

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.2
(123 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
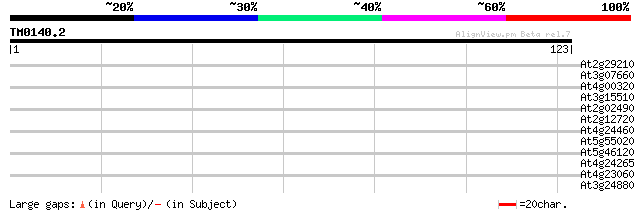
Score E
Sequences producing significant alignments: (bits) Value
At2g29210 proline-rich protein like 29 0.66
At3g07660 unknown protein 28 1.5
At4g00320 27 3.3
At3g15510 jasmonic acid regulatory protein like 27 3.3
At2g02490 hypothetical protein 27 3.3
At2g12720 putative protein 26 4.3
At4g24460 unknown protein 25 7.3
At5g55020 putative transcription factor MYB120 (MYB120) 25 9.5
At5g46120 putative protein 25 9.5
At4g24265 unknown protein 25 9.5
At4g23060 unknown protein 25 9.5
At3g24880 hypothetical protein 25 9.5
>At2g29210 proline-rich protein like
Length = 878
Score = 28.9 bits (63), Expect = 0.66
Identities = 20/76 (26%), Positives = 33/76 (43%), Gaps = 1/76 (1%)
Query: 48 PRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLRNTT 107
P P + +PP++ H S P R P +R P PP + + L + RN +
Sbjct: 343 PTPPARQRRSPSPPARRHRSPPPARRRRSPSPPARRRRSPSPPARRRRSPSPL-YRRNRS 401
Query: 108 QSDIHRTGSSDTTVRK 123
S ++R S + + K
Sbjct: 402 PSPLYRRNRSRSPLAK 417
>At3g07660 unknown protein
Length = 782
Score = 27.7 bits (60), Expect = 1.5
Identities = 16/50 (32%), Positives = 22/50 (44%), Gaps = 4/50 (8%)
Query: 60 PPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLRNTTQS 109
PP+ H + N A P+ GV+P PP TA L H + T +
Sbjct: 586 PPTMHQYLS----NGAYAQQPQASGVYPPPPGTATGGKYTLPHYKPGTNT 631
>At4g00320
Length = 773
Score = 26.6 bits (57), Expect = 3.3
Identities = 20/75 (26%), Positives = 33/75 (43%), Gaps = 7/75 (9%)
Query: 17 LASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAP 76
L +S ++L L + P CHLD +M TP + + T Q S + + +P
Sbjct: 196 LGKLSVVVKSLQRLTLKMSCP---CHLDGIMMNTPSLKYLKVTDERQESDSDNESDSDSP 252
Query: 77 GWFPEKRGVFPEPPK 91
+F + F + PK
Sbjct: 253 RYFYD----FEDMPK 263
>At3g15510 jasmonic acid regulatory protein like
Length = 364
Score = 26.6 bits (57), Expect = 3.3
Identities = 14/41 (34%), Positives = 23/41 (55%), Gaps = 1/41 (2%)
Query: 57 ASTPPSQHHHSCKPTVNRAPGWFP-EKRGVFPEPPKTALHN 96
AST QHHH+ ++N PG F G+F + T++++
Sbjct: 210 ASTGLHQHHHNVSRSMNFFPGKFSGGGYGIFSDGGNTSIYD 250
>At2g02490 hypothetical protein
Length = 415
Score = 26.6 bits (57), Expect = 3.3
Identities = 12/34 (35%), Positives = 17/34 (49%)
Query: 56 IASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEP 89
+ +P H + K +VN+ PG FP V P P
Sbjct: 112 VPPSPGHPPHQNAKISVNQYPGVFPIPHPVSPSP 145
Score = 25.4 bits (54), Expect = 7.3
Identities = 12/34 (35%), Positives = 17/34 (49%)
Query: 56 IASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEP 89
+ +P H + K +VN+ PG FP V P P
Sbjct: 83 VPPSPGHPPHQNTKISVNQYPGVFPIPHPVPPSP 116
>At2g12720 putative protein
Length = 819
Score = 26.2 bits (56), Expect = 4.3
Identities = 16/52 (30%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Query: 7 SSSAIDITTYLASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIAS 58
SS +D+ T AS A S H++++ + C L N + PC+ IA+
Sbjct: 671 SSRLLDVQTIDASRVQVAYEASLHVVNVDEKQCTCRLFN-KEKLPCIHAIAA 721
>At4g24460 unknown protein
Length = 431
Score = 25.4 bits (54), Expect = 7.3
Identities = 10/23 (43%), Positives = 17/23 (73%)
Query: 54 IMIASTPPSQHHHSCKPTVNRAP 76
+++A+TPP + H+ PTV R+P
Sbjct: 4 VLMATTPPIRCLHASIPTVFRSP 26
>At5g55020 putative transcription factor MYB120 (MYB120)
Length = 523
Score = 25.0 bits (53), Expect = 9.5
Identities = 15/54 (27%), Positives = 20/54 (36%)
Query: 23 NARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAP 76
NA++ S H L+ L P TP + PP C P N+ P
Sbjct: 199 NAKSSSSFTFHTTTANLLHPLSPHTPNTPSQLSSTPPPPPLSSPLCSPRNNQYP 252
>At5g46120 putative protein
Length = 83
Score = 25.0 bits (53), Expect = 9.5
Identities = 12/37 (32%), Positives = 15/37 (40%)
Query: 78 WFPEKRGVFPEPPKTALHNVTLLQHLRNTTQSDIHRT 114
W + EPP TA + L + S IHRT
Sbjct: 30 WLERNKRRHSEPPSTADQVIKFLDRIIRNKISSIHRT 66
>At4g24265 unknown protein
Length = 140
Score = 25.0 bits (53), Expect = 9.5
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 14/45 (31%)
Query: 61 PSQHHH-SCK-----------PTVNRAPGWFPEKRGVFPEPPKTA 93
PSQ HH SC+ PT P WF E R + PP+T+
Sbjct: 19 PSQPHHCSCEYSSTAVAVLVEPTAPPLPYWFDETRSLC--PPETS 61
>At4g23060 unknown protein
Length = 484
Score = 25.0 bits (53), Expect = 9.5
Identities = 12/37 (32%), Positives = 16/37 (42%)
Query: 45 NLMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPE 81
N +P TP + PPS H S + + P W E
Sbjct: 56 NQVPHTPSLPNSTPPPPSHHQSSPRRRRKQKPMWEDE 92
>At3g24880 hypothetical protein
Length = 1843
Score = 25.0 bits (53), Expect = 9.5
Identities = 15/43 (34%), Positives = 19/43 (43%), Gaps = 12/43 (27%)
Query: 36 QPMLVCHLDNLM-----PRTPCVIMIASTP-------PSQHHH 66
Q + H DN + P TPC I+ S+P PS H H
Sbjct: 1620 QQQMQLHSDNSIQGQSSPATPCNILSTSSPSIAPAVAPSNHQH 1662
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.131 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,820,073
Number of Sequences: 26719
Number of extensions: 114688
Number of successful extensions: 276
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 262
Number of HSP's gapped (non-prelim): 17
length of query: 123
length of database: 11,318,596
effective HSP length: 87
effective length of query: 36
effective length of database: 8,994,043
effective search space: 323785548
effective search space used: 323785548
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0140.2


