

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.4
(311 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
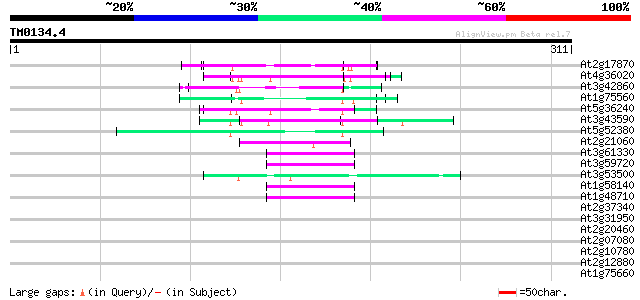
Score E
Sequences producing significant alignments: (bits) Value
At2g17870 putative glycine-rich, zinc-finger DNA-binding protein 69 2e-12
At4g36020 glycine-rich protein 64 1e-10
At3g42860 unknown protein (At3g42860) 60 2e-09
At1g75560 DNA-binding protein 59 3e-09
At5g36240 putative protein 52 5e-07
At3g43590 putative protein 49 3e-06
At5g52380 unknown protein 47 1e-05
At2g21060 glycine-rich protein (AtGRP2) 47 2e-05
At3g61330 copia-type polyprotein 41 7e-04
At3g59720 copia-type reverse transcriptase-like protein 41 7e-04
At3g53500 splicing factor - like protein 41 7e-04
At1g58140 hypothetical protein 41 7e-04
At1g48710 hypothetical protein 41 7e-04
At2g37340 unknown protein 40 0.002
At3g31950 hypothetical protein 38 0.008
At2g20460 putative retroelement pol polyprotein 37 0.010
At2g07080 putative gag-protease polyprotein 37 0.010
At2g10780 pseudogene 36 0.023
At2g12880 pseudogene 36 0.031
At1g75660 Dhp1-like protein 36 0.031
>At2g17870 putative glycine-rich, zinc-finger DNA-binding protein
Length = 301
Score = 69.3 bits (168), Expect = 2e-12
Identities = 41/127 (32%), Positives = 56/127 (43%), Gaps = 26/127 (20%)
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACL-----------GQGPRCFNCNQIGHLAVNCKTF 144
G + + E C+ CG GH+A C G G C++C ++GHLA +C+
Sbjct: 120 GGGGRRSGGEGECYMCGDVGHFARDCRQSGGGNSGGGGGGGRPCYSCGEVGHLAKDCR-- 177
Query: 145 QAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC--------SATGPRCFNCHRV 196
G N+ G RG+ CY CG +GHFA C G C+ C V
Sbjct: 178 -GGSGGNRYGG---GGGRGSGGD-GCYMCGGVGHFARDCRQNGGGNVGGGGSTCYTCGGV 232
Query: 197 GHLAVVC 203
GH+A VC
Sbjct: 233 GHIAKVC 239
Score = 66.2 bits (160), Expect = 2e-11
Identities = 40/120 (33%), Positives = 55/120 (45%), Gaps = 17/120 (14%)
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKG 155
G+ ++ EVT G N+ G G CFNC ++GH+A +C G S K G
Sbjct: 64 GSDGKTKAIEVTAPGGGSLNKKENSSRGSGGNCFNCGEVGHMAKDCD----GGSGGKSFG 119
Query: 156 KFPAIARGTEKQVSCYRCGEIGHFASRCSAT-----------GPRCFNCHRVGHLAVVCK 204
G E + CY CG++GHFA C + G C++C VGHLA C+
Sbjct: 120 GGGGRRSGGEGE--CYMCGDVGHFARDCRQSGGGNSGGGGGGGRPCYSCGEVGHLAKDCR 177
Score = 56.6 bits (135), Expect = 2e-08
Identities = 34/114 (29%), Positives = 45/114 (38%), Gaps = 32/114 (28%)
Query: 108 CFRCGKNGHYANACL--------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPA 159
C+ CG GH+A C G G C+ C +GH+A C + K P+
Sbjct: 198 CYMCGGVGHFARDCRQNGGGNVGGGGSTCYTCGGVGHIAKVCTS------------KIPS 245
Query: 160 IARGTEKQVSCYRCGEIGHFASRCSATGP----------RCFNCHRVGHLAVVC 203
G + +CY CG GH A C G +CF C + GH A C
Sbjct: 246 GGGGGGR--ACYECGGTGHLARDCDRRGSGSSGGGGGSNKCFICGKEGHFAREC 297
Score = 50.8 bits (120), Expect = 9e-07
Identities = 29/87 (33%), Positives = 38/87 (43%), Gaps = 20/87 (22%)
Query: 107 TCFRCGKNGHYANACL--------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFP 158
TC+ CG GH A C G G C+ C GHLA +C +G S
Sbjct: 225 TCYTCGGVGHIAKVCTSKIPSGGGGGGRACYECGGTGHLARDCDRRGSGSSGG------- 277
Query: 159 AIARGTEKQVSCYRCGEIGHFASRCSA 185
G+ K C+ CG+ GHFA C++
Sbjct: 278 --GGGSNK---CFICGKEGHFARECTS 299
Score = 31.6 bits (70), Expect = 0.58
Identities = 15/54 (27%), Positives = 21/54 (38%)
Query: 174 GEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAARGKVY 227
G + + +G CFNC VGH+A C S G+ G+ Y
Sbjct: 80 GSLNKKENSSRGSGGNCFNCGEVGHMAKDCDGGSGGKSFGGGGGRRSGGEGECY 133
>At4g36020 glycine-rich protein
Length = 299
Score = 63.9 bits (154), Expect = 1e-10
Identities = 39/128 (30%), Positives = 55/128 (42%), Gaps = 33/128 (25%)
Query: 108 CFRCGKNGHYANACLGQG------------PRCFNCNQIGHLAVNCKTFQAGPSDNKMKG 155
C+ CG GH+A C G C+ C +GH+A +C G D +
Sbjct: 134 CYNCGDTGHFARDCTSAGNGDQRGATKGGNDGCYTCGDVGHVARDCTQKSVGNGDQR--- 190
Query: 156 KFPAIARGTEKQVSCYRCGEIGHFASRCS---ATG---------PRCFNCHRVGHLAVVC 203
A+ G + CY CG++GHFA C+ A G C++C VGH+A C
Sbjct: 191 --GAVKGGND---GCYTCGDVGHFARDCTQKVAAGNVRSGGGGSGTCYSCGGVGHIARDC 245
Query: 204 KKPKVEPS 211
K +PS
Sbjct: 246 -ATKRQPS 252
Score = 55.5 bits (132), Expect = 4e-08
Identities = 30/98 (30%), Positives = 44/98 (44%), Gaps = 22/98 (22%)
Query: 123 GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASR 182
G G C+NC ++GH++ +C G + +RG E CY CG+ GHFA
Sbjct: 97 GGGSGCYNCGELGHISKDCGIGGGGGGGERR-------SRGGE---GCYNCGDTGHFARD 146
Query: 183 CSATG------------PRCFNCHRVGHLAVVCKKPKV 208
C++ G C+ C VGH+A C + V
Sbjct: 147 CTSAGNGDQRGATKGGNDGCYTCGDVGHVARDCTQKSV 184
Score = 55.1 bits (131), Expect = 5e-08
Identities = 29/98 (29%), Positives = 44/98 (44%), Gaps = 20/98 (20%)
Query: 108 CFRCGKNGHYANACL------------GQGPRCFNCNQIGHLAVNCKT--------FQAG 147
C+ CG GH+A C G C++C +GH+A +C T +Q G
Sbjct: 200 CYTCGDVGHFARDCTQKVAAGNVRSGGGGSGTCYSCGGVGHIARDCATKRQPSRGCYQCG 259
Query: 148 PSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSA 185
S + + + G +CY+CG+ GHFA CS+
Sbjct: 260 GSGHLARDCDQRGSGGGGNDNACYKCGKEGHFARECSS 297
Score = 54.7 bits (130), Expect = 6e-08
Identities = 37/136 (27%), Positives = 51/136 (37%), Gaps = 36/136 (26%)
Query: 108 CFRCGKNGHYANACL-----GQGPR-------CFNCNQIGHLAVNCKTFQAGPSDNKMKG 155
C+ CG+ GH + C G G R C+NC GH A +C + G KG
Sbjct: 102 CYNCGELGHISKDCGIGGGGGGGERRSRGGEGCYNCGDTGHFARDCTSAGNGDQRGATKG 161
Query: 156 KFPAIARGTEKQVSCYRCGEIGHFASRCSATG--------------PRCFNCHRVGHLAV 201
G + CY CG++GH A C+ C+ C VGH A
Sbjct: 162 -------GND---GCYTCGDVGHVARDCTQKSVGNGDQRGAVKGGNDGCYTCGDVGHFAR 211
Query: 202 VCKKPKVEPSVNTARG 217
C + +V + G
Sbjct: 212 DCTQKVAAGNVRSGGG 227
Score = 34.7 bits (78), Expect = 0.068
Identities = 15/47 (31%), Positives = 22/47 (45%), Gaps = 7/47 (14%)
Query: 104 PEVTCFRCGKNGHYANACLGQGP-------RCFNCNQIGHLAVNCKT 143
P C++CG +GH A C +G C+ C + GH A C +
Sbjct: 251 PSRGCYQCGGSGHLARDCDQRGSGGGGNDNACYKCGKEGHFARECSS 297
>At3g42860 unknown protein (At3g42860)
Length = 372
Score = 60.1 bits (144), Expect = 2e-09
Identities = 37/135 (27%), Positives = 54/135 (39%), Gaps = 35/135 (25%)
Query: 100 QSAHPEVTCFRCGKNGHYANACLGQ---GP--------RCFNCNQIGHLAVNCKTFQAGP 148
Q+A C++CGK GH+A C Q GP CF C + GH + +C P
Sbjct: 231 QNAKTGTPCYKCGKEGHWARDCTVQSDTGPVKSTSAAGDCFKCGKPGHWSRDCTAQSGNP 290
Query: 149 SDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC-----------------SATGPRCF 191
P + + CY+CG+ GH++ C S+TG C+
Sbjct: 291 KYE------PGQMKSSSSSGECYKCGKQGHWSRDCTGQSSNQQFQSGQAKSTSSTGD-CY 343
Query: 192 NCHRVGHLAVVCKKP 206
C + GH + C P
Sbjct: 344 KCGKAGHWSRDCTSP 358
Score = 43.5 bits (101), Expect = 1e-04
Identities = 24/91 (26%), Positives = 40/91 (43%), Gaps = 27/91 (29%)
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMK 154
PG + +S+ C++CGK GH++ C GQ + FQ+G
Sbjct: 294 PG-QMKSSSSSGECYKCGKQGHWSRDCTGQSSN--------------QQFQSGQ------ 332
Query: 155 GKFPAIARGTEKQVSCYRCGEIGHFASRCSA 185
A+ T CY+CG+ GH++ C++
Sbjct: 333 ------AKSTSSTGDCYKCGKAGHWSRDCTS 357
Score = 40.4 bits (93), Expect = 0.001
Identities = 22/69 (31%), Positives = 31/69 (44%), Gaps = 11/69 (15%)
Query: 170 CYRCGEIGHFASRC---SATGP--------RCFNCHRVGHLAVVCKKPKVEPSVNTARGK 218
CY+CG+ GH+A C S TGP CF C + GH + C P + K
Sbjct: 239 CYKCGKEGHWARDCTVQSDTGPVKSTSAAGDCFKCGKPGHWSRDCTAQSGNPKYEPGQMK 298
Query: 219 HPAARGKVY 227
++ G+ Y
Sbjct: 299 SSSSSGECY 307
Score = 27.7 bits (60), Expect = 8.3
Identities = 10/34 (29%), Positives = 18/34 (52%)
Query: 88 KSSTTATPGARSQSAHPEVTCFRCGKNGHYANAC 121
+SS +++S C++CGK GH++ C
Sbjct: 322 QSSNQQFQSGQAKSTSSTGDCYKCGKAGHWSRDC 355
>At1g75560 DNA-binding protein
Length = 257
Score = 59.3 bits (142), Expect = 3e-09
Identities = 37/127 (29%), Positives = 50/127 (39%), Gaps = 30/127 (23%)
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMK 154
PG ++ C CG GH A C + RC+NC + GH+A NC
Sbjct: 63 PGHFARDCSNVSVCNNCGLPGHIAAECTAES-RCWNCREPGHVASNC------------- 108
Query: 155 GKFPAIARGTEKQVSCYRCGEIGHFASRCSATGPR------CFNCHRVGHLAVVCKKPKV 208
+ C+ CG+ GH A CS + R C NC + GHLA C K
Sbjct: 109 ----------SNEGICHSCGKSGHRARDCSNSDSRAGDLRLCNNCFKQGHLAADCTNDKA 158
Query: 209 EPSVNTA 215
+ T+
Sbjct: 159 CKNCRTS 165
Score = 45.1 bits (105), Expect = 5e-05
Identities = 42/153 (27%), Positives = 51/153 (32%), Gaps = 43/153 (28%)
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQGPR------CFNCNQIGHLAVNCKTFQAGP 148
PG + + E C CGK+GH A C R C NC + GHLA +C +A
Sbjct: 101 PGHVASNCSNEGICHSCGKSGHRARDCSNSDSRAGDLRLCNNCFKQGHLAADCTNDKAC- 159
Query: 149 SDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC-------SATGPR------------ 189
+ + G IAR C C GH A C S G R
Sbjct: 160 KNCRTSGH---IARDCRNDPVCNICSISGHVARHCPKGDSNYSDRGSRVRDGGMQRGGLS 216
Query: 190 --------------CFNCHRVGHLAVVCKKPKV 208
C NC GH A C +V
Sbjct: 217 RMSRDREGVSAMIICHNCGGRGHRAYECPSARV 249
Score = 44.3 bits (103), Expect = 9e-05
Identities = 25/80 (31%), Positives = 31/80 (38%), Gaps = 24/80 (30%)
Query: 124 QGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
QG C NC + GH A +C C CG GH A+ C
Sbjct: 53 QGNLCNNCKRPGHFARDCSNVSV-----------------------CNNCGLPGHIAAEC 89
Query: 184 SATGPRCFNCHRVGHLAVVC 203
+A RC+NC GH+A C
Sbjct: 90 TAES-RCWNCREPGHVASNC 108
Score = 39.3 bits (90), Expect = 0.003
Identities = 25/87 (28%), Positives = 36/87 (40%), Gaps = 12/87 (13%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
C C +GH A C P C C+ GH+A +C + SD + + + RG +
Sbjct: 159 CKNCRTSGHIARDCRND-PVCNICSISGHVARHCPKGDSNYSDRGSRVRDGGMQRGGLSR 217
Query: 168 VS-----------CYRCGEIGHFASRC 183
+S C+ CG GH A C
Sbjct: 218 MSRDREGVSAMIICHNCGGRGHRAYEC 244
>At5g36240 putative protein
Length = 254
Score = 51.6 bits (122), Expect = 5e-07
Identities = 32/108 (29%), Positives = 42/108 (38%), Gaps = 30/108 (27%)
Query: 108 CFRCGKNGHYANACLGQ-------GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAI 160
C RCG GH C + +C+ CN +GHL + G + +
Sbjct: 28 CLRCGGFGHDMTLCKYEYSHEDLKNIKCYVCNSLGHLCC----IEPGHTQSWT------- 76
Query: 161 ARGTEKQVSCYRCGEIGHFASRC-----SATGPRCFNCHRVGHLAVVC 203
VSCYRCG++GH C + P CF C R GH C
Sbjct: 77 -------VSCYRCGQLGHTGLACGRHYDDSVSPSCFICGREGHFEHQC 117
Score = 43.1 bits (100), Expect = 2e-04
Identities = 29/96 (30%), Positives = 40/96 (41%), Gaps = 14/96 (14%)
Query: 106 VTCFRCGKNGHYANAC-----LGQGPRCFNCNQIGHLAVNCKT-----FQAGPSDNKMKG 155
V+C+RCG+ GH AC P CF C + GH C F S+++ +G
Sbjct: 77 VSCYRCGQLGHTGLACGRHYDDSVSPSCFICGREGHFEHQCHNSFSVCFPEDSSEDECQG 136
Query: 156 KFPAIARGTEKQVSCYRCGEIGHFASRCSATGPRCF 191
+ R E R E GHF +C + CF
Sbjct: 137 PDSSSVRFQENT----REEEEGHFEHQCPDSSSVCF 168
>At3g43590 putative protein
Length = 551
Score = 49.3 bits (116), Expect = 3e-06
Identities = 28/98 (28%), Positives = 39/98 (39%), Gaps = 31/98 (31%)
Query: 106 VTCFRCGKNGHYANAC--------------------LGQGPRCFNCNQIGHLAVNCKTFQ 145
V+C+RCG+ GH AC + C+ C + GH A C
Sbjct: 285 VSCYRCGQLGHSGLACGRHYEESNENDSATPERLFNSREASECYRCGEEGHFAREC---- 340
Query: 146 AGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
P+ + + + + G E Q CYRC GHFA C
Sbjct: 341 --PNSSSI-----STSHGRESQTLCYRCNGSGHFAREC 371
Score = 44.3 bits (103), Expect = 9e-05
Identities = 24/83 (28%), Positives = 34/83 (40%), Gaps = 6/83 (7%)
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC---- 183
C++C + GH + NC T + G A+ K CY C + GH A C
Sbjct: 168 CYSCGEQGHTSFNCPTPTKRRKPCFICGSLEHGAKQCSKGHDCYICKKTGHRAKDCPDKY 227
Query: 184 --SATGPRCFNCHRVGHLAVVCK 204
+ G C C GH ++CK
Sbjct: 228 KNGSKGAVCLRCGDFGHDMILCK 250
Score = 43.5 bits (101), Expect = 1e-04
Identities = 39/159 (24%), Positives = 58/159 (35%), Gaps = 19/159 (11%)
Query: 106 VTCFRCGKNGHYANACLGQGPR---CFNCNQIGHLAVNCK----TFQAGPSDNKMKGKFP 158
V+C+ CG+ GH + C R CF C + H A C + + ++ K P
Sbjct: 166 VSCYSCGEQGHTSFNCPTPTKRRKPCFICGSLEHGAKQCSKGHDCYICKKTGHRAK-DCP 224
Query: 159 AIARGTEKQVSCYRCGEIGHFASRC-------SATGPRCFNCHRVGHLAVVCKKPKVEPS 211
+ K C RCG+ GH C +C+ C GHL V + +
Sbjct: 225 DKYKNGSKGAVCLRCGDFGHDMILCKYEYSKEDLKDVQCYICKSFGHLCCVEPGNSLSWA 284
Query: 212 VNTAR----GKHPAARGKVYTMDGEEAEGADELVKGERK 246
V+ R G A G+ Y E E + R+
Sbjct: 285 VSCYRCGQLGHSGLACGRHYEESNENDSATPERLFNSRE 323
>At5g52380 unknown protein
Length = 268
Score = 47.4 bits (111), Expect = 1e-05
Identities = 35/160 (21%), Positives = 55/160 (33%), Gaps = 28/160 (17%)
Query: 60 PVRSGSQEYRQLGPYQHPKGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYAN 119
P + +++ ++ ++ K T R ++ ++ R P CF C H A
Sbjct: 28 PPKDPNKKKKKKSLFKKKKPGSSTDRPQRTGSSTRHPLRVPGMKPGEGCFICHSKTHIAK 87
Query: 120 AC-----LGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCG 174
C + C C + GH NC S+ K+ CY CG
Sbjct: 88 LCPEKSEWERNKICLQCRRRGHSLKNCPEKNNESSEKKL----------------CYNCG 131
Query: 175 EIGHFASRC-------SATGPRCFNCHRVGHLAVVCKKPK 207
+ GH S C CF C GH++ C + K
Sbjct: 132 DTGHSLSHCPYPMEDGGTKFASCFICKGQGHISKNCPENK 171
>At2g21060 glycine-rich protein (AtGRP2)
Length = 201
Score = 46.6 bits (109), Expect = 2e-05
Identities = 24/64 (37%), Positives = 32/64 (49%), Gaps = 2/64 (3%)
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ--VSCYRCGEIGHFASRCSA 185
CF C + GH+A C G S G++ + G +SCY CGE GHFA C++
Sbjct: 138 CFKCGEPGHMARECSQGGGGYSGGGGGGRYGSGGGGGGGGGGLSCYSCGESGHFARDCTS 197
Query: 186 TGPR 189
G R
Sbjct: 198 GGAR 201
Score = 31.2 bits (69), Expect = 0.75
Identities = 17/59 (28%), Positives = 21/59 (34%), Gaps = 24/59 (40%)
Query: 169 SCYRCGEIGHFASRCS------------------------ATGPRCFNCHRVGHLAVVC 203
SC++CGE GH A CS G C++C GH A C
Sbjct: 137 SCFKCGEPGHMARECSQGGGGYSGGGGGGRYGSGGGGGGGGGGLSCYSCGESGHFARDC 195
Score = 29.6 bits (65), Expect = 2.2
Identities = 9/22 (40%), Positives = 15/22 (67%)
Query: 106 VTCFRCGKNGHYANACLGQGPR 127
++C+ CG++GH+A C G R
Sbjct: 180 LSCYSCGESGHFARDCTSGGAR 201
>At3g61330 copia-type polyprotein
Length = 1352
Score = 41.2 bits (95), Expect = 7e-04
Identities = 18/49 (36%), Positives = 27/49 (54%)
Query: 143 TFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATGPRCF 191
T Q G + ++ +GK +R + V CY CG+ GH+AS C A + F
Sbjct: 254 TNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKF 302
Score = 40.0 bits (92), Expect = 0.002
Identities = 37/120 (30%), Positives = 53/120 (43%), Gaps = 19/120 (15%)
Query: 12 ATMIEALMN--QAAEDAYRRAEEAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEYR 69
A IE L+ QA E+ ++ E+ A + I + QS R G G VR R
Sbjct: 184 AMTIEQLLGSLQAYEEKKKKKEDIAEQVLNMQITKEENGQSYQRRGG--GQVRG-----R 236
Query: 70 QLGPYQHPKGRGFTPRSFKSSTTATPGARSQS-AHPE-------VTCFRCGKNGHYANAC 121
G Y + GRG+ P ++ +R + HP+ V C+ CGK GHYA+ C
Sbjct: 237 GRGGYGN--GRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
Score = 33.5 bits (75), Expect = 0.15
Identities = 13/30 (43%), Positives = 17/30 (56%)
Query: 177 GHFASRCSATGPRCFNCHRVGHLAVVCKKP 206
GH SR + +C+NC + GH A CK P
Sbjct: 268 GHPKSRYDKSSVKCYNCGKFGHYASECKAP 297
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 41.2 bits (95), Expect = 7e-04
Identities = 18/49 (36%), Positives = 27/49 (54%)
Query: 143 TFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATGPRCF 191
T Q G + ++ +GK +R + V CY CG+ GH+AS C A + F
Sbjct: 254 TNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKF 302
Score = 40.0 bits (92), Expect = 0.002
Identities = 27/101 (26%), Positives = 44/101 (42%), Gaps = 9/101 (8%)
Query: 29 RAEEAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEYRQLGPYQHPKGRGFTPRSFK 88
+A E +++ + ++E Q + G R G Q R G + GRG+ P
Sbjct: 195 QAYEEKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQ-VRGRGRGGYGNGRGWRPHEDN 253
Query: 89 SSTTATPGARSQS-AHPE-------VTCFRCGKNGHYANAC 121
++ +R + HP+ V C+ CGK GHYA+ C
Sbjct: 254 TNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
Score = 33.5 bits (75), Expect = 0.15
Identities = 13/30 (43%), Positives = 17/30 (56%)
Query: 177 GHFASRCSATGPRCFNCHRVGHLAVVCKKP 206
GH SR + +C+NC + GH A CK P
Sbjct: 268 GHPKSRYDKSSVKCYNCGKFGHYASECKAP 297
>At3g53500 splicing factor - like protein
Length = 243
Score = 41.2 bits (95), Expect = 7e-04
Identities = 35/150 (23%), Positives = 56/150 (37%), Gaps = 16/150 (10%)
Query: 108 CFRCGKNGHYANACLGQG--PRCFNCNQIGHLAVNCKTFQAGPSDNKMK-----GKFPAI 160
CF CG +GH+A C +C+ C + GH+ NCK PS K + + P
Sbjct: 60 CFNCGVDGHWARDCTAGDWKNKCYRCGERGHIERNCKN---SPSPKKARQGGSYSRSPVK 116
Query: 161 ARGTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHP 220
+R ++ S R S + P R + + PK + +G+
Sbjct: 117 SRSPRRRRSPSRSRSYSRGRSYSRSRSP----VRREKSVEDRSRSPKAMERSVSPKGRDQ 172
Query: 221 AARGKVYTMDGEEAEGADELVKGERKNDGN 250
+ +D G+D G K +GN
Sbjct: 173 SLSPDRKVIDASPKRGSD--YDGSPKENGN 200
Score = 38.5 bits (88), Expect = 0.005
Identities = 31/103 (30%), Positives = 34/103 (32%), Gaps = 23/103 (22%)
Query: 84 PRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGP---RCFNCNQIGHLAVN 140
PR + G + V R G N G P RCFNC GH A +
Sbjct: 13 PRDADDARYYLDGRDFDGSRITVEASRGAPRGSRDNGSRGPPPGSGRCFNCGVDGHWARD 72
Query: 141 CKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
C AG NK CYRCGE GH C
Sbjct: 73 CT---AGDWKNK-----------------CYRCGERGHIERNC 95
>At1g58140 hypothetical protein
Length = 1320
Score = 41.2 bits (95), Expect = 7e-04
Identities = 18/49 (36%), Positives = 27/49 (54%)
Query: 143 TFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATGPRCF 191
T Q G + ++ +GK +R + V CY CG+ GH+AS C A + F
Sbjct: 254 TNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKF 302
Score = 40.0 bits (92), Expect = 0.002
Identities = 27/101 (26%), Positives = 44/101 (42%), Gaps = 9/101 (8%)
Query: 29 RAEEAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEYRQLGPYQHPKGRGFTPRSFK 88
+A E +++ + ++E Q + G R G Q R G + GRG+ P
Sbjct: 195 QAYEEKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQ-VRGRGRGGYGNGRGWRPHEDN 253
Query: 89 SSTTATPGARSQS-AHPE-------VTCFRCGKNGHYANAC 121
++ +R + HP+ V C+ CGK GHYA+ C
Sbjct: 254 TNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
Score = 33.9 bits (76), Expect = 0.12
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 3/45 (6%)
Query: 177 GHFASRCSATGPRCFNCHRVGHLAVVCKKP---KVEPSVNTARGK 218
GH SR + +C+NC + GH A CK P K E N K
Sbjct: 268 GHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKANYVEEK 312
>At1g48710 hypothetical protein
Length = 1352
Score = 41.2 bits (95), Expect = 7e-04
Identities = 18/49 (36%), Positives = 27/49 (54%)
Query: 143 TFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATGPRCF 191
T Q G + ++ +GK +R + V CY CG+ GH+AS C A + F
Sbjct: 254 TNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKF 302
Score = 40.4 bits (93), Expect = 0.001
Identities = 28/101 (27%), Positives = 44/101 (42%), Gaps = 9/101 (8%)
Query: 29 RAEEAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEYRQLGPYQHPKGRGFTPRSFK 88
+A E +++ + +IE Q + G R G Q R G + GRG+ P
Sbjct: 195 QAYEEKKKKKEDIIEQVLNMQITKEENGQSYQRRGGGQ-VRGRGRGGYGNGRGWRPHEDN 253
Query: 89 SSTTATPGARSQS-AHPE-------VTCFRCGKNGHYANAC 121
++ +R + HP+ V C+ CGK GHYA+ C
Sbjct: 254 TNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
Score = 33.9 bits (76), Expect = 0.12
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 3/45 (6%)
Query: 177 GHFASRCSATGPRCFNCHRVGHLAVVCKKP---KVEPSVNTARGK 218
GH SR + +C+NC + GH A CK P K E N K
Sbjct: 268 GHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFEEKANYVEEK 312
>At2g37340 unknown protein
Length = 249
Score = 39.7 bits (91), Expect = 0.002
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Query: 108 CFRCGKNGHYANACLGQG--PRCFNCNQIGHLAVNCK 142
CF CG +GH+A C +C+ C + GH+ NCK
Sbjct: 60 CFNCGVDGHWARDCTAGDWKNKCYRCGERGHIERNCK 96
Score = 37.4 bits (85), Expect = 0.010
Identities = 22/57 (38%), Positives = 23/57 (39%), Gaps = 20/57 (35%)
Query: 127 RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
RCFNC GH A +C AG NK CYRCGE GH C
Sbjct: 59 RCFNCGVDGHWARDCT---AGDWKNK-----------------CYRCGERGHIERNC 95
Score = 33.9 bits (76), Expect = 0.12
Identities = 19/60 (31%), Positives = 26/60 (42%), Gaps = 9/60 (15%)
Query: 147 GPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG--PRCFNCHRVGHLAVVCK 204
G D +G P R C+ CG GH+A C+A +C+ C GH+ CK
Sbjct: 44 GSRDFDSRGPPPGAGR-------CFNCGVDGHWARDCTAGDWKNKCYRCGERGHIERNCK 96
>At3g31950 hypothetical protein
Length = 507
Score = 37.7 bits (86), Expect = 0.008
Identities = 32/158 (20%), Positives = 57/158 (35%), Gaps = 29/158 (18%)
Query: 31 EEAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEYRQLGPYQHPKGRGF-------- 82
E R + + + +A + ++ + RA +++ R +G H G G+
Sbjct: 206 EGNGRSKGKMIDKAGEDERQKGKRAAELEMELEKAKDQRIIGVNPHSLGIGYGINSKVKP 265
Query: 83 ----TPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGP---------RCF 129
T R ATP R C CG H CL P +C+
Sbjct: 266 RIRATARKSVGRKLATPAKRP--------CDICGHTDHLTEDCLYSSPTMPYMDNYTKCY 317
Query: 130 NCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
C +GH+++ C + ++G P + G E++
Sbjct: 318 CCRGLGHVSMYCPYVAPNAGERSLRGVGPLMTAGAEEK 355
Score = 30.8 bits (68), Expect = 0.98
Identities = 23/89 (25%), Positives = 37/89 (40%), Gaps = 20/89 (22%)
Query: 150 DNKMKGKFPAIAR-------GTEKQVSCYRCGEIGHFASRCSATGP---------RCFNC 193
++K+K + A AR T + C CG H C + P +C+ C
Sbjct: 260 NSKVKPRIRATARKSVGRKLATPAKRPCDICGHTDHLTEDCLYSSPTMPYMDNYTKCYCC 319
Query: 194 HRVGHLAVVCKKPKVEPSV--NTARGKHP 220
+GH+++ C P V P+ + RG P
Sbjct: 320 RGLGHVSMYC--PYVAPNAGERSLRGVGP 346
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 37.4 bits (85), Expect = 0.010
Identities = 36/120 (30%), Positives = 49/120 (40%), Gaps = 13/120 (10%)
Query: 150 DNKMKGKF-----PAIARGTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCK 204
DN KG F PA + +E S EI + S + P C C+RVGH+A C
Sbjct: 268 DNSQKGFFNVVAPPAAFQVSEVSHSPITSPEIMYVQSGPNKGRPTCSFCNRVGHIAERCY 327
Query: 205 KPKVEPSVNTARGK------HPAARGKVYTMDGEEAEGADELVKGERKND--GNLLTILS 256
K P T +GK P A T+ ++ G E + G D NL+ + S
Sbjct: 328 KKHGFPPGFTPKGKSSDKPPKPQAVAAQVTLSPDKMTGQLETLAGNFSPDQIQNLIALFS 387
Score = 28.5 bits (62), Expect = 4.9
Identities = 12/31 (38%), Positives = 15/31 (47%)
Query: 126 PRCFNCNQIGHLAVNCKTFQAGPSDNKMKGK 156
P C CN++GH+A C P KGK
Sbjct: 311 PTCSFCNRVGHIAERCYKKHGFPPGFTPKGK 341
>At2g07080 putative gag-protease polyprotein
Length = 627
Score = 37.4 bits (85), Expect = 0.010
Identities = 15/42 (35%), Positives = 19/42 (44%), Gaps = 3/42 (7%)
Query: 165 EKQVSCYRCGEIGHFASRCSAT---GPRCFNCHRVGHLAVVC 203
+K++ CY CG GH C T +C C VGH C
Sbjct: 259 KKEIQCYECGGFGHIKPECPITKRKEMKCLKCKGVGHTKFEC 300
Score = 32.7 bits (73), Expect = 0.26
Identities = 16/55 (29%), Positives = 24/55 (43%), Gaps = 9/55 (16%)
Query: 105 EVTCFRCGKNGHYANAC---LGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGK 156
E+ C+ CG GH C + +C C +GH C P+ +K+K K
Sbjct: 261 EIQCYECGGFGHIKPECPITKRKEMKCLKCKGVGHTKFEC------PNKSKLKEK 309
>At2g10780 pseudogene
Length = 1611
Score = 36.2 bits (82), Expect = 0.023
Identities = 30/125 (24%), Positives = 45/125 (36%), Gaps = 17/125 (13%)
Query: 61 VRSGSQEYRQLGPYQHP--KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYA 118
+ G +E R L + P + P K +S + E C CGKN ++
Sbjct: 334 IEEGIEEERYLNREKAPIRNNQSTKPADKKRKFDKVDNTKSDAKTGE--CVTCGKN--HS 389
Query: 119 NACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGH 178
C C C H +C + G S K+ G E+ +C+ CG+ GH
Sbjct: 390 GTCWKAIGACGRCGSKDHAIQSCPKMEPGQS--KVLG---------EETRTCFYCGKTGH 438
Query: 179 FASRC 183
C
Sbjct: 439 LKREC 443
Score = 33.9 bits (76), Expect = 0.12
Identities = 21/60 (35%), Positives = 25/60 (41%), Gaps = 15/60 (25%)
Query: 108 CFRCGKNGHYANAC----------LGQGPR-CFNCNQIGHLAVNCKTF----QAGPSDNK 152
C RCG H +C LG+ R CF C + GHL C QAG DN+
Sbjct: 399 CGRCGSKDHAIQSCPKMEPGQSKVLGEETRTCFYCGKTGHLKRECPKLTAEKQAGQRDNR 458
Score = 29.6 bits (65), Expect = 2.2
Identities = 26/90 (28%), Positives = 30/90 (32%), Gaps = 22/90 (24%)
Query: 169 SCYRCGEIGHFASRCSATGP-----------RCFNCHRVGHLAVVCKKPKVEPSVNTA-- 215
+C RCG H C P CF C + GHL C K E
Sbjct: 398 ACGRCGSKDHAIQSCPKMEPGQSKVLGEETRTCFYCGKTGHLKRECPKLTAEKQAGQRDN 457
Query: 216 RG--------KHPAARGKVYTMDGEEAEGA 237
RG K A +VY + EEA A
Sbjct: 458 RGGNGLPPPPKRQAVASRVYEL-SEEANDA 486
>At2g12880 pseudogene
Length = 119
Score = 35.8 bits (81), Expect = 0.031
Identities = 16/47 (34%), Positives = 24/47 (51%), Gaps = 6/47 (12%)
Query: 169 SCYRCGEIGHFASRCS-ATGP-----RCFNCHRVGHLAVVCKKPKVE 209
+CY+CG++GHFA C T P C+ C GH + C + +
Sbjct: 35 ACYKCGKLGHFARSCHVVTQPTTAYITCYFCSEEGHRSNGCPNKRTD 81
Score = 35.0 bits (79), Expect = 0.052
Identities = 26/101 (25%), Positives = 41/101 (39%), Gaps = 33/101 (32%)
Query: 98 RSQSAHPEVTCFRCGKNGHYANAC-LGQGP-----RCFNCNQIGHLAVNC---KTFQAGP 148
R ++ + C++CGK GH+A +C + P C+ C++ GH + C +T Q P
Sbjct: 26 RRRNDYDPRACYKCGKLGHFARSCHVVTQPTTAYITCYFCSEEGHRSNGCPNKRTDQVNP 85
Query: 149 SDNKMKGKFPAIARGTEKQVSCYRCGEIGH------FASRC 183
+ CY CG H + SRC
Sbjct: 86 KGH------------------CYWCGNQDHRFNLIIWRSRC 108
>At1g75660 Dhp1-like protein
Length = 1020
Score = 35.8 bits (81), Expect = 0.031
Identities = 18/36 (50%), Positives = 21/36 (58%), Gaps = 3/36 (8%)
Query: 123 GQGPRCFNCNQIGHLAVNCK---TFQAGPSDNKMKG 155
GQ RCF C Q+GH A NC+ +AG SD K G
Sbjct: 259 GQQERCFLCGQMGHFASNCEGKPKKRAGESDEKGDG 294
Score = 30.8 bits (68), Expect = 0.98
Identities = 10/18 (55%), Positives = 14/18 (77%)
Query: 166 KQVSCYRCGEIGHFASRC 183
+Q C+ CG++GHFAS C
Sbjct: 260 QQERCFLCGQMGHFASNC 277
Score = 28.1 bits (61), Expect = 6.4
Identities = 12/31 (38%), Positives = 18/31 (57%), Gaps = 7/31 (22%)
Query: 94 TPGARSQSAHPEVTCFRCGKNGHYANACLGQ 124
TPG + + CF CG+ GH+A+ C G+
Sbjct: 257 TPGQQER-------CFLCGQMGHFASNCEGK 280
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.134 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,117,869
Number of Sequences: 26719
Number of extensions: 301376
Number of successful extensions: 1352
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 59
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 948
Number of HSP's gapped (non-prelim): 307
length of query: 311
length of database: 11,318,596
effective HSP length: 99
effective length of query: 212
effective length of database: 8,673,415
effective search space: 1838763980
effective search space used: 1838763980
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0134.4


