

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.14
(277 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
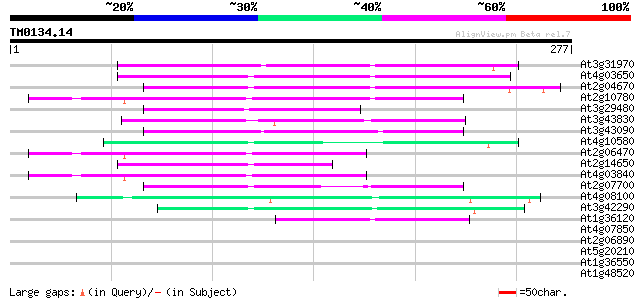
Score E
Sequences producing significant alignments: (bits) Value
At3g31970 hypothetical protein 87 1e-17
At4g03650 putative reverse transcriptase 85 4e-17
At2g04670 putative retroelement pol polyprotein 84 8e-17
At2g10780 pseudogene 70 2e-12
At3g29480 hypothetical protein 65 3e-11
At3g43830 putative protein 64 9e-11
At3g43090 putative protein 63 2e-10
At4g10580 putative reverse-transcriptase -like protein 62 3e-10
At2g06470 putative retroelement pol polyprotein 61 8e-10
At2g14650 putative retroelement pol polyprotein 60 2e-09
At4g03840 putative transposon protein 58 6e-09
At2g07700 putative retroelement pol polyprotein 56 2e-08
At4g08100 putative polyprotein 52 3e-07
At3g42290 putative protein 50 2e-06
At1g36120 putative reverse transcriptase gb|AAD22339.1 48 7e-06
At4g07850 putative polyprotein 38 0.007
At2g06890 putative retroelement integrase 38 0.007
At5g20210 putative protein 37 0.009
At1g36550 hypothetical protein 32 0.29
At1g48520 unknown protein 30 1.1
>At3g31970 hypothetical protein
Length = 1329
Score = 86.7 bits (213), Expect = 1e-17
Identities = 65/199 (32%), Positives = 88/199 (43%), Gaps = 5/199 (2%)
Query: 54 RGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWW 113
R + +R F GGT P+ AD +E F + +V LA + L GDA WW
Sbjct: 145 RMMEQLQRIGTRYFFGGTSPEEADSWKSRVEHNFGSSRCPAEYRVDLAVHFLEGDAHLWW 204
Query: 114 KGTRGIMRANHEEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESL 173
+ R H ++ F F KYFP A D EA+FL L QG +V EY + L
Sbjct: 205 RSVTARRRQAH--MSWADFVAEFNAKYFPQEALDRMEARFLELTQGERSVREYEREFNRL 262
Query: 174 AKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVELMKNRRMNG 233
+ D++ +RFL GLR D++ R L ALVE A EVE R++ G
Sbjct: 263 LVYAG--RGMQDDQAQMRRFLRGLRPDLRVRCRVLQYATKAALVETAAEVEEDLQRQVVG 320
Query: 234 AGTG-GPMRSSSQDNLGKG 251
P ++ Q KG
Sbjct: 321 VSPAVKPKKTQQQVAPSKG 339
>At4g03650 putative reverse transcriptase
Length = 839
Score = 85.1 bits (209), Expect = 4e-17
Identities = 59/194 (30%), Positives = 87/194 (44%), Gaps = 4/194 (2%)
Query: 54 RGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWW 113
R + +R F+GGT P+ AD ++E+ F + +V L + L GDA WW
Sbjct: 143 RMMKQLQRIGTEYFSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLEGDAHLWW 202
Query: 114 KGTRGIMRANHEEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESL 173
+ R +++ F F KYFP A D EA+FL L QG +V EY K L
Sbjct: 203 RSVTA--RRRQADMSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRL 260
Query: 174 AKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVELMKNRRMNG 233
+ D++ +RFL GLR +++ R ALVE A EVE R++ G
Sbjct: 261 LVYAG--RGMEDDQAQMRRFLRGLRPNLRVRCRVSQYATKAALVETAAEVEEDLQRQVVG 318
Query: 234 AGTGGPMRSSSQDN 247
+ + Q +
Sbjct: 319 VSPAVQPKKTQQQH 332
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 84.0 bits (206), Expect = 8e-17
Identities = 63/212 (29%), Positives = 93/212 (43%), Gaps = 10/212 (4%)
Query: 67 FTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANHEE 126
F+ GT P+ AD +++ F + +V LA + L GDA WW+ R +
Sbjct: 138 FSSGTSPEEADSWRSRVQRNFGSSRCPAEYRVDLAVHFLEGDAHLWWRSVTA--RRRQAD 195
Query: 127 VN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDE 186
++ F F KYFP A D EA+FL L QG +V EY + L + D+
Sbjct: 196 MSWADFVAEFNAKYFPQEALDRMEARFLELTQGEWSVREYDREFNRLLAYAG--RGMEDD 253
Query: 187 RYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVELMKNRRMNGAGTGGPMRSSSQ- 245
+ +RFL GLR D++ R ALVE A EVE R++ G +++ Q
Sbjct: 254 QAQMRRFLRGLRPDLRVQCRVSQYATKAALVETAAEVEEDLQRQVVGVSPAVQTKNTQQQ 313
Query: 246 ---DNLGKGRFQMKKPYQRP--MGEGYTLGQY 272
GK K+ + P G+G G +
Sbjct: 314 VTPSKGGKPAQGQKRKWDHPSRAGQGGRAGYF 345
>At2g10780 pseudogene
Length = 1611
Score = 69.7 bits (169), Expect = 2e-12
Identities = 60/220 (27%), Positives = 92/220 (41%), Gaps = 13/220 (5%)
Query: 10 QLAEMVATLVQAMTVQTNDNAHRRATEDTRELHRLQREVALDQNRG-----LNDFRRQDP 64
QLA + ++ V DN +D E+ + + + RG L R
Sbjct: 124 QLATLTKAVIADPVVLKVDNQK----DDDSEVVEVNPQPNHGRKRGDYLSLLEHVSRLGT 179
Query: 65 PKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANH 124
F G TDP AD +++ F+ + E + +A + L GDA WW R
Sbjct: 180 RHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVE--KRRGD 237
Query: 125 EEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHV 184
E + F F +KYFP A D E +L L QG+ TV EY + L ++
Sbjct: 238 EVRSFADFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVG--RELE 295
Query: 185 DERYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVE 224
+E+ +RF+ GLR +I++ LVE+A +E
Sbjct: 296 EEQAQLRRFIRGLRIEIRNHCLVRTFNSVSELVERAAMIE 335
>At3g29480 hypothetical protein
Length = 718
Score = 65.5 bits (158), Expect = 3e-11
Identities = 39/107 (36%), Positives = 50/107 (46%), Gaps = 2/107 (1%)
Query: 67 FTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANHEE 126
F+GGT P+ AD +E+ F + +V LA + L GDA WWK R
Sbjct: 51 FSGGTSPEEADSWRSRVERNFGSSRCPVEYRVDLAVHFLEGDAHLWWKSV--TTRRRQAN 108
Query: 127 VN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESL 173
++ F F KYFP A D EA FL L QG +V EY + L
Sbjct: 109 MSWADFVAEFNAKYFPQEALDRMEAHFLELTQGERSVREYDREFNRL 155
>At3g43830 putative protein
Length = 329
Score = 63.9 bits (154), Expect = 9e-11
Identities = 51/174 (29%), Positives = 77/174 (43%), Gaps = 12/174 (6%)
Query: 56 LNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKG 115
L R D F GGTDP A +E F+ ++ LA+ L G+A+ WW+
Sbjct: 105 LRFMRDLDIETFNGGTDPVKAYNWRNMLECKFKSMRCPVEFWRDLASCYLRGEAQEWWER 164
Query: 116 TRGIMRANHEEVN*----NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLE 171
+ E+V + F+ F +Y P D+ E +FL L QG+ TV +Y +
Sbjct: 165 VK-----QREQVGCVDQWSFFKEEFTRRYLPEETIDDLEMKFLRLQQGTKTVRKYEKEFH 219
Query: 172 SLAKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVEL 225
SL + F E + +F++GLR DI+ LVEKA +E+
Sbjct: 220 SLER---FERRKRGEHELIHKFISGLRVDIRLCCHVRDFDNMIDLVEKAASLEI 270
>At3g43090 putative protein
Length = 357
Score = 62.8 bits (151), Expect = 2e-10
Identities = 48/158 (30%), Positives = 66/158 (41%), Gaps = 3/158 (1%)
Query: 67 FTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANHEE 126
FTGG DP AD + +E FE + + + + L +A +WW G G H
Sbjct: 131 FTGGIDPVKADDWRKLLENNFENARCPVEFQKDIIVHFLREEASHWWDGVLGNTPVQH-V 189
Query: 127 VN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDE 186
+ FR F K+FP A D E F L Q + V E +L L++ E
Sbjct: 190 IFWEDFREEFNRKFFPQEAMDSLEDDFEELRQDTKKVRENERELSHLSRFSVRAGR--GE 247
Query: 187 RYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVE 224
+ M +R + GLR DI LVEKA +E
Sbjct: 248 QSMIRRLMRGLRPDILTRCASREFFSMIDLVEKAARIE 285
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 62.0 bits (149), Expect = 3e-10
Identities = 55/206 (26%), Positives = 78/206 (37%), Gaps = 32/206 (15%)
Query: 47 EVALDQNRGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLL 106
E L R + +R D F+GGT P+ AD + + F + +V LA + L
Sbjct: 132 EEVLSYLRMMEQLQRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVDLAVHFLE 191
Query: 107 GDAEYWWKGTRGIMRANHEEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEY 166
GDA WW+ R +++ F F KYFP A D Q +
Sbjct: 192 GDAHLWWRSVTA--RRRQTDMSWADFVAEFKAKYFPQEALDPYAGQGME----------- 238
Query: 167 ASKLESLAKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVELM 226
D++ +RFL GLR D++ R ALVE A EVE
Sbjct: 239 ------------------DDQAQMRRFLRGLRPDLRVRCRVSQYATKAALVETAAEVEED 280
Query: 227 KNRRMNGAG-TGGPMRSSSQDNLGKG 251
R++ G P ++ Q KG
Sbjct: 281 FQRQVVGVSPVVQPKKTQQQVTPSKG 306
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 60.8 bits (146), Expect = 8e-10
Identities = 50/172 (29%), Positives = 72/172 (41%), Gaps = 11/172 (6%)
Query: 10 QLAEMVATLVQAMTVQTNDNAHRRATEDTRELHRLQREVALDQNRG-----LNDFRRQDP 64
QLA + +V VQ DN +D E+ + + + RG L R
Sbjct: 89 QLATLTKAVVADPVVQRVDNQK----DDDSEVVEVNPQPNHGRKRGDYLSLLEHVSRLGT 144
Query: 65 PKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANH 124
F G TDP AD +++ F+ + E + +A + L GDA WW R
Sbjct: 145 RHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVE--KRRGD 202
Query: 125 EEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKH 176
E + F F +KYFP A D E +L L QG+ TV EY + L ++
Sbjct: 203 EVRSFADFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 59.7 bits (143), Expect = 2e-09
Identities = 35/106 (33%), Positives = 49/106 (46%), Gaps = 2/106 (1%)
Query: 54 RGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWW 113
R + +R F+GGT P+ AD +E+ F + ++ LA + L GDA WW
Sbjct: 143 RMMEQLQRIGTGYFSGGTSPEVADSWRSRVERNFGSSRCPAEYRIDLAVHFLEGDAHLWW 202
Query: 114 KGTRGIMRANHEEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQG 159
+ R +++ F F KYFP A D EA FL L QG
Sbjct: 203 RSVTA--RRRQADISWADFVAEFNAKYFPQEALDRMEAHFLELTQG 246
>At4g03840 putative transposon protein
Length = 973
Score = 57.8 bits (138), Expect = 6e-09
Identities = 49/172 (28%), Positives = 71/172 (40%), Gaps = 11/172 (6%)
Query: 10 QLAEMVATLVQAMTVQTNDNAHRRATEDTRELHRLQREVALDQNRG-----LNDFRRQDP 64
QLA + +V VQ DN +D E+ + + + RG L R
Sbjct: 89 QLATLTKAIVANPVVQRVDNQK----DDELEVVEVNPQPNHGRKRGDYLSLLEHVSRLGT 144
Query: 65 PKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANH 124
F G TDP AD +++ F+ + E + +A + L DA WW R
Sbjct: 145 RHFMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLERDAHNWWLTVE--KRRGD 202
Query: 125 EEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKH 176
E + F F +KYFP A D E +L L QG+ TV EY + L ++
Sbjct: 203 EVRSFADFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRY 254
>At2g07700 putative retroelement pol polyprotein
Length = 543
Score = 55.8 bits (133), Expect = 2e-08
Identities = 44/158 (27%), Positives = 68/158 (42%), Gaps = 23/158 (14%)
Query: 67 FTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANHEE 126
F G + AD ++++E FE+ + + K +A L DA W + R +
Sbjct: 53 FKGEHNTVVADKWLRDLEMNFEISRCPDNFKRQIAVNFLDKDARIWRESVTA--RNQGQM 110
Query: 127 VN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDE 186
++ +FR F +KYFP ARD E QF+ K + +N DE
Sbjct: 111 ISWEAFRREFEKKYFPPEARDRLEQQFM--------------------KRY-LYNGQDDE 149
Query: 187 RYMCKRFLNGLRADIKDSVRPLGIIRFQALVEKATEVE 224
M +RFL GLR +I ++ + L E+A VE
Sbjct: 150 ALMVRRFLRGLRLEIWGRLQAVKYDSVNELTERADNVE 187
>At4g08100 putative polyprotein
Length = 1054
Score = 52.0 bits (123), Expect = 3e-07
Identities = 55/235 (23%), Positives = 90/235 (37%), Gaps = 13/235 (5%)
Query: 34 ATEDTRELHRLQREVALDQNRGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTA 93
++ ++ HR +R D GL + P F G +PD Q+IE +F +
Sbjct: 70 SSHSSQRRHRRERPPPRDPLGGL----KLKIPAFHGTDNPDTYLEWEQKIELVFLCQECL 125
Query: 94 EGAKVSLATYLLLGDAEYWWKGTRGIMRANHEEV--N*NSFRTAFLEKYFPTSARDEREA 151
+ KV +A A WW R + N + +++ P+ E
Sbjct: 126 QSNKVKIAATKFYNYALSWWDQLVTSRRRTRDYPIKTWNQLKFVMRKRFVPSYYHRELHQ 185
Query: 152 QFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGII 211
+ L QGS TV EY ++E+L D + RF+ GL +I+D + +
Sbjct: 186 RLRNLVQGSKTVEEYFLEMETLMLRADL---QEDGEAVMSRFMGGLNREIQDRLETQHYV 242
Query: 212 RFQALVEKATEVELM---KNRRMNGAGTGGPMRSSSQDNLGKGRFQM-KKPYQRP 262
+ ++ KA E KN R + T S K +Q KP+ +P
Sbjct: 243 ELEEMLHKAVMFEQQIKRKNARSSHTKTNYSSGKPSYQKEEKFGYQNDSKPFVKP 297
>At3g42290 putative protein
Length = 462
Score = 49.7 bits (117), Expect = 2e-06
Identities = 50/192 (26%), Positives = 73/192 (37%), Gaps = 15/192 (7%)
Query: 74 DNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANHEEVN*NSFR 133
+ AD +++ F + +V LA + L DA WWK R +++ F
Sbjct: 161 EKADSWRSRVKRNFSSSRCLMEYRVDLAVHFLEEDAHLWWKSVTA--RRRQADMSWADFV 218
Query: 134 TAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCKRF 193
F KYF D E +FL L +V EY + + + + D + +RF
Sbjct: 219 VEFNAKYFLQDELDRMEVRFLELTLVERSVWEYDREFNRVLVYAGW--GMKDGQAELRRF 276
Query: 194 LNGLRADIKDSVRPLGIIRFQALVEKATEVELMKN-----------RRMNGAGTGGPMRS 242
+ GLR D++ R ALVE A EV K ++ AG GG
Sbjct: 277 MRGLRPDLRVRCRMSQYALKAALVETAAEVAPRKGGKPMQGKKRSWNHLSRAGQGGRAGC 336
Query: 243 SSQDNLGKGRFQ 254
S NL Q
Sbjct: 337 FSCGNLAHKNVQ 348
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 47.8 bits (112), Expect = 7e-06
Identities = 31/96 (32%), Positives = 45/96 (46%), Gaps = 2/96 (2%)
Query: 132 FRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCK 191
F F KYFP A D EA+FL L +G ++V EY + + L + D++ +
Sbjct: 157 FVAEFNAKYFPQEALDRMEARFLELTKGELSVREYGREFKRLLVYAG--RGMEDDQAQMR 214
Query: 192 RFLNGLRADIKDSVRPLGIIRFQALVEKATEVELMK 227
RFL GLR D++ R AL E +L +
Sbjct: 215 RFLRGLRPDLRVRCRVSQYATKAALTAAEVEEDLQR 250
>At4g07850 putative polyprotein
Length = 1138
Score = 37.7 bits (86), Expect = 0.007
Identities = 45/182 (24%), Positives = 69/182 (37%), Gaps = 22/182 (12%)
Query: 94 EGAKVSLATYLLLGDAEYWWKGTRGIMRANHEEV--N*NSFRTAFLEKYFPTSARDEREA 151
E KV +A A WW R ++ N + ++ P+ E
Sbjct: 92 EANKVKVAATEFYDYALSWWDQVVTTKRRLGDDSIETWNQLKNIMKRRFVPSHYHRELHQ 151
Query: 152 QFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCK----RFLNGLRADIKDSVRP 207
+ L QG+ TV EY ++E+L D + C+ RF+ GL DI D
Sbjct: 152 RLRNLVQGNRTVEEYFKEMETLML-------RADVQEECEATMSRFMGGLNRDILDRFEV 204
Query: 208 LGIIRFQALVEKATEVELMKNRRMNGAGTGGPMRSSSQDNL---GKGRFQMK-KPYQRPM 263
+ + L KA E RR + P +SS+ + K FQ + KP+ +P
Sbjct: 205 IHYENLEELFHKAVMFEKQIKRR-----SAKPSYNSSKPSYQREEKSGFQKEYKPFVKPK 259
Query: 264 GE 265
E
Sbjct: 260 VE 261
>At2g06890 putative retroelement integrase
Length = 1215
Score = 37.7 bits (86), Expect = 0.007
Identities = 31/124 (25%), Positives = 51/124 (41%), Gaps = 15/124 (12%)
Query: 138 EKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCKRFLNGL 197
+++ P+ E + L QG+ +V EY ++E+L D RFL L
Sbjct: 7 KRFVPSHYHRELHLKLRNLTQGNRSVEEYYKEMETLMLRADISE---DREATLSRFLGDL 63
Query: 198 RADIKDSVRPLGIIRFQALVEKATEVELMKNRRMNGAGTGGPMRSSSQDNLGKGRFQMKK 257
DI+D + ++ + ++ KA E R +SSS+ + G G K
Sbjct: 64 NRDIQDRLETQYYVQIEEMLHKAILFEQQVKR-----------KSSSRSSYGSGTI-AKP 111
Query: 258 PYQR 261
YQR
Sbjct: 112 TYQR 115
>At5g20210 putative protein
Length = 256
Score = 37.4 bits (85), Expect = 0.009
Identities = 43/164 (26%), Positives = 71/164 (43%), Gaps = 12/164 (7%)
Query: 72 DPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTRGIMRANHEEVN*NS 131
D D+ + ++ IE F + E KV A + GDA W + +R + + +
Sbjct: 71 DGDHLFIWLEIIEVYFTLQLAPEVCKVDTAFRSMEGDALLWLQ----CLRQENPTLTWSQ 126
Query: 132 FRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCK 191
+T +E++ E Q +L Q +V EYA++ + A Q N ++ +
Sbjct: 127 LKTELIEEFGGGDITASPEEQLASLRQTG-SVDEYANEFRARAA--QIGNK---DQLVKG 180
Query: 192 RFLNGLRADIKDSVRPLGIIRFQALVE--KATEVELMKNRRMNG 233
FLNGL+ +I+ RP + + KA E EL N R G
Sbjct: 181 MFLNGLKEEIRVMFRPNDAEGPEMAIRTAKAIERELNFNNRAKG 224
>At1g36550 hypothetical protein
Length = 353
Score = 32.3 bits (72), Expect = 0.29
Identities = 29/106 (27%), Positives = 42/106 (39%), Gaps = 15/106 (14%)
Query: 156 LGQGSMTVPEYASKLESLAKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGIIRFQA 215
L QG+ V EY ++E+L D RFL L DI+D + + +
Sbjct: 142 LTQGNRYVEEYYKEMETLMLRADISE---DREATMSRFLGDLNRDIQDRLETQYYVEIEE 198
Query: 216 LVEKATEVELMKNRRMNGAGTGGPMRSSSQDNLGKGRFQMKKPYQR 261
++ KA E R +SSS+ + G G K YQR
Sbjct: 199 MLHKAILFEQQVKR-----------KSSSRSSYGSGTV-AKPTYQR 232
>At1g48520 unknown protein
Length = 550
Score = 30.4 bits (67), Expect = 1.1
Identities = 16/54 (29%), Positives = 27/54 (49%), Gaps = 8/54 (14%)
Query: 165 EYASKLESLAKHFQFFNNHVDERYMCKRFLNGLRADIKDSVRPLGIIRFQALVE 218
EYA +++ +A++ N ++ E LR D+ S+RP+G F VE
Sbjct: 238 EYACEMQRIARYLGVSNGNMQE--------GSLRCDVNISIRPIGQAEFGTKVE 283
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.138 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,790,121
Number of Sequences: 26719
Number of extensions: 236593
Number of successful extensions: 699
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 655
Number of HSP's gapped (non-prelim): 31
length of query: 277
length of database: 11,318,596
effective HSP length: 98
effective length of query: 179
effective length of database: 8,700,134
effective search space: 1557323986
effective search space used: 1557323986
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0134.14


