

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.7
(841 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
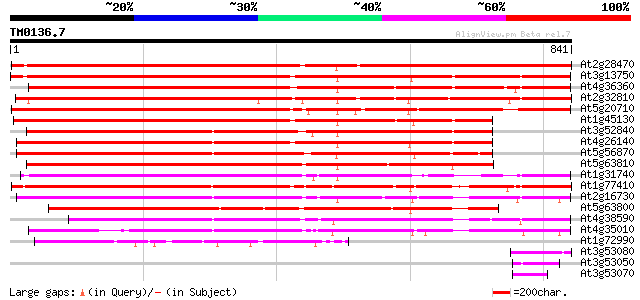
Score E
Sequences producing significant alignments: (bits) Value
At2g28470 beta-galactosidase like protein 1235 0.0
At3g13750 galactosidase, putative 936 0.0
At4g36360 beta-galactosidase like protein 926 0.0
At2g32810 putative beta-galactosidase (BGAL9) 915 0.0
At5g20710 beta-galactosidase like protein 827 0.0
At1g45130 beta-galactosidase like protein 823 0.0
At3g52840 beta-galactosidase precursor - like protein 815 0.0
At4g26140 putative beta-galactosidase 815 0.0
At5g56870 beta-galactosidase (emb|CAB64740.1) 813 0.0
At5g63810 beta-galactosidase (emb|CAB64746.1) 781 0.0
At1g31740 putative protein 744 0.0
At1g77410 unknown protein 726 0.0
At2g16730 putative beta-galactosidase 650 0.0
At5g63800 beta-galactosidase like protein 613 e-176
At4g38590 galactosidase like protein 547 e-155
At4g35010 beta-galactosidase - like protein 544 e-155
At1g72990 unknown protein 161 2e-39
At3g53080 unknown protein 73 8e-13
At3g53050 putative protein 49 9e-06
At3g53070 putative protein 48 3e-05
>At2g28470 beta-galactosidase like protein
Length = 839
Score = 1235 bits (3196), Expect = 0.0
Identities = 601/848 (70%), Positives = 691/848 (80%), Gaps = 21/848 (2%)
Query: 3 MRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDL 62
+R+ + +LLL+ ++ V A A ANVTYDHRALVIDGKR+VLISGSIHYPRSTPEMWP+L
Sbjct: 4 VRKMEMILLLILVIVVAATA---ANVTYDHRALVIDGKRKVLISGSIHYPRSTPEMWPEL 60
Query: 63 IQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACA 122
IQK+KDGGLDVIETYVFW+ HEP + +YNFEGR DLVKFVK A AGLYVHLRIGPY CA
Sbjct: 61 IQKSKDGGLDVIETYVFWSGHEPEKNKYNFEGRYDLVKFVKLAAKAGLYVHLRIGPYVCA 120
Query: 123 EWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIE 182
EWNYGGFP+WLHF+PGI+FRTDNEPFK EMQRFT KIVD+MKQE LYASQGGPIILSQIE
Sbjct: 121 EWNYGGFPVWLHFVPGIKFRTDNEPFKEEMQRFTTKIVDLMKQEKLYASQGGPIILSQIE 180
Query: 183 NEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTP 242
NEYGN++ AYG AA YI W+ASMA SLDTGVPW MCQQ +APDP+INTCNGFYCDQFTP
Sbjct: 181 NEYGNIDSAYGAAAKSYIKWSASMALSLDTGVPWNMCQQTDAPDPMINTCNGFYCDQFTP 240
Query: 243 NSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGR 302
NS KPK WTE ++GW L FG PYRPVEDLAFAVARF+Q GGT QNYYMYHGGTNF R
Sbjct: 241 NSNNKPKMWTENWSGWFLGFGDPSPYRPVEDLAFAVARFYQRGGTFQNYYMYHGGTNFDR 300
Query: 303 TSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLE 362
TSGGP ++TSYDYDA IDEYG +RQPKWGHL+D+HKAIKLCE+ALIATDPTITSLGSNLE
Sbjct: 301 TSGGPLISTSYDYDAPIDEYGLLRQPKWGHLRDLHKAIKLCEDALIATDPTITSLGSNLE 360
Query: 363 AAVYKTEA-ECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSES 421
AAVYKTE+ C AFLAN+D SDATV FNG SYNLPAWSVSILPDCKNV NTAK+ S
Sbjct: 361 AAVYKTESGSCAAFLANVDTKSDATVTFNGKSYNLPAWSVSILPDCKNVAFNTAKVKFNS 420
Query: 422 MISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYS 481
+ + S E+GS WS+I EPIGISKAD+F + GLLEQINTTAD+SDYLWYS
Sbjct: 421 ISKTPDGGSSAELGS------QWSYIKEPIGISKADAFLKPGLLEQINTTADKSDYLWYS 474
Query: 482 LRIDLED-----DAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVA 536
LR D++ D G++ VLHIESLG ++AFINGKLAGSG G + K++++IPI LV
Sbjct: 475 LRTDIKGDETFLDEGSKAVLHIESLGQVVYAFINGKLAGSGHGKQ---KISLDIPINLVT 531
Query: 537 GKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGED 596
G NTIDLLS+TVGL ++G FFD GAGITGPV LK K GS++D +S+QWTYQ+GLKGED
Sbjct: 532 GTNTIDLLSVTVGLANYGAFFDLVGAGITGPVTLKSAKGGSSIDLASQQWTYQVGLKGED 591
Query: 597 LGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGR 656
GL + S +W S+S LP QPL WYK F APSG++P+AIDFTG GKG AWVNGQSIGR
Sbjct: 592 TGLATVDSSEWVSKSPLPTKQPLIWYKTTFDAPSGSEPVAIDFTGTGKGIAWVNGQSIGR 651
Query: 657 YWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESG 716
YWPT I+ GCT SC+YRG Y ++KCLKNCGKPSQTLYHVPRSWL+P N LVLFEE G
Sbjct: 652 YWPTSIAGNGGCTESCDYRGSYRANKCLKNCGKPSQTLYHVPRSWLKPSGNILVLFEEMG 711
Query: 717 GDPTQISIAIKLIGSS-CSHVSESHPPPVDLWNSDTE-SDRS-GGPVLSLECPYPNEVIT 773
GDPTQIS A K GS+ C VS+SHPPPVD W SD++ S+R+ PVLSL+CP +VI
Sbjct: 712 GDPTQISFATKQTGSNLCLTVSQSHPPPVDTWTSDSKISNRNRTRPVLSLKCPISTQVIF 771
Query: 774 TIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTFGDPCGGVTKS 833
+IKFASFGTP G CG+F+ G C+S ++LS+VQKACIG SC++ VS FG+PC GV KS
Sbjct: 772 SIKFASFGTPKGTCGSFTQGHCNSSRSLSLVQKACIGLRSCNVEVSTRVFGEPCRGVVKS 831
Query: 834 LAVEASCT 841
LAVEASC+
Sbjct: 832 LAVEASCS 839
>At3g13750 galactosidase, putative
Length = 847
Score = 936 bits (2420), Expect = 0.0
Identities = 466/851 (54%), Positives = 583/851 (67%), Gaps = 27/851 (3%)
Query: 1 MAMRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWP 60
+AM L LL +L+C + +V+YD RA+ I+GKRR+LISGSIHYPRSTPEMWP
Sbjct: 12 VAMAAVSALFLLGFLVCSVS-----GSVSYDSRAITINGKRRILISGSIHYPRSTPEMWP 66
Query: 61 DLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYA 120
DLI+KAK+GGLDVI+TYVFWN HEP G+Y FEG DLVKFVK V +GLY+HLRIGPY
Sbjct: 67 DLIRKAKEGGLDVIQTYVFWNGHEPSPGKYYFEGNYDLVKFVKLVQQSGLYLHLRIGPYV 126
Query: 121 CAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQ 180
CAEWN+GGFP+WL +IPGI FRTDN PFKA+MQRFT KIV+MMK E L+ SQGGPIILSQ
Sbjct: 127 CAEWNFGGFPVWLKYIPGISFRTDNGPFKAQMQRFTTKIVNMMKAERLFESQGGPIILSQ 186
Query: 181 IENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQF 240
IENEYG +E G Y NWAA MA L TGVPWVMC+Q++APDPIIN CNGFYCD F
Sbjct: 187 IENEYGPMEYELGAPGRSYTNWAAKMAVGLGTGVPWVMCKQDDAPDPIINACNGFYCDYF 246
Query: 241 TPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNF 300
+PN KPK WTE + GW FGG VPYRP ED+AF+VARF Q GG+ NYYMYHGGTNF
Sbjct: 247 SPNKAYKPKMWTEAWTGWFTKFGGPVPYRPAEDMAFSVARFIQKGGSFINYYMYHGGTNF 306
Query: 301 GRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSN 360
GRT+GGPF+ATSYDYDA +DEYG RQPKWGHLKD+H+AIKLCE AL++ +PT LG+
Sbjct: 307 GRTAGGPFIATSYDYDAPLDEYGLERQPKWGHLKDLHRAIKLCEPALVSGEPTRMPLGNY 366
Query: 361 LEAAVYKTEA-ECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINS 419
EA VYK+++ C AFLAN + S A V F N YNLP WS+SILPDCKN V NTA++ +
Sbjct: 367 QEAHVYKSKSGACSAFLANYNPKSYAKVSFGNNHYNLPPWSISILPDCKNTVYNTARVGA 426
Query: 420 ESMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLW 479
+ T +K V W +E +SF+ GL+EQINTT D SDYLW
Sbjct: 427 Q-------TSRMKMVRVPVHGGLSWQAYNEDPSTYIDESFTMVGLVEQINTTRDTSDYLW 479
Query: 480 Y--SLRIDLEDD---AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITL 534
Y +++D + G L + S GHA+H FING+L+GS G+ D+ K+ + L
Sbjct: 480 YMTDVKVDANEGFLRNGDLPTLTVLSAGHAMHVFINGQLSGSAYGSLDSPKLTFRKGVNL 539
Query: 535 VAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKG 594
AG N I +LS+ VGL + GP F+TW AG+ GPV L GL NG D S ++WTY++GLKG
Sbjct: 540 RAGFNKIAILSIAVGLPNVGPHFETWNAGVLGPVSLNGL-NGGRRDLSWQKWTYKVGLKG 598
Query: 595 EDLGLPS---GTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNG 651
E L L S +S +W + + + QPLTWYK FSAP+G+ P+A+D MGKG+ W+NG
Sbjct: 599 ESLSLHSLSGSSSVEWAEGAFVAQKQPLTWYKTTFSAPAGDSPLAVDMGSMGKGQIWING 658
Query: 652 QSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVL 711
QS+GR+WP Y + G S C+Y G + KCL+NCG+ SQ YHVPRSWL+P N LV+
Sbjct: 659 QSLGRHWPAYKAV--GSCSECSYTGTFREDKCLRNCGEASQRWYHVPRSWLKPSGNLLVV 716
Query: 712 FEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDL-WNSDTESDRSGGPVLSLECPYPNE 770
FEE GGDP I++ + + S C+ + E V+ ++ + ++ P L+C P +
Sbjct: 717 FEEWGGDPNGITLVRREVDSVCADIYEWQSTLVNYQLHASGKVNKPLHPKAHLQCG-PGQ 775
Query: 771 VITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTF-GDPCGG 829
ITT+KFASFGTP G CG++ G C + + K C+G + CS+ V+ F GDPC
Sbjct: 776 KITTVKFASFGTPEGTCGSYRQGSCHAHHSYDAFNKLCVGQNWCSVTVAPEMFGGDPCPN 835
Query: 830 VTKSLAVEASC 840
V K LAVEA C
Sbjct: 836 VMKKLAVEAVC 846
>At4g36360 beta-galactosidase like protein
Length = 856
Score = 926 bits (2392), Expect = 0.0
Identities = 447/829 (53%), Positives = 577/829 (68%), Gaps = 32/829 (3%)
Query: 28 VTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQ 87
VTYD +AL+I+G+RR+L SGSIHYPRSTP+MW DLIQKAKDGG+DVIETYVFWNLHEP
Sbjct: 33 VTYDRKALLINGQRRILFSGSIHYPRSTPDMWEDLIQKAKDGGIDVIETYVFWNLHEPSP 92
Query: 88 GQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNEP 147
G+Y+FEGR DLV+FVK + AGLY HLRIGPY CAEWN+GGFP+WL ++PGI FRTDNEP
Sbjct: 93 GKYDFEGRNDLVRFVKTIHKAGLYAHLRIGPYVCAEWNFGGFPVWLKYVPGISFRTDNEP 152
Query: 148 FKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMA 207
FK M+ FT +IV++MK ENL+ SQGGPIILSQIENEYG G Y+ WAA MA
Sbjct: 153 FKRAMKGFTERIVELMKSENLFESQGGPIILSQIENEYGRQGQLLGAEGHNYMTWAAKMA 212
Query: 208 TSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYNGWLLYFGGAVP 267
+ +TGVPWVMC++++APDP+INTCNGFYCD F PN KP WTE ++GW FGG +
Sbjct: 213 IATETGVPWVMCKEDDAPDPVINTCNGFYCDSFAPNKPYKPLIWTEAWSGWFTEFGGPMH 272
Query: 268 YRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFIRQ 327
+RPV+DLAF VARF Q GG+ NYYMYHGGTNFGRT+GGPFV TSYDYDA IDEYG IRQ
Sbjct: 273 HRPVQDLAFGVARFIQKGGSFVNYYMYHGGTNFGRTAGGPFVTTSYDYDAPIDEYGLIRQ 332
Query: 328 PKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTEA-ECVAFLANIDNTSDAT 386
PK+GHLK++H+AIK+CE+AL++ DP +TS+G+ +A VY E+ +C AFLAN D S A
Sbjct: 333 PKYGHLKELHRAIKMCEKALVSADPVVTSIGNKQQAHVYSAESGDCSAFLANYDTESAAR 392
Query: 387 VKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGW-S 445
V FN YNLP WS+SILPDC+N V NTAK+ ++ S E+ + + W S
Sbjct: 393 VLFNNVHYNLPPWSISILPDCRNAVFNTAKVGVQT--------SQMEMLPTDTKNFQWES 444
Query: 446 WISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDD-----AGAQTVLHIES 500
++ + + + +F+ GLLEQIN T D SDYLWY +D+ D G L I+S
Sbjct: 445 YLEDLSSLDDSSTFTTHGLLEQINVTRDTSDYLWYMTSVDIGDSESFLHGGELPTLIIQS 504
Query: 501 LGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTW 560
GHA+H F+NG+L+GS G R N + + I L +G N I LLS+ VGL + G F++W
Sbjct: 505 TGHAVHIFVNGQLSGSAFGTRQNRRFTYQGKINLHSGTNRIALLSVAVGLPNVGGHFESW 564
Query: 561 GAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGE--DLGLPSGTS--GQWNSQSTLPKN 616
GI GPV L GL G +D S ++WTYQ+GLKGE +L P+ T G ++ T+ K
Sbjct: 565 NTGILGPVALHGLSQGK-MDLSWQKWTYQVGLKGEAMNLAFPTNTPSIGWMDASLTVQKP 623
Query: 617 QPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRG 676
QPLTW+K F AP GN+P+A+D GMGKG+ WVNG+SIGRYW + + G S C+Y G
Sbjct: 624 QPLTWHKTYFDAPEGNEPLALDMEGMGKGQIWVNGESIGRYWTAFAT---GDCSHCSYTG 680
Query: 677 PYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHV 736
Y +KC CG+P+Q YHVPR+WL+P N LV+FEE GG+P+ +S+ + + C+ V
Sbjct: 681 TYKPNKCQTGCGQPTQRWYHVPRAWLKPSQNLLVIFEELGGNPSTVSLVKRSVSGVCAEV 740
Query: 737 SESHPPPVDLWNSDTESDRSG----GPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSH 792
SE HP ++ N ES G P + L+C P + I +IKFASFGTP G CG++
Sbjct: 741 SEYHP---NIKNWQIESYGKGQTFHRPKVHLKCS-PGQAIASIKFASFGTPLGTCGSYQQ 796
Query: 793 GDCSSKQALSIVQKACIGSSSCSIGVSINTFG-DPCGGVTKSLAVEASC 840
G+C + + +I+++ C+G + C++ +S + FG DPC V K L VEA C
Sbjct: 797 GECHAATSYAILERKCVGKARCAVTISNSNFGKDPCPNVLKRLTVEAVC 845
>At2g32810 putative beta-galactosidase (BGAL9)
Length = 887
Score = 915 bits (2364), Expect = 0.0
Identities = 463/873 (53%), Positives = 580/873 (66%), Gaps = 49/873 (5%)
Query: 9 LLLLLWLLCVYAPACFCA-----NVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLI 63
+L L+ L VY P + NV+YDHRAL+I GKRR+L+S IHYPR+TPEMW DLI
Sbjct: 14 ILSLIIALLVYFPILSGSYFKPFNVSYDHRALIIAGKRRMLVSAGIHYPRATPEMWSDLI 73
Query: 64 QKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAE 123
K+K+GG DV++TYVFWN HEPV+GQYNFEGR DLVKFVK + ++GLY+HLRIGPY CAE
Sbjct: 74 AKSKEGGADVVQTYVFWNGHEPVKGQYNFEGRYDLVKFVKLIGSSGLYLHLRIGPYVCAE 133
Query: 124 WNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIEN 183
WN+GGFP+WL IPGI+FRTDNEPFK EMQ+F KIVD+M++ L+ QGGPII+ QIEN
Sbjct: 134 WNFGGFPVWLRDIPGIEFRTDNEPFKKEMQKFVTKIVDLMREAKLFCWQGGPIIMLQIEN 193
Query: 184 EYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPN 243
EYG+VE +YG Y+ WAASMA L GVPWVMC+Q +AP+ II+ CNG+YCD F PN
Sbjct: 194 EYGDVEKSYGQKGKDYVKWAASMALGLGAGVPWVMCKQTDAPENIIDACNGYYCDGFKPN 253
Query: 244 SKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRT 303
S+ KP WTE ++GW +GG++P+RP EDLAFAVARF+Q GG+ QNYYMY GGTNFGRT
Sbjct: 254 SRTKPVLWTEDWDGWYTKWGGSLPHRPAEDLAFAVARFYQRGGSFQNYYMYFGGTNFGRT 313
Query: 304 SGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATD-PTITSLGSNLE 362
SGGPF TSYDYDA +DEYG +PKWGHLKD+H AIKLCE AL+A D P LGS E
Sbjct: 314 SGGPFYITSYDYDAPLDEYGLRSEPKWGHLKDLHAAIKLCEPALVAADAPQYRKLGSKQE 373
Query: 363 AAVYKTEAE-----CVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKI 417
A +Y + E C AFLANID A VKFNG SY LP WSVSILPDC++V NTAK+
Sbjct: 374 AHIYHGDGETGGKVCAAFLANIDEHKSAHVKFNGQSYTLPPWSVSILPDCRHVAFNTAKV 433
Query: 418 NSESMISSFTTESLK-EVGSL------------EGPSPGWSWISEPIGISKADSFSRFGL 464
+++ + T ES + +GS+ S W + EPIGI ++F+ GL
Sbjct: 434 GAQTSVK--TVESARPSLGSMSILQKVVRQDNVSYISKSWMALKEPIGIWGENNFTFQGL 491
Query: 465 LEQINTTADRSDYLWYSLRIDLEDD-------AGAQTVLHIESLGHALHAFINGKLAGSG 517
LE +N T DRSDYLW+ RI + +D G + + I+S+ L F+N +LAGS
Sbjct: 492 LEHLNVTKDRSDYLWHKTRISVSEDDISFWKKNGPNSTVSIDSMRDVLRVFVNKQLAGSI 551
Query: 518 KGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGS 577
G+ V P+ + G N + LL+ TVGLQ++G F + GAG G L G KNG
Sbjct: 552 VGH----WVKAVQPVRFIQGNNDLLLLTQTVGLQNYGAFLEKDGAGFRGKAKLTGFKNGD 607
Query: 578 TLDFSSKQWTYQIGLKGED---LGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDP 634
LD S WTYQ+GLKGE + +W++ T WYK F P+G DP
Sbjct: 608 -LDLSKSSWTYQVGLKGEADKIYTVEHNEKAEWSTLETDASPSIFMWYKTYFDPPAGTDP 666
Query: 635 IAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTL 694
+ ++ MG+G+AWVNGQ IGRYW IS DGC +C+YRG Y+S KC NCGKP+QT
Sbjct: 667 VVLNLESMGRGQAWVNGQHIGRYW-NIISQKDGCDRTCDYRGAYNSDKCTTNCGKPTQTR 725
Query: 695 YHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWN-----S 749
YHVPRSWL+P SN LVLFEE+GG+P +IS+ G C VSESH PP+ W+ +
Sbjct: 726 YHVPRSWLKPSSNLLVLFEETGGNPFKISVKTVTAGILCGQVSESHYPPLRKWSTPDYIN 785
Query: 750 DTESDRSGGPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACI 809
T S S P + L C VI++I+FAS+GTP G+C FS G C + +LSIV +AC
Sbjct: 786 GTMSINSVAPEVHLHCE-DGHVISSIEFASYGTPRGSCDGFSIGKCHASNSLSIVSEACK 844
Query: 810 GSSSCSIGVSINTF-GDPCGGVTKSLAVEASCT 841
G +SC I VS F DPC G K+LAV + C+
Sbjct: 845 GRNSCFIEVSNTAFISDPCSGTLKTLAVMSRCS 877
>At5g20710 beta-galactosidase like protein
Length = 824
Score = 827 bits (2137), Expect = 0.0
Identities = 417/859 (48%), Positives = 544/859 (62%), Gaps = 56/859 (6%)
Query: 3 MRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDL 62
M+ LL L ++L V++D RA+ I+GKRR+L+SGSIHYPRST +MWPDL
Sbjct: 1 MKHFTRLLSLFFILITSLSLAKSTIVSHDERAITINGKRRILLSGSIHYPRSTADMWPDL 60
Query: 63 IQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACA 122
I KAKDGGLD IETYVFWN HEP + +Y+F G D+V+F+K + AGLY LRIGPY CA
Sbjct: 61 INKAKDGGLDAIETYVFWNAHEPKRREYDFSGNLDVVRFIKTIQDAGLYSVLRIGPYVCA 120
Query: 123 EWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIE 182
EWNYGGFP+WLH +P ++FRT N F EMQ FT KIV MMK+E L+ASQGGPIIL+QIE
Sbjct: 121 EWNYGGFPVWLHNMPNMKFRTVNPSFMNEMQNFTTKIVKMMKEEKLFASQGGPIILAQIE 180
Query: 183 NEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTP 242
NEYGNV +YG YI+W A+MA SLD GVPW+MCQQ NAP P++ TCNGFYCDQ+ P
Sbjct: 181 NEYGNVISSYGAEGKAYIDWCANMANSLDIGVPWLMCQQPNAPQPMLETCNGFYCDQYEP 240
Query: 243 NSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGR 302
+ + PK WTE + GW +GG PYR EDLAF+VARFFQ GGT QNYYMYHGGTNFGR
Sbjct: 241 TNPSTPKMWTENWTGWFKNWGGKHPYRTAEDLAFSVARFFQTGGTFQNYYMYHGGTNFGR 300
Query: 303 TSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLE 362
+GGP++ TSYDY A +DE+G + QPKWGHLK +H +K E++L + + LG++++
Sbjct: 301 VAGGPYITTSYDYHAPLDEFGNLNQPKWGHLKQLHTVLKSMEKSLTYGNISRIDLGNSIK 360
Query: 363 AAVYKTEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESM 422
A +Y T+ F+ N++ T+DA V F G Y++PAWSVS+LPDC NTAK+N++
Sbjct: 361 ATIYTTKEGSSCFIGNVNATADALVNFKGKDYHVPAWSVSVLPDCDKEAYNTAKVNTQ-- 418
Query: 423 ISSFTTESLKEVGSLEGPSPGWSW---ISEPIGISKADSFSRFGLLEQINTTADRSDYLW 479
+S TE + LE W+W ++ + + + GL++Q + T D SDYLW
Sbjct: 419 -TSIMTEDSSKPERLE-----WTWRPESAQKMILKGSGDLIAKGLVDQKDVTNDASDYLW 472
Query: 480 YSLRIDLEDDA---GAQTVLHIESLGHALHAFINGKLAGS-----GKGN-RDNSKVNVEI 530
Y R+ L+ L + S H LHA++NGK G+ GK + R KVN
Sbjct: 473 YMTRLHLDKKDPLWSRNMTLRVHSNAHVLHAYVNGKYVGNQFVKDGKFDYRFERKVN--- 529
Query: 531 PITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTL--DFSSKQWTY 588
LV G N I LLS++VGLQ++GPFF++ GI GPV L G K T+ D S QW Y
Sbjct: 530 --HLVHGTNHISLLSVSVGLQNYGPFFESGPTGINGPVSLVGYKGEETIEKDLSQHQWDY 587
Query: 589 QIGLKGED---LGLPSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKG 645
+IGL G + + S +W ++ LP + LTWYK F AP G +P+ +D G+GKG
Sbjct: 588 KIGLNGYNDKLFSIKSVGHQKWANEK-LPTGRMLTWYKAKFKAPLGKEPVIVDLNGLGKG 646
Query: 646 EAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPD 705
EAW+NGQSIGRYWP++ S DGC C+YRG Y S KC CGKP+Q YHVPRS+L
Sbjct: 647 EAWINGQSIGRYWPSFNSSDDGCKDECDYRGAYGSDKCAFMCGKPTQRWYHVPRSFLNAS 706
Query: 706 S-NTLVLFEESGGDPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLSLE 764
NT+ LFEE GG+P+ ++ ++G+ C+ E + +E
Sbjct: 707 GHNTITLFEEMGGNPSMVNFKTVVVGTVCARAHEHN---------------------KVE 745
Query: 765 CPYPNEVITTIKFASFGTPHGNCGNFSHGDC-SSKQALSIVQKACIGSSSCSIGVSINTF 823
N I+ +KFASFG P G+CG+F+ G C K A V K C+G +C++ VS +TF
Sbjct: 746 LSCHNRPISAVKFASFGNPLGHCGSFAVGTCQGDKDAAKTVAKECVGKLNCTVNVSSDTF 805
Query: 824 GD--PCGGVTKSLAVEASC 840
G CG K LAVE C
Sbjct: 806 GSTLDCGDSPKKLAVELEC 824
>At1g45130 beta-galactosidase like protein
Length = 732
Score = 823 bits (2125), Expect = 0.0
Identities = 400/729 (54%), Positives = 498/729 (67%), Gaps = 21/729 (2%)
Query: 6 TQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQK 65
++ L LL + + + C++VTYD +A+VI+G RR+L+SGSIHYPRSTPEMW DLI+K
Sbjct: 9 SKILTFLLTTMLIGSSVIQCSSVTYDKKAIVINGHRRILLSGSIHYPRSTPEMWEDLIKK 68
Query: 66 AKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWN 125
AKDGGLDVI+TYVFWN HEP G YNFEGR DLV+F+K + GLYVHLRIGPY CAEWN
Sbjct: 69 AKDGGLDVIDTYVFWNGHEPSPGTYNFEGRYDLVRFIKTIQEVGLYVHLRIGPYVCAEWN 128
Query: 126 YGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEY 185
+GGFP+WL ++ GI FRTDN PFK+ MQ FT KIV MMK+ +ASQGGPIILSQIENE+
Sbjct: 129 FGGFPVWLKYVDGISFRTDNGPFKSAMQGFTEKIVQMMKEHRFFASQGGPIILSQIENEF 188
Query: 186 GNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSK 245
GPA Y+NWAA MA L+TGVPWVMC++++APDPIINTCNGFYCD FTPN
Sbjct: 189 EPDLKGLGPAGHSYVNWAAKMAVGLNTGVPWVMCKEDDAPDPIINTCNGFYCDYFTPNKP 248
Query: 246 AKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSG 305
KP WTE ++GW FGG VP RPVEDLAF VARF Q GG+ NYYMYHGGTNFGRT+G
Sbjct: 249 YKPTMWTEAWSGWFTEFGGTVPKRPVEDLAFGVARFIQKGGSYINYYMYHGGTNFGRTAG 308
Query: 306 GPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAV 365
GPF+ TSYDYDA IDEYG +++PK+ HLK +H+AIK CE AL+++DP +T LG+ EA V
Sbjct: 309 GPFITTSYDYDAPIDEYGLVQEPKYSHLKQLHQAIKQCEAALVSSDPHVTKLGNYEEAHV 368
Query: 366 YKT-EAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMIS 424
+ + CVAFL N + A V FN Y LPAWS+SILPDC+NVV NTA + ++
Sbjct: 369 FTAGKGSCVAFLTNYHMNAPAKVVFNNRHYTLPAWSISILPDCRNVVFNTATVAAK---- 424
Query: 425 SFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRI 484
T ++ V S + + + + GLLEQ+N T D +DYLWY+ +
Sbjct: 425 ---TSHVQMVPSGSILYSVARYDEDIATYGNRGTITARGLLEQVNVTRDTTDYLWYTTSV 481
Query: 485 DLEDD-----AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKN 539
D++ G L ++S GHA+H F+NG GS G R+N K + + L G N
Sbjct: 482 DIKASESFLRGGKWPTLTVDSAGHAVHVFVNGHFYGSAFGTRENRKFSFSSQVNLRGGAN 541
Query: 540 TIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGL 599
I LLS+ VGL + GP F+TW GI G V+L GL G+ D S ++WTYQ GL+GE + L
Sbjct: 542 KIALLSVAVGLPNVGPHFETWATGIVGSVVLHGLDEGNK-DLSWQKWTYQAGLRGESMNL 600
Query: 600 PSGT---SGQWNSQSTLPKN-QPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIG 655
S T S W S +N QPLTWYK F AP GN+P+A+D MGKG+AW+NGQSIG
Sbjct: 601 VSPTEDSSVDWIKGSLAKQNKQPLTWYKAYFDAPRGNEPLALDLKSMGKGQAWINGQSIG 660
Query: 656 RYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEES 715
RYW + G SCNY G Y +KC CG+P+Q YHVPRSWL+P N LVLFEE
Sbjct: 661 RYWMAFAK---GDCGSCNYAGTYRQNKCQSGCGEPTQRWYHVPRSWLKPKGNLLVLFEEL 717
Query: 716 GGDPTQISI 724
GGD +++S+
Sbjct: 718 GGDISKVSV 726
>At3g52840 beta-galactosidase precursor - like protein
Length = 727
Score = 815 bits (2105), Expect = 0.0
Identities = 402/710 (56%), Positives = 498/710 (69%), Gaps = 26/710 (3%)
Query: 26 ANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEP 85
A VTYDH+AL+I+G+RR+LISGSIHYPRSTPEMWPDLI+KAK+GGLDVI+TYVFWN HEP
Sbjct: 27 AVVTYDHKALIINGQRRILISGSIHYPRSTPEMWPDLIKKAKEGGLDVIQTYVFWNGHEP 86
Query: 86 VQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDN 145
G Y F+ R DLVKF K V AGLY+ LRIGPY CAEWN+GGFP+WL ++PG+ FRTDN
Sbjct: 87 SPGNYYFQDRYDLVKFTKLVHQAGLYLDLRIGPYVCAEWNFGGFPVWLKYVPGMVFRTDN 146
Query: 146 EPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAAS 205
EPFK MQ+FT KIVDMMK+E L+ +QGGPIILSQIENEYG ++ G A Y W A
Sbjct: 147 EPFKIAMQKFTKKIVDMMKEEKLFETQGGPIILSQIENEYGPMQWEMGAAGKAYSKWTAE 206
Query: 206 MATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYNGWLLYFGGA 265
MA L TGVPW+M +QE+AP PII+TCNGFYC+ F PNS KPK WTE + GW FGGA
Sbjct: 207 MALGLSTGVPWIMSKQEDAPYPIIDTCNGFYCEGFKPNSDNKPKLWTENWTGWFTEFGGA 266
Query: 266 VPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFI 325
+P RPVED+AF+VARF Q GG+ NYYMY+GGTNF RT+ G F+ATSYDYDA IDEYG +
Sbjct: 267 IPNRPVEDIAFSVARFIQNGGSFMNYYMYYGGTNFDRTA-GVFIATSYDYDAPIDEYGLL 325
Query: 326 RQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTEAECVAFLANIDNTSDA 385
R+PK+ HLK++HK IKLCE AL++ DPTITSLG E V+K++ C AFL+N D +S A
Sbjct: 326 REPKYSHLKELHKVIKLCEPALVSVDPTITSLGDKQEIHVFKSKTSCAAFLSNYDTSSAA 385
Query: 386 TVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGWS 445
V F G Y+LP WSVSILPDCK NTAKI + +++ S K +S
Sbjct: 386 RVMFRGFPYDLPPWSVSILPDCKTEYYNTAKIRAPTILMKMIPTSTK-----------FS 434
Query: 446 WISEPIG---ISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDD-----AGAQTVLH 497
W S G ++A +F + GL+EQI+ T D++DY WY I + D G +L
Sbjct: 435 WESYNEGSPSSNEAGTFVKDGLVEQISMTRDKTDYFWYFTDITIGSDESFLKTGDNPLLT 494
Query: 498 IESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFF 557
I S GHALH F+NG LAG+ G NSK+ I L G N + LLS VGL + G +
Sbjct: 495 IFSAGHALHVFVNGLLAGTSYGALSNSKLTFSQNIKLSVGINKLALLSTAVGLPNAGVHY 554
Query: 558 DTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLP--SGTSG-QWNSQSTLP 614
+TW GI GPV LKG+ +G T D S +W+Y+IGL+GE + L +G+S +W + +
Sbjct: 555 ETWNTGILGPVTLKGVNSG-TWDMSKWKWSYKIGLRGEAMSLHTLAGSSAVKWWIKGFVV 613
Query: 615 KNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNY 674
K QPLTWYK +F P GN+P+A+D MGKG+ WVNG +IGR+WP Y + G CNY
Sbjct: 614 KKQPLTWYKSSFDTPRGNEPLALDMNTMGKGQVWVNGHNIGRHWPAYTA--RGNCGRCNY 671
Query: 675 RGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISI 724
G Y+ KCL +CG+PSQ YHVPRSWL+P N LV+FEE GGDP+ IS+
Sbjct: 672 AGIYNEKKCLSHCGEPSQRWYHVPRSWLKPFGNLLVIFEEWGGDPSGISL 721
>At4g26140 putative beta-galactosidase
Length = 729
Score = 815 bits (2104), Expect = 0.0
Identities = 396/723 (54%), Positives = 501/723 (68%), Gaps = 19/723 (2%)
Query: 11 LLLWLLCVYAPACFC-ANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDG 69
+LL +LC + C A VTYD +A++I+G+RR+L+SGSIHYPRSTPEMWPDLIQKAKDG
Sbjct: 11 ILLGILCCSSLICSVKAIVTYDRKAVIINGQRRILLSGSIHYPRSTPEMWPDLIQKAKDG 70
Query: 70 GLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGF 129
GLDVI+TYVFWN HEP GQY FE R DLVKF+K V AGLYVHLRIGPY CAEWN+GGF
Sbjct: 71 GLDVIQTYVFWNGHEPSPGQYYFEDRYDLVKFIKVVQQAGLYVHLRIGPYVCAEWNFGGF 130
Query: 130 PLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVE 189
P+WL ++PG+ FRTDNEPFKA MQ+FT KIV MMK+E L+ +QGGPIILSQIENEYG +E
Sbjct: 131 PVWLKYVPGMVFRTDNEPFKAAMQKFTEKIVRMMKEEKLFETQGGPIILSQIENEYGPIE 190
Query: 190 GAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPK 249
G Y W A MA L TGVPW+MC+Q++AP+ IINTCNGFYC+ F PNS KPK
Sbjct: 191 WEIGAPGKAYTKWVAEMAQGLSTGVPWIMCKQDDAPNSIINTCNGFYCENFKPNSDNKPK 250
Query: 250 FWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFV 309
WTE + GW FGGAVPYRP ED+A +VARF Q GG+ NYYMYHGGTNF RT+ G F+
Sbjct: 251 MWTENWTGWFTEFGGAVPYRPAEDIALSVARFIQNGGSFINYYMYHGGTNFDRTA-GEFI 309
Query: 310 ATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTE 369
ATSYDYDA +DEYG R+PK+ HLK +HK IKLCE AL++ DPT+TSLG EA V+K++
Sbjct: 310 ATSYDYDAPLDEYGLPREPKYSHLKRLHKVIKLCEPALVSADPTVTSLGDKQEAHVFKSK 369
Query: 370 AECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTE 429
+ C AFL+N + +S A V F G++Y+LP WSVSILPDCK NTAK+ T+
Sbjct: 370 SSCAAFLSNYNTSSAARVLFGGSTYDLPPWSVSILPDCKTEYYNTAKVQVR------TSS 423
Query: 430 SLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDD 489
++ P S+ E + +FS+ GL+EQI+ T D++DY WY I + D
Sbjct: 424 IHMKMVPTNTPFSWGSYNEEIPSANDNGTFSQDGLVEQISITRDKTDYFWYLTDITISPD 483
Query: 490 ----AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLS 545
G +L I S GHALH F+NG+LAG+ G+ + K+ I L AG N + LLS
Sbjct: 484 EKFLTGEDPLLTIGSAGHALHVFVNGQLAGTAYGSLEKPKLTFSQKIKLHAGVNKLALLS 543
Query: 546 LTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTY-QIGLKGEDLG---LPS 601
GL + G ++TW G+ GPV L G+ +G T D + +W+Y QIG KGE L L
Sbjct: 544 TAAGLPNVGVHYETWNTGVLGPVTLNGVNSG-TWDMTKWKWSYKQIGTKGEALSVHTLAG 602
Query: 602 GTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTY 661
++ +W S + K QPLTWYK F +P+GN+P+A+D MGKG+ W+NGQ+IGR+WP Y
Sbjct: 603 SSTVEWKEGSLVAKKQPLTWYKSTFDSPTGNEPLALDMNTMGKGQMWINGQNIGRHWPAY 662
Query: 662 ISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQ 721
+ G C+Y G + KCL NCG+ SQ YHVPRSWL+P +N +++ EE GG+P
Sbjct: 663 TA--RGKCERCSYAGTFTEKKCLSNCGEASQRWYHVPRSWLKPTNNLVIVLEEWGGEPNG 720
Query: 722 ISI 724
IS+
Sbjct: 721 ISL 723
>At5g56870 beta-galactosidase (emb|CAB64740.1)
Length = 724
Score = 813 bits (2099), Expect = 0.0
Identities = 401/724 (55%), Positives = 505/724 (69%), Gaps = 24/724 (3%)
Query: 11 LLLWLLCVYAPACFC-ANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDG 69
+ L +LC + +C A+V+YD +A++I+G+RR+L+SGSIHYPRSTPEMWP LIQKAK+G
Sbjct: 11 IFLAILCCLSLSCIVKASVSYDRKAVIINGQRRILLSGSIHYPRSTPEMWPGLIQKAKEG 70
Query: 70 GLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGF 129
GLDVIETYVFWN HEP GQY F R DLVKF+K V AGLYV+LRIGPY CAEWN+GGF
Sbjct: 71 GLDVIETYVFWNGHEPSPGQYYFGDRYDLVKFIKLVHQAGLYVNLRIGPYVCAEWNFGGF 130
Query: 130 PLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVE 189
P+WL F+PG+ FRTDNEPFKA M++FT KIV MMK E L+ +QGGPIIL+QIENEYG VE
Sbjct: 131 PVWLKFVPGMAFRTDNEPFKAAMKKFTEKIVWMMKAEKLFQTQGGPIILAQIENEYGPVE 190
Query: 190 GAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPK 249
G Y W A MA L TGVPW+MC+QE+AP PII+TCNG+YC+ F PNS KPK
Sbjct: 191 WEIGAPGKAYTKWVAQMALGLSTGVPWIMCKQEDAPGPIIDTCNGYYCEDFKPNSINKPK 250
Query: 250 FWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFV 309
WTE + GW FGGAVPYRPVED+A++VARF Q GG+L NYYMYHGGTNF RT+ G F+
Sbjct: 251 MWTENWTGWYTDFGGAVPYRPVEDIAYSVARFIQKGGSLVNYYMYHGGTNFDRTA-GEFM 309
Query: 310 ATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTE 369
A+SYDYDA +DEYG R+PK+ HLK +HKAIKL E AL++ D T+TSLG+ EA V+ ++
Sbjct: 310 ASSYDYDAPLDEYGLPREPKYSHLKALHKAIKLSEPALLSADATVTSLGAKQEAYVFWSK 369
Query: 370 AECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTE 429
+ C AFL+N D S A V F G Y+LP WSVSILPDCK V NTAK+N+ S+ +
Sbjct: 370 SSCAAFLSNKDENSAARVLFRGFPYDLPPWSVSILPDCKTEVYNTAKVNAPSVHRNMVPT 429
Query: 430 SLK-EVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLED 488
K GS +P ++A +F+R GL+EQI+ T D+SDY WY I +
Sbjct: 430 GTKFSWGSFNEATP---------TANEAGTFARNGLVEQISMTWDKSDYFWYITDITIGS 480
Query: 489 -----DAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDL 543
G +L + S GHALH F+NG+L+G+ G D+ K+ I L AG N I L
Sbjct: 481 GETFLKTGDSPLLTVMSAGHALHVFVNGQLSGTAYGGLDHPKLTFSQKIKLHAGVNKIAL 540
Query: 544 LSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPSGT 603
LS+ VGL + G F+ W G+ GPV LKG+ +G T D S +W+Y+IG+KGE L L + T
Sbjct: 541 LSVAVGLPNVGTHFEQWNKGVLGPVTLKGVNSG-TWDMSKWKWSYKIGVKGEALSLHTNT 599
Query: 604 SG---QWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPT 660
+W S + K QPLTWYK F+ P+GN+P+A+D MGKG+ W+NG++IGR+WP
Sbjct: 600 ESSGVRWTQGSFVAKKQPLTWYKSTFATPAGNEPLALDMNTMGKGQVWINGRNIGRHWPA 659
Query: 661 YISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPT 720
Y G CNY G +D+ KCL NCG+ SQ YHVPRSWL+ N +V+FEE GGDP
Sbjct: 660 Y--KAQGSCGRCNYAGTFDAKKCLSNCGEASQRWYHVPRSWLK-SQNLIVVFEELGGDPN 716
Query: 721 QISI 724
IS+
Sbjct: 717 GISL 720
>At5g63810 beta-galactosidase (emb|CAB64746.1)
Length = 741
Score = 781 bits (2017), Expect = 0.0
Identities = 379/712 (53%), Positives = 487/712 (68%), Gaps = 19/712 (2%)
Query: 26 ANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEP 85
ANV+YDHR+L I +R+++IS +IHYPRS P MWP L+Q AK+GG + IE+YVFWN HEP
Sbjct: 30 ANVSYDHRSLTIGNRRQLIISAAIHYPRSVPAMWPSLVQTAKEGGCNAIESYVFWNGHEP 89
Query: 86 VQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDN 145
G+Y F GR ++VKF+K V AG+++ LRIGP+ AEWNYGG P+WLH++PG FR DN
Sbjct: 90 SPGKYYFGGRYNIVKFIKIVQQAGMHMILRIGPFVAAEWNYGGVPVWLHYVPGTVFRADN 149
Query: 146 EPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAAS 205
EP+K M+ FT IV+++KQE L+A QGGPIILSQ+ENEYG E YG Y W+AS
Sbjct: 150 EPWKHYMESFTTYIVNLLKQEKLFAPQGGPIILSQVENEYGYYEKDYGEGGKRYAQWSAS 209
Query: 206 MATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYNGWLLYFGGA 265
MA S + GVPW+MCQQ +AP +I+TCNGFYCDQFTPN+ KPK WTE + GW FGG
Sbjct: 210 MAVSQNIGVPWMMCQQWDAPPTVISTCNGFYCDQFTPNTPDKPKIWTENWPGWFKTFGGR 269
Query: 266 VPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFI 325
P+RP ED+A++VARFF GG++ NYYMYHGGTNFGRTSGGPF+ TSYDY+A IDEYG
Sbjct: 270 DPHRPAEDVAYSVARFFGKGGSVHNYYMYHGGTNFGRTSGGPFITTSYDYEAPIDEYGLP 329
Query: 326 RQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVY-KTEAECVAFLANIDNTSD 384
R PKWGHLKD+HKAI L E LI+ + +LG +LEA VY + C AFL+N+D+ +D
Sbjct: 330 RLPKWGHLKDLHKAIMLSENLLISGEHQNFTLGHSLEADVYTDSSGTCAAFLSNLDDKND 389
Query: 385 ATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGW 444
V F SY+LPAWSVSILPDCK V NTAK+ S+S E LK L+ W
Sbjct: 390 KAVMFRNTSYHLPAWSVSILPDCKTEVFNTAKVTSKSSKVEMLPEDLKSSSGLK-----W 444
Query: 445 SWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDD-----AGAQTVLHIE 499
SE GI A F + L++ INTT D +DYLWY+ I + ++ G+ VL IE
Sbjct: 445 EVFSEKPGIWGAADFVKNELVDHINTTKDTTDYLWYTTSITVSENEAFLKKGSSPVLFIE 504
Query: 500 SLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDT 559
S GH LH FIN + G+ GN + ++ P+ L AG+N IDLLS+TVGL + G F++
Sbjct: 505 SKGHTLHVFINKEYLGTATGNGTHVPFKLKKPVALKAGENNIDLLSMTVGLANAGSFYEW 564
Query: 560 WGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGL-PSGTSG--QWNSQSTLPKN 616
GAG+T V +KG G TL+ ++ +W+Y++G++GE L L G SG +W + PK
Sbjct: 565 VGAGLTS-VSIKGFNKG-TLNLTNSKWSYKLGVEGEHLELFKPGNSGAVKWTVTTKPPKK 622
Query: 617 QPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYI---SPIDGCTSSCN 673
QPLTWYK+ PSG++P+ +D MGKG AW+NG+ IGRYWP SP D C C+
Sbjct: 623 QPLTWYKVVIEPPSGSEPVGLDMISMGKGMAWLNGEEIGRYWPRIARKNSPNDECVKECD 682
Query: 674 YRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIA 725
YRG + KCL CG+PSQ YHVPRSW + N LV+FEE GG+P +I ++
Sbjct: 683 YRGKFMPDKCLTGCGEPSQRWYHVPRSWFKSSGNELVIFEEKGGNPMKIKLS 734
>At1g31740 putative protein
Length = 757
Score = 744 bits (1922), Expect = 0.0
Identities = 391/836 (46%), Positives = 507/836 (59%), Gaps = 100/836 (11%)
Query: 17 CVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIET 76
C YA V++D RA+ IDG RRVL+SGSIHYPRST EMWPDLI+K K+G LD IET
Sbjct: 10 CAYATI-----VSHDGRAITIDGHRRVLLSGSIHYPRSTTEMWPDLIKKGKEGSLDAIET 64
Query: 77 YVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFI 136
YVFWN HEP + QY+F G DL++F+K + G+Y LRIGPY CAEWNYGGFP+WLH +
Sbjct: 65 YVFWNAHEPTRRQYDFSGNLDLIRFLKTIQNEGMYGVLRIGPYVCAEWNYGGFPVWLHNM 124
Query: 137 PGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAA 196
PG++FRT N F EMQ FT IV+M+K+E L+ASQGGPIIL+QIENEYGNV G+YG A
Sbjct: 125 PGMEFRTTNTAFMNEMQNFTTMIVEMVKKEKLFASQGGPIILAQIENEYGNVIGSYGEAG 184
Query: 197 VPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYN 256
YI W A+MA SLD GVPW+MCQQ++AP P++NTCNG+YCD F+PN+ PK WTE +
Sbjct: 185 KAYIQWCANMANSLDVGVPWIMCQQDDAPQPMLNTCNGYYCDNFSPNNPNTPKMWTENWT 244
Query: 257 GWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYD 316
GW +GG P+R ED+AFAVARFFQ GT QNYYMYHGGTNF RT+GGP++ T+YDYD
Sbjct: 245 GWYKNWGGKDPHRTTEDVAFAVARFFQKEGTFQNYYMYHGGTNFDRTAGGPYITTTYDYD 304
Query: 317 AAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTEAECVAFL 376
A +DE+G + QPK+GHLK +H + E+ L + + G+ + A VY+TE F+
Sbjct: 305 APLDEFGNLNQPKYGHLKQLHDVLHAMEKTLTYGNISTVDFGNLVTATVYQTEEGSSCFI 364
Query: 377 ANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSE-SMISSFTTESLKEVG 435
N++ TSDA + F G SY++PAWSVSILPDCK NTAKIN++ S++ E+ E
Sbjct: 365 GNVNETSDAKINFQGTSYDVPAWSVSILPDCKTETYNTAKINTQTSVMVKKANEAENEPS 424
Query: 436 SLEGPSPGWSWISEPIGI----SKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDD-- 489
+L+ WSW E I K +S R L +Q + D SDYLWY ++L++
Sbjct: 425 TLK-----WSWRPENIDSVLLKGKGESTMR-QLFDQKVVSNDESDYLWYMTTVNLKEQDP 478
Query: 490 -AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTV 548
G L I S H LHAF+NG+ G+ + E G N I LLS+TV
Sbjct: 479 VLGKNMSLRINSTAHVLHAFVNGQHIGNYRVENGKFHYVFEQDAKFNPGANVITLLSITV 538
Query: 549 GLQHFGPFFDTWGAGITGPVILKGLKNGSTL--DFSSKQWTYQIGLKGEDLGLPSGTSGQ 606
GL ++G FF+ + AGITGPV + G T+ D S+ +W+Y+ GL G + L S S
Sbjct: 539 GLPNYGAFFENFSAGITGPVFIIGRNGDETIVKDLSTHKWSYKTGLSGFENQLFSSES-- 596
Query: 607 WNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPID 666
P TW SAP G++P+ +D G+GKG AW+NG +IGRYWP ++S ID
Sbjct: 597 -----------PSTW-----SAPLGSEPVVVDLLGLGKGTAWINGNNIGRYWPAFLSDID 640
Query: 667 GCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAI 726
G NTLVLFEE GG+P+ ++
Sbjct: 641 G--------------------------------------DNTLVLFEEIGGNPSLVNFQT 662
Query: 727 KLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLSLECPYPNEVITTIKFASFGTPHGN 786
+GS C++V E + VL L C + I+ IKFASFG P G+
Sbjct: 663 IGVGSVCANVYEKN-------------------VLELSC--NGKPISAIKFASFGNPGGD 701
Query: 787 CGNFSHGDC-SSKQALSIVQKACIGSSSCSIGVSINTFG-DPCGGVTKSLAVEASC 840
CG+F G C +S A +I+ + C+G CSI VS + FG CG + K LAVEA C
Sbjct: 702 CGSFEKGTCEASNNAAAILTQECVGKEKCSIDVSEDKFGAAECGALAKRLAVEAIC 757
>At1g77410 unknown protein
Length = 815
Score = 726 bits (1874), Expect = 0.0
Identities = 402/855 (47%), Positives = 514/855 (60%), Gaps = 56/855 (6%)
Query: 3 MRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDL 62
M Q+ L+ L L+ V A ANVTYD R+L+IDG+ ++L SGSIHY RSTP+MWP L
Sbjct: 1 MTTFQYSLVFLVLMAVIV-AGDVANVTYDGRSLIIDGEHKILFSGSIHYTRSTPQMWPSL 59
Query: 63 IQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACA 122
I KAK GG+DV++TYVFWN+HEP QGQ++F G D+VKF+K V GLYV LRIGP+
Sbjct: 60 IAKAKSGGIDVVDTYVFWNVHEPQQGQFDFSGSRDIVKFIKEVKNHGLYVCLRIGPFIQG 119
Query: 123 EWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIE 182
EW+YGG P WLH + GI FRTDNEPFK M+R+ IV +MK ENLYASQGGPIILSQIE
Sbjct: 120 EWSYGGLPFWLHNVQGIVFRTDNEPFKYHMKRYAKMIVKLMKSENLYASQGGPIILSQIE 179
Query: 183 NEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQF-- 240
NEYG V A+ Y+ W A +A LDTGVPWVMC+Q++APDP++N CNG C +
Sbjct: 180 NEYGMVGRAFRQEGKSYVKWTAKLAVELDTGVPWVMCKQDDAPDPLVNACNGRQCGETFK 239
Query: 241 TPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNF 300
PNS KP WTE + + +G R ED+AF VA F G+ NYYMYHGGTNF
Sbjct: 240 GPNSPNKPAIWTENWTSFYQTYGEEPLIRSAEDIAFHVALFIAKNGSFVNYYMYHGGTNF 299
Query: 301 GRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSN 360
GR + FV TSY A +DEYG +RQPKWGHLK++H A+KLCEE L++ T SLG
Sbjct: 300 GR-NASQFVITSYYDQAPLDEYGLLRQPKWGHLKELHAAVKLCEEPLLSGLQTTISLGKL 358
Query: 361 LEAAVYKTEAE-CVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINS 419
A V+ +A C A L N D ++TV+F +SY L SVS+LPDCKNV NTAK+N+
Sbjct: 359 QTAFVFGKKANLCAAILVNQDK-CESTVQFRNSSYRLSPKSVSVLPDCKNVAFNTAKVNA 417
Query: 420 ESMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLW 479
+ + T + K +L P W +E + S LLE +NTT D SDYLW
Sbjct: 418 Q-----YNTRTRKARQNLSSPQM-WEEFTETVPSFSETSIRSESLLEHMNTTQDTSDYLW 471
Query: 480 YSLRIDLEDDAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKN 539
+ R + GA +VL + LGHALHAF+NG+ GS G + +E ++L G N
Sbjct: 472 QTTR--FQQSEGAPSVLKVNHLGHALHAFVNGRFIGSMHGTFKAHRFLLEKNMSLNNGTN 529
Query: 540 TIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGL 599
+ LLS+ VGL + G + G I G L F++ W YQ+GLKGE +
Sbjct: 530 NLALLSVMVGLPNSGAHLERRVVGSRSVKIWNGRYQ---LYFNNYSWGYQVGLKGEKFHV 586
Query: 600 ---PSGTSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGR 656
QW Q K+QPLTWYK +F P G DP+A++ MGKGEAWVNGQSIGR
Sbjct: 587 YTEDGSAKVQW-KQYRDSKSQPLTWYKASFDTPEGEDPVALNLGSMGKGEAWVNGQSIGR 645
Query: 657 YWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEES- 715
YW ++ + Y+ G PSQ YH+PRS+L+P+SN LV+ EE
Sbjct: 646 YWVSFHT----------YK------------GNPSQIWYHIPRSFLKPNSNLLVILEEER 683
Query: 716 GGDPTQISIAIKLIGSSCSHVSESHPPPV--------DLWNSDTESDRSGGPVLSLECPY 767
G+P I+I + C HVS ++P PV + N DR P + L+CP
Sbjct: 684 EGNPLGITIDTVSVTEVCGHVSNTNPHPVISPRKKGLNRKNLTYRYDRK--PKVQLQCP- 740
Query: 768 PNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTF-GDP 826
I+ I FASFGTP+G+CG++S G C S +L++VQKAC+ S CS+ V TF GD
Sbjct: 741 TGRKISKILFASFGTPNGSCGSYSIGSCHSPNSLAVVQKACLKKSRCSVPVWSKTFGGDS 800
Query: 827 CGGVTKSLAVEASCT 841
C KSL V A C+
Sbjct: 801 CPHTVKSLLVRAQCS 815
>At2g16730 putative beta-galactosidase
Length = 832
Score = 650 bits (1677), Expect = 0.0
Identities = 358/855 (41%), Positives = 499/855 (57%), Gaps = 64/855 (7%)
Query: 11 LLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGG 70
LLL +L + ++TYD +L+I+G R +L SGSIHYPRSTPEMWP++I++AK GG
Sbjct: 11 LLLAVLVILLSFSGALSITYDGTSLIINGNRELLYSGSIHYPRSTPEMWPNIIKRAKQGG 70
Query: 71 LDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFP 130
L+ I+TYVFWN+HEP QG++NF GR DLVKF+K + GLYV LR+GP+ AEW +GG P
Sbjct: 71 LNTIQTYVFWNVHEPEQGKFNFSGRADLVKFIKLIEKNGLYVTLRLGPFIQAEWTHGGLP 130
Query: 131 LWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEG 190
WL +PGI FRTDNEPFK +R+ ++DMMK+E L+ASQGGPIIL QIENEY V+
Sbjct: 131 YWLREVPGIFFRTDNEPFKEHTERYVKVVLDMMKEEKLFASQGGPIILGQIENEYSAVQR 190
Query: 191 AYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYC-DQFT-PNSKAKP 248
AY + YI WA+ + S+D G+PWVMC+Q +APDP+IN CNG +C D F PN KP
Sbjct: 191 AYKEDGLNYIKWASKLVHSMDLGIPWVMCKQNDAPDPMINACNGRHCGDTFPGPNKDNKP 250
Query: 249 KFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPF 308
WTE + FG R VED+A++VARFF GT NYYMYHGGTNFGRTS +
Sbjct: 251 SLWTENWTTQFRVFGDPPAQRSVEDIAYSVARFFSKNGTHVNYYMYHGGTNFGRTS-AHY 309
Query: 309 VATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKT 368
V T Y DA +DE+G R+PK+GHLK +H A+ LC++AL+ P + + E Y+
Sbjct: 310 VTTRYYDDAPLDEFGLEREPKYGHLKHLHNALNLCKKALLWGQPRVEKPSNETEIRYYEQ 369
Query: 369 EAE--CVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSF 426
C AFLAN + + +KF G Y +P S+SILPDCK VV NT +I S +F
Sbjct: 370 PGTKVCAAFLANNNTEAAEKIKFRGKEYLIPHRSISILPDCKTVVYNTGEIISHHTSRNF 429
Query: 427 TTESLKEVGSLEGPSPGWSWISEPIGIS-KADSFSRFGLLEQINTTADRSDYLWYSLRID 485
+S K + + + +E + K DSF +E T D SDY WY+
Sbjct: 430 -MKSKKANKNFD-----FKVFTESVPSKIKGDSFIP---VELYGLTKDESDYGWYTTSFK 480
Query: 486 LEDD-----AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNT 540
++D+ G + L I SLGHALH ++NG+ G+G G+ + + P+TL G+N
Sbjct: 481 IDDNDLSKKKGGKPNLRIASLGHALHVWLNGEYLGNGHGSHEEKSFVFQKPVTLKEGENH 540
Query: 541 IDLLSLTVGLQHFGPFFDTWGAGITGP--VILKGLKNGSTLDFSSK-QWTYQIGLKGEDL 597
+ +L + G G + + TGP V + GL +G TLD + + +W ++G++GE L
Sbjct: 541 LTMLGVLTGFPDSGSYME---HRYTGPRSVSILGLGSG-TLDLTEENKWGNKVGMEGERL 596
Query: 598 GLPSG---TSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSI 654
G+ + +W S K +TWY+ F AP AI GMGKG WVNG+ +
Sbjct: 597 GIHAEEGLKKVKWEKAS--GKEPGMTWYQTYFDAPESQSAAAIRMNGMGKGLIWVNGEGV 654
Query: 655 GRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEE 714
GRYW +++SP+ G+P+Q YH+PRS+L+P N LV+FEE
Sbjct: 655 GRYWMSFLSPL----------------------GQPTQIEYHIPRSFLKPKKNLLVIFEE 692
Query: 715 SGG-DPTQISIAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGP----VLSLECPYPN 769
P I I + CS++ E++ P V W + ++ +L+C
Sbjct: 693 EPNVKPELIDFVIVNRDTVCSYIGENYTPSVRHWTRKNDQVQAITDDVHLTANLKCS-GT 751
Query: 770 EVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTF----GD 825
+ I+ ++FASFG P+G CGNF+ G C++ + +V+K C+G + C I V+ +TF D
Sbjct: 752 KKISAVEFASFGNPNGTCGNFTLGSCNAPVSKKVVEKYCLGKAECVIPVNKSTFEQDKKD 811
Query: 826 PCGGVTKSLAVEASC 840
C V K LAV+ C
Sbjct: 812 SCPKVEKKLAVQVKC 826
>At5g63800 beta-galactosidase like protein
Length = 657
Score = 613 bits (1582), Expect = e-176
Identities = 320/683 (46%), Positives = 417/683 (60%), Gaps = 41/683 (6%)
Query: 58 MWPDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIG 117
MWP LI+K K+GG+DVI+TYVFWNLHEP GQY+F GR DLVKF+K + + GLYV LRIG
Sbjct: 1 MWPSLIKKTKEGGIDVIQTYVFWNLHEPKLGQYDFSGRNDLVKFIKEIRSQGLYVCLRIG 60
Query: 118 PYACAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPII 177
P+ AEWNYGG P WL +PG+ +RTDNEPFK MQ+FTAKIVD+MK E LYASQGGPII
Sbjct: 61 PFIEAEWNYGGLPFWLRDVPGMVYRTDNEPFKFHMQKFTAKIVDLMKSEGLYASQGGPII 120
Query: 178 LSQIENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYC 237
LSQIENEY NVEGA+ YI WA MA L TGVPW+MC+ +APDP+INTCNG C
Sbjct: 121 LSQIENEYANVEGAFHEKGASYIKWAGQMAVGLKTGVPWIMCKSPDAPDPVINTCNGMKC 180
Query: 238 DQF--TPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYH 295
+ PNS KPK WTE + + +G R ED+AF A F G+ NYYMYH
Sbjct: 181 GETFPGPNSPNKPKMWTEDWTSFFQVYGKEPYIRSAEDIAFHAALFVAKNGSYINYYMYH 240
Query: 296 GGTNFGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTIT 355
GGTNFGRTS F+ YD A +DEYG +RQPK+GHLK++H AIK L+ TI
Sbjct: 241 GGTNFGRTSSSYFITGYYD-QAPLDEYGLLRQPKYGHLKELHAAIKSSANPLLQGKQTIL 299
Query: 356 SLGSNLEAAVYK-TEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNT 414
SLG +A V++ CVAFL N D + ++F N+Y+L S+ IL +CKN++ T
Sbjct: 300 SLGPMQQAYVFEDANNGCVAFLVNND-AKASQIQFRNNAYSLSPKSIGILQNCKNLIYET 358
Query: 415 AKINSESMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADR 474
AK+N + T + V W+ E I S LLE N T D+
Sbjct: 359 AKVNVKMNTRVTTPVQVFNV------PDNWNLFRETIPAFPGTSLKTNALLEHTNLTKDK 412
Query: 475 SDYLWYSLRIDLEDDAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITL 534
+DYLWY+ L D ++ ES GH +H F+N LAGSG G+RD V ++ P++L
Sbjct: 413 TDYLWYTSSFKL-DSPCTNPSIYTESSGHVVHVFVNNALAGSGHGSRDIRVVKLQAPVSL 471
Query: 535 VAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKG 594
+ G+N I +LS VGL G + + G+T I G +D S QW Y +GL G
Sbjct: 472 INGQNNISILSGMVGLPDSGAYMERRSYGLTKVQISCG--GTKPIDLSRSQWGYSVGLLG 529
Query: 595 EDLGL---PSGTSGQWN-SQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVN 650
E + L + +W+ +++ L KN+PL WYK F P+G+ P+ + + MGKGE WVN
Sbjct: 530 EKVRLYQWKNLNRVKWSMNKAGLIKNRPLAWYKTTFDGPNGDGPVGLHMSSMGKGEIWVN 589
Query: 651 GQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLV 710
G+SIGRYW ++++P G+PSQ++YH+PR++L+P N LV
Sbjct: 590 GESIGRYWVSFLTP----------------------AGQPSQSIYHIPRAFLKPSGNLLV 627
Query: 711 LFEESGGDPTQISI-AIKLIGSS 732
+FEE GGDP IS+ I ++GSS
Sbjct: 628 VFEEEGGDPLGISLNTISVVGSS 650
>At4g38590 galactosidase like protein
Length = 1036
Score = 547 bits (1409), Expect = e-155
Identities = 311/771 (40%), Positives = 429/771 (55%), Gaps = 54/771 (7%)
Query: 89 QYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNEPF 148
QY+F+GR DLVKF+K + GLYV LR+GP+ AEWN+GG P WL +P + FRT+NEPF
Sbjct: 80 QYDFKGRFDLVKFIKLIHEKGLYVTLRLGPFIQAEWNHGGLPYWLREVPDVYFRTNNEPF 139
Query: 149 KAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMAT 208
K +R+ KI+ MMK+E L+ASQGGPIIL QIENEY V+ AY YI WAA++
Sbjct: 140 KEHTERYVRKILGMMKEEKLFASQGGPIILGQIENEYNAVQLAYKENGEKYIKWAANLVE 199
Query: 209 SLDTGVPWVMCQQENAPDPIINTCNGFYC-DQFT-PNSKAKPKFWTEGYNGWLLYFGGAV 266
S++ G+PWVMC+Q +AP +IN CNG +C D F PN KP WTE + FG
Sbjct: 200 SMNLGIPWVMCKQNDAPGNLINACNGRHCGDTFPGPNRHDKPSLWTENWTTQFRVFGDPP 259
Query: 267 PYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFIR 326
R VED+AF+VAR+F G+ NYYMYHGGTNFGRTS FV T Y DA +DE+G +
Sbjct: 260 TQRTVEDIAFSVARYFSKNGSHVNYYMYHGGTNFGRTS-AHFVTTRYYDDAPLDEFGLEK 318
Query: 327 QPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTEAE--CVAFLANIDNTSD 384
PK+GHLK VH+A++LC++AL +LG + E Y+ C AFL+N +
Sbjct: 319 APKYGHLKHVHRALRLCKKALFWGQLRAQTLGPDTEVRYYEQPGTKVCAAFLSNNNTRDT 378
Query: 385 ATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGW 444
T+KF G Y LP+ S+SILPDCK VV NTA+I ++ F G +
Sbjct: 379 NTIKFKGQDYVLPSRSISILPDCKTVVYNTAQIVAQHSWRDFVKSEKTSKGL------KF 432
Query: 445 SWISEPI-GISKADSFSRFGLLEQINTTADRSDYLWYSLRI-----DLEDDAGAQTVLHI 498
SE I + DS E T D++DY WY+ + D D G +T+L +
Sbjct: 433 EMFSENIPSLLDGDSLIPG---ELYYLTKDKTDYAWYTTSVKIDEDDFPDQKGLKTILRV 489
Query: 499 ESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFD 558
SLGHAL ++NG+ AG G + P+ G N I +L + GL G + +
Sbjct: 490 ASLGHALIVYVNGEYAGKAHGRHEMKSFEFAKPVNFKTGDNRISILGVLTGLPDSGSYME 549
Query: 559 TWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPSGTSGQWNSQSTLPKNQP 618
AG I+ GLK+G+ + +W + GL+GE + + + K +P
Sbjct: 550 HRFAGPRAISII-GLKSGTRDLTENNEWGHLAGLEGEKKEVYTEEGSKKVKWEKDGKRKP 608
Query: 619 LTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPY 678
LTWYK F P G + +AI MGKG WVNG +GRYW +++SP+
Sbjct: 609 LTWYKTYFETPEGVNAVAIRMKAMGKGLIWVNGIGVGRYWMSFLSPL------------- 655
Query: 679 DSSKCLKNCGKPSQTLYHVPRSWLQPD--SNTLVLFEESGGDPTQISIAIKLIG--SSCS 734
G+P+QT YH+PRS+++ + N LV+ EE G + SI L+ + CS
Sbjct: 656 ---------GEPTQTEYHIPRSFMKGEKKKNMLVILEEEPGVKLE-SIDFVLVNRDTICS 705
Query: 735 HVSESHPPPVDLWNSDTESDRSGGPVLSLE----CPYPNEVITTIKFASFGTPHGNCGNF 790
+V E +P V W + S + L+ CP P + + ++FASFG P G CGNF
Sbjct: 706 NVGEDYPVSVKSWKREGPKIVSRSKDMRLKAVMRCP-PEKQMVEVQFASFGDPTGTCGNF 764
Query: 791 SHGDCSSKQALSIVQKACIGSSSCSIGVSINTFGDP-CGGVTKSLAVEASC 840
+ G CS+ ++ +V+K C+G + CSI V+ TFGD C + K+LAV+ C
Sbjct: 765 TMGKCSASKSKEVVEKECLGRNYCSIVVARETFGDKGCPEIVKTLAVQVKC 815
>At4g35010 beta-galactosidase - like protein
Length = 831
Score = 544 bits (1401), Expect = e-155
Identities = 326/844 (38%), Positives = 458/844 (53%), Gaps = 103/844 (12%)
Query: 28 VTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQ 87
VTYD +L+IDGKR +L SGSIHYPRSTPEMWP +I++AK GGL+ I+TYVFWN+HEP Q
Sbjct: 54 VTYDGTSLIIDGKRELLYSGSIHYPRSTPEMWPSIIKRAKQGGLNTIQTYVFWNVHEPQQ 113
Query: 88 GQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNEP 147
G++NF GR DLVKF+K + G+YV LR+GP+ AEW +G + H
Sbjct: 114 GKFNFSGRADLVKFIKLIQKNGMYVTLRLGPFIQAEWTHGYITRYDH------------- 160
Query: 148 FKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMA 207
+N+ + +IENEY V+ AY + YI WA+++
Sbjct: 161 ------------------KNIAGAY------RKIENEYSAVQRAYKQDGLNYIKWASNLV 196
Query: 208 TSLDTGVPWVMCQQENAPDPIINTCNGFYC-DQFT-PNSKAKPKFWTEGYNGWLLYFGGA 265
S+ G+PWVMC+Q +APDP+IN CNG +C D F PN + KP WTE + FG
Sbjct: 197 DSMKLGIPWVMCKQNDAPDPMINACNGRHCGDTFPGPNRENKPSLWTENWTTQFRVFGDP 256
Query: 266 VPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFI 325
R VED+A++VARFF GT NYYMYHGGTNFGRTS +V T Y DA +DEYG
Sbjct: 257 PTQRSVEDIAYSVARFFSKNGTHVNYYMYHGGTNFGRTS-AHYVTTRYYDDAPLDEYGLE 315
Query: 326 RQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTEA--ECVAFLANIDNTS 383
++PK+GHLK +H A+ LC++ L+ P G + E Y+ C AFLAN + +
Sbjct: 316 KEPKYGHLKHLHNALNLCKKPLLWGQPKTEKPGKDTEIRYYEQPGTKTCAAFLANNNTEA 375
Query: 384 DATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPG 443
T+KF G Y + S+SILPDCK VV NTA+I S+ +F +S K +
Sbjct: 376 AETIKFKGREYVIAPRSISILPDCKTVVYNTAQIVSQHTSRNF-MKSKKANKKFD----- 429
Query: 444 WSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSL-----RIDLEDDAGAQTVLHI 498
+ +E + SK + S + +E T D++DY WY+ + L G +T + I
Sbjct: 430 FKVFTETLP-SKLEGNS-YIPVELYGLTKDKTDYGWYTTSFKVHKNHLPTKKGVKTFVRI 487
Query: 499 ESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFD 558
SLGHALHA++NG+ GSG G+ + + +TL AG+N + +L + G G + +
Sbjct: 488 ASLGHALHAWLNGEYLGSGHGSHEEKSFVFQKQVTLKAGENHLVMLGVLTGFPDSGSYME 547
Query: 559 TWGAGITGPVILKGLKNGSTLDFS-SKQWTYQIGLKGEDLGLPSG---TSGQWNSQSTLP 614
G G IL GL +G TLD + S +W +IG++GE LG+ + +W +
Sbjct: 548 HRYTGPRGISIL-GLTSG-TLDLTESSKWGNKIGMEGEKLGIHTEEGLKKVEW--KKFTG 603
Query: 615 KNQPLTWY----------KINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISP 664
K LTWY + F AP I GMGKG WVNG+ +GRYW +++SP
Sbjct: 604 KAPGLTWYQKFSKECETLQTYFDAPESVSAATIRMHGMGKGLIWVNGEGVGRYWQSFLSP 663
Query: 665 IDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGG-DPTQIS 723
+ G+P+Q YH+PRS+L+P N LV+FEE P +
Sbjct: 664 L----------------------GQPTQIEYHIPRSFLKPKKNLLVIFEEEPNVKPELMD 701
Query: 724 IAIKLIGSSCSHVSESHPPPVDLWNSDTESDRSGGPVLSLECPYP---NEVITTIKFASF 780
AI + CS+V E++ P V W + ++ +SL + I ++FASF
Sbjct: 702 FAIVNRDTVCSYVGENYTPSVRHWTRKKDQVQAITDNVSLTATLKCSGTKKIAAVEFASF 761
Query: 781 GTPHGNCGNFSHGDCSSKQALSIVQKACIGSSSCSIGVSINTF----GDPCGGVTKSLAV 836
G P G CGNF+ G C++ + +++K C+G + C I V+ +TF D C V K LAV
Sbjct: 762 GNPIGVCGNFTLGTCNAPVSKQVIEKHCLGKAECVIPVNKSTFQQDKKDSCKNVVKMLAV 821
Query: 837 EASC 840
+ C
Sbjct: 822 QVKC 825
>At1g72990 unknown protein
Length = 697
Score = 161 bits (407), Expect = 2e-39
Identities = 150/513 (29%), Positives = 221/513 (42%), Gaps = 69/513 (13%)
Query: 38 DGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGD 97
DG R +I G +HY R PE W D + +A GL+ I+ YV WNLHEP G+ FEG GD
Sbjct: 73 DGNRFQIIGGDLHYFRVLPEYWEDRLLRANALGLNTIQVYVPWNLHEPKPGKMVFEGIGD 132
Query: 98 LVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFI-PGIQFRTDNEPFKAEMQRFT 156
LV F+K V LR GPY C EW+ GGFP WL + P +Q RT + + ++R+
Sbjct: 133 LVSFLKLCEKLDFLVMLRAGPYICGEWDLGGFPAWLLAVKPRLQLRTSDPVYLKLVERWW 192
Query: 157 AKIVDMMKQENLYASQGGPIILSQIENEYGN--------------VEGAYGPAAVPYINW 202
V + K L S GGP+I+ QIENEYG+ G G + Y
Sbjct: 193 D--VLLPKVFPLLYSNGGPVIMVQIENEYGSYGNDKAYLRKLVSMARGHLGDDIIVYTTD 250
Query: 203 AASMATSLDTG-VP------WVMCQQENAPDPIINTCNGFYCDQFTPNSKAK-PKFWTEG 254
+ T LD G VP V + P PI F N+ + P +E
Sbjct: 251 GGTKET-LDKGTVPVADVYSAVDFSTGDDPWPIFKLQKKF-------NAPGRSPPLSSEF 302
Query: 255 YNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVA---- 310
Y GWL ++G + E A ++ + G+ YM HGGTNFG +G +
Sbjct: 303 YTGWLTHWGEKITKTDAEFTAASLEKILSRNGSAV-LYMVHGGTNFGFYNGANTGSEESD 361
Query: 311 -----TSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGS------ 359
TSYDYDA I E G I PK+ L+ V K + ++ + GS
Sbjct: 362 YKPDLTSYDYDAPIKESGDIDNPKFQALQRVIKKYNASPHPISPSNKQRKAYGSIKMQMT 421
Query: 360 -NLEAAVYKTEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKIN 418
+L V T+ V I + + +++ G + + S + L K++
Sbjct: 422 TSLFDLVRMTDPADV-----ITSANPISMESVGQMFGFLLYESSYIAKKSGNTLRIPKVH 476
Query: 419 SESMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKAD---SFSRFGLLEQINTTADRS 475
+ + +VG L W ++PI + + + S F L+E + R
Sbjct: 477 DRAQVFVSCLSQDVDVGVLRYIGTTERWNNQPISLPTIECTTNTSLFILVENMG----RV 532
Query: 476 DYLWYSLRIDLEDDAGAQTVLHIESLGHALHAF 508
+Y Y + DD G + ++++ G LH +
Sbjct: 533 NYGPY-----IFDDKGILSSVYLD--GQILHGW 558
Score = 37.4 bits (85), Expect = 0.035
Identities = 26/77 (33%), Positives = 34/77 (43%), Gaps = 28/77 (36%)
Query: 637 IDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYH 696
+ F G GKG A+VN +IGRYWP+ + P CN +
Sbjct: 619 LSFNGWGKGVAFVNEFNIGRYWPS-VGP------QCN---------------------LY 650
Query: 697 VPRSWLQPDSNTLVLFE 713
VP L+ NTLV+FE
Sbjct: 651 VPAPLLKRGKNTLVVFE 667
>At3g53080 unknown protein
Length = 155
Score = 72.8 bits (177), Expect = 8e-13
Identities = 36/92 (39%), Positives = 50/92 (54%), Gaps = 2/92 (2%)
Query: 751 TESDRSGGPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIG 810
T + GP+ + C P VIT I FA +G P G CG+F G+C ++ + IV+K C+G
Sbjct: 63 TNEEPDLGPLTRISCNEPGYVITKINFADYGNPTGTCGHFRRGNCGARATMRIVKKNCLG 122
Query: 811 SSSCSIGVSINTFG-DPCGGVTKSLAVEASCT 841
C + V+ FG C G LAVE +CT
Sbjct: 123 KEKCHLLVTDEMFGPSKCKG-APMLAVETTCT 153
>At3g53050 putative protein
Length = 142
Score = 49.3 bits (116), Expect = 9e-06
Identities = 24/71 (33%), Positives = 36/71 (49%), Gaps = 2/71 (2%)
Query: 755 RSGGPV-LSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQKACIGSSS 813
+ G P+ L +C VI+ I +A +G G+CG F G+C + L+IV K C+
Sbjct: 66 KDGKPIALDFDCEQ-GYVISKITYADYGQSTGSCGKFKRGNCGASNTLNIVNKKCLRKEK 124
Query: 814 CSIGVSINTFG 824
C + V FG
Sbjct: 125 CKLFVPDKIFG 135
>At3g53070 putative protein
Length = 105
Score = 47.8 bits (112), Expect = 3e-05
Identities = 21/53 (39%), Positives = 29/53 (54%)
Query: 754 DRSGGPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHGDCSSKQALSIVQK 806
+RS PVL +C V + I FA +G G+CGNF G C + L +V+K
Sbjct: 46 NRSRAPVLMFDCKEKGYVFSKINFADYGHSSGDCGNFRRGTCGAPDTLRLVKK 98
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,117,122
Number of Sequences: 26719
Number of extensions: 995836
Number of successful extensions: 2254
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 2094
Number of HSP's gapped (non-prelim): 33
length of query: 841
length of database: 11,318,596
effective HSP length: 108
effective length of query: 733
effective length of database: 8,432,944
effective search space: 6181347952
effective search space used: 6181347952
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0136.7


