

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.12
(245 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
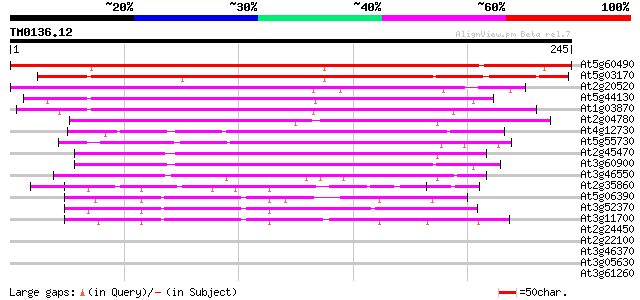
At5g60490 fasciclin-like arabinogalactan protein FLA12 233 7e-62
At5g03170 fasciclin-like arabinogalactan protein FLA11 212 1e-55
At2g20520 fasciclin-like arabinogalactan-protein 6 (Fla6) 159 1e-39
At5g44130 predicted GPI-anchored protein 155 2e-38
At1g03870 predicted GPI-anchored protein 146 1e-35
At2g04780 GPI-anchored protein (Fla7) 117 6e-27
At4g12730 fasciclin-like arabinogalactan protein FLA2 95 4e-20
At5g55730 GPI-anchored protein (Fla1) 91 4e-19
At2g45470 GPI-anchored protein (Fla8) 87 1e-17
At3g60900 GPI-anchored protein (Fla10) 80 1e-15
At3g46550 predicted GPI-anchored protein 80 1e-15
At2g35860 unknown protein 47 7e-06
At5g06390 unknown protein 46 2e-05
At3g52370 unknown protein 45 4e-05
At3g11700 unknown protein 44 8e-05
At2g24450 predicted GPI-anchored protein 32 0.24
At2g22100 putative RNA-binding protein 32 0.40
At3g46370 receptor-like protein kinase homolog 31 0.69
At3g05630 putative phospholipase D 31 0.69
At3g61260 putative DNA-binding protein 30 0.90
>At5g60490 fasciclin-like arabinogalactan protein FLA12
Length = 249
Score = 233 bits (594), Expect = 7e-62
Identities = 126/248 (50%), Positives = 171/248 (68%), Gaps = 5/248 (2%)
Query: 1 MKHQLFTLPLLALMFFFHSTTTLAASSPAPAPSS-SPTDIIRILKKAGGYTTLIRLLQTT 59
M+H L L L+ + L+ SPA AP+ PT++ +IL+KAG +T IRLL++T
Sbjct: 1 MEHSLIILLFTVLLLLTTTPGILSQPSPAVAPAPPGPTNVTKILEKAGQFTVFIRLLKST 60
Query: 60 QVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSN 119
VA Q+ QL NS+ G+T FAP+D++F+ LK G LNSL D+Q+ ELIQFH++P++VS SN
Sbjct: 61 GVANQLYGQLNNSDNGITIFAPSDSSFTGLKAGTLNSLTDEQQVELIQFHVIPSYVSSSN 120
Query: 120 FDTLSNPVRTQAGENPD-RLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKV 178
F T+SNP+RTQAG++ D LNVT+SGNTVN+T+G+ N TV G+VYSD QLAVYQVDKV
Sbjct: 121 FQTISNPLRTQAGDSADGHFPLNVTTSGNTVNITSGVTNTTVSGNVYSDGQLAVYQVDKV 180
Query: 179 LLPRDFFVAKPPAPAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPL-AVAMV 237
LLP+ F +PPAPAPAP +K+ KKK DD++ + D A S R V AV+
Sbjct: 181 LLPQQVFDPRPPAPAPAPSVSKSKKKK--DDSDSSSDDSPADASFALRNVGSVCDAVSFC 238
Query: 238 VVAAFSSW 245
V++ +W
Sbjct: 239 VMSVMLAW 246
>At5g03170 fasciclin-like arabinogalactan protein FLA11
Length = 246
Score = 212 bits (540), Expect = 1e-55
Identities = 123/234 (52%), Positives = 159/234 (67%), Gaps = 7/234 (2%)
Query: 13 LMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINS 72
L FF T +PAP PS PT+I IL+KAG +T IRLL++TQ + QIN QL +S
Sbjct: 12 LFIFFLVIATTYGQAPAPGPSG-PTNITAILEKAGQFTLFIRLLKSTQASDQINTQLNSS 70
Query: 73 NA-GLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQA 131
++ GLT FAP DNAF++LK G LNSL+DQQK +L+QFH+LPT ++M F T+SNP+RTQA
Sbjct: 71 SSNGLTVFAPTDNAFNSLKSGTLNSLSDQQKVQLVQFHVLPTLITMPQFQTVSNPLRTQA 130
Query: 132 GENPD-RLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPP 190
G+ + + LN+TSSGN VN+TTG+V+ TV SVYSD QLAVYQVD+VLLP F
Sbjct: 131 GDGQNGKFPLNITSSGNQVNITTGVVSATVANSVYSDKQLAVYQVDQVLLPLAMF-GSSV 189
Query: 191 APAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPLAVAMVVVAAFSS 244
APAPAPEK + K SA + G G DS S +R +G + + VAA ++
Sbjct: 190 APAPAPEKGGSVSKGSA--SGGDDGGDSTDSSDAERTGFG-FGIRITTVAAIAA 240
>At2g20520 fasciclin-like arabinogalactan-protein 6 (Fla6)
Length = 247
Score = 159 bits (403), Expect = 1e-39
Identities = 96/232 (41%), Positives = 136/232 (58%), Gaps = 12/232 (5%)
Query: 1 MKHQLFTLPLLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQ 60
M LF+ +L + F ++PAP SP ++ IL+ +TTLI+LL TTQ
Sbjct: 1 MSSSLFSYVVLLIFLFTIPYIQSQPTAPAPTTEKSPINLTAILEAGHQFTTLIQLLNTTQ 60
Query: 61 VATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNF 120
V Q++ QL +S+ G+T FAP DNAF+ LKPG LNSL QQ+ +L+ +H++P + S+S+
Sbjct: 61 VGFQVSVQLNSSDQGMTIFAPTDNAFNKLKPGTLNSLTYQQQIQLMLYHIIPKYYSLSDL 120
Query: 121 DTLSNPVRTQA-GENPDRLALNVT--SSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDK 177
SNPVRTQA G++ LN T + N VN++TG+V + ++ LAVY VD
Sbjct: 121 LLASNPVRTQATGQDGGVFGLNFTGQAQSNQVNVSTGVVETRINNALRQQFPLAVYVVDS 180
Query: 178 VLLPRDFFVAK-PPAPAPAPEKAKASKKKSADDTEGPAGDD---SAAVSVKQ 225
VLLP + F K P APAP+ + S+ D + PA DD SA SVK+
Sbjct: 181 VLLPEELFGTKTTPTGAPAPKSS-----TSSSDADSPAADDEHKSAGSSVKR 227
>At5g44130 predicted GPI-anchored protein
Length = 247
Score = 155 bits (391), Expect = 2e-38
Identities = 91/218 (41%), Positives = 128/218 (57%), Gaps = 14/218 (6%)
Query: 7 TLPLLALMF---FFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVAT 63
T PLL L+ F + T ++PAP P+ P +I IL+K G + TLIRLL TTQ+
Sbjct: 3 TTPLLLLLLTAVFLSTEITAQRAAPAPGPAG-PINITAILEKGGQFVTLIRLLNTTQIGN 61
Query: 64 QINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTL 123
QIN Q+ +S+ G+T AP DNAF NLKPG LN L+ + +LI +H+ P F ++ + ++
Sbjct: 62 QINIQINSSSEGMTVLAPTDNAFQNLKPGTLNKLSPDDQVKLILYHVSPKFYTLEDLLSV 121
Query: 124 SNPVRTQAG--ENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLP 181
SNPVRTQA + LN T GN VN++TG+V + S+ + LAVY VD VLLP
Sbjct: 122 SNPVRTQASGRDVGGVYGLNFTGQGNQVNVSTGVVETRLSTSLRQERPLAVYVVDMVLLP 181
Query: 182 RDFFVAKPPAPAPAPEKAKA--------SKKKSADDTE 211
+ F + +P P K+K+ S KK+A +E
Sbjct: 182 EEMFGERKISPMAPPPKSKSPDVSDDSESSKKAAAPSE 219
>At1g03870 predicted GPI-anchored protein
Length = 247
Score = 146 bits (368), Expect = 1e-35
Identities = 88/230 (38%), Positives = 129/230 (55%), Gaps = 4/230 (1%)
Query: 4 QLFTLPLLALMFFFHST-TTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVA 62
+L PLL + +T T ++PAP P+ P ++ IL+K G +TT I LL TQV
Sbjct: 5 RLTLAPLLLIAAVLLATKATAQPAAPAPEPAG-PINLTAILEKGGQFTTFIHLLNITQVG 63
Query: 63 TQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDT 122
+Q+N Q+ +S+ G+T FAP DNAF NLKPG LN L+ + +LI +H+ P + SM + +
Sbjct: 64 SQVNIQVNSSSEGMTVFAPTDNAFQNLKPGTLNQLSPDDQVKLILYHVSPKYYSMDDLLS 123
Query: 123 LSNPVRTQA-GENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLP 181
+SNPVRTQA G + LN T N +N++TG V + S+ LAVY VD VLLP
Sbjct: 124 VSNPVRTQASGRDNGVYGLNFTGQTNQINVSTGYVETRISNSLRQQRPLAVYVVDMVLLP 183
Query: 182 RDFFVAKPPAP-APAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWG 230
+ F +P APAP+ + T+ A + S ++++ G
Sbjct: 184 GEMFGEHKLSPIAPAPKSKSGGVTDDSGSTKKAASPSDKSGSGEKKVGLG 233
>At2g04780 GPI-anchored protein (Fla7)
Length = 254
Score = 117 bits (293), Expect = 6e-27
Identities = 75/215 (34%), Positives = 112/215 (51%), Gaps = 8/215 (3%)
Query: 27 SPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAF 86
+PAPAP+ ++ +L AG + T + L +T V Q N+ G+T F P D+AF
Sbjct: 35 TPAPAPAPENVNLTELLSVAGPFHTFLDYLLSTGVIETFQNQANNTEEGITIFVPKDDAF 94
Query: 87 SNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTL--SNPVRTQAGENPDRLALNVTS 144
K L++L Q +L+ FH LP + S+S F L S PV T AG + +L T
Sbjct: 95 KAQKNPPLSNLTKDQLKQLVLFHALPHYYSLSEFKNLSQSGPVSTFAG---GQYSLKFTD 151
Query: 145 SGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF---VAKPPAPAPAPEKAKA 201
TV + + V SV+S + +AVYQV++VLLP F V PAPAPAP +
Sbjct: 152 VSGTVRIDSLWTRTKVSSSVFSTDPVAVYQVNRVLLPEAIFGTDVPPMPAPAPAPIVSAP 211
Query: 202 SKKKSADDTEGPAGDDSAAVSVKQRLVWGPLAVAM 236
S S D+EG + S+ + Q+L+ P+++ +
Sbjct: 212 SDSPSVADSEGASSPKSSHKNSGQKLLLAPISMVI 246
>At4g12730 fasciclin-like arabinogalactan protein FLA2
Length = 403
Score = 94.7 bits (234), Expect = 4e-20
Identities = 62/193 (32%), Positives = 99/193 (51%), Gaps = 8/193 (4%)
Query: 26 SSPAPAPSSSPTDII--RILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPND 83
S A AP++SP+D+I IL+K G +L++T + + GLT F P+D
Sbjct: 176 SPEAEAPTASPSDLILTTILEKQG-CKAFSDILKSTGADKTFQDTV---DGGLTVFCPSD 231
Query: 84 NAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVT 143
+A P F SL+ K L+ +H +P + S+ + + V T A E ++ V
Sbjct: 232 SAVGKFMPKF-KSLSPANKTALVLYHGMPVYQSLQMLRSGNGAVNTLATEGNNKFDFTVQ 290
Query: 144 SSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKASK 203
+ G V + T +V V G++ L VY++DKVLLPR+ + A + APAP+ +K
Sbjct: 291 NDGEDVTLETDVVTAKVMGTLKDQEPLIVYKIDKVLLPREIYKAVKTS-APAPKSSKKKP 349
Query: 204 KKSADDTEGPAGD 216
K + D +GP+ D
Sbjct: 350 KNAEADADGPSAD 362
Score = 29.6 bits (65), Expect = 1.5
Identities = 33/152 (21%), Positives = 64/152 (41%), Gaps = 24/152 (15%)
Query: 34 SSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGF 93
S+ +I RIL K ++T L T +A +IN + +T A +++A S++
Sbjct: 24 SNAHNITRILAKDPDFSTFNHYLSATHLADEINRR-----QTITVLAVDNSAMSSI---L 75
Query: 94 LNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNTVNMTT 153
N + Q ++ H+L + +++ + A ++ S + T+
Sbjct: 76 SNGYSLYQIRNILSLHVLVDYFGTKKLHQITDGSTSTA---------SMFQSTGSATGTS 126
Query: 154 GIVNVT--VGGSVY-----SDNQLAVYQVDKV 178
G +N+T GG V D++L + V V
Sbjct: 127 GYINITDIKGGKVAFGVQDDDSKLTAHYVKSV 158
>At5g55730 GPI-anchored protein (Fla1)
Length = 424
Score = 91.3 bits (225), Expect = 4e-19
Identities = 66/215 (30%), Positives = 101/215 (46%), Gaps = 25/215 (11%)
Query: 22 TLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAP 81
T AA +PAPA + + + A G L T A++ + + G+T F P
Sbjct: 174 TAAAPTPAPAEMN-----LTGIMSAHGCKVFAETLLTNPGASKTYQESLEG--GMTVFCP 226
Query: 82 NDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALN 141
D+A P + N L +K + F +PT+ SM+ + + P+ T A + ++ L
Sbjct: 227 GDDAMKGFLPKYKN-LTAPKKEAFLDFLAVPTYYSMAMLKSNNGPMNTLATDGANKFELT 285
Query: 142 VTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVA---KPPAPAPAPE- 197
V + G V + T I V + ++ + LA+Y DKVLLP++ F A + PAPAPAPE
Sbjct: 286 VQNDGEKVTLKTRINTVKIVDTLIDEQPLAIYATDKVLLPKELFKASAVEAPAPAPAPED 345
Query: 198 ------------KAKASKKKSADDTEG-PAGDDSA 219
KAK KKK+A + P GD +
Sbjct: 346 GDVADSPKAAKGKAKGKKKKAAPSPDNDPFGDSDS 380
>At2g45470 GPI-anchored protein (Fla8)
Length = 420
Score = 86.7 bits (213), Expect = 1e-17
Identities = 58/181 (32%), Positives = 89/181 (49%), Gaps = 5/181 (2%)
Query: 29 APAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSN 88
APAPS+S ++I +L+KAG T L+ + + T +A GLT FAP+D AF
Sbjct: 180 APAPSASLSNITGLLEKAGCKTFANLLVSSGVLKTYESAV----EKGLTVFAPSDEAFKA 235
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNT 148
L L + L+++H L + + T N + T A + L ++SG+
Sbjct: 236 EGVPDLTKLTQAEVVSLLEYHALAEYKPKGSLKTNKNNISTLATNGAGKFDLTTSTSGDE 295
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF-VAKPPAPAPAPEKAKASKKKSA 207
V + TG+ + +V + ++ VD VLLP + F +K P+PAPAPE A A
Sbjct: 296 VILHTGVAPSRLADTVLDATPVVIFTVDNVLLPAELFGKSKSPSPAPAPEPVTAPTPSPA 355
Query: 208 D 208
D
Sbjct: 356 D 356
>At3g60900 GPI-anchored protein (Fla10)
Length = 422
Score = 80.1 bits (196), Expect = 1e-15
Identities = 56/189 (29%), Positives = 88/189 (45%), Gaps = 7/189 (3%)
Query: 29 APAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSN 88
APAPSS+ I L + G T LL ++ V + + GLT FAP+D AF
Sbjct: 180 APAPSSAGVSNITGLLEKAGCKTFANLLVSSGVIKTFESTV---EKGLTVFAPSDEAFKA 236
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVRTQAGENPDRLALNVTSSGNT 148
L +L + L+++H L + + T + + T A + L ++SG+
Sbjct: 237 RGVPDLTNLTQAEVVSLLEYHALAEYKPKGSLKTNKDAISTLATNGAGKYDLTTSTSGDE 296
Query: 149 VNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKA---SKKK 205
V + TG+ + +V + + ++ VD VLLP + F K +PAPAPE A + K
Sbjct: 297 VILHTGVGPSRLADTVVDETPVVIFTVDNVLLPAELF-GKSSSPAPAPEPVSAPTPTPAK 355
Query: 206 SADDTEGPA 214
S E P+
Sbjct: 356 SPSPVEAPS 364
>At3g46550 predicted GPI-anchored protein
Length = 420
Score = 80.1 bits (196), Expect = 1e-15
Identities = 57/196 (29%), Positives = 106/196 (54%), Gaps = 10/196 (5%)
Query: 20 TTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFF 79
T T +S + +P + ++ +IL + + LL + V T+ AG+T F
Sbjct: 189 TLTPPPTSTSLSPPPAGINLTQILINGHNFNVALSLLVASGVITEFEND--ERGAGITVF 246
Query: 80 APNDNAFSNLKPGF-LNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNPVR-TQAGEN--P 135
P D+AFS+L L SL +QK +++FH+L ++ ++ + ++++NPV+ T A E
Sbjct: 247 VPTDSAFSDLPSNVNLQSLPAEQKAFVLKFHVLHSYYTLGSLESITNPVQPTLATEEMGA 306
Query: 136 DRLALNVTS-SGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFF--VAKPPAP 192
LN++ +G+ V + +G+V V + + N ++V+ V KVLLP++ F +P A
Sbjct: 307 GSYTLNISRVNGSIVTINSGVVLAVVTQTAFDQNPVSVFGVSKVLLPKELFPKSGQPVAT 366
Query: 193 APAPEKAKASKKKSAD 208
AP P++ S + S++
Sbjct: 367 AP-PQEISLSPESSSE 381
Score = 29.3 bits (64), Expect = 2.0
Identities = 38/153 (24%), Positives = 64/153 (40%), Gaps = 12/153 (7%)
Query: 61 VATQINAQLINSNAGLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLPTFVSMSNF 120
V++ I A+L N+ LT A ++ FS+ L +L++FH+L F+S S+
Sbjct: 49 VSSGIAAELSGRNS-LTLLAVPNSQFSSASLDLTRRLPPSALADLLRFHVLLQFLSDSDL 107
Query: 121 DTL--SNPVRTQAGENPDRL-----ALNVT---SSGNTVNMTTGIVNVTVGGSVYS-DNQ 169
+ S T E R ++NVT +SG+ + NVTV + +
Sbjct: 108 RRIPPSGSAVTTLYEASGRTFFGSGSVNVTRDPASGSVTIGSPATKNVTVLKLLETKPPN 167
Query: 170 LAVYQVDKVLLPRDFFVAKPPAPAPAPEKAKAS 202
+ V VD +++P + P P S
Sbjct: 168 ITVLTVDSLIVPTGIDITASETLTPPPTSTSLS 200
>At2g35860 unknown protein
Length = 445
Score = 47.4 bits (111), Expect = 7e-06
Identities = 49/194 (25%), Positives = 82/194 (42%), Gaps = 18/194 (9%)
Query: 25 ASSPAPAPS---------SSPTDIIRILKKAGGYTTLIRLL-QTTQVATQINAQLINSNA 74
A +PAP P + D I L GGY + +L T +AT++ +L++
Sbjct: 237 APAPAPGPGGPRGHFNGDAQVKDFIHTLLHYGGYNEMADILVNLTSLATEMG-RLVSEGY 295
Query: 75 GLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLP---TFVSMSNFDTLSNPVRTQA 131
LT APND A + L L+ + +++ +H++P T SM N V+ +
Sbjct: 296 VLTVLAPNDEAMAKLTTDQLSEPGAPE--QIMYYHIIPEYQTEESMYNAVRRFGKVKYDS 353
Query: 132 GENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPA 191
P ++ G + +Y+D +++V +D VL P++ A
Sbjct: 354 LRFPHKVLAQEADGSVKFGHGDGSAYL-FDPDIYTDGRISVQGIDGVLFPKEETPATEIK 412
Query: 192 PAPAPEKAKASKKK 205
PA AP K SK +
Sbjct: 413 PA-APVVKKVSKSR 425
Score = 43.5 bits (101), Expect = 1e-04
Identities = 42/178 (23%), Positives = 83/178 (46%), Gaps = 17/178 (9%)
Query: 10 LLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQL 69
LL L T L + P P +S + ++ +L YT L L++ + + +
Sbjct: 11 LLLLFLTTSIATALPDNKPVPGQINSNSVLVALLDSH--YTELAELVEKALLLQTLEEAV 68
Query: 70 INSNAGLTFFAPNDNAFS-NLKPGFLNSL----NDQQKNELIQFHLLPTFVSMSNFDTLS 124
N +T FAP ++A NL P F + L N + L+ FH+LP ++ + +LS
Sbjct: 69 GKHN--ITIFAPRNDALERNLDPLFKSFLLEPRNLKSLQSLLMFHILPKRITSPQWPSLS 126
Query: 125 NPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPR 182
+ RT + ++ L++T NT+ + + + + + D ++ ++++L+PR
Sbjct: 127 HHHRTLSNDH-----LHLTVDVNTLKVDSAEI-IRPDDVIRPDG--IIHGIERLLIPR 176
>At5g06390 unknown protein
Length = 458
Score = 45.8 bits (107), Expect = 2e-05
Identities = 52/203 (25%), Positives = 84/203 (40%), Gaps = 41/203 (20%)
Query: 25 ASSPAPAPSSSP---------TDIIRILKKAGGYTTLIRLL-QTTQVATQINAQLINSNA 74
A +PAP P D I L GGY + +L T +AT++ +L++
Sbjct: 248 APAPAPGPGGKQHHFDGEAQVKDFIHTLLHYGGYNEMADILVNLTSLATEMG-RLVSEGY 306
Query: 75 GLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLP---TFVSMSN---------FDT 122
LT APND A + L L+ + +++ +H++P T SM N FDT
Sbjct: 307 VLTVLAPNDEAMAKLTTDQLSEPGAPE--QIVYYHIIPEYQTEESMYNSVRRFGKVKFDT 364
Query: 123 LSNPVRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTV-GGSVYSDNQLAVYQVDKVLLP 181
L P + A E + +V G + + +Y+D +++V +D VL P
Sbjct: 365 LRFPHKVAAKE-----------ADGSVKFGDGEKSAYLFDPDIYTDGRISVQGIDGVLFP 413
Query: 182 RD----FFVAKPPAPAPAPEKAK 200
++ V KP P + K
Sbjct: 414 QEEEVVESVKKPVKKIVQPRRGK 436
>At3g52370 unknown protein
Length = 436
Score = 45.1 bits (105), Expect = 4e-05
Identities = 44/193 (22%), Positives = 79/193 (40%), Gaps = 17/193 (8%)
Query: 25 ASSPAPAPS---------SSPTDIIRILKKAGGYTTLIRLL-QTTQVATQINAQLINSNA 74
A +PAP P + D I L GGY + +L T +AT++ +L++
Sbjct: 229 APAPAPGPGGPRHHFNGEAQVKDFIHTLLHYGGYNEMADILVNLTSLATEMG-RLVSEGY 287
Query: 75 GLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLP---TFVSMSNFDTLSNPVRTQA 131
LT APND A + L L+ + +++ +H++P T SM N +R +
Sbjct: 288 VLTVLAPNDEAMAKLTTDQLSEPGAPE--QIMYYHIIPEYQTEESMYNSVRRFGKIRYDS 345
Query: 132 GENPDRLALNVTSSGNTVNMTTGIVNVTVGGSVYSDNQLAVYQVDKVLLPRDFFVAKPPA 191
P ++ G + +Y+D +++V +D VL P + +
Sbjct: 346 LRFPHKVEAQEADGSVKFGHGDGSAYL-FDPDIYTDGRISVQGIDGVLFPEEKTPVEKKT 404
Query: 192 PAPAPEKAKASKK 204
P +KA ++
Sbjct: 405 GVPVVKKAPKPRR 417
Score = 37.4 bits (85), Expect = 0.007
Identities = 50/208 (24%), Positives = 83/208 (39%), Gaps = 26/208 (12%)
Query: 10 LLALMFFFHSTTTLAASSPAPAPSSSPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQL 69
LL + S TT P +S + ++ +L YT L L++ + + +
Sbjct: 7 LLFFLLLTISITTALPDKPGSGQINSNSVLVALLDSH--YTELAELVEKALLLQTLEEAV 64
Query: 70 INSNAGLTFFAPNDNAFS-NLKPGFLNSL----NDQQKNELIQFHLLPTFVSMSNFDTLS 124
N +T FAP ++A NL P F + L N + L+ FH+LP ++ F +
Sbjct: 65 GQHN--ITIFAPRNDALEKNLDPEFKSFLLQPKNLKSLQSLLMFHILPKRITSPQFSSAV 122
Query: 125 NPVRTQAGENPDRLALNVTSSGNTV--------NMTTGIVNVTVGGSVYSD--NQLAVYQ 174
RT + ++ V S+ T + GI + + SV D + ++
Sbjct: 123 VSHRTLSNDHLHFTNGKVNSAEITKPDDLTRPDGIIHGIERLLIPRSVQEDFNRRRSLRS 182
Query: 175 VDKVLL-------PRDFFVAKPPAPAPA 195
+ VL PR + K PAP PA
Sbjct: 183 IAAVLPEGAPEVDPRTHRLKKKPAPIPA 210
>At3g11700 unknown protein
Length = 462
Score = 43.9 bits (102), Expect = 8e-05
Identities = 52/212 (24%), Positives = 89/212 (41%), Gaps = 23/212 (10%)
Query: 25 ASSPAPAPSSSPT---------DIIRILKKAGGYTTLIRLL-QTTQVATQINAQLINSNA 74
A +PAP P + D I L GGY + +L T +AT++ +L++
Sbjct: 251 APAPAPGPGGAHKHFNGDAQVKDFIHTLLHYGGYNEMADILVNLTSLATEMG-RLVSEGY 309
Query: 75 GLTFFAPNDNAFSNLKPGFLNSLNDQQKNELIQFHLLP---TFVSMSNFDTLSNPVRTQA 131
LT APND A L L+ + +++ +H++P T SM N V+ +
Sbjct: 310 VLTVLAPNDEAMGKLTTDQLSEPGAPE--QIMYYHIIPEYQTEESMYNSVRRFGKVKYET 367
Query: 132 GENPDRLALNVTSSGNTVNMTTGIVNVTV-GGSVYSDNQLAVYQVDKVLLP---RDFFVA 187
P + + + +V +G + + +Y+D +++V +D VL P + V
Sbjct: 368 LRFPHK--VGAKEADGSVKFGSGDRSAYLFDPDIYTDGRISVQGIDGVLFPEEKEEETVK 425
Query: 188 KPPAPAPAPEKAKASK-KKSADDTEGPAGDDS 218
KP P + + K + A G G DS
Sbjct: 426 KPTGPVKKVVQPRRGKLLEVACSMLGAIGKDS 457
>At2g24450 predicted GPI-anchored protein
Length = 280
Score = 32.3 bits (72), Expect = 0.24
Identities = 16/43 (37%), Positives = 25/43 (57%)
Query: 191 APAPAPEKAKASKKKSADDTEGPAGDDSAAVSVKQRLVWGPLA 233
+PAPAP+K+ K +AD+ E P+ + +S LV G +A
Sbjct: 233 SPAPAPDKSGKKKMAAADEAEPPSSASNTGLSFGAVLVLGFVA 275
Score = 32.0 bits (71), Expect = 0.31
Identities = 45/202 (22%), Positives = 80/202 (39%), Gaps = 28/202 (13%)
Query: 35 SPTDIIRILKKAGGYTTLIRLLQTTQVATQINAQLINSNAGLTFFAPNDNAFSNLKPGFL 94
S +I R+L+K ++T+ LL T++ +IN +T A N++A ++
Sbjct: 23 SAVNITRVLEKYPEFSTMTELLAKTELTP-----IINKRQTITVLALNNDAIGSIS---- 73
Query: 95 NSLNDQQKNELIQFHLLPTF-----VSMSNFDTLSNPVRTQAGENPDRLA-LNVTSSGNT 148
++ KN L+ +L F ++ TL + G + LN T S
Sbjct: 74 GRPEEEVKNILMNHVVLDYFDELKLKALKEKSTLLTTLYQSTGLGQQQNGFLNCTKSNGK 133
Query: 149 VNMTTGIVNV--------TVGGSVYSDNQLAVYQVDKVLLPRDFF--VAKPPAPAPAPEK 198
+ +G+ TV + Y+ L+V Q+ ++ V PP P +
Sbjct: 134 IYFGSGVKGAPQTAEYITTVFRNPYN---LSVVQISMPIVAPGLGSPVKVPPPPPMSSPP 190
Query: 199 AKASKKKSADDTEGPAGDDSAA 220
A + KK +A PA + A
Sbjct: 191 APSPKKGAATPAPAPADEGDYA 212
>At2g22100 putative RNA-binding protein
Length = 382
Score = 31.6 bits (70), Expect = 0.40
Identities = 14/27 (51%), Positives = 22/27 (80%), Gaps = 3/27 (11%)
Query: 188 KPPAPAPAPEKAKASKKK---SADDTE 211
K P+P+P+P+K+K SKKK S+D++E
Sbjct: 68 KSPSPSPSPKKSKESKKKHKRSSDESE 94
>At3g46370 receptor-like protein kinase homolog
Length = 793
Score = 30.8 bits (68), Expect = 0.69
Identities = 40/130 (30%), Positives = 58/130 (43%), Gaps = 17/130 (13%)
Query: 127 VRTQAGENPDRLALNVTSSGNTVNMTTGIVNVTVGGSV-YSDNQLAVYQVD-----KVLL 180
+ T P ++LN++SSG T N+ TGI N+T + S+N L + K LL
Sbjct: 314 IDTNVSTPPRIISLNLSSSGLTGNIATGIQNLTKLQKLDLSNNNLTGVVPEFLANMKSLL 373
Query: 181 PRDFFVAKPPAPAPAPEKAKASKKKSAD---DTEGPAGDDSAAVS------VKQRLVWGP 231
D + K P+ KKK D + GDD+ +S +K L+
Sbjct: 374 FID--LRKNKLNGSIPKTLLDRKKKGLQLFVDGDDDKGDDNKCLSGSCVPKMKFPLMIVA 431
Query: 232 LAVAMVVVAA 241
LAV+ VVV A
Sbjct: 432 LAVSAVVVIA 441
>At3g05630 putative phospholipase D
Length = 1039
Score = 30.8 bits (68), Expect = 0.69
Identities = 22/67 (32%), Positives = 29/67 (42%), Gaps = 2/67 (2%)
Query: 89 LKPGFLNSLNDQQKNELIQFHLLPTFVSMSNFDTLSNP-VRTQAGE-NPDRLALNVTSSG 146
LKPGFL L D +L+ + T ++ P + Q E NP R VTS
Sbjct: 259 LKPGFLALLEDPFSGKLLDIMVFDTLGLQGTKESSEQPRLAEQVKEHNPLRFGFKVTSGD 318
Query: 147 NTVNMTT 153
TV + T
Sbjct: 319 RTVRLRT 325
>At3g61260 putative DNA-binding protein
Length = 212
Score = 30.4 bits (67), Expect = 0.90
Identities = 18/49 (36%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Query: 187 AKPPAPAPAPEKAKASKKKSADDTEGPAG----DDSAAVSVKQRLVWGP 231
A+ PAPAPAP A +K + + + P DDS A++V ++ V P
Sbjct: 31 AEIPAPAPAPTPADVTKDVAEEKIQNPPPEQIFDDSKALTVVEKPVEEP 79
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.130 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,239,084
Number of Sequences: 26719
Number of extensions: 219957
Number of successful extensions: 1297
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 1192
Number of HSP's gapped (non-prelim): 85
length of query: 245
length of database: 11,318,596
effective HSP length: 97
effective length of query: 148
effective length of database: 8,726,853
effective search space: 1291574244
effective search space used: 1291574244
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0136.12


