

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.12
(414 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
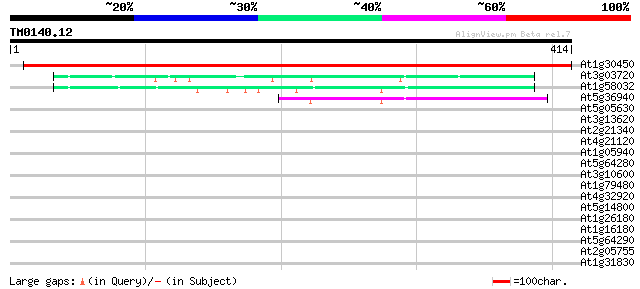
Score E
Sequences producing significant alignments: (bits) Value
At1g30450 cation-chloride co-transporter, putative 689 0.0
At3g03720 putative cationic amino acid transporter 50 2e-06
At1g58032 unknown protein 48 1e-05
At5g36940 cationic amino acid transporter -like protein 43 3e-04
At5g05630 unknown protein 37 0.020
At3g13620 unknown protein 37 0.026
At2g21340 unknown protein 36 0.034
At4g21120 amino acid transport protein AAT1 34 0.13
At1g05940 unknown protein (At1g05940) 34 0.13
At5g64280 2-oxoglutarate/malate translocator 33 0.29
At3g10600 putative amino acid transporter 32 0.64
At1g79480 hypothetical protein 31 1.1
At4g32920 unknown protein 30 3.2
At5g14800 pyrroline-5-carboxylate reductase 29 4.2
At1g26180 unknown protein (At1g26180) 29 4.2
At1g16180 29 4.2
At5g64290 2-oxoglutarate/malate translocator 29 5.4
At2g05755 unknown protein 28 9.3
At1g31830 unknown protein 28 9.3
>At1g30450 cation-chloride co-transporter, putative
Length = 1321
Score = 689 bits (1778), Expect = 0.0
Identities = 332/404 (82%), Positives = 370/404 (91%)
Query: 11 IVGMGGIGGTLLLVALCGTCTFLTAISLSAIATNGAMKGGGPYYLIGRALGPEVGVSIGL 70
IVGM GIG L+LV LCG CTFLT ISLSAIATNGAMKGGGPYYLIGRALGPEVG+SIGL
Sbjct: 158 IVGMAGIGQGLVLVFLCGLCTFLTTISLSAIATNGAMKGGGPYYLIGRALGPEVGISIGL 217
Query: 71 CFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETITQVNGTTIAQPIESPSSHDLQIYGI 130
CFFLGNAVAGALYVLGAVETFLKA PAAGIFRETIT+VNGT +++ I+SP+SHDLQ+YGI
Sbjct: 218 CFFLGNAVAGALYVLGAVETFLKAFPAAGIFRETITKVNGTAVSESIQSPNSHDLQVYGI 277
Query: 131 VVTIVLCFIVFGGVKMINRVAPAFLIPVLFSLICIYLGILLAREDHPAEGITGLSLETLK 190
VVTI+LCFIVFGGVKMINRVAPAFL+PVL S+ CI++GI LA+ D P GITGL L++ K
Sbjct: 278 VVTILLCFIVFGGVKMINRVAPAFLVPVLLSIFCIFIGIFLAKTDDPDNGITGLRLKSFK 337
Query: 191 DNWGSEYQKTNDAGIPEPDGSVSWNFNALVGLFFPAVTGIMAGSNRSSSLKDTQRSIPLG 250
DNWGS YQ TNDAGIP+P G W+FN LVGLFFPAVTGIMAGSNRS+SLKDTQ+SIP+G
Sbjct: 338 DNWGSAYQMTNDAGIPDPTGGTYWSFNELVGLFFPAVTGIMAGSNRSASLKDTQKSIPVG 397
Query: 251 TLAATLVTTFMYLVSVIMFGALATREKLLTDRLLTATVAWPFPSLIKIGIILSTMGAALQ 310
TLAATL TT +YL+SV+ FGA+ATR+KLLTDRLLTAT+AWPFP+++ +GIILST+GAALQ
Sbjct: 398 TLAATLTTTSLYLISVLFFGAVATRDKLLTDRLLTATIAWPFPAIVHVGIILSTLGAALQ 457
Query: 311 SLTGAPRLLAAIANDDILPILKYFKVADGSEPHVATLFTAFLCSGCVVIGNLDLITPTVT 370
SLTGAPRLLAAIANDDILPIL YFKVAD SEPH+ATLFTAF+C GCVVIGNLDLITPTVT
Sbjct: 458 SLTGAPRLLAAIANDDILPILNYFKVADTSEPHIATLFTAFICIGCVVIGNLDLITPTVT 517
Query: 371 MFFLLCYAGVNLSCFLLDLLDAPSWRPRWKFHHWSLSLVGALLC 414
MF+LLCY+GVNLSCFLLDLLDAPSWRPRWK+HHWSLS VGA LC
Sbjct: 518 MFYLLCYSGVNLSCFLLDLLDAPSWRPRWKYHHWSLSFVGASLC 561
>At3g03720 putative cationic amino acid transporter
Length = 614
Score = 50.4 bits (119), Expect = 2e-06
Identities = 81/400 (20%), Positives = 159/400 (39%), Gaps = 55/400 (13%)
Query: 33 LTAISLSAIATNGAMKGGGPYYLIGRALGPEVGVSIGLCFFLGNAVAGALYVLGAVETFL 92
L+A+ L AI G G G Y L+G G ++ + FF+ VA AL E
Sbjct: 28 LSAVDLVAIGV-GTTIGAGVYILVGTVAREHTGPALAVSFFIAG-VAAALSACCYAELAS 85
Query: 93 KAVPAAGIFRETIT------------------QVNGTTIAQPIESPS---SHDLQIYGIV 131
+ A + + G+ IA+ I +P+ + +L ++G
Sbjct: 86 RCPSAGSAYHYAYICLGEGIAWLVGWALVLDYTIGGSAIARGI-TPNLVFAFELYVFGFS 144
Query: 132 -----------VTIVLCFIVFGGVKMINRVAPAFLIPVLFSLICIYLGILLAREDHPAEG 180
+ + L GV ++ A LI ++ L+C + +E +
Sbjct: 145 QEASFFGGLDNLPVFLARQTIPGVGIVVDPCAALLIMIVTILLCFGI-----KESSTVQA 199
Query: 181 I-TGLSLETLKDN---WGSEYQKTNDAGIPEPDGSVSWNFNALVG---LFFPAVTGIMAG 233
I T +++ TL G KT G P G + N ++ + F + G
Sbjct: 200 IVTSVNVCTLVFIIVVGGYLACKTGWVGYDLPSGYFPFGLNGILAGSAVVFFSYIGFDTV 259
Query: 234 SNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATREKLLTDRLLTAT-----V 288
++ + +K+ QR +PLG A L+ +Y++ ++ L L D +++ +
Sbjct: 260 TSTAEEVKNPQRDLPLGIGIALLICCILYMLLSVVIVGLVPYYSLNPDTPISSAFGDSGM 319
Query: 289 AWPFPSLIKIGIILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKVADGSE-PHVATL 347
W ++ G I + + L SL PR+ A+A D +LP + +++ ++ P +T+
Sbjct: 320 QWA-AYILTTGAITALCASLLGSLLAQPRIFMAMARDGLLPAF-FSEISPRTQVPVKSTI 377
Query: 348 FTAFLCSGCVVIGNLDLITPTVTMFFLLCYAGVNLSCFLL 387
L + ++ ++ V++ L+ + V + +L
Sbjct: 378 AIGVLAAALAFFMDVAQLSEMVSVGTLMAFTAVAVCVLVL 417
>At1g58032 unknown protein
Length = 635
Score = 47.8 bits (112), Expect = 1e-05
Identities = 88/381 (23%), Positives = 150/381 (39%), Gaps = 31/381 (8%)
Query: 33 LTAISLSAIATNGAMKGGGPYYLIGRALGPEVGVSIGLCFFLGNAVAGALYVLGAVETFL 92
LT L AI GA G G Y L+G G S+ L F + AG L E
Sbjct: 44 LTVPHLVAIGV-GATIGAGVYILVGTVAREHSGPSLALSFLIAGIAAG-LSAFCYAELSS 101
Query: 93 KAVPAAGIFRETITQVNGTTIAQPIESPSSHDLQIYGIVVTIVLC---FIVFGGVKMINR 149
+ A + + V G +A I + I G V + ++FGG +
Sbjct: 102 RCPSAGSAYHYSYICV-GEGVAWIIGWALILEYTIGGSAVARGISPNLALIFGGEDGLPA 160
Query: 150 VAPAFLIPVL--------FSLICIYLGILLA--REDHPAEGIT---GLSLETLKDNWGSE 196
+ IP L L+ + G+L +E A+GI + + GS
Sbjct: 161 ILARHQIPGLDIVVDPCAAILVFVVTGLLCMGIKESTFAQGIVTAVNVCVLLFVIVAGSY 220
Query: 197 YQ-KTNDAGIPEPDG----SVSWNFNALVGLFFPAVTGIMAGSNRSSSLKDTQRSIPLGT 251
KT G P G V F +FF A G + ++ + +++ QR +P+G
Sbjct: 221 LGFKTGWPGYELPTGFFPFGVDGMFAGSATVFF-AFIGFDSVASTAEEVRNPQRDLPIGI 279
Query: 252 LAATLVTTFMYL-VSVIMFGALA----TREKLLTDRLLTATVAWPFPSLIKIGIILSTMG 306
A L+ +Y+ VS+++ G + + ++ + + W LI +G +++
Sbjct: 280 GLALLLCCSLYMMVSIVIVGLIPYYAMDPDTPISSAFASHDMQWAV-YLITLGAVMALCS 338
Query: 307 AALQSLTGAPRLLAAIANDDILPILKYFKVADGSEPHVATLFTAFLCSGCVVIGNLDLIT 366
A + +L PR+L A+A D +LP + P AT+ T + ++ +
Sbjct: 339 ALMGALLPQPRILMAMARDGLLPSIFSDINKRTQVPVKATVATGLCAATLAFFMDVSQLA 398
Query: 367 PTVTMFFLLCYAGVNLSCFLL 387
V++ LL + V +S +L
Sbjct: 399 GMVSVGTLLAFTMVAISVLIL 419
>At5g36940 cationic amino acid transporter -like protein
Length = 609
Score = 43.1 bits (100), Expect = 3e-04
Identities = 42/207 (20%), Positives = 88/207 (42%), Gaps = 9/207 (4%)
Query: 199 KTNDAGIPEPDGSVSWNFNALV---GLFFPAVTGIMAGSNRSSSLKDTQRSIPLGTLAAT 255
KT G P G + + ++ F A G ++ + +K+ +R +PLG +
Sbjct: 213 KTGWVGYELPTGYFPYGVDGMLTGSATVFFAYIGFDTVASMAEEVKNPRRDLPLGIGISL 272
Query: 256 LVTTFMYL-VSVIMFGALA----TREKLLTDRLLTATVAWPFPSLIKIGIILSTMGAALQ 310
L+ +Y+ VSV++ G + + ++ + + W LI +G +++ A +
Sbjct: 273 LLCCLLYMMVSVVIVGLVPYYAMDPDTPISSAFSSHGIQWA-AYLINLGAVMALCSALMG 331
Query: 311 SLTGAPRLLAAIANDDILPILKYFKVADGSEPHVATLFTAFLCSGCVVIGNLDLITPTVT 370
S+ PR+L A+A D +LP + P T+ T + ++ + V+
Sbjct: 332 SILPQPRILMAMARDGLLPSYFSYVNQRTQVPINGTITTGVCAAILAFFMDVSQLAGMVS 391
Query: 371 MFFLLCYAGVNLSCFLLDLLDAPSWRP 397
+ L+ + V +S ++ + P P
Sbjct: 392 VGTLVAFTMVAISLLIVRYVVPPDEVP 418
>At5g05630 unknown protein
Length = 490
Score = 37.0 bits (84), Expect = 0.020
Identities = 68/342 (19%), Positives = 127/342 (36%), Gaps = 64/342 (18%)
Query: 48 KGGGPYYLIGRALGPEVGVSIGLCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETITQ 107
+ GG + A+GP G G +L + ALY + ++ +P G +
Sbjct: 113 ENGGYVVWVTLAMGPYWGFQQGWVKWLSGVIDNALYPILFLDYLKSGIPILGSGIPRVAA 172
Query: 108 VNGTTIAQPIESPSSHDLQIYGIVVTIVLCFIVFGGVKMINRVAPAFLIPVLFSLICIYL 167
+ +V+T+ L ++ + G+ ++ A L+ V L + +
Sbjct: 173 I---------------------LVLTVALTYLNYRGLSIVG--VAAVLLGVFSILPFVVM 209
Query: 168 GILLAREDHPAEGITGLSLETLKDNWGSEYQKTNDAGIPEPDGSVSWNFNALVGLFFPAV 227
+ + P+ + +S + NW S Y T + WN N ++ +V
Sbjct: 210 SFMSIPKLKPSRWLV-VSKKMKGVNW-SLYLNT-----------LFWNLN-----YWDSV 251
Query: 228 TGIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIM-FGALATREKLLTDRLLT- 285
S + +++ +++P A L+ F Y+ V+ GA+A +KL TD
Sbjct: 252 ------STLTGEVENPSKTLPRALFYALLLVVFSYIFPVLTGTGAIALDQKLWTDGYFAD 305
Query: 286 -------ATVAWPFPSLIKIGIILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKVAD 338
+ W I+ S MG L ++ L +A +LP + + K +
Sbjct: 306 IGKVIGGVWLGW----WIQAAAATSNMGMFLAEMSSDSFQLLGMAERGMLPEV-FAKRSR 360
Query: 339 GSEPHVATLFTAFLCSGCVVIGNLDLITPTVTMFFLLCYAGV 380
P V LF+A SG +++ L L C+ V
Sbjct: 361 YRTPWVGILFSA---SGVIILSWLSFQEIVAAENLLYCFGMV 399
>At3g13620 unknown protein
Length = 478
Score = 36.6 bits (83), Expect = 0.026
Identities = 72/339 (21%), Positives = 130/339 (38%), Gaps = 63/339 (18%)
Query: 46 AMKGGGPYYLIG-RALGPEVGVSIGLCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRET 104
A G G + + RA G VG +G FL + A + + V K P
Sbjct: 87 AFPGNGGFVIWAHRAFGSFVGSMMGSLKFLSGVINVASFPVLCVTYLDKLFPV------- 139
Query: 105 ITQVNGTTIAQPIESPSSHDLQIYGIVVTIVLCFIVFGGVKMINRVAPAFLIPVLFSLIC 164
+ES ++ I+ T+VL F+ + G+ ++ A V+ L+
Sbjct: 140 ------------LESGWPRNVCIFAS--TVVLSFLNYTGLAIVGYAA------VVLGLVS 179
Query: 165 IYLGILLAREDHPAEGITGLSLETLKDN-WGSEYQKTNDAGIPEPDGSVSWNFNALVGLF 223
+ P ++ +++ +K + WGS K D + ++ WN N F
Sbjct: 180 L----------SPFLVMSAMAIPKIKPHRWGSLGTKKKDWNLYF--NTLFWNLN-----F 222
Query: 224 FPAVTGIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATREKLLTDRL 283
+ V S + + + Q++ PL L A + T YL+ + + ++ +
Sbjct: 223 WDNV------STLAGEVDEPQKTFPLALLIAVIFTCVAYLIPLFAVTGAVSVDQSRWENG 276
Query: 284 LTATVA------WPFPSLIKIGIILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKVA 337
A A W I+IG +LS++G L+ + L +A LP K+F V
Sbjct: 277 FHAEAAEMIAGKW-LKIWIEIGAVLSSIGLFEAQLSSSAYQLEGMAELGFLP--KFFGVR 333
Query: 338 DG--SEPHVATLFTAFLCSGCVVIGNLDLITPTVTMFFL 374
+ P V L +A + G + D+I+ ++ L
Sbjct: 334 SKWFNTPWVGILISALMSLGLSYMNFTDIISSANFLYTL 372
>At2g21340 unknown protein
Length = 414
Score = 36.2 bits (82), Expect = 0.034
Identities = 63/300 (21%), Positives = 113/300 (37%), Gaps = 70/300 (23%)
Query: 20 TLLLVALCGTCTFLTAISLSAIATNGAMKGGG---PYYLIG-----------RALGPEVG 65
T++ LC T FL+ + + +AT+ A + G P LIG + GP
Sbjct: 155 TVICDYLCYTFMFLSVATSNLVATSLARQIRGLAWPAVLIGWVAQSASLGMKDSWGPLKA 214
Query: 66 VSIG----------LCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETITQVNGTTIAQ 115
+++ LC FLG +AGA + T + V AA + + + + + +
Sbjct: 215 LAVASAINGVGDVVLCTFLGYGIAGAAWA-----TMVSQVVAAYMMMDALNKKGYSAFSF 269
Query: 116 PIESPSSHDLQIYGIVVTIVLCFIVFGGVKMINRVAPAFLIPVLFSLICIYLGILLARED 175
+ SPS L I+G+ + + + VLF + +Y +
Sbjct: 270 CVPSPSEL-LTIFGLAAPVFI----------------TMMSKVLFYTLLVYFATSMGTNI 312
Query: 176 HPAEGITGLSLETLKDNWGSEYQKTNDAGIPEPDGSVSWNF-NALVGLFFPAVTGIMAGS 234
A + L + T+ WG +T + +PE ++ N A V L + G
Sbjct: 313 IAAHQVM-LQIYTMSTVWGEPLSQTAQSFMPELLFGINRNLPKARVLLKSLVIIG----- 366
Query: 235 NRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATREKLLTDRLLTATVAWPFPS 294
LG + T+ T +L F + TR+K++T + + + FP+
Sbjct: 367 ------------ATLGIVVGTIGTAVPWL-----FPGIFTRDKVVTSEVYLYSFSHMFPT 409
>At4g21120 amino acid transport protein AAT1
Length = 594
Score = 34.3 bits (77), Expect = 0.13
Identities = 27/118 (22%), Positives = 49/118 (40%), Gaps = 5/118 (4%)
Query: 216 FNALVGLFFPAVTGIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATR 275
F + LFF A G A S + K+ R IP+G + + +VTT Y + + +
Sbjct: 268 FKSAAVLFF-AYIGFDAVSTMAEETKNPGRDIPIGLVGSMVVTTVCYCLMAVTLCLMQPY 326
Query: 276 EKLLTD---RLLTATVAWPFPS-LIKIGIILSTMGAALQSLTGAPRLLAAIANDDILP 329
+++ D + + V W + ++ G + L G R + IA ++P
Sbjct: 327 QQIDPDAPFSVAFSAVGWDWAKYIVAFGALKGMTTVLLVGAIGQARYMTHIARAHMMP 384
>At1g05940 unknown protein (At1g05940)
Length = 569
Score = 34.3 bits (77), Expect = 0.13
Identities = 28/110 (25%), Positives = 46/110 (41%), Gaps = 4/110 (3%)
Query: 224 FPAVTGIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATREKLLTDRL 283
F + G A +N + K+ QR +P+G + + LV +Y+ ++ + L D
Sbjct: 259 FFSYVGFDAVANSAEESKNPQRDLPIGIMGSLLVCISLYIGVCLVLTGMVPFSLLSEDAP 318
Query: 284 LT---ATVAWPFPS-LIKIGIILSTMGAALQSLTGAPRLLAAIANDDILP 329
L ++ F S LI IG + L L RL + D +LP
Sbjct: 319 LAEAFSSKGMKFVSILISIGAVAGLTTTLLVGLYVQSRLYLGLGRDGLLP 368
>At5g64280 2-oxoglutarate/malate translocator
Length = 549
Score = 33.1 bits (74), Expect = 0.29
Identities = 37/149 (24%), Positives = 66/149 (43%), Gaps = 19/149 (12%)
Query: 205 IPEPDGSVSWNFNALVGLFFPAVTGIMAGS---------NRSSSLKDTQRSIPLGTLAAT 255
IP P+ S + L+ +F ++G++ G ++S+ +++P T A
Sbjct: 99 IPRPEQVTSQGWQ-LLSIFLFTISGLVLGPLPVGAWAFIGLTASI--VTKTLPFSTAFAA 155
Query: 256 LVTTFMYLVSVIMFGALATREKLLTDRLLTATVAWPFPSLIKIGIILSTMGAALQSLTG- 314
++L+++ F A + L DR+ T V W L K + LS A ++L G
Sbjct: 156 FTNELIWLIAISFFFARGFIKTGLGDRIATYFVKW----LGKSTLGLSYGLAFCETLMGL 211
Query: 315 -APRLLAAIANDDILPILKYFKVADGSEP 342
P +A A LP++K ++ GS P
Sbjct: 212 IMPSTMAR-AGGVFLPVIKSLAISAGSYP 239
>At3g10600 putative amino acid transporter
Length = 584
Score = 32.0 bits (71), Expect = 0.64
Identities = 77/365 (21%), Positives = 130/365 (35%), Gaps = 39/365 (10%)
Query: 45 GAMKGGGPYYLIGRA----LGPEVGVSI---GLCFFLG----NAVAGALYVLGAVETFLK 93
G M G G + GRA GP + VS GLC L A L V G ++++
Sbjct: 71 GGMIGAGVFVTTGRASRLYAGPSIVVSYAIAGLCALLSAFCYTEFAVHLPVAGGAFSYIR 130
Query: 94 AVPAAGIFRETITQVNGTTIAQPIESPSSHDLQIY-----GIVVTIVLCFIVFGGVKMIN 148
G F IT N + S Y GI T FIV G N
Sbjct: 131 IT--FGEFPAFITGANLIMDYVLSNAAVSRGFTAYLGSAFGIS-TSEWRFIVSGLPNGFN 187
Query: 149 RVAPAFLIPVLFSLICIYLGILLAREDHPAEGI-TGLSLETLKDNWGSEYQKTNDAGIPE 207
+ P +I VL I RE + T L + + + K + +
Sbjct: 188 EIDPIAVIVVLAVTFVICYS---TRESSKVNMVLTALHIAFIVFVIVMGFSKGDVKNLTR 244
Query: 208 PDG----------SVSWNFNALVGLFFPAVTGIMAGSNRSSSLKDTQRSIPLGTLAATLV 257
PD VS FN ++ + G A S + +KD + IP+G + +
Sbjct: 245 PDNPENPSGFFPFGVSGVFNGAAMVYLSYI-GYDAVSTMAEEVKDPVKDIPMGISGSVAI 303
Query: 258 TTFMYLVSVIMFGALATREKLLTDRLLTATVA----WPFPS-LIKIGIILSTMGAALQSL 312
+Y + I L + + + +A + W + + ++ IG + + + ++
Sbjct: 304 VIVLYCLMAISMSMLLPYDLIDAEAPYSAAFSKSEGWEWVTRVVGIGASFGILTSLIVAM 363
Query: 313 TGAPRLLAAIANDDILPILKYFKVADGSEPHVATLFTAFLCSGCVVIGNLDLITPTVTMF 372
G R + I ++PI S P A+ F + + +L+++ V++
Sbjct: 364 LGQARYMCVIGRSRVVPIWFAKVHPKTSTPVNASAFLGIFTAVLALFTDLNVLLNLVSIG 423
Query: 373 FLLCY 377
L +
Sbjct: 424 TLFVF 428
>At1g79480 hypothetical protein
Length = 356
Score = 31.2 bits (69), Expect = 1.1
Identities = 15/45 (33%), Positives = 28/45 (61%), Gaps = 3/45 (6%)
Query: 92 LKAVPAAGIFRETITQVNGTTIAQPIESPSSHDLQIYGIVVTIVL 136
+KA+ + ++ +TQ+N +AQP++ SS + Q YG+ T+ L
Sbjct: 2 IKALKHTSLLQKMMTQLN---LAQPLDYSSSSNTQPYGVSTTLTL 43
>At4g32920 unknown protein
Length = 1365
Score = 29.6 bits (65), Expect = 3.2
Identities = 17/37 (45%), Positives = 22/37 (58%), Gaps = 2/37 (5%)
Query: 15 GGIGGTLLLVALCGTCTFLTAISLSAIATNGAMKGGG 51
GG GGT+LL T + LS+I NG++KGGG
Sbjct: 639 GGSGGTVLL--FLRTLEIGRSAILSSIGGNGSLKGGG 673
>At5g14800 pyrroline-5-carboxylate reductase
Length = 276
Score = 29.3 bits (64), Expect = 4.2
Identities = 22/69 (31%), Positives = 33/69 (46%), Gaps = 2/69 (2%)
Query: 47 MKGGGPYYL-IGRALGPEVGVSIGLCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETI 105
+ G GP Y+ + + GV+ GL L ++A VLGA K G+ ++ +
Sbjct: 178 LSGSGPAYIFLAIEALADGGVAAGLPRELALSLASQT-VLGAATMVSKTGKHPGVLKDDV 236
Query: 106 TQVNGTTIA 114
T GTTIA
Sbjct: 237 TSPGGTTIA 245
>At1g26180 unknown protein (At1g26180)
Length = 289
Score = 29.3 bits (64), Expect = 4.2
Identities = 35/119 (29%), Positives = 57/119 (47%), Gaps = 15/119 (12%)
Query: 55 LIGRALGP--EVGVSIGLCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETITQVNGTT 112
L+G ALG +G S L FL V AL ++G V T ++ + RE I V
Sbjct: 168 LVGLALGSLWSMGASWRLSIFLCTMVR-ALGLIGYVLT------SSFLIRENILAV---- 216
Query: 113 IAQPIESPSSHDLQIYG--IVVTIVLCFIVFGGVKMINRVAPAFLIPVLFSLICIYLGI 169
I I + S+ G I+ ++ L +++FG V ++N L+ +L+S+ LG+
Sbjct: 217 ITINIHASLSYVFTAMGLNIMPSMSLIYMIFGTVLLLNSGFFVLLLHLLYSIFLTRLGM 275
>At1g16180
Length = 412
Score = 29.3 bits (64), Expect = 4.2
Identities = 21/68 (30%), Positives = 34/68 (49%), Gaps = 6/68 (8%)
Query: 119 SPSSHD--LQIYGIVVTIVLCFIVFGGVKMINRVAPAFLIPVLFSLICIYL---GILLAR 173
+PS HD L + I++T++ F VF V + V + L + SL C+YL G+
Sbjct: 206 TPSGHDCGLNTFFIIMTLIFVF-VFAIVVLHPTVGGSILPASVISLYCMYLCYSGLASEP 264
Query: 174 EDHPAEGI 181
D+ G+
Sbjct: 265 RDYECNGL 272
>At5g64290 2-oxoglutarate/malate translocator
Length = 563
Score = 28.9 bits (63), Expect = 5.4
Identities = 39/182 (21%), Positives = 74/182 (40%), Gaps = 14/182 (7%)
Query: 205 IPEPDGSVSWNFNALVGLFFPAVTGIM-------AGSNRSSSLKDTQRSIPLGTLAATLV 257
+P P+G + L+ +F + G++ A + + +++ +
Sbjct: 113 VPVPEGVTPQGWQ-LLSIFLSTIAGLVLSPLPVGAWAFIGLTASIVTKTLSFSAAFSAFT 171
Query: 258 TTFMYLVSVIMFGALATREKLLTDRLLTATVAWPFPSLIKIGIILSTMGAALQSLTGAPR 317
+ ++L+ + F A + L DR+ T V W S + + L T+ AL + A
Sbjct: 172 SEVIWLIVISFFFARGFVKTGLGDRIATYFVKWLGKSTLGLSYGL-TLSEAL--IAPAMP 228
Query: 318 LLAAIANDDILPILKYFKVADGSEPHVAT---LFTAFLCSGCVVIGNLDLITPTVTMFFL 374
A A LPI+K ++ GS+P+ ++ L + + S GN + T L
Sbjct: 229 STTARAGGIFLPIIKSLSLSAGSKPNDSSSRKLGSYLIQSQFQCAGNSSALFLTAAAQNL 288
Query: 375 LC 376
LC
Sbjct: 289 LC 290
>At2g05755 unknown protein
Length = 401
Score = 28.1 bits (61), Expect = 9.3
Identities = 32/120 (26%), Positives = 51/120 (41%), Gaps = 17/120 (14%)
Query: 219 LVGLFFPAVTG------IMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMY---------- 262
L+GLF ++TG I A + S T S L AT + F +
Sbjct: 256 LLGLF-SSITGGITYCLIKAAAKASEQPVITVLSFGLVACPATAICMFSFESFVLPAFDT 314
Query: 263 LVSVIMFGALATREKLLTDRLLTATVAWPFPSLIKIGIILSTMGAALQSLTGAPRLLAAI 322
LVS+I+ G LA ++L R L +++ I ++LS + TG+P L + +
Sbjct: 315 LVSMIVLGLLAFCAEVLLARGLQLEKISKAANVLYIEVVLSQLWLVSTGKTGSPGLFSRL 374
>At1g31830 unknown protein
Length = 495
Score = 28.1 bits (61), Expect = 9.3
Identities = 23/104 (22%), Positives = 41/104 (39%), Gaps = 21/104 (20%)
Query: 48 KGGGPYYLIGRALGPEVGVSIGLCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETITQ 107
+ GG + ALGP G G +L + ALY + ++ VPA G +
Sbjct: 109 ENGGYVVWVSSALGPFWGFQQGWMKWLSGVIDNALYPVLFLDYLKSGVPALGSGLPRVAS 168
Query: 108 VNGTTIAQPIESPSSHDLQIYGIVVTIVLCFIVFGGVKMINRVA 151
+ +V+TI+L ++ + G+ ++ VA
Sbjct: 169 I---------------------LVLTILLTYLNYRGLTIVGWVA 191
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.142 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,840,459
Number of Sequences: 26719
Number of extensions: 365057
Number of successful extensions: 1127
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 1111
Number of HSP's gapped (non-prelim): 27
length of query: 414
length of database: 11,318,596
effective HSP length: 102
effective length of query: 312
effective length of database: 8,593,258
effective search space: 2681096496
effective search space used: 2681096496
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0140.12


