

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.9
(1610 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
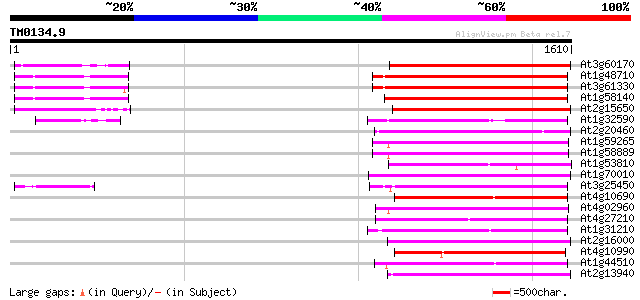
Score E
Sequences producing significant alignments: (bits) Value
At3g60170 putative protein 488 e-137
At1g48710 hypothetical protein 480 e-135
At3g61330 copia-type polyprotein 477 e-134
At1g58140 hypothetical protein 475 e-134
At2g15650 putative retroelement pol polyprotein 454 e-127
At1g32590 hypothetical protein, 5' partial 414 e-115
At2g20460 putative retroelement pol polyprotein 404 e-112
At1g59265 polyprotein, putative 400 e-111
At1g58889 polyprotein, putative 399 e-111
At1g53810 395 e-109
At1g70010 hypothetical protein 393 e-109
At3g25450 hypothetical protein 392 e-109
At4g10690 retrotransposon like protein 392 e-108
At4g02960 putative polyprotein of LTR transposon 391 e-108
At4g27210 putative protein 391 e-108
At1g31210 putative reverse transcriptase 390 e-108
At2g16000 putative retroelement pol polyprotein 389 e-108
At4g10990 putative retrotransposon polyprotein 386 e-107
At1g44510 polyprotein, putative 381 e-105
At2g13940 putative retroelement pol polyprotein 379 e-104
>At3g60170 putative protein
Length = 1339
Score = 488 bits (1256), Expect = e-137
Identities = 246/521 (47%), Positives = 351/521 (67%), Gaps = 3/521 (0%)
Query: 1091 EETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTK 1150
E + L +++ +P D+A++DK W AME E+ KN+ W L P+ IG K
Sbjct: 812 ENLSVMLLMMMTEADPIQFDDAVKDKIWREAMEHEIESIVKNNTWELTTLPKGFTPIGVK 871
Query: 1151 WVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVL 1210
WV++ KLNE G+V + KARLVA+GY+Q GIDYTE FAPVARL+ +R +++ S N +
Sbjct: 872 WVYKTKLNEDGEVDKYKARLVAKGYAQCYGIDYTEVFAPVARLDTVRTILAISSQFNWEI 931
Query: 1211 HQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFL 1270
Q+DVKSAFL+G + EEVYV QP GF E + + V+KL+K+LYGLKQAPRAWY R+ ++
Sbjct: 932 FQLDVKSAFLHGELKEEVYVRQPEGFIREGEEEKVYKLRKALYGLKQAPRAWYSRIEAYF 991
Query: 1271 LENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMG 1330
L+ EF R + TLF KT +ILIV +YVDD+IF +++++C EF + M EFEMS +G
Sbjct: 992 LKEEFERCPSEHTLFTKTRVGNILIVSLYVDDLIFTGSDKAMCDEFKKSMMLEFEMSDLG 1051
Query: 1331 ELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVC 1390
++K+FLGI+V Q+ G +I Q +Y +E+L +F M ES K P+ P L K++ KV
Sbjct: 1052 KMKHFLGIEVKQSDGGIFICQRRYAREVLARFGMDESNAVKNPIVPGTKLTKDENGEKVD 1111
Query: 1391 QKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMY 1450
+ +++ ++GSL+YLT +RPD+++ V L +RF S+PR +H A KRILRYLKGT LG+ Y
Sbjct: 1112 ETMFKQLVGSLMYLTVTRPDLMYGVCLISRFMSNPRMSHWLAAKRILRYLKGTVELGIFY 1171
Query: 1451 --KKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYI 1508
+K KL + D+DYAGD +R+STSG + S + WASK+Q +ALST EAEYI
Sbjct: 1172 RRRKNRSLKLMAFTDSDYAGDLNDRRSTSGFVFLMASGAICWASKKQPVVALSTTEAEYI 1231
Query: 1509 SAAICSTQMLWMKHQLEDYQILE-SNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIR 1567
+AA C+ Q +W++ LE E S I CDN++ I LSK+P+LH ++KHIEV++H++R
Sbjct: 1232 AAAFCACQCVWLRKVLEKLGAEEKSATVINCDNSSTIQLSKHPVLHGKSKHIEVRFHYLR 1291
Query: 1568 DYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
D V V+ L++ T+ Q ADIFTKPL ++F + L M
Sbjct: 1292 DLVNGDVVKLEYCPTEDQVADIFTKPLKLEQFEKLRALLGM 1332
Score = 90.1 bits (222), Expect = 9e-18
Identities = 72/331 (21%), Positives = 143/331 (42%), Gaps = 46/331 (13%)
Query: 13 PMFDGQRFEYWKDRLESFFLGFDADLWDIIVDGYERPVDADGKKIPRSEMTAEQKKLYSQ 72
P FDG +++W +E+F +LW ++ +G V E+ KL +
Sbjct: 13 PRFDGY-YDFWSMTMENFLRS--RELWRLVEEGIPAIVVGTTPVSEAQRSAVEEAKL--K 67
Query: 73 HHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIM 132
K + L AI E + I D+ +K I++S+K ++G+ KVK ++ +L +++E M
Sbjct: 68 DLKVKNFLFQAIDREILETILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAM 127
Query: 133 EPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENM 192
+ E I+ R +V ++ + V +++R L + +V SIE + D+ +
Sbjct: 128 KEGEKIDTFLGRTLTVVNKMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEESNDLSTL 187
Query: 193 SLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGE 252
S++EL L HE + + V++ +AL+ EE G
Sbjct: 188 SIDELHGSLLVHEQRLN------------------GHVQEEQALKVTHEERPSQGRGRG- 228
Query: 253 DELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCP 312
F+GS +G+ G+ ++ V C++C GH++ +CP
Sbjct: 229 -------------------VFRGS---RGRGRGRGRSGTNRAIVECYKCHNLGHFQYECP 266
Query: 313 KLKKDKKPKKHFKTKKSLMVTFDESESEDVD 343
+ +K+ + + ++ L++ + E + D
Sbjct: 267 EWEKNANYAELEEEEELLLMAYVEQNQANRD 297
>At1g48710 hypothetical protein
Length = 1352
Score = 480 bits (1236), Expect = e-135
Identities = 252/564 (44%), Positives = 349/564 (61%), Gaps = 8/564 (1%)
Query: 1041 EPEEDEPEEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETL----LS 1096
E +E EP E PS T + T++ +E + + R RS E T L+
Sbjct: 772 EEDEPEPTREEPPSEEPTTPPTSPTSSQIEE---SSSERTPRFRSIQELYEVTENQENLT 828
Query: 1097 LKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNK 1156
L L + EP EA++ K W AM+EE+ KND W L P IG KWV++ K
Sbjct: 829 LFCLFAECEPMDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKTIGVKWVYKAK 888
Query: 1157 LNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVK 1216
N KG+V R KARLVA+GY Q+ GIDY E FAPVARLE +RL+IS + + +HQMDVK
Sbjct: 889 KNSKGEVERYKARLVAKGYIQRAGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVK 948
Query: 1217 SAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFV 1276
SAFLNG + EEVY+ QP G+ + + D V +LKK+LYGLKQAPRAW R+ + E +F+
Sbjct: 949 SAFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFI 1008
Query: 1277 RGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFL 1336
+ + L+ K K+DILI +YVDD+IF N S+ +EF + M EFEM+ +G + Y+L
Sbjct: 1009 KCPYEHALYIKIQKEDILIACLYVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYL 1068
Query: 1337 GIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRG 1396
GI+V Q G +I Q Y KE+LKKF M +S TPM L K+++ V ++
Sbjct: 1069 GIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKS 1128
Query: 1397 MIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EY 1456
++GSL YLT +RPDIL++V + +R+ P TH A KRILRY+KGT N GL Y T +Y
Sbjct: 1129 LVGSLRYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDY 1188
Query: 1457 KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQ 1516
KL GY D+D+ GD +RKSTSG ++G +W SK+Q + LST EAEY++A C
Sbjct: 1189 KLVGYSDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCH 1248
Query: 1517 MLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVL 1575
+W+++ L++ + E I+ DN +AI+L+KNP+ H R+KHI+ +YH+IR+ V K +
Sbjct: 1249 AIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDV 1308
Query: 1576 LLKFVDTDHQWADIFTKPLAEDRF 1599
L++V T Q ADIFTKPL + F
Sbjct: 1309 QLEYVKTHDQVADIFTKPLKREDF 1332
Score = 94.7 bits (234), Expect = 4e-19
Identities = 72/341 (21%), Positives = 158/341 (46%), Gaps = 32/341 (9%)
Query: 13 PMFDGQRFEYWKDRLESFFLGFDADLWDIIVDGYERPVDADGKKIPRSEMTAEQKKLYSQ 72
P+ ++ W R+++ LG D+W+I+ G+ P + + + + +K +
Sbjct: 11 PVLTKSNYDNWSLRMKAI-LGAH-DVWEIVEKGFIEPENEGSLSQTQKDGLRDSRK---R 65
Query: 73 HHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIM 132
KA ++ + + ++K+ + AK ++ L+ S++G +VK+ + +L ++E+ M
Sbjct: 66 DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 133 EPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENM 192
+ E + + FSR + ++ + + +V+R L + +VT IE T+D+E M
Sbjct: 126 KEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAM 185
Query: 193 SLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGE 252
++E+L+ L+ +E K+ + +D+ +E+ +Q +EE+ ++ + G
Sbjct: 186 TIEQLLGSLQAYEEKKKKKEDI---------------IEQVLNMQITKEENGQSYQRRGG 230
Query: 253 DELTLISK---RLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSI--REVTCFECKESGHY 307
++ + R W+ + G+ + G KS V C+ C + GHY
Sbjct: 231 GQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHY 290
Query: 308 KSDC--PKLKKDKKPKKHFKTK-----KSLMVTFDESESED 341
S+C P KK ++ + + K LM ++ + E E+
Sbjct: 291 ASECKAPSNKKFEEKANYVEEKIQEEDMLLMASYKKDEQEE 331
>At3g61330 copia-type polyprotein
Length = 1352
Score = 477 bits (1228), Expect = e-134
Identities = 250/564 (44%), Positives = 348/564 (61%), Gaps = 8/564 (1%)
Query: 1041 EPEEDEPEEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETL----LS 1096
E +E EP E PS T + T++ +E + + R RS E T L+
Sbjct: 772 EEDEPEPTREEPPSEEPTTPPTSPTSSQIEE---SSSERTPRFRSIQELYEVTENQENLT 828
Query: 1097 LKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNK 1156
L L + EP +A++ K W AM+EE+ KND W L P IG KWV++ K
Sbjct: 829 LFCLFAECEPMDFQKAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAK 888
Query: 1157 LNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVK 1216
N KG+V R KARLVA+GYSQ+ GIDY E FAPVARLE +RL+IS + + +HQMDVK
Sbjct: 889 KNSKGEVERYKARLVAKGYSQRVGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVK 948
Query: 1217 SAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFV 1276
SAFLNG + EEVY+ QP G+ + + D V +LKK LYGLKQAPRAW R+ + E +F+
Sbjct: 949 SAFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFI 1008
Query: 1277 RGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFL 1336
+ + L+ K K+DILI +YVDD+IF N S+ +EF + M EFEM+ +G + Y+L
Sbjct: 1009 KCPYEHALYIKIQKEDILIACLYVDDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYL 1068
Query: 1337 GIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRG 1396
GI+V Q G +I Q Y KE+LKKF + +S TPM L K+++ V ++
Sbjct: 1069 GIEVKQEDNGIFITQEGYAKEVLKKFKIDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKS 1128
Query: 1397 MIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EY 1456
++GSL YLT +RPDIL++V + +R+ P TH A KRILRY+KGT N GL Y T +Y
Sbjct: 1129 LVGSLRYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDY 1188
Query: 1457 KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQ 1516
KL GY D+D+ GD +RKSTSG ++G +W SK+Q + LST EAEY++A C
Sbjct: 1189 KLVGYSDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCH 1248
Query: 1517 MLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVL 1575
+W+++ L++ + E I+ DN +AI+L+KNP+ H R+KHI+ +YH+IR+ V K +
Sbjct: 1249 AIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDV 1308
Query: 1576 LLKFVDTDHQWADIFTKPLAEDRF 1599
L++V T Q AD FTKPL + F
Sbjct: 1309 QLEYVKTHDQVADFFTKPLKRENF 1332
Score = 92.8 bits (229), Expect = 1e-18
Identities = 73/342 (21%), Positives = 158/342 (45%), Gaps = 34/342 (9%)
Query: 13 PMFDGQRFEYWKDRLESFFLGFDADLWDIIVDGYERPVDADGKKIPRSEMTAEQKKLYSQ 72
P+ ++ W R+++ LG D+W+I+ G+ P + + + + +K +
Sbjct: 11 PVLTKSNYDNWSLRMKAI-LGAH-DVWEIVEKGFIEPENEGSLSQTQKDGLRDSRK---R 65
Query: 73 HHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIM 132
KA ++ + + ++K+ + AK ++ L+ S++G +VK+ + +L ++E+ M
Sbjct: 66 DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 133 EPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENM 192
+ E + + FSR + ++ + + +V+R L + +VT IE T+D+E M
Sbjct: 126 KEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAM 185
Query: 193 SLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGE 252
++E+L+ L+ +E K+ + +D+ E+ +Q +EE+ ++ + G
Sbjct: 186 TIEQLLGSLQAYEEKKKKKEDI---------------AEQVLNMQITKEENGQSYQRRGG 230
Query: 253 DELTLISK---RLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSI--REVTCFECKESGHY 307
++ + R W+ + G+ + G KS V C+ C + GHY
Sbjct: 231 GQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHY 290
Query: 308 KSDCPKLKKDKK--PKKHFKTKK------SLMVTFDESESED 341
S+C K +KK K H+ +K LM ++ + E ++
Sbjct: 291 ASEC-KAPSNKKFEEKAHYVEEKIQEEDMLLMASYKKDEQKE 331
>At1g58140 hypothetical protein
Length = 1320
Score = 475 bits (1223), Expect = e-134
Identities = 244/529 (46%), Positives = 336/529 (63%), Gaps = 7/529 (1%)
Query: 1077 KDEPVRTRSAFRPSEE-----TLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSK 1131
+D+P TR PSEE T + + EP EA++ K W AM+EE+ K
Sbjct: 773 EDKPEPTREE-PPSEEPTTPPTSPTSSQIEEKCEPMDFQEAIEKKTWRNAMDEEIKSIQK 831
Query: 1132 NDVWSLVKKPENVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVA 1191
ND W L P IG KWV++ K N KG+V R KARLVA+GYSQ+ GIDY E FAPVA
Sbjct: 832 NDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRAGIDYDEVFAPVA 891
Query: 1192 RLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKS 1251
RLE +RL+IS + + +HQMDVKSAFLNG + EEVY+ QP G+ + + D V +LKK+
Sbjct: 892 RLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKA 951
Query: 1252 LYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQS 1311
LYGLKQAPRAW R+ + E +F++ + L+ K K+DILI +YVDD+IF N S
Sbjct: 952 LYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIFTGNNPS 1011
Query: 1312 LCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAK 1371
+ +EF + M EFEM+ +G + Y+LGI+V Q G +I Q Y KE+LKKF M +S
Sbjct: 1012 MFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVC 1071
Query: 1372 TPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLT 1431
TPM L K+++ V ++ ++GSL YLT +RPDIL++V + +R+ P TH
Sbjct: 1072 TPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHPTTTHFK 1131
Query: 1432 AVKRILRYLKGTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWA 1491
A KRILRY+KGT N GL Y T +YKL GY D+D+ GD +RKSTSG ++G +W
Sbjct: 1132 AAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGDTAFTWM 1191
Query: 1492 SKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNP 1550
SK+Q + LST EAEY++A C +W+++ L++ + E I+ DN +AI+L+KNP
Sbjct: 1192 SKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNP 1251
Query: 1551 ILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 1599
+ H R+KHI+ +YH+IR+ V K + L++V T Q ADIFTKPL + F
Sbjct: 1252 VFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIFTKPLKREDF 1300
Score = 95.1 bits (235), Expect = 3e-19
Identities = 73/341 (21%), Positives = 158/341 (45%), Gaps = 32/341 (9%)
Query: 13 PMFDGQRFEYWKDRLESFFLGFDADLWDIIVDGYERPVDADGKKIPRSEMTAEQKKLYSQ 72
P+ ++ W R+++ LG D+W+I+ G+ P + + + + +K +
Sbjct: 11 PVLTKSNYDNWSLRMKAI-LGAH-DVWEIVEKGFIEPENEGSLSQTQKDGLRDSRK---R 65
Query: 73 HHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIM 132
KA ++ + + ++K+ + AK ++ L+ S++G +VK+ + +L ++E+ M
Sbjct: 66 DKKALCLIYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 133 EPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENM 192
+ E + + FSR + ++ + + +V+R L + +VT IE T+D+E M
Sbjct: 126 KEGELVSDYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAM 185
Query: 193 SLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGE 252
++E+L+ L+ +E K+ + +D+ VE+ +Q +EE+ ++ + G
Sbjct: 186 TIEQLLGSLQAYEEKKKKKEDI---------------VEQVLNMQITKEENGQSYQRRGG 230
Query: 253 DELTLISK---RLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSI--REVTCFECKESGHY 307
++ + R W+ + G+ + G KS V C+ C + GHY
Sbjct: 231 GQVRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHY 290
Query: 308 KSDC--PKLKKDKKPKKHFKTK-----KSLMVTFDESESED 341
S+C P KK ++ + + K LM ++ + E E+
Sbjct: 291 ASECKAPSNKKFEEKANYVEEKIQEEDMLLMASYKKDEQEE 331
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 454 bits (1169), Expect = e-127
Identities = 230/513 (44%), Positives = 329/513 (63%), Gaps = 5/513 (0%)
Query: 1100 LVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNKLNE 1159
LV+ EP++ DEA DK+W AM EE+ KN W LV KPE +VI KW+++ K +
Sbjct: 831 LVANEEPQTYDEARGDKEWEEAMNEEIKVIEKNRTWKLVDKPEKKNVISVKWIYKIKTDA 890
Query: 1160 KGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAF 1219
G+ V++KARLVA+G+SQ+ GIDY ETFAPV+R + IR L++++ L+QMDVKSAF
Sbjct: 891 SGNHVKHKARLVARGFSQEYGIDYLETFAPVSRYDTIRALLAYAAQMKWRLYQMDVKSAF 950
Query: 1220 LNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGK 1279
LNG + EEVYV QPPGF E K + V +L K+LYGLKQAPRAWYER+ S+ ++N F R
Sbjct: 951 LNGELEEEVYVTQPPGFVIEGKEEKVLRLYKALYGLKQAPRAWYERIDSYFIQNGFARSM 1010
Query: 1280 VDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQ 1339
D L+ K +D+LIV +YVDD+I N L F + M+ EFEM+ +G L YFLG++
Sbjct: 1011 NDAALYSKKKGEDVLIVSLYVDDLIITGNNTHLINTFKKNMKDEFEMTDLGLLNYFLGME 1070
Query: 1340 VDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCI---LEKEDKSGKVCQKLYRG 1396
V+Q G ++ Q KY +L+ KF M ES TP+ P +E +DK K YR
Sbjct: 1071 VNQDDSGIFLSQEKYANKLIDKFGMKESKSVSTPLTPQGKRKGVEGDDKEFADPTK-YRR 1129
Query: 1397 MIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EY 1456
++G LLYL ASRPD++++ +R+ S P H KR+LRY+KGT+N G+++
Sbjct: 1130 IVGGLLYLCASRPDVMYASSYLSRYMSSPSIQHYQEAKRVLRYVKGTSNFGVLFTSKETP 1189
Query: 1457 KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQ 1516
+L GY D+D+ G ++KST+G LG + W S +Q T+A STAEAEYI+ + Q
Sbjct: 1190 RLVGYSDSDWGGSLEDKKSTTGYVFTLGLAMFCWQSCKQQTVAQSTAEAEYIAVCAATNQ 1249
Query: 1517 MLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVL 1575
+W++ ED+ + + IPI CDN +AI++ +NP+ H R KHIE+KYHF+R+ KG++
Sbjct: 1250 AIWLQRLFEDFGLKFKEGIPILCDNKSAIAIGRNPVQHRRTKHIEIKYHFVREAEHKGLI 1309
Query: 1576 LLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
L++ + Q AD+ TK L+ RF + + L +
Sbjct: 1310 QLEYCKGEDQLADVLTKALSVSRFEGLRRKLGV 1342
Score = 92.4 bits (228), Expect = 2e-18
Identities = 72/335 (21%), Positives = 157/335 (46%), Gaps = 39/335 (11%)
Query: 13 PMFDGQRFEYWKDRLESFFLGFDADLWDIIVDGYE-RPVDADGKKIPRSEMTAEQKKLYS 71
P+FDG+++++W ++ + F LW ++ +G PV A+ T ++ + +
Sbjct: 10 PIFDGEKYDFWSIKMATIFR--TRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAV-T 66
Query: 72 QHHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFI 131
A IL +A++ + + +I +K +D LK ++G+ +V+ K SL ++YE+
Sbjct: 67 NDTMALQILQTAVTDQIFSRIAAASSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENLK 126
Query: 132 MEPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVEN 191
M N++I+ + +L + + T + +++ LP + +V+ +E TRD++
Sbjct: 127 MYDNDNIKTFTDKLIVLEIQLTYHGEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLDA 186
Query: 192 MSLEELISILKCHELKRSEMQDLRKK-SIALKSKSEKAKVEKSKALQAEEEESEEASEDS 250
+++ EL+ ILK E + + ++ K+ + ++SK ++ ++ ++ +
Sbjct: 187 LTMSELLGILKAQEARVTAREESTKEGAFYVRSKGRESGFKQDNTNNRVNQDKKWCG--- 243
Query: 251 GEDELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSD 310
++ KH + + + K K + G+ K S + C++C + GHY ++
Sbjct: 244 -----------FHKSSKHTEEEC----REKPKNDDHGKNKRS--NIKCYKCGKIGHYANE 286
Query: 311 CPKLKKDKKPKKHFKTKKSLMVTFDESESEDVDSD 345
C K K+ VT +E EDV+ D
Sbjct: 287 C-----------RSKNKERAHVTLEE---EDVNED 307
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 414 bits (1065), Expect = e-115
Identities = 224/578 (38%), Positives = 337/578 (57%), Gaps = 44/578 (7%)
Query: 1027 GFNVSDKGKAPEEVEPEEDEPEEEAGPSNSQTLK----KSRITAAHPKELILGNKDEPVR 1082
G ++ G+ +E EE+E E N + + R K+ ++GN +
Sbjct: 711 GPEINHNGQQDQEETEEEEETVAETVHQNLPAVGTGGVRQRQQPVWMKDYVVGNARVLIT 770
Query: 1083 TRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPE 1142
+ E+ +L+L + +P +EA Q + W AME E+ +N+ W LV+ PE
Sbjct: 771 -----QDEEDEVLAL--FIGPDDPVCFEEAAQLEVWRKAMEAEITSIEENNTWELVELPE 823
Query: 1143 NVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISF 1202
VIG KW+F+ K NEKG+V + KARLVA+GY Q+ G+D+ E FAPVA+ + IRL++
Sbjct: 824 EAKVIGLKWIFKTKFNEKGEVDKFKARLVAKGYHQRYGVDFYEVFAPVAKWDTIRLILGL 883
Query: 1203 SVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAW 1262
+ + Q+DVKSAFL+G + E+V+V QP GFE E++ V+KLKK+LYGLKQAPRAW
Sbjct: 884 AAEKGWSVFQLDVKSAFLHGDLKEDVFVEQPKGFEVEEESSKVYKLKKALYGLKQAPRAW 943
Query: 1263 YERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQA 1322
Y R+ F + F + + TLF K + D L+V +YVDD+I+ ++ + + F M
Sbjct: 944 YSRIEEFFGKEGFEKCYCEHTLFVKKERSDFLVVSVYVDDLIYTGSSMEMIEGFKNSMME 1003
Query: 1323 EFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEK 1382
EF M+ +G++KYFLG++V Q G +I+Q KY E++KK+ M K P+ P +K
Sbjct: 1004 EFAMTDLGKMKYFLGVEVIQDERGIFINQRKYAAEIIKKYGMEGCNSVKNPIVPG---QK 1060
Query: 1383 EDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKG 1442
K+G V +R+ P E HL AVKRILRY++G
Sbjct: 1061 LTKAGAV-----------------------------SRYMESPNEQHLLAVKRILRYVQG 1091
Query: 1443 TTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALST 1502
T +LG+ Y++ +L G+ D+DYAGD +RKSTSG LG ++WASK+Q + LST
Sbjct: 1092 TLDLGIQYERGGATELVGFVDSDYAGDVDDRKSTSGYVFMLGGGAIAWASKKQPIVTLST 1151
Query: 1503 AEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEV 1561
EAE++SA+ + Q +W+++ LE+ E ++CDN++ I LSKNP+LH R+KHI V
Sbjct: 1152 TEAEFVSASYGACQAVWLRNVLEEIGCRQEGGTLVFCDNSSTIKLSKNPVLHGRSKHIHV 1211
Query: 1562 KYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 1599
+YHF+R+ V++G + L + T Q ADI TK + + F
Sbjct: 1212 RYHFLRELVKEGTIRLDYCTTTDQVADIMTKAVKREVF 1249
Score = 80.5 bits (197), Expect = 7e-15
Identities = 55/244 (22%), Positives = 111/244 (44%), Gaps = 38/244 (15%)
Query: 74 HKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIME 133
HK + L ++I + I +E +K +++S+K ++GN +V+ ++ L + +E M+
Sbjct: 26 HKVKNYLFASIDKTILKTILQKETSKDLWESMKRKYQGNDRVQSAQLQRLRRSFEVLEMK 85
Query: 134 PNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLPESWMPLVTSIELTRDVENMS 193
E+I FSR + +R L + V +++R L E + +V +IE + +++ ++
Sbjct: 86 IGETITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTLVEKFTYVVCAIEESNNIKELT 145
Query: 194 LEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEESEEASEDSGED 253
++ L S L HE Q+L + + + + L+AE + + G
Sbjct: 146 VDGLQSSLMVHE------QNLSRHDV------------EERVLKAETQWRPDGGRGRGGS 187
Query: 254 ELTLISKRLNRIWKHRQSKFKGSGKAKGKYESSGQKKSSIREVTCFECKESGHYKSDCPK 313
G+ +G Y+ G+ + V CF+C + GHYK++CP
Sbjct: 188 --------------------PSRGRGRGGYQGRGRGYVNRDTVECFKCHKMGHYKAECPS 227
Query: 314 LKKD 317
+K+
Sbjct: 228 WEKE 231
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 404 bits (1038), Expect = e-112
Identities = 219/574 (38%), Positives = 338/574 (58%), Gaps = 15/574 (2%)
Query: 1047 PEEEAGPSNSQTLKKSRITA--AHPKEL--ILGNKDE--PVRTRSAFRP-SEETLLSLKG 1099
P + PS + K RIT AH ++ NKD+ P+ + ++ S +L +
Sbjct: 883 PSPQISPSTQ--ISKRRITKFPAHLQDYHCYFVNKDDSHPISSSLSYSQISPSHMLYINN 940
Query: 1100 LVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNKLNE 1159
+ + P+S EA K+W A+++E+ + D W + P +G KWVF K +
Sbjct: 941 ISKIPIPQSYHEAKDSKEWCGAIDQEIGAMERTDTWEITSLPPGKKAVGCKWVFTVKFHA 1000
Query: 1160 KGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAF 1219
G + R KAR+VA+GY+Q+EG+DYTETF+PVA++ ++LL+ S + L+Q+D+ +AF
Sbjct: 1001 DGSLERFKARIVAKGYTQKEGLDYTETFSPVAKMATVKLLLKVSASKKWYLNQLDISNAF 1060
Query: 1220 LNGYISEEVYVHQPPGFEDEK----KPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEF 1275
LNG + E +Y+ P G+ D K P+ V +LKKS+YGLKQA R W+ + S+ LL F
Sbjct: 1061 LNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLKKSIYGLKQASRQWFLKFSNSLLALGF 1120
Query: 1276 VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYF 1335
+ D TLF + + +++ +YVDDI+ S + + +E ++A F++ +G LKYF
Sbjct: 1121 EKQHGDHTLFVRCIGSEFIVLLVYVDDIVIASTTEQAAQSLTEALKASFKLRELGPLKYF 1180
Query: 1336 LGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYR 1395
LG++V +T EG + Q KY ELL +ML+ + PM P L K D +++YR
Sbjct: 1181 LGLEVARTSEGISLSQRKYALELLTSADMLDCKPSSIPMTPNIRLSKNDGLLLEDKEMYR 1240
Query: 1396 GMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*E 1455
++G L+YLT +RPDI F+V+ +F S PR HL AV ++L+Y+KGT GL Y +
Sbjct: 1241 RLVGKLMYLTITRPDITFAVNKLCQFSSAPRTAHLAAVYKVLQYIKGTVGQGLFYSAEDD 1300
Query: 1456 YKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICST 1515
L GY DAD+ R+ST+G F+GS+L+SW SK+Q T++ S+AEAEY + A+ S
Sbjct: 1301 LTLKGYTDADWGTCPDSRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASC 1360
Query: 1516 QMLWMKHQLEDYQILESNIPI-YCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGV 1574
+M W+ L ++ S +PI Y D+TAA+ ++ NP+ H R KHIE+ H +R+ + G
Sbjct: 1361 EMAWLSTLLLALRV-HSGVPILYSDSTAAVYIATNPVFHERTKHIEIDCHTVREKLDNGQ 1419
Query: 1575 LLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
L L V T Q ADI TKPL +F +L +++
Sbjct: 1420 LKLLHVKTKDQVADILTKPLFPYQFAHLLSKMSI 1453
>At1g59265 polyprotein, putative
Length = 1466
Score = 400 bits (1029), Expect = e-111
Identities = 225/579 (38%), Positives = 330/579 (56%), Gaps = 16/579 (2%)
Query: 1041 EPEEDEPEEEAGPSNSQTLKKSRITAAHPK----ELILGNKDEPVRTRSA--------FR 1088
+ P S+S T HP +++ N P+ T S +
Sbjct: 880 QSSSSSPSPTTSASSSSTSPTPPSILIHPPPPLAQIVNNNNQAPLNTHSMGTRAKAGIIK 939
Query: 1089 PSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKP-ENVHVI 1147
P+ + L++ L + EP++ +AL+D+ W AM E+N N W LV P +V ++
Sbjct: 940 PNPKYSLAVS-LAAESEPRTAIQALKDERWRNAMGSEINAQIGNHTWDLVPPPPSHVTIV 998
Query: 1148 GTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHN 1207
G +W+F K N G + R KARLVA+GY+Q+ G+DY ETF+PV + +IR+++ +V+ +
Sbjct: 999 GCRWIFTKKYNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRS 1058
Query: 1208 IVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1267
+ Q+DV +AFL G ++++VY+ QPPGF D+ +P++V KL+K+LYGLKQAPRAWY L
Sbjct: 1059 WPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVCKLRKALYGLKQAPRAWYVELR 1118
Query: 1268 SFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMS 1327
++LL FV DT+LF I+ + +YVDDI+ + +L + + F +
Sbjct: 1119 NYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILITGNDPTLLHNTLDNLSQRFSVK 1178
Query: 1328 MMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSG 1387
EL YFLGI+ + P G ++ Q +Y +LL + NM+ + TPM P+ L +
Sbjct: 1179 DHEELHYFLGIEAKRVPTGLHLSQRRYILDLLARTNMITAKPVTTPMAPSPKLSLYSGTK 1238
Query: 1388 KVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1447
YRG++GSL YL +RPDI ++V+ ++F P E HL A+KRILRYL GT N G
Sbjct: 1239 LTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMPTEEHLQALKRILRYLAGTPNHG 1298
Query: 1448 LMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEY 1507
+ KK L Y DAD+AGD+ + ST+G +LG + +SW+SK+Q + S+ EAEY
Sbjct: 1299 IFLKKGNTLSLHAYSDADWAGDKDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEY 1358
Query: 1508 ISAAICSTQMLWMKHQLEDYQILESNIP-IYCDNTAAISLSKNPILHSRAKHIEVKYHFI 1566
S A S++M W+ L + I + P IYCDN A L NP+ HSR KHI + YHFI
Sbjct: 1359 RSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNVGATYLCANPVFHSRMKHIAIDYHFI 1418
Query: 1567 RDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF-NFILK 1604
R+ VQ G L + V T Q AD TKPL+ F NF K
Sbjct: 1419 RNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQNFASK 1457
>At1g58889 polyprotein, putative
Length = 1466
Score = 399 bits (1025), Expect = e-111
Identities = 224/579 (38%), Positives = 329/579 (56%), Gaps = 16/579 (2%)
Query: 1041 EPEEDEPEEEAGPSNSQTLKKSRITAAHPK----ELILGNKDEPVRTRSA--------FR 1088
+ P S+S T HP +++ N P+ T S +
Sbjct: 880 QSSSSSPSPTTSASSSSTSPTPPSILIHPPPPLAQIVNNNNQAPLNTHSMGTRAKAGIIK 939
Query: 1089 PSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKP-ENVHVI 1147
P+ + L++ L + EP++ +AL+D+ W AM E+N N W LV P +V ++
Sbjct: 940 PNPKYSLAVS-LAAESEPRTAIQALKDERWRNAMGSEINAQIGNHTWDLVPPPPSHVTIV 998
Query: 1148 GTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHN 1207
G +W+F K N G + R KAR VA+GY+Q+ G+DY ETF+PV + +IR+++ +V+ +
Sbjct: 999 GCRWIFTKKYNSDGSLNRYKARFVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRS 1058
Query: 1208 IVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1267
+ Q+DV +AFL G ++++VY+ QPPGF D+ +P++V KL+K+LYGLKQAPRAWY L
Sbjct: 1059 WPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVCKLRKALYGLKQAPRAWYVELR 1118
Query: 1268 SFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMS 1327
++LL FV DT+LF I+ + +YVDDI+ + +L + + F +
Sbjct: 1119 NYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILITGNDPTLLHNTLDNLSQRFSVK 1178
Query: 1328 MMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSG 1387
EL YFLGI+ + P G ++ Q +Y +LL + NM+ + TPM P+ L +
Sbjct: 1179 DHEELHYFLGIEAKRVPTGLHLSQRRYILDLLARTNMITAKPVTTPMAPSPKLSLYSGTK 1238
Query: 1388 KVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1447
YRG++GSL YL +RPDI ++V+ ++F P E HL A+KRILRYL GT N G
Sbjct: 1239 LTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMPTEEHLQALKRILRYLAGTPNHG 1298
Query: 1448 LMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEY 1507
+ KK L Y DAD+AGD+ + ST+G +LG + +SW+SK+Q + S+ EAEY
Sbjct: 1299 IFLKKGNTLSLHAYSDADWAGDKDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEY 1358
Query: 1508 ISAAICSTQMLWMKHQLEDYQILESNIP-IYCDNTAAISLSKNPILHSRAKHIEVKYHFI 1566
S A S++M W+ L + I + P IYCDN A L NP+ HSR KHI + YHFI
Sbjct: 1359 RSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNVGATYLCANPVFHSRMKHIAIDYHFI 1418
Query: 1567 RDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF-NFILK 1604
R+ VQ G L + V T Q AD TKPL+ F NF K
Sbjct: 1419 RNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQNFASK 1457
>At1g53810
Length = 1522
Score = 395 bits (1014), Expect = e-109
Identities = 208/535 (38%), Positives = 320/535 (58%), Gaps = 15/535 (2%)
Query: 1088 RPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVI 1147
+P++ +L L VS+ EPK++ EAL+ W AM+EE+ + + W+LV N++V+
Sbjct: 899 KPNKRYVL-LTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVL 957
Query: 1148 GTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHN 1207
G+ WVFR KL+ G + + KARLVA+G+ Q+EGIDY ET++PV R +RL++ +
Sbjct: 958 GSMWVFRTKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVATVLK 1017
Query: 1208 IVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1267
L QMDVK+AFL+G ++E VY+ QP GF D+ KPDHV L KSLYGLKQ+PRAW++R S
Sbjct: 1018 WELKQMDVKNAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYGLKQSPRAWFDRFS 1077
Query: 1268 SFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMS 1327
+FLLE F+ D +LF + +D++++ +YVDD++ N + EF M
Sbjct: 1078 NFLLEFGFICSLFDPSLFVYSSNNDVILLLLYVDDMVITGNNSQSLTHLLAALNKEFRMK 1137
Query: 1328 MMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSG 1387
MG++ YFLGIQ+ G ++ Q KY ++LL +M + TP+ L++
Sbjct: 1138 DMGQVHYFLGIQIQTYDGGLFMSQQKYAEDLLITASMANCSPMPTPL--PLQLDRVSNQD 1195
Query: 1388 KVCQ--KLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTN 1445
+V +R + G L YLT +RPDI F+V+ + P + +KRILRY+KGT +
Sbjct: 1196 EVFSDPTYFRSLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLKRILRYIKGTVS 1255
Query: 1446 LGLMYKKT---------*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQS 1496
+G+ Y +Y LS Y D+DYA + R+S G C F+G N++SW+SK+Q
Sbjct: 1256 MGIQYNSNSSSVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFMGQNIISWSSKKQP 1315
Query: 1497 TIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIP-IYCDNTAAISLSKNPILHSR 1555
T++ S+ EAEY S + ++++ WM L + + + P ++CDN +A+ L+ NP H+R
Sbjct: 1316 TVSRSSTEAEYRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLSAVYLTANPAFHAR 1375
Query: 1556 AKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNMDF 1610
KH +V +H+IR+ V L++K + Q ADIFTK L + F + L +DF
Sbjct: 1376 TKHFDVDHHYIRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRLRFKLGVDF 1430
>At1g70010 hypothetical protein
Length = 1315
Score = 393 bits (1010), Expect = e-109
Identities = 218/588 (37%), Positives = 330/588 (56%), Gaps = 8/588 (1%)
Query: 1029 NVSDKGKAPE---EVEPEEDEPEEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRS 1085
N SD + E P + PE S+ + K + + + ++ E + S
Sbjct: 721 NPSDSSSSVEILPSANPTNNVPEPSVQTSHRKAKKPAYLQDYYCHSVVSSTPHEIRKFLS 780
Query: 1086 AFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVH 1145
R ++ L L L EP + EA + + W AM E + W + P +
Sbjct: 781 YDRINDPYLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDFLEGTHTWEVCSLPADKR 840
Query: 1146 VIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVN 1205
IG +W+F+ K N G V R KARLVAQGY+Q+EGIDY ETF+PVA+L +++LL+ +
Sbjct: 841 CIGCRWIFKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFSPVAKLNSVKLLLGVAAR 900
Query: 1206 HNIVLHQMDVKSAFLNGYISEEVYVHQPPGFE----DEKKPDHVFKLKKSLYGLKQAPRA 1261
+ L Q+D+ +AFLNG + EE+Y+ P G+ D P+ V +LKKSLYGLKQA R
Sbjct: 901 FKLSLTQLDISNAFLNGDLDEEIYMRLPQGYASRQGDSLPPNAVCRLKKSLYGLKQASRQ 960
Query: 1262 WYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQ 1321
WY + SS LL F++ D T F K L V +Y+DDII S N + M+
Sbjct: 961 WYLKFSSTLLGLGFIQSYCDHTCFLKISDGIFLCVLVYIDDIIIASNNDAAVDILKSQMK 1020
Query: 1322 AEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILE 1381
+ F++ +GELKYFLG+++ ++ +G +I Q KY +LL + L + PM P+ +
Sbjct: 1021 SFFKLRDLGELKYFLGLEIVRSDKGIHISQRKYALDLLDETGQLGCKPSSIPMDPSMVFA 1080
Query: 1382 KEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLK 1441
+ V YR +IG L+YL +RPDI F+V+ A+F PR+ HL AV +IL+Y+K
Sbjct: 1081 HDSGGDFVEVGPYRRLIGRLMYLNITRPDITFAVNKLAQFSMAPRKAHLQAVYKILQYIK 1140
Query: 1442 GTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALS 1501
GT GL Y T E +L Y +ADY R R+STSG C FLG +L+ W S++Q ++ S
Sbjct: 1141 GTIGQGLFYSATSELQLKVYANADYNSCRDSRRSTSGYCMFLGDSLICWKSRKQDVVSKS 1200
Query: 1502 TAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIE 1560
+AEAEY S ++ + +++W+ + L++ Q+ L ++CDN AAI ++ N + H R KHIE
Sbjct: 1201 SAEAEYRSLSVATDELVWLTNFLKELQVPLSKPTLLFCDNEAAIHIANNHVFHERTKHIE 1260
Query: 1561 VKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
H +R+ + KG+ L ++T+ Q AD FTKPL F+ ++ + +
Sbjct: 1261 SDCHSVRERLLKGLFELYHINTELQIADPFTKPLYPSHFHRLISKMGL 1308
>At3g25450 hypothetical protein
Length = 1343
Score = 392 bits (1007), Expect = e-109
Identities = 215/584 (36%), Positives = 340/584 (57%), Gaps = 24/584 (4%)
Query: 1032 DKGKAPEEVEPEEDEPEEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRT--RSAFRP 1089
+ G ++ E +E EE + + + T H + + +PVR R RP
Sbjct: 739 NNGVTENDISTEPEETEEAEINGEDENIIEEAETEEHDQSQ---EEPQPVRRSQRQVIRP 795
Query: 1090 S-------------EETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWS 1136
+ E LL++ EP EA + K+W A +EE+ KN WS
Sbjct: 796 NYLKDYVLCAEIEAEHLLLAVND-----EPWDFKEANKSKEWRDACKEEIQSIEKNRTWS 850
Query: 1137 LVKKPENVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAI 1196
LV P IG KWVF+ K N G + + KARLVA+GY Q+ G+D+ E FAPVAR+E +
Sbjct: 851 LVDLPVGSKAIGVKWVFKLKHNSDGSINKYKARLVAKGYVQRHGVDFEEVFAPVARIETV 910
Query: 1197 RLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLK 1256
RL+I+ + ++ +H +DVK+AFL+G + E+VYV QP GF +++ + V+KL K+LYGL+
Sbjct: 911 RLIIALAASNGWEIHHLDVKTAFLHGELREDVYVSQPEGFTNKESKEKVYKLHKALYGLR 970
Query: 1257 QAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEF 1316
QAPRAW +L+ L E +F + + +L+ K ++IL+V +YVDD++ +N + F
Sbjct: 971 QAPRAWNTKLNEILKELKFEKCHKEPSLYRKQEGENILVVAVYVDDLLVTGSNLDIILNF 1030
Query: 1317 SEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHP 1376
+ M +FEMS +G+L Y+LGI+V Q+ +G + Q +Y K++L++ M + TPM
Sbjct: 1031 KKGMVGKFEMSDLGKLTYYLGIEVLQSKDGITLKQERYAKKILEEAGMSKCNTVNTPMIA 1090
Query: 1377 TCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRI 1436
+ L K ++ + YR IG L YL +RPD+ ++V + +R+ +PRE+H A+K+I
Sbjct: 1091 SLELSKAQDEKRIDETDYRRNIGCLRYLLHTRPDLSYNVGILSRYLQEPRESHGAALKQI 1150
Query: 1437 LRYLKGTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQS 1496
LRYL+GTT+ GL +KK L GY D+ + D + KST G+ +L ++W S++Q
Sbjct: 1151 LRYLQGTTSHGLYFKKGENAGLIGYSDSSHNVDLDDGKSTGGHIFYLNDCPITWCSQKQQ 1210
Query: 1497 TIALSTAEAEYISAAICSTQMLWMKHQLEDYQILE-SNIPIYCDNTAAISLSKNPILHSR 1555
+ LS+ EAE+++A + Q +W++ L + E + I DN +AI+L+KNP+ H R
Sbjct: 1211 VVTLSSCEAEFMAATEAAKQAIWLQELLAEVIGTECEKVTIRVDNKSAIALTKNPVFHGR 1270
Query: 1556 AKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 1599
+KHI +YHFIR+ V+ G + ++ V Q ADI TK L + +F
Sbjct: 1271 SKHIHRRYHFIRECVENGQIEVEHVPGVRQKADILTKALGKIKF 1314
Score = 53.5 bits (127), Expect = 1e-06
Identities = 45/231 (19%), Positives = 107/231 (45%), Gaps = 27/231 (11%)
Query: 13 PMFDGQRFEYWKDRLESFFLGFDADLWDIIVDGYERPVDADGKKIPRSEMTAEQKKLYSQ 72
PM + + W R+E+ LW I G +A+++K
Sbjct: 23 PMLNSVNYTVWTMRMEAVLRVHK--LWGTIEPG-----------------SADEEK---- 59
Query: 73 HHKARAILLSAISYEEYQKITDREFAKGIFDSLKMSHEGNKKVKESKALSLIQKYESFIM 132
+ ARA+L +I ++ ++ + +++++K + G ++VKE++ +L+ +++ M
Sbjct: 60 NDMARALLFQSIPESLILQVGKQKTSSAVWEAIKSRNLGAERVKEARLQTLMAEFDKLKM 119
Query: 133 EPNESIEEMFSRFQLLVAGIRPLNKSYTTKDHVIRVIRCLP-ESWMPLVTSIELTRDVEN 191
+ +E+I++ R + L + V + ++ LP + ++ +V ++E D++
Sbjct: 120 KDSETIDDYVGRISEITTKAAALGEDIEESKIVKKFLKSLPRKKYIHIVAALEQVLDLKT 179
Query: 192 MSLEELISILKCHELKRSEMQDLRKKSIALKSKSEKAKVEKSKALQAEEEE 242
+ E++ +K +E + + D + L ++ E+ V+ L+ EEEE
Sbjct: 180 TTFEDIAGRIKTYEDRVWDDDDSHEDQGKLMTEVEEEVVDD---LEEEEEE 227
>At4g10690 retrotransposon like protein
Length = 1515
Score = 392 bits (1006), Expect = e-108
Identities = 202/496 (40%), Positives = 302/496 (60%), Gaps = 3/496 (0%)
Query: 1105 EPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNKLNEKGDVV 1164
EPKS+ EAL+D+ W AM EE+ + D W LV ++G KWVF+ KLN G +
Sbjct: 919 EPKSVKEALKDEGWTNAMGEEMGTMHETDTWDLVPPEMVDRLLGCKWVFKTKLNSDGSLD 978
Query: 1165 RNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYI 1224
R KARLVA+GY Q+EG+DY ET++PV R +R ++ + + L Q+DVK+AFL+ +
Sbjct: 979 RLKARLVARGYEQEEGVDYVETYSPVVRSATVRSILHVATINKWSLKQLDVKNAFLHDEL 1038
Query: 1225 SEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTL 1284
E V++ QPPGFED +PD+V KLKK++Y LKQAPRAW+++ SS+LL+ F+ D +L
Sbjct: 1039 KETVFMTQPPGFEDPSRPDYVCKLKKAIYDLKQAPRAWFDKFSSYLLKYGFICSFSDPSL 1098
Query: 1285 FCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTP 1344
F D++ + +YVDD+I N L ++ ++ EF M MG L YFLGIQ
Sbjct: 1099 FVYLKGRDVMFLLLYVDDMILTGNNDVLLQQLLNILSTEFRMKDMGALHYFLGIQAHYHN 1158
Query: 1345 EGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYL 1404
+G ++ Q KYT +LL M + + TP+ +L+ +K +R + G L YL
Sbjct: 1159 DGLFLSQEKYTSDLLVNAGMSDCSSMPTPLQ-LDLLQGNNKPFPE-PTYFRRLAGKLQYL 1216
Query: 1405 TASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EYKLSGYCDA 1464
T +RPDI F+V+ + P + +KRIL YLKGT +G+ + L Y D+
Sbjct: 1217 TLTRPDIQFAVNFVCQKMHAPTMSDFHLLKRILHYLKGTMTMGINLSSNTDSVLRCYSDS 1276
Query: 1465 DYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQL 1524
D+AG + R+ST G C FLG N++SW++KR T++ S+ EAEY + + ++++ W+ L
Sbjct: 1277 DWAGCKDTRRSTGGFCTFLGYNIISWSAKRHPTVSKSSTEAEYRTLSFAASEVSWIGFLL 1336
Query: 1525 EDYQILESNIP-IYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTD 1583
++ + + IP +YCDN +A+ LS NP LHSR+KH +V Y+++R+ V G L +K +
Sbjct: 1337 QEIGLPQQQIPEMYCDNLSAVYLSANPALHSRSKHFQVDYYYVRERVALGALTVKHIPAS 1396
Query: 1584 HQWADIFTKPLAEDRF 1599
Q ADIFTK L + F
Sbjct: 1397 QQLADIFTKSLPQAPF 1412
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 391 bits (1005), Expect = e-108
Identities = 219/566 (38%), Positives = 321/566 (56%), Gaps = 10/566 (1%)
Query: 1049 EEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRS-------AFRPSEETLLSLKGLV 1101
E PS+S T +I N PV T S R + L
Sbjct: 875 EPNSPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAKDGIRKPNQKYSYATSLA 934
Query: 1102 SLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPE-NVHVIGTKWVFRNKLNEK 1160
+ EP++ +A++D W AM E+N N W LV P +V ++G +W+F K N
Sbjct: 935 ANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPPPPSVTIVGCRWIFTKKFNSD 994
Query: 1161 GDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1220
G + R KARLVA+GY+Q+ G+DY ETF+PV + +IR+++ +V+ + + Q+DV +AFL
Sbjct: 995 GSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFL 1054
Query: 1221 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1280
G +++EVY+ QPPGF D+ +PD+V +L+K++YGLKQAPRAWY L ++LL FV
Sbjct: 1055 QGTLTDEVYMSQPPGFVDKDRPDYVCRLRKAIYGLKQAPRAWYVELRTYLLTVGFVNSIS 1114
Query: 1281 DTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQV 1340
DT+LF I+ + +YVDDI+ + L K + + F + +L YFLGI+
Sbjct: 1115 DTSLFVLQRGRSIIYMLVYVDDILITGNDTVLLKHTLDALSQRFSVKEHEDLHYFLGIEA 1174
Query: 1341 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGS 1400
+ P+G ++ Q +YT +LL + NML + TPM + L + YRG++GS
Sbjct: 1175 KRVPQGLHLSQRRYTLDLLARTNMLTAKPVATPMATSPKLTLHSGTKLPDPTEYRGIVGS 1234
Query: 1401 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EYKLSG 1460
L YL +RPD+ ++V+ +++ P + H A+KR+LRYL GT + G+ KK L
Sbjct: 1235 LQYLAFTRPDLSYAVNRLSQYMHMPTDDHWNALKRVLRYLAGTPDHGIFLKKGNTLSLHA 1294
Query: 1461 YCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWM 1520
Y DAD+AGD + ST+G +LG + +SW+SK+Q + S+ EAEY S A S+++ W+
Sbjct: 1295 YSDADWAGDTDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSELQWI 1354
Query: 1521 KHQLEDYQILESNIP-IYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKF 1579
L + I S+ P IYCDN A L NP+ HSR KHI + YHFIR+ VQ G L +
Sbjct: 1355 CSLLTELGIQLSHPPVIYCDNVGATYLCANPVFHSRMKHIALDYHFIRNQVQSGALRVVH 1414
Query: 1580 VDTDHQWADIFTKPLAEDRF-NFILK 1604
V T Q AD TKPL+ F NF K
Sbjct: 1415 VSTHDQLADTLTKPLSRVAFQNFSRK 1440
>At4g27210 putative protein
Length = 1318
Score = 391 bits (1004), Expect = e-108
Identities = 208/552 (37%), Positives = 324/552 (58%), Gaps = 4/552 (0%)
Query: 1050 EAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSI 1109
+AG + +T++++ + H R + + L VS EPK++
Sbjct: 659 QAGSDSEETIQQASVNV-HQTHASTNVHPMVTRAKVGISKPNPRYVFLSHKVSYPEPKTV 717
Query: 1110 DEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNKLNEKGDVVRNKAR 1169
AL+ W AM EE+ S+ WSLV ++HV+G+KWVFR KL+ G + + KAR
Sbjct: 718 TAALKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVFRTKLHADGTLNKLKAR 777
Query: 1170 LVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVY 1229
+VA+G+ Q+EGIDY ET++PV R +RL++ + N + QMDVK+AFL+G + E VY
Sbjct: 778 IVAKGFLQEEGIDYLETYSPVVRTPTVRLVLHLATALNWDIKQMDVKNAFLHGDLKETVY 837
Query: 1230 VHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTY 1289
+ QP GF D KPDHV L KS+YGLKQ+PRAW+++ S+FLLE F K D +LF +
Sbjct: 838 MTQPAGFVDPSKPDHVCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCSKSDPSLFIYAH 897
Query: 1290 KDDILIVQIYVDD-IIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTY 1348
++++++ +YVDD +I G+++Q+L + + EF M+ MG+L YFLGIQV + G +
Sbjct: 898 NNNLILLLLYVDDMVITGNSSQTLTSLLA-ALNKEFRMTDMGQLHYFLGIQVQRQQNGLF 956
Query: 1349 IHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASR 1408
+ Q KY ++LL +M T TP+ + +R + G L YLT +R
Sbjct: 957 MSQQKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEELFSDPTYFRSIAGKLQYLTLTR 1016
Query: 1409 PDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EYKLSGYCDADYAG 1468
PDI F+V+ + P + +KRILRY+KGT +G+ Y + L Y D+D+
Sbjct: 1017 PDIQFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITMGISYSRDSPTLLQAYSDSDWGN 1076
Query: 1469 DRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQ 1528
+ R+S G C F+G+NLVSW+SK+ T++ S+ EAEY S + ++++LW+ L + +
Sbjct: 1077 CKQTRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLSDAASEILWLSTLLRELR 1136
Query: 1529 ILESNIP-IYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWA 1587
I + P ++CDN +A+ L+ NP H+R KH ++ +HF+R+ V L++K + Q A
Sbjct: 1137 IPLPDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHFVRERVALKALVVKHIPGSEQIA 1196
Query: 1588 DIFTKPLAEDRF 1599
DIFTK L + F
Sbjct: 1197 DIFTKSLPYEAF 1208
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 390 bits (1003), Expect = e-108
Identities = 206/574 (35%), Positives = 331/574 (56%), Gaps = 12/574 (2%)
Query: 1027 GFNVSDKGKAPEEVEPEEDEPEEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSA 1086
G V+++ PE E +EE P + A +E ++ + R+++
Sbjct: 790 GSQVTEQLTDPEPTSNNEGS-DEEVNPVAEEI--------AANQEQVINSHAMTTRSKAG 840
Query: 1087 FRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHV 1146
+ + ++ EPK++ A++ W A+ EE+N+ WSLV +++++
Sbjct: 841 IQKPNTRYALITSRMNTAEPKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNI 900
Query: 1147 IGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNH 1206
+ +KWVF+ KL+ G + + KARLVA+G+ Q+EG+DY ETF+PV R IRL++ S +
Sbjct: 901 LSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSK 960
Query: 1207 NIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERL 1266
+ Q+DV +AFL+G + E V+++QP GF D +KP HV +L K++YGLKQAPRAW++
Sbjct: 961 GWPIKQLDVSNAFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTF 1020
Query: 1267 SSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEM 1326
S+FLL+ FV K D +LF IL + +YVDDI+ ++QSL ++ + ++ F M
Sbjct: 1021 SNFLLDYGFVCSKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSM 1080
Query: 1327 SMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKS 1386
+G +YFLGIQ++ G ++HQ+ Y ++L++ M + TP+ L+ +
Sbjct: 1081 KDLGPPRYFLGIQIEDYANGLFLHQTAYATDILQQAGMSDCNPMPTPLPQQ--LDNLNSE 1138
Query: 1387 GKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNL 1446
+R + G L YLT +RPDI F+V+ + P + +KRILRY+KGT +
Sbjct: 1139 LFAEPTYFRSLAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGM 1198
Query: 1447 GLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAE 1506
GL K+ LS Y D+D+AG + R+ST+G C LGSNL+SW++KRQ T++ S+ EAE
Sbjct: 1199 GLPIKRNSTLTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAE 1258
Query: 1507 YISAAICSTQMLWMKHQLEDYQILE-SNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHF 1565
Y + + ++ W+ L D I + +YCDN +A+ LS NP LH+R+KH + YH+
Sbjct: 1259 YRALTYAAREITWISFLLRDLGIPQYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHY 1318
Query: 1566 IRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 1599
IR+ V G++ + + Q AD+FTK L F
Sbjct: 1319 IREQVALGLIETQHISATFQLADVFTKSLPRRAF 1352
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 389 bits (1000), Expect = e-108
Identities = 203/530 (38%), Positives = 314/530 (58%), Gaps = 4/530 (0%)
Query: 1083 TRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPE 1142
T S + S + + + + P + EA K+W A++ E+ K + W + P+
Sbjct: 917 TISYSKISPSHMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPK 976
Query: 1143 NVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISF 1202
+G KWVF K G++ R KARLVA+GY+Q+EG+DYT+TF+PVA++ I+LL+
Sbjct: 977 GKKAVGCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKV 1036
Query: 1203 SVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEK----KPDHVFKLKKSLYGLKQA 1258
S + L Q+DV +AFLNG + EE+++ P G+ + K + V +LK+S+YGLKQA
Sbjct: 1037 SASKKWFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGLKQA 1096
Query: 1259 PRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSE 1318
R W+++ SS LL F + D TLF K Y + +IV +YVDDI+ S +++ + +E
Sbjct: 1097 SRQWFKKFSSSLLSLGFKKTHGDHTLFLKMYDGEFVIVLVYVDDIVIASTSEAAAAQLTE 1156
Query: 1319 MMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTC 1378
+ F++ +G+LKYFLG++V +T G I Q KY ELL+ ML PM P
Sbjct: 1157 ELDQRFKLRDLGDLKYFLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSVPMIPNL 1216
Query: 1379 ILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILR 1438
+ K+D + YR ++G L+YLT +RPDI F+V+ +F S PR THLTA R+L+
Sbjct: 1217 KMRKDDGDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAYRVLQ 1276
Query: 1439 YLKGTTNLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTI 1498
Y+KGT GL Y + + L G+ D+D+A + R+ST+ F+G +L+SW SK+Q T+
Sbjct: 1277 YIKGTVGQGLFYSASSDLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTV 1336
Query: 1499 ALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKH 1558
+ S+AEAEY + A+ + +M+W+ L Q +Y D+TAAI ++ NP+ H R KH
Sbjct: 1337 SRSSAEAEYRALALATCEMVWLFTLLVSLQASPPVPILYSDSTAAIYIATNPVFHERTKH 1396
Query: 1559 IEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
I++ H +R+ + G L L V T+ Q ADI TKPL +F + +++
Sbjct: 1397 IKLDCHTVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSI 1446
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 386 bits (992), Expect = e-107
Identities = 197/497 (39%), Positives = 302/497 (60%), Gaps = 9/497 (1%)
Query: 1105 EPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGTKWVFRNKLNEKGDVV 1164
EPK+ +A++ + W A EEL+ +N W + E +V+G KWVF K N G +
Sbjct: 524 EPKAFTQAMKSEKWTRAANEELHALEQNKTWIVESLTEGKNVVGCKWVFTIKYNPDGSIE 583
Query: 1165 RNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYI 1224
R KARLVAQG++QQEGIDY ETF+PVA+ +++LL+ + L QMDV +AFL+G +
Sbjct: 584 RYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLGLAAATGWSLTQMDVSNAFLHGEL 643
Query: 1225 SEEVYVHQPPGFED------EKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRG 1278
EE+Y+ P G+ KP V +L KSLYGLKQA R WY+RLSS L F++
Sbjct: 644 DEEIYMSLPQGYTPPTGISLPSKP--VCRLLKSLYGLKQASRQWYKRLSSVFLGANFIQS 701
Query: 1279 KVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGI 1338
D T+F K I++V +YVDD++ S + S + E++++EF++ +G ++FLG+
Sbjct: 702 PADNTMFVKVSCTSIIVVLVYVDDLMIASNDSSAVENLKELLRSEFKIKDLGPARFFLGL 761
Query: 1339 QVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMI 1398
++ ++ EG + Q KY + LL+ + + PM P L KE + YR ++
Sbjct: 762 EIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDPNLHLTKEMGTLLPNATSYRELV 821
Query: 1399 GSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKT*EYKL 1458
G LLYL +RPDI F+VH ++F S P + H+ A ++LRYLKG GLMY + E L
Sbjct: 822 GRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKVLRYLKGNPGQGLMYSASSELCL 881
Query: 1459 SGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQML 1518
+G+ DAD+ + R+S +G C +LG++L++W SK+QS ++ S+ E+EY S A + +++
Sbjct: 882 NGFSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYRSLAQATCEII 941
Query: 1519 WMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLL 1577
W++ L+D + + ++CDN +A+ L+ NP+ H R KHIE+ H +RD ++ G L
Sbjct: 942 WLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVRDQIKAGKLKT 1001
Query: 1578 KFVDTDHQWADIFTKPL 1594
V T +Q ADI TKPL
Sbjct: 1002 LHVPTGNQLADILTKPL 1018
>At1g44510 polyprotein, putative
Length = 1459
Score = 381 bits (979), Expect = e-105
Identities = 218/574 (37%), Positives = 326/574 (55%), Gaps = 24/574 (4%)
Query: 1047 PEEEAGPSNS---QTLKKSRITAAHPKELILG----NKDEP------VRTRSA---FRPS 1090
P E PS+S + + KS TA P + + ++ +P ++TRS +P
Sbjct: 874 PNPETNPSSSIEQRPVDKSTTTALPPNQTTIAATSNSRSQPPKNNHQMKTRSKNNITKPK 933
Query: 1091 EETLLSLK-GLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENVHVIGT 1149
+T L++ L EP ++ +AL+DK W AM +E + +N W LV H++G
Sbjct: 934 TKTSLTVALTQPHLSEPNTVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPPNPTQHLVGC 993
Query: 1150 KWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIV 1209
+WVF+ K G + + KARLVA+G++QQ G+DY ETF+PV + IR+++ +V N
Sbjct: 994 RWVFKLKYLPNGLIDKYKARLVAKGFNQQYGVDYAETFSPVIKATTIRVVLDVAVKKNWP 1053
Query: 1210 LHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSF 1269
L Q+DV +AFL G ++EEVY+ QPPGF D+ +P HV +L+K++YGLKQAPRAWY L
Sbjct: 1054 LKQLDVNNAFLQGTLTEEVYMAQPPGFVDKDRPSHVCRLRKAIYGLKQAPRAWYMELKQH 1113
Query: 1270 LLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIF-GSANQSLCKEFSEMMQAEFEMSM 1328
LL FV DT+LF ++ +L + +YVDDII GS ++S+ S + + F +
Sbjct: 1114 LLNIGFVNSLADTSLFIYSHGTTLLYLLVYVDDIIVTGSDHKSVSAVLSSLAE-RFSIKD 1172
Query: 1329 MGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPM--HPTCILEKEDKS 1386
+L YFLGI+ +T G ++ Q KY +LL K NML++ TP+ P L K
Sbjct: 1173 PTDLHYFLGIEATRTNTGLHLMQRKYMTDLLAKHNMLDAKPVATPLPTSPKLTLHGGTKL 1232
Query: 1387 GKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNL 1446
+ YR ++GSL YL +RPDI F+V+ ++F P H A KR+LRYL GTT
Sbjct: 1233 NDASE--YRSVVGSLQYLAFTRPDIAFAVNRLSQFMHQPTSDHWQAAKRVLRYLAGTTTH 1290
Query: 1447 GLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAE 1506
G+ + L + DAD+AGD + ST+ +LG N +SW+SK+Q ++ S+ E+E
Sbjct: 1291 GIFLNSSSPIHLHAFSDADWAGDSADYVSTNAYVIYLGRNPISWSSKKQRGVSRSSTESE 1350
Query: 1507 YISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHF 1565
Y + A ++++ W+ L + I L I+CDN A + NP+ HSR KHI + YHF
Sbjct: 1351 YRAVANAASEIRWLCSLLTELHIRLPHGPTIFCDNIGATYICANPVFHSRMKHIALDYHF 1410
Query: 1566 IRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 1599
+R +Q L + V T+ Q AD TK L+ F
Sbjct: 1411 VRGMIQSRALRVSHVSTNDQLADALTKSLSRPHF 1444
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 379 bits (972), Expect = e-104
Identities = 206/525 (39%), Positives = 305/525 (57%), Gaps = 4/525 (0%)
Query: 1085 SAFRPSEETLLSLKGLVSLIEPKSIDEALQDKDWILAMEEELNQFSKNDVWSLVKKPENV 1144
+AF S L+ + +EPK EA+Q K W AM E++ N W +V P
Sbjct: 973 AAFSSSHRAYLA--AITDNVEPKHFKEAVQIKVWNDAMFTEVDALEINKTWDIVDLPPGK 1030
Query: 1145 HVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSV 1204
IG++WVF+ K N G V R KARLV QG Q EG DY ETFAPV R+ +R L+
Sbjct: 1031 VAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQVEGEDYKETFAPVVRMTTVRTLLRNVA 1090
Query: 1205 NHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYE 1264
+ ++QMDV +AFL+G + EEVY+ PPGF PD V +L+KSLYGLKQAPR W++
Sbjct: 1091 ANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFR-HSHPDKVCRLRKSLYGLKQAPRCWFK 1149
Query: 1265 RLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEF 1324
+LS LL FV+ D +LF T + L V IYVDD++ + + ++F + + F
Sbjct: 1150 KLSDSLLRFGFVQSYEDYSLFSYTRNNIELRVLIYVDDLLICGNDGYMLQKFKDYLSRCF 1209
Query: 1325 EMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKED 1384
M +G+LKYFLGI+V + PEG ++ Q KY +++ L S A TP+ L +D
Sbjct: 1210 SMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALDVIADSGNLGSRPAHTPLEQNHHLASDD 1269
Query: 1385 KSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTT 1444
K YR ++G LLYL +RP++ +SVH+ A+F +PRE H A R++RYLKG+
Sbjct: 1270 GPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVLAQFMQNPREAHFDAALRVVRYLKGSP 1329
Query: 1445 NLGLMYKKT*EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAE 1504
G++ + L YCD+D+ R+S S LG + +SW +K+Q T++ S+AE
Sbjct: 1330 GQGILLNADPDLTLEVYCDSDWQSCPLTRRSISAYVVLLGGSPISWKTKKQDTVSHSSAE 1389
Query: 1505 AEYISAAICSTQMLWMKHQLEDYQILESN-IPIYCDNTAAISLSKNPILHSRAKHIEVKY 1563
AEY + + ++ W++ L++ I +S +YCD+ AAI ++ NP+ H R KHIE
Sbjct: 1390 AEYRAMSYALKEIKWLRKLLKELGIEQSTPARLYCDSKAAIHIAANPVFHERTKHIESDC 1449
Query: 1564 HFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNLNM 1608
H +RD V+ G++ + V T Q AD+FTK L ++F +++ L +
Sbjct: 1450 HSVRDAVRDGIITTQHVRTTEQLADVFTKALGRNQFLYLMSKLGV 1494
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.335 0.145 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,417,863
Number of Sequences: 26719
Number of extensions: 1473971
Number of successful extensions: 12989
Number of sequences better than 10.0: 428
Number of HSP's better than 10.0 without gapping: 142
Number of HSP's successfully gapped in prelim test: 296
Number of HSP's that attempted gapping in prelim test: 11110
Number of HSP's gapped (non-prelim): 1456
length of query: 1610
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1497
effective length of database: 8,299,349
effective search space: 12424125453
effective search space used: 12424125453
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0134.9


