: 425 DTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRG 484
K +C + R ++ GCPSKFG+ A+V L LYII ++PGMGTVPW++NSEIYPLRYRG
Sbjct: 433 ADMKALCHSKGRTFFKDGCPSKFGYLAIVFLGLYIIVYAPGMGTVPWIVNSEIYPLRYRG 492
Query: 485 VCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPM 544
+ GGIA+ + W+SNL+VS++FL+LT +G++ TF++F + V +FF+ + VPETKG+
Sbjct: 493 LAGGIAAVSNWMSNLVVSETFLTLTNAVGSSGTFLLFAGSSAVGLFFIWLLVPETKGLQF 552
Query: 545 EEVEKMLE 552
EEVEK+LE
Sbjct: 553 EEVEKLLE 560
>At2g43330 membrane transporter like protein
Length = 509
Score = 397 bits (1020), Expect = e-111
Identities = 201/359 (55%), Positives = 264/359 (72%), Gaps = 11/359 (3%)
Query: 6 PEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKT 65
PE MS F N Y+L L +AGIGGLLFGYDTGVISGALLYI+DDF+ V + +
Sbjct: 19 PERRMSYFG--------NSYILGLTVTAGIGGLLFGYDTGVISGALLYIKDDFEVVKQSS 70
Query: 66 WLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFG 125
+LQE IVS A+ GA+IGAA GGWIND +GR+ A + AD +F G+++MAAAP+P +L+ G
Sbjct: 71 FLQETIVSMALVGAMIGAAAGGWINDYYGRKKATLFADVVFAAGAIVMAAAPDPYVLISG 130
Query: 126 RVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTW 185
R+ VG GVG+AS+ +P+YI+EASP+ VRG LVS N +ITGGQFLSYL+N AFT PGTW
Sbjct: 131 RLLVGLGVGVASVTAPVYIAEASPSEVRGGLVSTNVLMITGGQFLSYLVNSAFTQVPGTW 190
Query: 186 RWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKES 245
RWMLGV+ PA+IQ +LML +PESPRWL+ K R+ E+ +L + Y ++ EI+ L +
Sbjct: 191 RWMLGVSGVPAVIQFILMLFMPESPRWLFMKNRKAEAIQVLARTYDISRLEDEIDHLSAA 250
Query: 246 VESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFA 305
E E + +T + + + ++ +R AG GLQ FQQF GINTVMYYSPTIVQ+AGF
Sbjct: 251 EEEEKQRKRT--VGYLDVFRSKELRLAFLAGAGLQAFQQFTGINTVMYYSPTIVQMAGFH 308
Query: 306 SNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVA-FRESE 363
SN+ AL LSLI + +NA G+++ IYFID GRKKLAL SL GV++SL +L+V+ F++SE
Sbjct: 309 SNQLALFLSLIVAAMNAAGTVVGIYFIDHCGRKKLALSSLFGVIISLLILSVSFFKQSE 367
Score = 143 bits (360), Expect = 3e-34
Identities = 61/109 (55%), Positives = 89/109 (80%)
Query: 447 FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFL 506
+GW A++GLALYI+ F+PGMG VPW +NSEIYP +YRG+CGG+++T WISNLIV+Q+FL
Sbjct: 375 YGWLAVLGLALYIVFFAPGMGPVPWTVNSEIYPQQYRGICGGMSATVNWISNLIVAQTFL 434
Query: 507 SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
++ + GT TF++ +AV+A+ FVIVFVPET+G+ EVE++ ++R+
Sbjct: 435 TIAEAAGTGMTFLILAGIAVLAVIFVIVFVPETQGLTFSEVEQIWKERA 483
>At3g18830 sugar transporter like protein
Length = 539
Score = 207 bits (528), Expect = 1e-53
Identities = 123/350 (35%), Positives = 195/350 (55%), Gaps = 20/350 (5%)
Query: 22 KNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAI-AGAI 80
+N Y A A + +L GYD GV+SGA++YI+ D K D LQ I++ ++ ++
Sbjct: 32 RNNYAFACAILASMTSILLGYDIGVMSGAMIYIKRDLKIND----LQIGILAGSLNIYSL 87
Query: 81 IGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMAS 140
IG+ G +D GRR I++A +F G+++M +PN A L+FGR G GVG A M +
Sbjct: 88 IGSCAAGRTSDWIGRRYTIVLAGAIFFAGAILMGLSPNYAFLMFGRFIAGIGVGYALMIA 147
Query: 141 PLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAP--GTWRWMLGVAAAPAII 198
P+Y +E SP RG L S I G L Y+ NLAF+N P WR MLG+ A P++I
Sbjct: 148 PVYTAEVSPASSRGFLNSFPEVFINAGIMLGYVSNLAFSNLPLKVGWRLMLGIGAVPSVI 207
Query: 199 QIVLMLSLPESPRWLYRKGREEESKSILKKIY-APEDVDAEIEALKESV-------ESEI 250
+ +L++PESPRWL +GR ++K +L K +P + +E +K + + +
Sbjct: 208 LAIGVLAMPESPRWLVMQGRLGDAKRVLDKTSDSPTEATLRLEDIKHAAGIPADCHDDVV 267
Query: 251 EESKTSKIS-----MMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFA 305
+ S+ + + + T VRR + A +G+ FFQQ GI+ V+ +SP I + AG
Sbjct: 268 QVSRRNSHGEGVWRELLIRPTPAVRRVMIAAIGIHFFQQASGIDAVVLFSPRIFKTAGLK 327
Query: 306 SNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
++ LL ++ + +++ + +D+ GR+ L L S+ G+VLSLA L
Sbjct: 328 TDHQQLLATVAVGVVKTSFILVATFLLDRIGRRPLLLTSVGGMVLSLAAL 377
Score = 75.1 bits (183), Expect = 1e-13
Identities = 45/127 (35%), Positives = 69/127 (53%), Gaps = 10/127 (7%)
Query: 446 KFGWAALVGLAL---YIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVS 502
K WA +V +A Y+ TFS G G + WV +SEI+PLR R + +++ ++S
Sbjct: 390 KVMWAVVVAIATVMTYVATFSIGAGPITWVYSSEIFPLRLRSQGSSMGVVVNRVTSGVIS 449
Query: 503 QSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQ 562
SFL +++ + T F +FG +A VA F F+PET+G +MLE F ++
Sbjct: 450 ISFLPMSKAMTTGGAFYLFGGIATVAWVFFYTFLPETQG-------RMLEDMDELFSGFR 502
Query: 563 KRDSGSE 569
RDS S+
Sbjct: 503 WRDSKSK 509
>At4g36670 sugar transporter like protein
Length = 493
Score = 202 bits (513), Expect = 5e-52
Identities = 120/348 (34%), Positives = 190/348 (54%), Gaps = 18/348 (5%)
Query: 23 NPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIG 82
N + L+ A A I ++FGYDTGV+SGA+++I +D K D + + I++ A++G
Sbjct: 14 NRFALQCAIVASIVSIIFGYDTGVMSGAMVFIEEDLKTNDVQIEVLTGILNLC---ALVG 70
Query: 83 AAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPL 142
+ + G +D GRR I++A LF++GS++M PN +LL GR G GVG A M +P+
Sbjct: 71 SLLAGRTSDIIGRRYTIVLASILFMLGSILMGWGPNYPVLLSGRCTAGLGVGFALMVAPV 130
Query: 143 YISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPG--TWRWMLGVAAAPAIIQI 200
Y +E + RG L SL I+ G L Y++N F+ P WR MLG+AA P+++
Sbjct: 131 YSAEIATASHRGLLASLPHLCISIGILLGYIVNYFFSKLPMHIGWRLMLGIAAVPSLVLA 190
Query: 201 VLMLSLPESPRWLYRKGREEESKSILKKI-YAPEDVDAEIEALKESVESE---------I 250
+L +PESPRWL +GR +E K IL+ + +PE+ + + +K + + +
Sbjct: 191 FGILKMPESPRWLIMQGRLKEGKEILELVSNSPEEAELRFQDIKAAAGIDPKCVDDVVKM 250
Query: 251 EESKTSKISM---MTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASN 307
E KT + + L T VRR L +G+ FFQ GI V+ Y P I + AG +
Sbjct: 251 EGKKTHGEGVWKELILRPTPAVRRVLLTALGIHFFQHASGIEAVLLYGPRIFKKAGITTK 310
Query: 308 RTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
L+++ + + +DK GR+KL L S+ G+V++L +L
Sbjct: 311 DKLFLVTIGVGIMKTTFIFTATLLLDKVGRRKLLLTSVGGMVIALTML 358
Score = 77.4 bits (189), Expect = 2e-14
Identities = 40/111 (36%), Positives = 60/111 (54%), Gaps = 3/111 (2%)
Query: 446 KFGWAALVGLAL---YIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVS 502
K WA ++ + ++ FS G+G + WV +SE++PL+ R + + N VS
Sbjct: 371 KLAWALVLSIVAAYSFVAFFSIGLGPITWVYSSEVFPLKLRAQGASLGVAVNRVMNATVS 430
Query: 503 QSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQ 553
SFLSLT I T F MF +A VA F +PETKG +EE+E + ++
Sbjct: 431 MSFLSLTSAITTGGAFFMFAGVAAVAWNFFFFLLPETKGKSLEEIEALFQR 481
>At5g16150 sugar transporter like protein
Length = 546
Score = 196 bits (498), Expect = 3e-50
Identities = 125/355 (35%), Positives = 187/355 (52%), Gaps = 13/355 (3%)
Query: 26 VLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAAV 85
VL A +G +LFGY GV++GAL Y+ D + T LQ IVS+ +AGA +G+
Sbjct: 105 VLPFVGVACLGAILFGYHLGVVNGALEYLAKDL-GIAENTVLQGWIVSSLLAGATVGSFT 163
Query: 86 GGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYIS 145
GG + D+FGR + IG+ + A A + ++ GR+ G G+G++S PLYIS
Sbjct: 164 GGALADKFGRTRTFQLDAIPLAIGAFLCATAQSVQTMIVGRLLAGIGIGISSAIVPLYIS 223
Query: 146 EASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLS 205
E SPT +RGAL S+N I G + + L P WR M GVA P+++ + M
Sbjct: 224 EISPTEIRGALGSVNQLFICIGILAALIAGLPLAANPLWWRTMFGVAVIPSVLLAIGMAF 283
Query: 206 LPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIE-ESKTSKISMMTLL 264
PESPRWL ++G+ E++ +K +Y E V + L S + E E+ +
Sbjct: 284 SPESPRWLVQQGKVSEAEKAIKTLYGKERVVELVRDLSASGQGSSEPEAGWFDLFSSRYW 343
Query: 265 KTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFG 324
K +V G L FQQ GIN V+YYS ++ + AG S+ A S + N FG
Sbjct: 344 KVVSV------GAALFLFQQLAGINAVVYYSTSVFRSAGIQSDVAA---SALVGASNVFG 394
Query: 325 SILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFRESEL--HAPTVSAIETSHY 377
+ ++ +DK GRK L L S G+ LS+ LL+++F L ++ T++ + T Y
Sbjct: 395 TAVASSLMDKMGRKSLLLTSFGGMALSMLLLSLSFTWKALAAYSGTLAVVGTVLY 449
Score = 68.2 bits (165), Expect = 1e-11
Identities = 36/104 (34%), Positives = 58/104 (55%)
Query: 448 GWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLS 507
G A+VG LY+++FS G G VP ++ EI+ R R ++ WISN ++ FLS
Sbjct: 439 GTLAVVGTVLYVLSFSLGAGPVPALLLPEIFASRIRAKAVALSLGMHWISNFVIGLYFLS 498
Query: 508 LTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKML 551
+ G + ++ F + V+A+ ++ V ETKG +EE+E L
Sbjct: 499 VVTKFGISSVYLGFAGVCVLAVLYIAGNVVETKGRSLEEIELAL 542
>At2g18480 putative sugar transporter
Length = 508
Score = 196 bits (497), Expect = 4e-50
Identities = 128/356 (35%), Positives = 189/356 (52%), Gaps = 24/356 (6%)
Query: 23 NPYVLRLAFS----AGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAG 78
NP++ + AF A I ++FGYDTGV+SGA ++IRDD K D + + I++
Sbjct: 15 NPHMNKFAFGCAIVASIISIIFGYDTGVMSGAQIFIRDDLKINDTQIEVLAGILNLC--- 71
Query: 79 AIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASM 138
A++G+ G +D GRR I ++ +FL+GSV+M PN +L+ GR G GVG A M
Sbjct: 72 ALVGSLTAGKTSDVIGRRYTIALSAVIFLVGSVLMGYGPNYPVLMVGRCIAGVGVGFALM 131
Query: 139 ASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAF--TNAPGTWRWMLGVAAAPA 196
+P+Y +E S RG L SL I+ G L Y+ N F WR MLG+AA P+
Sbjct: 132 IAPVYSAEISSASHRGFLTSLPELCISLGILLGYVSNYCFGKLTLKLGWRLMLGIAAFPS 191
Query: 197 IIQIVLMLSLPESPRWLYRKGREEESKSILKKI-YAPEDVDAEIEALKESVESEIEESK- 254
+I + +PESPRWL +GR EE+K I+ + E+ + + + E ++ E K
Sbjct: 192 LILAFGITRMPESPRWLVMQGRLEEAKKIMVLVSNTEEEAEERFRDILTAAEVDVTEIKE 251
Query: 255 -----------TSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAG 303
S + + VR L A +G+ FF+ GI V+ YSP I + AG
Sbjct: 252 VGGGVKKKNHGKSVWRELVIKPRPAVRLILIAAVGIHFFEHATGIEAVVLYSPRIFKKAG 311
Query: 304 FASNRTALLLSLITSGL-NAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVA 358
S + LLL+ + GL AF I++ + +DK GR+KL L S G+V +L L V+
Sbjct: 312 VVS-KDKLLLATVGVGLTKAFFIIIATFLLDKVGRRKLLLTSTGGMVFALTSLAVS 366
Score = 84.0 bits (206), Expect = 2e-16
Identities = 44/108 (40%), Positives = 63/108 (57%), Gaps = 3/108 (2%)
Query: 446 KFGWA---ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVS 502
+ WA ++V ++ FS G+G + WV +SEI+PLR R I I N VS
Sbjct: 375 RLAWALSLSIVSTYAFVAFFSIGLGPITWVYSSEIFPLRLRAQGASIGVAVNRIMNATVS 434
Query: 503 QSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKM 550
SFLS+T+ I T F +F +AV A +F +PETKG+P+EE+EK+
Sbjct: 435 MSFLSMTKAITTGGVFFVFAGIAVAAWWFFFFMLPETKGLPLEEMEKL 482
>At2g20780 putative sugar transporter
Length = 547
Score = 183 bits (464), Expect = 3e-46
Identities = 116/356 (32%), Positives = 178/356 (49%), Gaps = 28/356 (7%)
Query: 25 YVLRLAFSAGIGGLLFGY---------------------DTGVISGALLYIRDDFKAVDR 63
YV+ AF A + +L GY D GV+SGA+L+I+ D K +
Sbjct: 54 YVMACAFFASLNNVLLGYGRFYLYNRILLLLLYFVDLQKDVGVMSGAVLFIQQDLKITEV 113
Query: 64 KTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILL 123
+T E ++ + ++ G+ GG +D GR+ + +A +F G+ +MA AP+ +L+
Sbjct: 114 QT---EVLIGSLSIISLFGSLAGGRTSDSIGRKWTMALAALVFQTGAAVMAVAPSFEVLM 170
Query: 124 FGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPG 183
GR G G+G+ M +P+YI+E SPT RG S I G L Y+ N AF+
Sbjct: 171 IGRTLAGIGIGLGVMIAPVYIAEISPTVARGFFTSFPEIFINLGILLGYVSNYAFSGLSV 230
Query: 184 --TWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEA 241
+WR ML V P++ + +PESPRWL KGR + ++ +L K +D E A
Sbjct: 231 HISWRIMLAVGILPSVFIGFALCVIPESPRWLVMKGRVDSAREVLMKTNERDDEAEERLA 290
Query: 242 LKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQL 301
+ + E S+ + L + VR+ L G G+Q FQQ GI+ +YYSP I++
Sbjct: 291 EIQLAAAHTEGSEDRPVWRELLSPSPVVRKMLIVGFGIQCFQQITGIDATVYYSPEILKE 350
Query: 302 AGFASNRTALLLSLITSGLNAFGSIL-SIYFIDKTGRKKLALISLCGVVLSLALLT 356
AG + T LL + + G+ IL + + ID GRK L +S G+ L L L+
Sbjct: 351 AGI-QDETKLLAATVAVGVTKTVFILFATFLIDSVGRKPLLYVSTIGMTLCLFCLS 405
Score = 75.5 bits (184), Expect = 8e-14
Identities = 37/102 (36%), Positives = 62/102 (60%)
Query: 451 ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQ 510
AL+ + + FS GMG V WV+ SEI+PLR R + + + + +V+ SFLS+++
Sbjct: 421 ALLFVCGNVAFFSIGMGPVCWVLTSEIFPLRLRAQASALGAVGNRVCSGLVAMSFLSVSR 480
Query: 511 TIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLE 552
I TF +F +++ +++ FV V VPET G +E++E M +
Sbjct: 481 AITVGGTFFVFSLVSALSVIFVYVLVPETSGKSLEQIELMFQ 522
>At1g07340 hexose transporter like protein
Length = 383
Score = 182 bits (461), Expect = 6e-46
Identities = 104/326 (31%), Positives = 171/326 (51%), Gaps = 21/326 (6%)
Query: 33 AGIGGLLFGYDTGVISGAL---LYIRDDFKAVDRKTW-------------LQEAIVSTAI 76
A +GGL+FGYD G+ G ++ D F V K L + S+
Sbjct: 30 AAVGGLMFGYDIGISGGVTSMDTFLLDFFPHVYEKKHRVHENNYCKFDDQLLQLFTSSLY 89
Query: 77 AGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMA 136
I + + +++ FGR+ I++A FL+G+++ +A +L+ GR+ +GFG+G
Sbjct: 90 LAGIFASFISSYVSRAFGRKPTIMLASIFFLVGAILNLSAQELGMLIGGRILLGFGIGFG 149
Query: 137 SMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPA 196
+ PL+ISE +P R RG L + FLIT G + +N + WR+ LG AA PA
Sbjct: 150 NQTVPLFISEIAPARYRGGLNVMFQFLITIGILAASYVNYLTSTLKNGWRYSLGGAAVPA 209
Query: 197 IIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTS 256
+I ++ + E+P L +G++E+ K +L+KI ED++ E +K + E +
Sbjct: 210 LILLIGSFFIHETPASLIERGKDEKGKQVLRKIRGIEDIELEFNEIKYATEVATKVKSPF 269
Query: 257 KISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLI 316
K L + R L G LQFFQQF GIN VM+Y+P + Q G N +L+ +++
Sbjct: 270 K----ELFTKSENRPPLVCGTLLQFFQQFTGINVVMFYAPVLFQTMGSGDN-ASLISTVV 324
Query: 317 TSGLNAFGSILSIYFIDKTGRKKLAL 342
T+G+NA +++S+ +D GR+ L +
Sbjct: 325 TNGVNAIATVISLLVVDFAGRRCLLM 350
>At5g26250 hexose transporter - like protein
Length = 507
Score = 179 bits (453), Expect = 5e-45
Identities = 119/353 (33%), Positives = 184/353 (51%), Gaps = 30/353 (8%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAV-DRKTWLQE----------- 69
YV A +GGL+FGYD G+ G ++++ F +V +RK E
Sbjct: 21 YVFICVIIAAVGGLIFGYDIGISGGVTAMDDFLKEFFPSVYERKKHAHENNYCKYDNQFL 80
Query: 70 -AIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVF 128
S+ A++ + + GRR + +A FLIG + A A N +L+ GR+
Sbjct: 81 QLFTSSLYLAALVASFFASATCSKLGRRPTMQLASIFFLIGVGLAAGAVNIYMLIIGRIL 140
Query: 129 VGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNA--PGTWR 186
+GFGVG + A PL++SE +P R+RG L + ++T G ++ ++N FT++ P WR
Sbjct: 141 LGFGVGFGNQAVPLFLSEIAPARLRGGLNIVFQLMVTIGILIANIVNY-FTSSIHPYGWR 199
Query: 187 WMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESV 246
LG A PA+I + L + E+P L + + +E K LKKI EDVD E ES+
Sbjct: 200 IALGGAGIPALILLFGSLLICETPTSLIERNKTKEGKETLKKIRGVEDVDEEY----ESI 255
Query: 247 ESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFAS 306
+ ++ K L+K + R GM LQFFQQF GIN +M+Y+P + Q GF
Sbjct: 256 VHACDIARQVKDPYTKLMKPAS-RPPFVIGMLLQFFQQFTGINAIMFYAPVLFQTVGF-G 313
Query: 307 NRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALIS-----LCGVVLSLAL 354
N ALL +++T +N + + I+ +DKTGR+ L L S +C +V+ + L
Sbjct: 314 NDAALLSAVVTGTINVLSTFVGIFLVDKTGRRFLLLQSSVHMLICQLVIGIIL 366
Score = 52.0 bits (123), Expect = 9e-07
Identities = 27/96 (28%), Positives = 54/96 (56%), Gaps = 1/96 (1%)
Query: 452 LVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQT 511
++ + +Y++ F+ G + W+I SE +PL R +A + +++Q+FLS+
Sbjct: 385 VIFVCVYVMGFAWSWGPLGWLIPSETFPLETRTEGFALAVSCNMFFTFVIAQAFLSMLCA 444
Query: 512 IGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEV 547
+ + F G + V+ + F + FVPETKGV ++++
Sbjct: 445 MKSGIFFFFSGWIVVMGL-FALFFVPETKGVSIDDM 479
>At3g05960 putative hexose transporter
Length = 507
Score = 178 bits (452), Expect = 6e-45
Identities = 117/352 (33%), Positives = 182/352 (51%), Gaps = 28/352 (7%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAV-DRKTWLQE----------- 69
YV A +GGL+FGYD G+ G ++++ F AV +RK + E
Sbjct: 20 YVFICVMIAAVGGLIFGYDIGISGGVSAMDDFLKEFFPAVWERKKHVHENNYCKYDNQFL 79
Query: 70 -AIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVF 128
S+ A++ + V + GRR + A FLIG + A A N +L+ GR+F
Sbjct: 80 QLFTSSLYLAALVASFVASATCSKLGRRPTMQFASIFFLIGVGLTAGAVNLVMLIIGRLF 139
Query: 129 VGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLIN-LAFTNAPGTWRW 187
+GFGVG + A PL++SE +P ++RG L + ++T G ++ ++N T P WR
Sbjct: 140 LGFGVGFGNQAVPLFLSEIAPAQLRGGLNIVFQLMVTIGILIANIVNYFTATVHPYGWRI 199
Query: 188 MLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVE 247
LG A PA+I + L + E+P L + + EE K L+KI +D++ E ES+
Sbjct: 200 ALGGAGIPAVILLFGSLLIIETPTSLIERNKNEEGKEALRKIRGVDDINDEY----ESIV 255
Query: 248 SEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASN 307
+ + K LLK + R GM LQ FQQF GIN +M+Y+P + Q GF S+
Sbjct: 256 HACDIASQVKDPYRKLLKPAS-RPPFIIGMLLQLFQQFTGINAIMFYAPVLFQTVGFGSD 314
Query: 308 RTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALIS-----LCGVVLSLAL 354
ALL ++IT +N + + IY +D+TGR+ L L S +C +++ + L
Sbjct: 315 -AALLSAVITGSINVLATFVGIYLVDRTGRRFLLLQSSVHMLICQLIIGIIL 365
Score = 51.6 bits (122), Expect = 1e-06
Identities = 25/98 (25%), Positives = 54/98 (54%), Gaps = 1/98 (1%)
Query: 452 LVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQT 511
++ + +Y++ F+ G + W+I SE +PL R +A + +++Q+FLS+
Sbjct: 384 VIFVCVYVMGFAWSWGPLGWLIPSETFPLETRSAGFAVAVSCNMFFTFVIAQAFLSMLCG 443
Query: 512 IGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEK 549
+ + F G + V+ + F F+PETKG+ ++++ +
Sbjct: 444 MRSGIFFFFSGWIIVMGL-FAFFFIPETKGIAIDDMRE 480
>At2g16120 putative sugar transporter
Length = 511
Score = 178 bits (452), Expect = 6e-45
Identities = 112/351 (31%), Positives = 179/351 (50%), Gaps = 22/351 (6%)
Query: 22 KNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAII 81
++ Y A A + ++ GYD GV+SGA ++I+DD K D + + I++ +++
Sbjct: 22 RSRYAFACAILASMTSIILGYDIGVMSGASIFIKDDLKLSDVQLEILMGILNIY---SLV 78
Query: 82 GAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASP 141
G+ G +D GRR I++A F G+++M A N ++ GR G GVG A M +P
Sbjct: 79 GSGAAGRTSDWLGRRYTIVLAGAFFFCGALLMGFATNYPFIMVGRFVAGIGVGYAMMIAP 138
Query: 142 LYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPG--TWRWMLGVAAAPAIIQ 199
+Y +E +P RG L S I G L Y+ N F+ P WR+MLGV A P++
Sbjct: 139 VYTAEVAPASSRGFLTSFPEIFINIGILLGYVSNYFFSKLPEHLGWRFMLGVGAVPSVFL 198
Query: 200 IVLMLSLPESPRWLYRKGREEESKSILKKI-YAPEDVDAEIEALKESVESEIEESKTSKI 258
+ +L++PESPRWL +GR ++ +L K E+ + ++ +K +V I + T +
Sbjct: 199 AIGVLAMPESPRWLVLQGRLGDAFKVLDKTSNTKEEAISRLDDIKRAV--GIPDDMTDDV 256
Query: 259 SMMTLLK--------------TTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGF 304
++ K T +VR L A +G+ F QQ GI+ V+ YSPTI AG
Sbjct: 257 IVVPNKKSAGKGVWKDLLVRPTPSVRHILIACLGIHFAQQASGIDAVVLYSPTIFSKAGL 316
Query: 305 ASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
S LL ++ + ++ +D+ GR+ L L S+ G+ LSL L
Sbjct: 317 KSKNDQLLATVAVGVVKTLFIVVGTCVVDRFGRRALLLTSMGGMFLSLTAL 367
Score = 69.7 bits (169), Expect = 4e-12
Identities = 38/107 (35%), Positives = 61/107 (56%), Gaps = 7/107 (6%)
Query: 449 WAALVGLAL-----YIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQ 503
WA +GLA+ ++ TFS G G V WV SEI+P+R R + + + I+
Sbjct: 384 WA--IGLAVTTVMTFVATFSIGAGPVTWVYCSEIFPVRLRAQGASLGVMLNRLMSGIIGM 441
Query: 504 SFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKM 550
+FLSL++ + F++F +A A F F+PET+G+P+EE+E +
Sbjct: 442 TFLSLSKGLTIGGAFLLFAGVAAAAWVFFFTFLPETRGIPLEEMETL 488
>At1g50310 hexose transporter, putative
Length = 517
Score = 177 bits (450), Expect = 1e-44
Identities = 104/336 (30%), Positives = 181/336 (52%), Gaps = 23/336 (6%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRK--------------TWL 67
+V+ A +GGLLFGYD G+ G ++ F VD++ L
Sbjct: 24 FVIMTCIVAAMGGLLFGYDLGISGGVTSMEEFLSKFFPEVDKQMHEARRETAYCKFDNQL 83
Query: 68 QEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRV 127
+ S+ A+ + V + ++GR++++ + FLIGS+ A A N A+L+ GR+
Sbjct: 84 LQLFTSSLYLAALASSFVASAVTRKYGRKISMFVGGVAFLIGSLFNAFATNVAMLIVGRL 143
Query: 128 FVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN-APGTWR 186
+G GVG A+ ++P+Y+SE +P ++RGAL IT G ++ LIN + A WR
Sbjct: 144 LLGVGVGFANQSTPVYLSEMAPAKIRGALNIGFQMAITIGILIANLINYGTSQMAKNGWR 203
Query: 187 WMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESV 246
LG+AA PA+I ++ LP++P + +G+ E+++ +L+KI ++VD E + L ++
Sbjct: 204 VSLGLAAVPAVIMVIGSFVLPDTPNSMLERGKYEQAREMLQKIRGADNVDEEFQDLCDAC 263
Query: 247 ESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFAS 306
E+ +K + + R L + FFQQ GIN +M+Y+P + + GFA
Sbjct: 264 EA----AKKVDNPWKNIFQQAKYRPALVFCSAIPFFQQITGINVIMFYAPVLFKTLGFAD 319
Query: 307 NRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLAL 342
+ +L+ ++IT +N +++SIY +D+ GR+ L L
Sbjct: 320 D-ASLISAVITGAVNVVSTLVSIYAVDRYGRRILFL 354
Score = 55.8 bits (133), Expect = 6e-08
Identities = 34/120 (28%), Positives = 61/120 (50%), Gaps = 6/120 (5%)
Query: 444 PSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQ 503
P+ W L + LY+ F+ G + W++ SEI PL R I + ++ Q
Sbjct: 385 PATADWI-LAFICLYVAGFAWSWGPLGWLVPSEICPLEIRPAGQAINVSVNMFFTFLIGQ 443
Query: 504 SFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 563
FL++ + + G++AV+ + F+ +PETKGVP+EE+ ++ +Q FW++
Sbjct: 444 FFLTMLCHMKFGLFYFFGGMVAVMTV-FIYFLLPETKGVPIEEMGRVWKQH----PFWKR 498
Score = 29.6 bits (65), Expect = 4.7
Identities = 43/171 (25%), Positives = 70/171 (40%), Gaps = 45/171 (26%)
Query: 70 AIVSTAIAGAI--IGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRV 127
+++S I GA+ + V + DR+GRR+ LFL G + M + L G
Sbjct: 322 SLISAVITGAVNVVSTLVSIYAVDRYGRRI-------LFLEGGIQMIVSQIVVGTLIGMK 374
Query: 128 FVGFGVGMASMASP--------LYI---------------SEASPTRVRGA----LVSLN 160
F G G + A+ LY+ SE P +R A VS+N
Sbjct: 375 FGTTGSGTLTPATADWILAFICLYVAGFAWSWGPLGWLVPSEICPLEIRPAGQAINVSVN 434
Query: 161 SF--LITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLSLPES 209
F + G FL+ L ++ F G + + G+ A++ + + LPE+
Sbjct: 435 MFFTFLIGQFFLTMLCHMKF----GLFYFFGGMV---AVMTVFIYFLLPET 478
>At5g59250 D-xylose-H+ symporter - like protein
Length = 558
Score = 175 bits (444), Expect = 5e-44
Identities = 122/362 (33%), Positives = 192/362 (52%), Gaps = 25/362 (6%)
Query: 18 SLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEA------I 71
S SW + +L F A +GGLLFGYD G SGA L ++ A+ TW + +
Sbjct: 92 SFSWSS-VILPFIFPA-LGGLLFGYDIGATSGATLSLQSP--ALSGTTWFNFSPVQLGLV 147
Query: 72 VSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGF 131
VS ++ GA++G+ + D GRR +IIA L+L+GS+I AP+ ILL GR+ GF
Sbjct: 148 VSGSLYGALLGSISVYGVADFLGRRRELIIAAVLYLLGSLITGCAPDLNILLVGRLLYGF 207
Query: 132 GVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGV 191
G+G+A +PLYI+E P+++RG L+SL I G L + + + G WR+M G
Sbjct: 208 GIGLAMHGAPLYIAETCPSQIRGTLISLKELFIVLGILLGFSVGSFQIDVVGGWRYMYGF 267
Query: 192 AAAPAIIQIVLMLSLPESPRWL-----YRKGREEESKS----ILKKIYAPEDVDAEIEAL 242
A++ + M SLP SPRWL KG+ +E K L K+ D E L
Sbjct: 268 GTPVALLMGLGMWSLPASPRWLLLRAVQGKGQLQEYKEKAMLALSKLRGRPPGDKISEKL 327
Query: 243 KE----SVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTI 298
+ SV++ E+ K+ + + + + + + L G GL FQQ G +V+YY+ +I
Sbjct: 328 VDDAYLSVKTAYEDEKSGG-NFLEVFQGPNL-KALTIGGGLVLFQQITGQPSVLYYAGSI 385
Query: 299 VQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVA 358
+Q AGF++ A +S+I + +++ +D GR+ L + + G+ LSL LL+
Sbjct: 386 LQTAGFSAAADATRVSVIIGVFKLLMTWVAVAKVDDLGRRPLLIGGVSGIALSLFLLSAY 445
Query: 359 FR 360
++
Sbjct: 446 YK 447
Score = 77.4 bits (189), Expect = 2e-14
Identities = 41/105 (39%), Positives = 65/105 (61%)
Query: 447 FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFL 506
F A+ L LY+ + G + W++ SEI+PLR RG +A T + SN IV+ +F
Sbjct: 452 FPLVAVGALLLYVGCYQISFGPISWLMVSEIFPLRTRGRGISLAVLTNFGSNAIVTFAFS 511
Query: 507 SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKML 551
L + +G F++FG +A+V++ FVI+ VPETKG+ +EE+E +
Sbjct: 512 PLKEFLGAENLFLLFGGIALVSLLFVILVVPETKGLSLEEIESKI 556
>At2g16130 putative sugar transporter
Length = 511
Score = 174 bits (440), Expect = 2e-43
Identities = 110/351 (31%), Positives = 176/351 (49%), Gaps = 22/351 (6%)
Query: 22 KNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAII 81
++ + A A + ++ GYD GV+SGA ++I+DD K D + + I++ ++I
Sbjct: 22 RSRFAFACAILASMTSIILGYDIGVMSGAAIFIKDDLKLSDVQLEILMGILNIY---SLI 78
Query: 82 GAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASP 141
G+ G +D GRR I++A F G+++M A N ++ GR G GVG A M +P
Sbjct: 79 GSGAAGRTSDWIGRRYTIVLAGFFFFCGALLMGFATNYPFIMVGRFVAGIGVGYAMMIAP 138
Query: 142 LYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPG--TWRWMLGVAAAPAIIQ 199
+Y +E +P RG L S I G L Y+ N F P WR+MLG+ A P++
Sbjct: 139 VYTTEVAPASSRGFLSSFPEIFINIGILLGYVSNYFFAKLPEHIGWRFMLGIGAVPSVFL 198
Query: 200 IVLMLSLPESPRWLYRKGREEESKSILKKI-YAPEDVDAEIEALKESVESEIEESKTSKI 258
+ +L++PESPRWL +GR ++ +L K E+ + + +K +V I + T +
Sbjct: 199 AIGVLAMPESPRWLVMQGRLGDAFKVLDKTSNTKEEAISRLNDIKRAV--GIPDDMTDDV 256
Query: 259 SMMTLLK--------------TTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGF 304
++ K T +VR L A +G+ F QQ GI+ V+ YSPTI AG
Sbjct: 257 IVVPNKKSAGKGVWKDLLVRPTPSVRHILIACLGIHFSQQASGIDAVVLYSPTIFSRAGL 316
Query: 305 ASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
S LL ++ + ++ +D+ GR+ L L S+ G+ SL L
Sbjct: 317 KSKNDQLLATVAVGVVKTLFIVVGTCLVDRFGRRALLLTSMGGMFFSLTAL 367
Score = 73.6 bits (179), Expect = 3e-13
Identities = 42/116 (36%), Positives = 64/116 (54%), Gaps = 7/116 (6%)
Query: 449 WAALVGLAL-----YIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQ 503
WA +GLA+ ++ TFS G G V WV SEI+P+R R + + + I+
Sbjct: 384 WA--IGLAVTTVMTFVATFSLGAGPVTWVYASEIFPVRLRAQGASLGVMLNRLMSGIIGM 441
Query: 504 SFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFK 559
+FLSL++ + F++F +AV A F F+PET+GVP+EE+E + S K
Sbjct: 442 TFLSLSKGLTIGGAFLLFAGVAVAAWVFFFTFLPETRGVPLEEIESLFGSYSANKK 497
>At3g19940 putative monosaccharide transport protein, STP4
Length = 514
Score = 173 bits (438), Expect = 3e-43
Identities = 106/336 (31%), Positives = 179/336 (52%), Gaps = 24/336 (7%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRK--------------TWL 67
+V+ A +GGLLFGYD G+ G ++ F V+ + +
Sbjct: 24 FVIMTCIVAAMGGLLFGYDLGISGGVTSMEEFLTKFFPQVESQMKKAKHDTAYCKFDNQM 83
Query: 68 QEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRV 127
+ S+ A++ + + I + GR+V++ I FLIG++ A A N ++L+ GR+
Sbjct: 84 LQLFTSSLYLAALVASFMASVITRKHGRKVSMFIGGLAFLIGALFNAFAVNVSMLIIGRL 143
Query: 128 FVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN-APGTWR 186
+G GVG A+ ++P+Y+SE +P ++RGAL IT G ++ LIN + A WR
Sbjct: 144 LLGVGVGFANQSTPVYLSEMAPAKIRGALNIGFQMAITIGILVANLINYGTSKMAQHGWR 203
Query: 187 WMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESV 246
LG+AA PA++ ++ LP++P + +G+ EE+K +LKKI ++VD E + L ++V
Sbjct: 204 VSLGLAAVPAVVMVIGSFILPDTPNSMLERGKNEEAKQMLKKIRGADNVDHEFQDLIDAV 263
Query: 247 ESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFAS 306
E+ + K M + R L + FFQQ GIN +M+Y+P + + GF
Sbjct: 264 EAAKKVENPWKNIM-----ESKYRPALIFCSAIPFFQQITGINVIMFYAPVLFKTLGFGD 318
Query: 307 NRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLAL 342
+ AL+ ++IT +N + +SIY +D+ GR+ L L
Sbjct: 319 D-AALMSAVITGVVNMLSTFVSIYAVDRYGRRLLFL 353
Score = 52.8 bits (125), Expect = 5e-07
Identities = 32/120 (26%), Positives = 60/120 (49%), Gaps = 6/120 (5%)
Query: 444 PSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQ 503
P+ W L + +Y+ F+ G + W++ SEI PL R I + ++ Q
Sbjct: 384 PATADWI-LAFICVYVAGFAWSWGPLGWLVPSEICPLEIRPAGQAINVSVNMFFTFLIGQ 442
Query: 504 SFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 563
FL++ + + ++A++ +F + +PETKGVP+EE+ ++ +Q FW+K
Sbjct: 443 FFLTMLCHMKFGLFYFFASMVAIMTVF-IYFLLPETKGVPIEEMGRVWKQH----WFWKK 497
>At5g26340 hexose transporter - like protein
Length = 526
Score = 167 bits (424), Expect = 1e-41
Identities = 112/340 (32%), Positives = 170/340 (49%), Gaps = 28/340 (8%)
Query: 24 PYVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRKT--------------- 65
P V+ A GGL+FGYD GV G ++ F V RK
Sbjct: 21 PIVIISCIMAATGGLMFGYDVGVSGGVTSMPDFLEKFFPVVYRKVVAGADKDSNYCKYDN 80
Query: 66 -WLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLF 124
LQ S +AG + + GRR+ ++IA F+IG + A A + A+L+
Sbjct: 81 QGLQLFTSSLYLAG-LTATFFASYTTRTLGRRLTMLIAGVFFIIGVALNAGAQDLAMLIA 139
Query: 125 GRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT 184
GR+ +G GVG A+ A PL++SE +PTR+RG L L +T G + L+N G
Sbjct: 140 GRILLGCGVGFANQAVPLFLSEIAPTRIRGGLNILFQLNVTIGILFANLVNYGTAKIKGG 199
Query: 185 WRW--MLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEAL 242
W W LG+A PA++ V L + E+P L +GR +E K++L++I ++V+ E L
Sbjct: 200 WGWRLSLGLAGIPALLLTVGALLVTETPNSLVERGRLDEGKAVLRRIRGTDNVEPEFADL 259
Query: 243 KESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLA 302
E+ +K K LL+ R L + LQ FQQ GIN +M+Y+P +
Sbjct: 260 LEASRL----AKEVKHPFRNLLQRRN-RPQLVIAVALQIFQQCTGINAIMFYAPVLFSTL 314
Query: 303 GFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLAL 342
GF S+ +L +++T +N +++SIY +DK GR+ L L
Sbjct: 315 GFGSD-ASLYSAVVTGAVNVLSTLVSIYSVDKVGRRVLLL 353
Score = 52.0 bits (123), Expect = 9e-07
Identities = 32/117 (27%), Positives = 55/117 (46%), Gaps = 4/117 (3%)
Query: 447 FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFL 506
F +V + Y+ F+ G + W+I SE +PL R + + I++Q+FL
Sbjct: 385 FAILVVVMICTYVAAFAWSWGPLGWLIPSETFPLETRSAGQSVTVCVNLLFTFIIAQAFL 444
Query: 507 SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 563
S+ F+ F ++ FV+ +PETK +P+EE M E+ + FW +
Sbjct: 445 SMLCHFKFG-IFIFFSAWVLIMSVFVMFLLPETKNIPIEE---MTERVWKKHWFWAR 497
>At1g11260 glucose transporter
Length = 522
Score = 167 bits (424), Expect = 1e-41
Identities = 106/338 (31%), Positives = 179/338 (52%), Gaps = 25/338 (7%)
Query: 24 PYVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRKTWLQEA---------- 70
P+VL A +GGL+FGYD G+ G +++ F +V RK +
Sbjct: 21 PFVLFTCVVAAMGGLIFGYDIGISGGVTSMPSFLKRFFPSVYRKQQEDASTNQYCQYDSP 80
Query: 71 ----IVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGR 126
S+ A+I + V + +FGRR++++ LF G++I A + +L+ GR
Sbjct: 81 TLTMFTSSLYLAALISSLVASTVTRKFGRRLSMLFGGILFCAGALINGFAKHVWMLIVGR 140
Query: 127 VFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWR 186
+ +GFG+G A+ A PLY+SE +P + RGAL IT G ++ ++N F G W
Sbjct: 141 ILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVAEVLNYFFAKIKGGWG 200
Query: 187 W--MLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKE 244
W LG A PA+I + L LP++P + +G+ EE+K+ L++I +DV E + L
Sbjct: 201 WRLSLGGAVVPALIITIGSLVLPDTPNSMIERGQHEEAKTKLRRIRGVDDVSQEFDDL-- 258
Query: 245 SVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGF 304
+ +ES++ + LL+ R L + + FFQQ GIN +M+Y+P + GF
Sbjct: 259 --VAASKESQSIEHPWRNLLR-RKYRPHLTMAVMIPFFQQLTGINVIMFYAPVLFNTIGF 315
Query: 305 ASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLAL 342
++ +L+ +++T +N +++SIY +D+ GR+ L L
Sbjct: 316 TTD-ASLMSAVVTGSVNVAATLVSIYGVDRWGRRFLFL 352
Score = 64.3 bits (155), Expect = 2e-10
Identities = 36/123 (29%), Positives = 64/123 (51%), Gaps = 4/123 (3%)
Query: 449 WAALVG---LALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSF 505
W A+V + +Y+ F+ G + W++ SEI+PL R I + I I++Q F
Sbjct: 385 WYAIVVVTFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSITVSVNMIFTFIIAQIF 444
Query: 506 LSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRD 565
L++ + F++F VV FV +F+PETKG+P+EE+ ++ +F + +
Sbjct: 445 LTMLCHLKFG-LFLVFAFFVVVMSIFVYIFLPETKGIPIEEMGQVWRSHWYWSRFVEDGE 503
Query: 566 SGS 568
G+
Sbjct: 504 YGN 506
>At5g17010 sugar transporter - like protein
Length = 503
Score = 166 bits (421), Expect = 2e-41
Identities = 112/345 (32%), Positives = 180/345 (51%), Gaps = 19/345 (5%)
Query: 26 VLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEA------IVSTAIAGA 79
+L F A +GGLL+GY+ G S A + ++ ++ +W + + S ++ GA
Sbjct: 48 ILPFLFPA-LGGLLYGYEIGATSCATISLQSP--SLSGISWYNLSSVDVGLVTSGSLYGA 104
Query: 80 IIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMA 139
+ G+ V I D GRR +I+A L+L+G+++ A AP ++L+ GRV G VG+A A
Sbjct: 105 LFGSIVAFTIADVIGRRKELILAALLYLVGALVTALAPTYSVLIIGRVIYGVSVGLAMHA 164
Query: 140 SPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQ 199
+P+YI+E +P+ +RG LVSL F I G Y I N WR+M + A+I
Sbjct: 165 APMYIAETAPSPIRGQLVSLKEFFIVLGMVGGYGIGSLTVNVHSGWRYMYATSVPLAVIM 224
Query: 200 IVLMLSLPESPRWL-----YRKGREEESKSILKK----IYAPEDVDAEIEALKESVESEI 250
+ M LP SPRWL KG E + K + P VD+ E + E +
Sbjct: 225 GIGMWWLPASPRWLLLRVIQGKGNVENQREAAIKSLCCLRGPAFVDSAAEQVNEILAELT 284
Query: 251 EESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTA 310
+ +++ L + + + L G GL FQQ G +V+YY+P+I+Q AGF++ A
Sbjct: 285 FVGEDKEVTFGELFQGKCL-KALIIGGGLVLFQQITGQPSVLYYAPSILQTAGFSAAGDA 343
Query: 311 LLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
+S++ L + +++ ID+ GR+ L L + G+V+SL LL
Sbjct: 344 TRVSILLGLLKLIMTGVAVVVIDRLGRRPLLLGGVGGMVVSLFLL 388
Score = 70.5 bits (171), Expect = 2e-12
Identities = 34/98 (34%), Positives = 59/98 (59%)
Query: 451 ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQ 510
A+V L LY+ + G + W++ SEI+PL+ RG +A + +N +V+ +F L +
Sbjct: 402 AVVALLLYVGCYQLSFGPIGWLMISEIFPLKLRGRGLSLAVLVNFGANALVTFAFSPLKE 461
Query: 511 TIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVE 548
+G F FG++ V+++ F+ VPETKG+ +EE+E
Sbjct: 462 LLGAGILFCGFGVICVLSLVFIFFIVPETKGLTLEEIE 499
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.138 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,069,605
Number of Sequences: 26719
Number of extensions: 491758
Number of successful extensions: 1874
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 1487
Number of HSP's gapped (non-prelim): 171
length of query: 571
length of database: 11,318,596
effective HSP length: 105
effective length of query: 466
effective length of database: 8,513,101
effective search space: 3967105066
effective search space used: 3967105066
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0140.7


