

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0137.5
(214 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
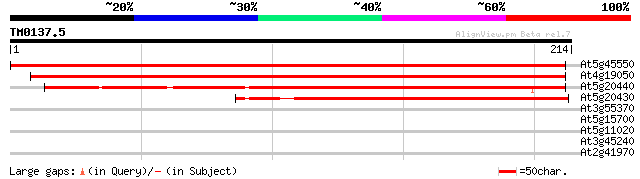
Score E
Sequences producing significant alignments: (bits) Value
At5g45550 unknown protein 406 e-114
At4g19050 putative protein 389 e-109
At5g20440 putative protein 248 2e-66
At5g20430 putative protein 170 6e-43
At3g55370 zinc finger protein OBP3 28 4.7
At5g15700 DNA-directed RNA polymerase (mitochondrial) 27 6.2
At5g11020 ser/thr specific protein kinase-like protein 27 8.1
At3g45240 protein kinase like protein 27 8.1
At2g41970 putative protein kinase 27 8.1
>At5g45550 unknown protein
Length = 215
Score = 406 bits (1043), Expect = e-114
Identities = 191/212 (90%), Positives = 205/212 (96%)
Query: 1 MSLFGLGRNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVN 60
MSLFGLGRNQKTFRPKKSAPSGSKGAQL+KHIDATLGSGNLREAV+LPPGED NEWLAVN
Sbjct: 1 MSLFGLGRNQKTFRPKKSAPSGSKGAQLRKHIDATLGSGNLREAVRLPPGEDANEWLAVN 60
Query: 61 TVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMD 120
TVDFFNQVN+L+GTLTEFCTP NCP+MTAGPKYEYRWADGV IKKPIEVSAPKYVEYLMD
Sbjct: 61 TVDFFNQVNLLYGTLTEFCTPDNCPTMTAGPKYEYRWADGVQIKKPIEVSAPKYVEYLMD 120
Query: 121 WIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN 180
WIE+QLDDET+FPQ+LGAPFP NF+DVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN
Sbjct: 121 WIETQLDDETLFPQRLGAPFPQNFKDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN 180
Query: 181 TCFKHFVLFTWEFRLIDKAELAPLEDLVDSII 212
TCFKHF+LFT EF LIDK ELAPL++L++SII
Sbjct: 181 TCFKHFILFTHEFGLIDKKELAPLQELIESII 212
>At4g19050 putative protein
Length = 1405
Score = 389 bits (998), Expect = e-109
Identities = 183/204 (89%), Positives = 197/204 (95%)
Query: 9 NQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQV 68
NQKTFRPKKSAPSG+KGA+L+KHIDATLGSGNLREAVKLPPGED+NEWLAVNTVDFFNQV
Sbjct: 1199 NQKTFRPKKSAPSGTKGAELRKHIDATLGSGNLREAVKLPPGEDLNEWLAVNTVDFFNQV 1258
Query: 69 NILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDD 128
N+LFGTLTEFCTP NC +MTAGPKYEYRWADGV IKKPIEVSAPKYVEYLMDWIE+QLDD
Sbjct: 1259 NLLFGTLTEFCTPENCSTMTAGPKYEYRWADGVQIKKPIEVSAPKYVEYLMDWIETQLDD 1318
Query: 129 ETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVL 188
ETIFPQKLGA FPPNF++VVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHF+L
Sbjct: 1319 ETIFPQKLGAAFPPNFKEVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFIL 1378
Query: 189 FTWEFRLIDKAELAPLEDLVDSII 212
FT EF LIDK ELAPL++L++SII
Sbjct: 1379 FTHEFVLIDKKELAPLQELIESII 1402
>At5g20440 putative protein
Length = 217
Score = 248 bits (633), Expect = 2e-66
Identities = 118/200 (59%), Positives = 157/200 (78%), Gaps = 5/200 (2%)
Query: 14 RPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQVNILFG 73
+ +K SKG+Q+++ I + S NLREAV+LP G DINEW A+N +FFNQ+++L+
Sbjct: 20 KKRKHPEYKSKGSQIRELISG-IRSDNLREAVRLPQGVDINEWFAMN--NFFNQISLLYA 76
Query: 74 TLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFP 133
TL EFCT + CP M AG +YEYRWADG TI KP VSAPKYVEYL+DW+E+++D+E IFP
Sbjct: 77 TLEEFCTQTTCPVMNAG-RYEYRWADGTTITKPKTVSAPKYVEYLIDWVETEIDNEAIFP 135
Query: 134 QKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVLFTWEF 193
+ G PFPPNF D VK I ++LFRVYAHIY+SHF +IV+L E+AHLNTCFKHF+LF EF
Sbjct: 136 KNPGEPFPPNFEDFVKRILRKLFRVYAHIYYSHFHEIVALNEQAHLNTCFKHFLLFVSEF 195
Query: 194 RLIDK-AELAPLEDLVDSII 212
+L+DK E+AP++ LV++++
Sbjct: 196 QLVDKEKEMAPIKSLVETML 215
>At5g20430 putative protein
Length = 122
Score = 170 bits (430), Expect = 6e-43
Identities = 79/127 (62%), Positives = 102/127 (80%), Gaps = 6/127 (4%)
Query: 87 MTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLGAPFPPNFRD 146
M AG +YEYRWADG T+ VSAP+YVE LM+WIE+Q+D+E IFP+K G PFPPNF D
Sbjct: 1 MKAG-RYEYRWADGTTM-----VSAPEYVELLMNWIETQIDNEHIFPKKTGEPFPPNFED 54
Query: 147 VVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVLFTWEFRLIDKAELAPLED 206
VK I ++LFRVYAHIYHSHF KIV+L E+AHLNTCF ++LF EF+L+DK E+ P++
Sbjct: 55 FVKRILRKLFRVYAHIYHSHFPKIVTLNEQAHLNTCFHRYLLFVSEFQLVDKEEMVPIQK 114
Query: 207 LVDSIIQ 213
LV++I++
Sbjct: 115 LVETILK 121
>At3g55370 zinc finger protein OBP3
Length = 323
Score = 27.7 bits (60), Expect = 4.7
Identities = 17/77 (22%), Positives = 32/77 (41%), Gaps = 1/77 (1%)
Query: 25 GAQLQKHIDATLGSGNLREAVKLPPGEDINEW-LAVNTVDFFNQVNILFGTLTEFCTPSN 83
G Q+ I SG + +A ++PP + ++ +NT N L+ L + +
Sbjct: 200 GTQISNMISGMSSSGGILDAWRIPPSQQAQQFPFLINTTGLVQSSNALYPLLEGGVSATQ 259
Query: 84 CPSMTAGPKYEYRWADG 100
++ A + R DG
Sbjct: 260 TRNVKAEENDQDRGRDG 276
>At5g15700 DNA-directed RNA polymerase (mitochondrial)
Length = 1011
Score = 27.3 bits (59), Expect = 6.2
Identities = 14/63 (22%), Positives = 26/63 (41%), Gaps = 2/63 (3%)
Query: 84 CPSMTAGPKYEYRWAD--GVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLGAPFP 141
C + A RW G+ + +P K V+ + + Q + + + ++ FP
Sbjct: 854 CAKIIASENETVRWTTPLGLPVVQPYHQMGTKLVKTSLQTLSLQHETDQVIVRRQRTAFP 913
Query: 142 PNF 144
PNF
Sbjct: 914 PNF 916
>At5g11020 ser/thr specific protein kinase-like protein
Length = 402
Score = 26.9 bits (58), Expect = 8.1
Identities = 18/57 (31%), Positives = 27/57 (46%), Gaps = 6/57 (10%)
Query: 104 KKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYA 160
KKP+E AP + ++ W L D T KL + P +D + K L++V A
Sbjct: 306 KKPVEKLAPGECQSIITWAMPYLTDRT----KLPSVIDPAIKDTMD--LKHLYQVAA 356
>At3g45240 protein kinase like protein
Length = 396
Score = 26.9 bits (58), Expect = 8.1
Identities = 19/72 (26%), Positives = 30/72 (41%), Gaps = 4/72 (5%)
Query: 78 FCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLG 137
F P C +T + WA GVT+ I P + L D + + + I P+ L
Sbjct: 280 FTAPECCLGITYSGRSADTWAVGVTLYCMILGQYPFLGDTLQDTYDKIVHNPLIIPEGLN 339
Query: 138 APFPPNFRDVVK 149
P RD+++
Sbjct: 340 ----PRLRDLIE 347
>At2g41970 putative protein kinase
Length = 365
Score = 26.9 bits (58), Expect = 8.1
Identities = 13/45 (28%), Positives = 24/45 (52%), Gaps = 3/45 (6%)
Query: 101 VTIKKPIEVSAPKYVEYLMDWIESQLDDETI---FPQKLGAPFPP 142
+T +KP++ + PK + L+ W +L ++ + KL FPP
Sbjct: 275 LTGRKPVDHTMPKGQQSLVTWATPRLSEDKVKQCIDPKLNNDFPP 319
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.139 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,056,636
Number of Sequences: 26719
Number of extensions: 215007
Number of successful extensions: 468
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 461
Number of HSP's gapped (non-prelim): 9
length of query: 214
length of database: 11,318,596
effective HSP length: 95
effective length of query: 119
effective length of database: 8,780,291
effective search space: 1044854629
effective search space used: 1044854629
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0137.5


