

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.6
(162 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
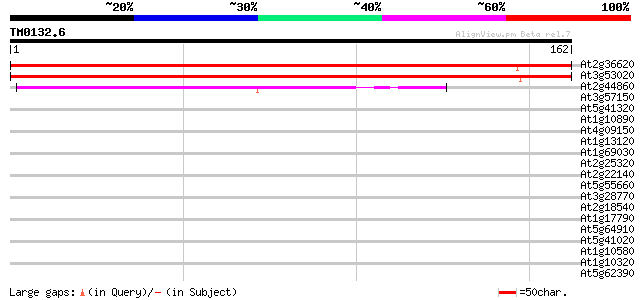
Score E
Sequences producing significant alignments: (bits) Value
At2g36620 60S ribosomal protein L24 272 7e-74
At3g53020 60S ribosomal protein - like 268 1e-72
At2g44860 60S ribosomal protein L30 80 4e-16
At3g57150 putative pseudouridine synthase (NAP57) 39 0.001
At5g41320 unknown protein 36 0.010
At1g10890 unknown protein 35 0.018
At4g09150 unknown protein 35 0.023
At1g13120 unknown protein 34 0.030
At1g69030 hypothetical protein 34 0.040
At2g25320 unknown protein 33 0.052
At2g22140 hypothetical protein 33 0.068
At5g55660 putative protein 33 0.089
At3g28770 hypothetical protein 33 0.089
At2g18540 putative vicilin storage protein (globulin-like) 33 0.089
At1g17790 unknown protein (At1g17790) 33 0.089
At5g64910 unknown protein 32 0.12
At5g41020 putative protein 32 0.12
At1g10580 putative pre-mRNA splicing factor 32 0.12
At1g10320 unknown protein 32 0.12
At5g62390 KED - like protein 32 0.15
>At2g36620 60S ribosomal protein L24
Length = 164
Score = 272 bits (695), Expect = 7e-74
Identities = 135/164 (82%), Positives = 148/164 (89%), Gaps = 2/164 (1%)
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSG KIYPGRGIRFIRSDSQVFLF+NSKCKRYFHN+LKPSKL WTAM+RKQ
Sbjct: 1 MVLKTELCRFSGQKIYPGRGIRFIRSDSQVFLFLNSKCKRYFHNKLKPSKLCWTAMYRKQ 60
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKD AQEA +++RR+TKKPYSRSIVGATLEVIQK+R EKPEVRDAAREAALREIKER+K
Sbjct: 61 HKKDAAQEAVKRRRRATKKPYSRSIVGATLEVIQKKRAEKPEVRDAAREAALREIKERIK 120
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGG--KVAAPKGPKLGGGGGKR 162
KTKDEK+AKK E +K QKSQ KG K AAPK K+GGGGG+R
Sbjct: 121 KTKDEKKAKKVEYASKQQKSQVKGNIPKSAAPKAAKMGGGGGRR 164
>At3g53020 60S ribosomal protein - like
Length = 163
Score = 268 bits (684), Expect = 1e-72
Identities = 133/163 (81%), Positives = 146/163 (88%), Gaps = 1/163 (0%)
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSG KIYPGRGIRFIRSDSQVFLF+NSKCKRYFHN+LKPSKL WTAM+RKQ
Sbjct: 1 MVLKTELCRFSGQKIYPGRGIRFIRSDSQVFLFLNSKCKRYFHNKLKPSKLAWTAMYRKQ 60
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKD AQEA +++RR+TKKPYSRSIVGATLEVIQK+R EKPEVRDAAREAALREIKER+K
Sbjct: 61 HKKDAAQEAVKRRRRATKKPYSRSIVGATLEVIQKKRAEKPEVRDAAREAALREIKERIK 120
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKGGK-VAAPKGPKLGGGGGKR 162
KTKDEK+AKK E +K QK + K AA KGPK+GGGGGKR
Sbjct: 121 KTKDEKKAKKVEFASKQQKVKANFPKAAAASKGPKVGGGGGKR 163
>At2g44860 60S ribosomal protein L30
Length = 159
Score = 80.5 bits (197), Expect = 4e-16
Identities = 44/126 (34%), Positives = 71/126 (55%), Gaps = 9/126 (7%)
Query: 3 LKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHK 62
++ E C F + IYPG GI+F+R+D+++F F SKC + F + P K+ WT FR H
Sbjct: 1 MRLEKCWFCSSTIYPGHGIQFVRNDAKIFRFCRSKCHKNFKMKRNPRKVKWTKAFRAAHG 60
Query: 63 KDIAQEAA--RKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
KD+ ++ +K+R+ + Y R++ TL I+K + AREA + I+ R+K
Sbjct: 61 KDMTKDTTFEFEKKRNRPERYDRNVTENTLMAIKKIAKIR-----TAREA--KHIENRLK 113
Query: 121 KTKDEK 126
K +K
Sbjct: 114 PNKQKK 119
>At3g57150 putative pseudouridine synthase (NAP57)
Length = 565
Score = 38.9 bits (89), Expect = 0.001
Identities = 27/94 (28%), Positives = 47/94 (49%), Gaps = 5/94 (5%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAARE-----AAL 112
+K K++ E A +K +S+KK + E + EK + +D E A+
Sbjct: 457 KKSKTKEVEGEEAEEKVKSSKKKKKKDKEEEKEEEAGSEKKEKKKKKDKKEEVIEEVASP 516
Query: 113 REIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGK 146
+ K++ KK+KD + A AE ++ A+KS+ K K
Sbjct: 517 KSEKKKKKKSKDTEAAVDAEDESAAEKSEKKKKK 550
Score = 29.3 bits (64), Expect = 0.98
Identities = 18/84 (21%), Positives = 41/84 (48%), Gaps = 3/84 (3%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKE 117
+K +++ +EA +K+ KK + V +E + ++EK + + + A + ++
Sbjct: 481 KKDKEEEKEEEAGSEKKEKKKKKDKKEEV---IEEVASPKSEKKKKKKSKDTEAAVDAED 537
Query: 118 RVKKTKDEKRAKKAEVQTKAQKSQ 141
K EK+ KK + + K + S+
Sbjct: 538 ESAAEKSEKKKKKKDKKKKNKDSE 561
>At5g41320 unknown protein
Length = 515
Score = 35.8 bits (81), Expect = 0.010
Identities = 24/65 (36%), Positives = 34/65 (51%), Gaps = 5/65 (7%)
Query: 85 IVGATLEVIQKRRTEKPEVRDAAREAALREIKERV---KKTKDEKRAKKAEVQTKAQKSQ 141
+ GA E + R P R A AA R +K+R+ KK ++EK KK E +TK + Q
Sbjct: 133 LCGAPNENVASR--PPPNKRGVAAVAAKRPVKKRMTKKKKEEEEKMKKKEEEETKESEKQ 190
Query: 142 GKGGK 146
K G+
Sbjct: 191 SKPGE 195
>At1g10890 unknown protein
Length = 592
Score = 35.0 bits (79), Expect = 0.018
Identities = 27/74 (36%), Positives = 36/74 (48%), Gaps = 6/74 (8%)
Query: 71 RKKRRSTKKPYSRSIVGATLEVIQKRRTEK-----PEVRDAAREAALREIKERVKKTKDE 125
+K R ST P RS ATLE + R EK E + REA L+ I+E K +E
Sbjct: 34 QKSRSSTPSPAKRS-PAATLESAKNRNGEKLKREEEERKRRQREAELKLIEEETVKRVEE 92
Query: 126 KRAKKAEVQTKAQK 139
KK E +++K
Sbjct: 93 AIRKKVEESLQSEK 106
Score = 31.6 bits (70), Expect = 0.20
Identities = 18/92 (19%), Positives = 42/92 (45%)
Query: 55 AMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALRE 114
A ++ + + + +++R +K I L+ +++ + ++ R E RE
Sbjct: 128 AQLEEEKEASLIEAKEKEEREQQEKEERERIAEENLKRVEEAQRKEAMERQRKEEERYRE 187
Query: 115 IKERVKKTKDEKRAKKAEVQTKAQKSQGKGGK 146
++E ++ ++ R KKAE + + K GK
Sbjct: 188 LEELQRQKEEAMRRKKAEEEEERLKQMKLLGK 219
>At4g09150 unknown protein
Length = 1097
Score = 34.7 bits (78), Expect = 0.023
Identities = 32/106 (30%), Positives = 54/106 (50%), Gaps = 17/106 (16%)
Query: 49 SKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSI------VGATLEVIQKRRT---E 99
SKL + R+Q+ + ++ +A K R P S SI + + L +++R E
Sbjct: 52 SKLREADLRRQQYYESLSSKARPKMR----SPRSGSIEELSQRLESKLNAAEQKRLSILE 107
Query: 100 KPEVR----DAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQ 141
K R D AR+AA +++RV+K +DE +K E KA+K++
Sbjct: 108 KELARLAKMDEARQAAKNGLEQRVEKERDELESKVEERVLKAEKNR 153
>At1g13120 unknown protein
Length = 611
Score = 34.3 bits (77), Expect = 0.030
Identities = 26/79 (32%), Positives = 43/79 (53%), Gaps = 5/79 (6%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRT--EKPEV-RDAAREAALRE 114
RK ++ +EA RK+R ++ + A +++ K R EK EV R AARE A +E
Sbjct: 219 RKIRSEEAQEEARRKERAHQEEKIRQEKARAEAQMLAKIRAEEEKKEVERKAAREVAEKE 278
Query: 115 IKERVKKTKDEKRAKKAEV 133
+ +R K ++K A++ V
Sbjct: 279 VADR--KAAEQKLAEQKAV 295
>At1g69030 hypothetical protein
Length = 320
Score = 33.9 bits (76), Expect = 0.040
Identities = 23/77 (29%), Positives = 38/77 (48%), Gaps = 3/77 (3%)
Query: 63 KDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKT 122
K A+EAA+K + T + S + V E +K K +A+++A + E V +T
Sbjct: 9 KSFAEEAAKKSQTITLQSSSTTFVNLVTETAKK---SKEFALEASKKADSINVSEFVAET 65
Query: 123 KDEKRAKKAEVQTKAQK 139
+ + AEV TKA +
Sbjct: 66 AKKSKEFAAEVSTKADQ 82
>At2g25320 unknown protein
Length = 1660
Score = 33.5 bits (75), Expect = 0.052
Identities = 33/132 (25%), Positives = 61/132 (46%), Gaps = 5/132 (3%)
Query: 21 IRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKP 80
+ +IRS+ Q + S K+ +RL ++ T A+ +K K+D ++ ++K T+K
Sbjct: 1420 LEWIRSERQDEIDKLSSEKKTLLDRLHEAE-TQLAL-QKTRKRDELKKVGKEKNALTEKL 1477
Query: 81 YSRSIVGATLEVIQKRRTEKPEVRDAAR---EAALREIKERVKKTKDEKRAKKAEVQTKA 137
E KR + R+ R E +R++ + V +TK+EKR K+ ++
Sbjct: 1478 KVTEAARKRFEEELKRYATENVTREELRKSLEDQIRQLTQTVGQTKEEKREKEDQIARCE 1537
Query: 138 QKSQGKGGKVAA 149
G K+ A
Sbjct: 1538 AYIDGMESKLQA 1549
>At2g22140 hypothetical protein
Length = 506
Score = 33.1 bits (74), Expect = 0.068
Identities = 27/88 (30%), Positives = 44/88 (49%), Gaps = 11/88 (12%)
Query: 60 QHKKDIAQE--AARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKE 117
Q K+DI+ E +KK R+T P E + K+R K + A E LR+ +E
Sbjct: 153 QEKEDISVEKIGRKKKIRTTTLPVPG-------EALPKKRQSKEDKTSAMEEKKLRKEQE 205
Query: 118 RVKK--TKDEKRAKKAEVQTKAQKSQGK 143
R++K +K E+ +K + K + +GK
Sbjct: 206 RLEKAASKAEEAERKRLEKEKKKWEKGK 233
>At5g55660 putative protein
Length = 759
Score = 32.7 bits (73), Expect = 0.089
Identities = 18/93 (19%), Positives = 46/93 (49%), Gaps = 4/93 (4%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEV----IQKRRTEKPEVRDAAREAALR 113
+++++ ++A+E K K+ V A +V ++ ++TE + + E
Sbjct: 196 KEKNEAELAEEEETNKGEEVKEANKEDDVEADTKVAEPEVEDKKTESKDENEDKEEEKED 255
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGK 146
E ++ +++ D+K KK +++ ++ +GK K
Sbjct: 256 EKEDEKEESNDDKEDKKEDIKKSNKRGKGKTEK 288
>At3g28770 hypothetical protein
Length = 2081
Score = 32.7 bits (73), Expect = 0.089
Identities = 31/132 (23%), Positives = 59/132 (44%), Gaps = 16/132 (12%)
Query: 34 VNSKCKRYFHNRLKPSKLTWTAMFRKQH--KKDIAQEAARKKRRSTKKPYSRSIVGATLE 91
+N+ K+ ++ K K + + +K+ KK+ KK+ KK ++S + L+
Sbjct: 933 INTSSKQKGKDKKKKKKESKNSNMKKKEEDKKEYVNNEL-KKQEDNKKETTKS-ENSKLK 990
Query: 92 VIQKRRTEKPEVRDAA------------REAALREIKERVKKTKDEKRAKKAEVQTKAQK 139
K EK E D+A + E K+ KK++D+KR +K + K++K
Sbjct: 991 EENKDNKEKKESEDSASKNREKKEYEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKK 1050
Query: 140 SQGKGGKVAAPK 151
+ + + A K
Sbjct: 1051 EKEESRDLKAKK 1062
Score = 30.0 bits (66), Expect = 0.57
Identities = 24/110 (21%), Positives = 52/110 (46%), Gaps = 5/110 (4%)
Query: 58 RKQHKKDIAQEAARK-KRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAR--EAALRE 114
+ QH K + +E+ +K K+ + +K ++ I + + + + EK +D + E ++E
Sbjct: 1140 KSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVDKKEKKSSKDQQKKKEKEMKE 1199
Query: 115 IKERV--KKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
+E+ K +D K+ E K ++++ + K K GGK+
Sbjct: 1200 SEEKKLKKNEEDRKKQTSVEENKKQKETKKEKNKPKDDKKNTTKQSGGKK 1249
Score = 29.3 bits (64), Expect = 0.98
Identities = 26/100 (26%), Positives = 41/100 (41%), Gaps = 3/100 (3%)
Query: 47 KPSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPY--SRSIVGATLEVIQKRRTEKPEVR 104
K SK A K+ +D +E + R +KK SR + E K + E +
Sbjct: 1018 KKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHK 1077
Query: 105 DAAREAALR-EIKERVKKTKDEKRAKKAEVQTKAQKSQGK 143
+E E + +KK +D+K KK E +K + K
Sbjct: 1078 SKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDK 1117
Score = 29.3 bits (64), Expect = 0.98
Identities = 22/100 (22%), Positives = 42/100 (42%)
Query: 47 KPSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDA 106
K +K T T R ++K++ Q ++ + K S ++E ++ E RD
Sbjct: 756 KENKKTKTNENRVRNKEENVQGNKKESEKVEKGEKKESKDAKSVETKDNKKLSSTENRDE 815
Query: 107 AREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGK 146
A+E + + KE +++KD + + E G K
Sbjct: 816 AKERSGEDNKEDKEESKDYQSVEAKEKNENGGVDTNVGNK 855
Score = 28.9 bits (63), Expect = 1.3
Identities = 22/105 (20%), Positives = 51/105 (47%), Gaps = 5/105 (4%)
Query: 39 KRYFHNRLKPSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRT 98
K Y +N LK + + ++ K + K+++ ++ S++ E +K+
Sbjct: 964 KEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDSASKNREKKEYE--EKKSK 1021
Query: 99 EKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGK 143
K E A +E + K+R +K +E+++KK + +++ K++ K
Sbjct: 1022 TKEE---AKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKK 1063
Score = 26.2 bits (56), Expect = 8.3
Identities = 13/46 (28%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Query: 98 TEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGK 143
T++ D ++ ++ E K++ K+TK+EK K + + ++S GK
Sbjct: 1426 TQRNNEEDRKKQTSVAENKKQ-KETKEEKNKPKDDKKNTTEQSGGK 1470
>At2g18540 putative vicilin storage protein (globulin-like)
Length = 699
Score = 32.7 bits (73), Expect = 0.089
Identities = 18/84 (21%), Positives = 41/84 (48%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKE 117
RK+ ++ +E K+R ++ R V + Q+R+ E+ + +E +E +E
Sbjct: 555 RKREEERKREEEMAKRREQERQRKEREEVERKIREEQERKREEEMAKRREQERQKKEREE 614
Query: 118 RVKKTKDEKRAKKAEVQTKAQKSQ 141
+K ++E+ K+ E K ++ +
Sbjct: 615 MERKKREEEARKREEEMAKIREEE 638
Score = 31.6 bits (70), Expect = 0.20
Identities = 25/96 (26%), Positives = 44/96 (45%), Gaps = 3/96 (3%)
Query: 59 KQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKER 118
++ +K +E RKKR + + E Q++ E E R E A+R +ER
Sbjct: 605 QERQKKEREEMERKKREEEARKREEEMAKIREEERQRKEREDVE-RKRREEEAMRREEER 663
Query: 119 VKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPK 154
++ + KRA+ E + K ++ + K PK P+
Sbjct: 664 KREEEAAKRAE--EERRKKEEEEEKRRWPPQPKPPE 697
Score = 31.6 bits (70), Expect = 0.20
Identities = 20/86 (23%), Positives = 41/86 (47%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKE 117
R++ ++ +E +KRR ++ R E +KR E + R+ R+ RE E
Sbjct: 525 REEERQRKEREEVERKRREEQERKRREEEARKREEERKREEEMAKRREQERQRKEREEVE 584
Query: 118 RVKKTKDEKRAKKAEVQTKAQKSQGK 143
R + + E++ ++ + + Q+ Q K
Sbjct: 585 RKIREEQERKREEEMAKRREQERQKK 610
Score = 31.2 bits (69), Expect = 0.26
Identities = 20/82 (24%), Positives = 40/82 (48%), Gaps = 5/82 (6%)
Query: 67 QEAARKKRRSTKKPYSRSIVGATLEVIQKRRTE-----KPEVRDAAREAALREIKERVKK 121
+E ARK+ + ++ + E +K+R E + E R E A R +ER K+
Sbjct: 442 EEEARKREEAKRREEEEAKRREEEETERKKREEEEARKREEERKREEEEAKRREEERKKR 501
Query: 122 TKDEKRAKKAEVQTKAQKSQGK 143
++ ++A+K E + + ++ K
Sbjct: 502 EEEAEQARKREEEREKEEEMAK 523
Score = 30.4 bits (67), Expect = 0.44
Identities = 24/90 (26%), Positives = 43/90 (47%), Gaps = 7/90 (7%)
Query: 59 KQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAAL-----R 113
++ K +E +K+R ++ R E KRR E+ + R+ E A R
Sbjct: 456 EEEAKRREEEETERKKREEEEARKREEERKREEEEAKRREEERKKREEEAEQARKREEER 515
Query: 114 EIKERVKKTKDEKRAKK--AEVQTKAQKSQ 141
E +E + K ++E+R +K EV+ K ++ Q
Sbjct: 516 EKEEEMAKKREEERQRKEREEVERKRREEQ 545
Score = 30.4 bits (67), Expect = 0.44
Identities = 21/88 (23%), Positives = 43/88 (48%), Gaps = 4/88 (4%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAR----EAALR 113
R++ K+ +EA R++ T++ E +KR E+ + R+ R E A +
Sbjct: 448 REEAKRREEEEAKRREEEETERKKREEEEARKREEERKREEEEAKRREEERKKREEEAEQ 507
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQ 141
K ++ K+E+ AKK E + + ++ +
Sbjct: 508 ARKREEEREKEEEMAKKREEERQRKERE 535
Score = 30.4 bits (67), Expect = 0.44
Identities = 22/86 (25%), Positives = 42/86 (48%), Gaps = 5/86 (5%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKE 117
RK+ ++ +E KKR ++ R V Q+R+ + E R E R+ +E
Sbjct: 509 RKREEEREKEEEMAKKREEERQRKEREEVERKRREEQERKRREEEARKREEE---RKREE 565
Query: 118 RVKKTKDEKRAKK--AEVQTKAQKSQ 141
+ K ++++R +K EV+ K ++ Q
Sbjct: 566 EMAKRREQERQRKEREEVERKIREEQ 591
Score = 28.5 bits (62), Expect = 1.7
Identities = 19/81 (23%), Positives = 36/81 (43%), Gaps = 6/81 (7%)
Query: 67 QEAARKKRRSTKKPYSRSIVGATLEVIQKRR------TEKPEVRDAAREAALREIKERVK 120
QE RK+R ++ E + KRR E+ E+ RE R+ +E +
Sbjct: 573 QERQRKEREEVERKIREEQERKREEEMAKRREQERQKKEREEMERKKREEEARKREEEMA 632
Query: 121 KTKDEKRAKKAEVQTKAQKSQ 141
K ++E+R +K + ++ +
Sbjct: 633 KIREEERQRKEREDVERKRRE 653
>At1g17790 unknown protein (At1g17790)
Length = 487
Score = 32.7 bits (73), Expect = 0.089
Identities = 18/53 (33%), Positives = 27/53 (49%), Gaps = 2/53 (3%)
Query: 112 LREIKERVKKTKDEKRA--KKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGKR 162
+R +K ++K DE R+ K+ + + S K G V K K G GGGK+
Sbjct: 68 VRNLKRKLKSELDEVRSLIKRFDPEANPGGSMAKSGVVGRSKKVKTGNGGGKK 120
>At5g64910 unknown protein
Length = 487
Score = 32.3 bits (72), Expect = 0.12
Identities = 22/83 (26%), Positives = 39/83 (46%), Gaps = 3/83 (3%)
Query: 59 KQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKER 118
K+ K++ +EAAR+ + ++ + E + + +P++R R+ R K
Sbjct: 97 KEDKEEEKEEAAREDKEEEEEAVKPDESASQKEEAKGASSSEPQLRRGKRK---RGTKTE 153
Query: 119 VKKTKDEKRAKKAEVQTKAQKSQ 141
+K RAKK TKAQ S+
Sbjct: 154 AEKKVSTPRAKKRAKTTKAQASE 176
Score = 26.9 bits (58), Expect = 4.9
Identities = 17/70 (24%), Positives = 34/70 (48%), Gaps = 3/70 (4%)
Query: 93 IQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKG 152
++ + E+ + A+E E +E ++ K+E+ ++ +QK + KG A+
Sbjct: 82 VESKAAEEGGNEEEAKEDKEEEKEEAAREDKEEEEEAVKPDESASQKEEAKG---ASSSE 138
Query: 153 PKLGGGGGKR 162
P+L G KR
Sbjct: 139 PQLRRGKRKR 148
>At5g41020 putative protein
Length = 588
Score = 32.3 bits (72), Expect = 0.12
Identities = 22/84 (26%), Positives = 38/84 (45%), Gaps = 5/84 (5%)
Query: 72 KKRRSTKKPYSRSIV-----GATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDEK 126
K++ + KKP S V +T + +KR+ +K E L K+ K+ K +K
Sbjct: 185 KRKNNKKKPSVDSDVEDINLDSTNDGKKKRKKKKQSEDSETEENGLNSTKDAKKRRKKKK 244
Query: 127 RAKKAEVQTKAQKSQGKGGKVAAP 150
+ K++EV +KS + P
Sbjct: 245 KKKQSEVSEAEEKSDKSDEDLTTP 268
Score = 32.0 bits (71), Expect = 0.15
Identities = 27/112 (24%), Positives = 46/112 (40%), Gaps = 25/112 (22%)
Query: 72 KKRRSTKKPYSRSIVGATLEVI---------------QKRRTEKP-EVRDAAREAALREI 115
KKR+ K+ G +E+ +K++++KP + A +A ++
Sbjct: 37 KKRKKNKRENKDGFTGEDMEITGRESEKLGDEVFIVKKKKKSKKPIRIDSEAVDAVKKKS 96
Query: 116 KERVKKTKD---------EKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGG 158
K+R K+TK EK++K+ +TK G K K K GG
Sbjct: 97 KKRSKETKADSEAEDDGVEKKSKEKSKETKVDSEAHDGVKRKKKKSKKESGG 148
Score = 31.6 bits (70), Expect = 0.20
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 2/78 (2%)
Query: 72 KKRRSTKKPY--SRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAK 129
KK++ +KKP V A + +KR E +A + ++ KE+ K+TK + A
Sbjct: 73 KKKKKSKKPIRIDSEAVDAVKKKSKKRSKETKADSEAEDDGVEKKSKEKSKETKVDSEAH 132
Query: 130 KAEVQTKAQKSQGKGGKV 147
+ K + + GG V
Sbjct: 133 DGVKRKKKKSKKESGGDV 150
>At1g10580 putative pre-mRNA splicing factor
Length = 616
Score = 32.3 bits (72), Expect = 0.12
Identities = 29/101 (28%), Positives = 47/101 (45%), Gaps = 11/101 (10%)
Query: 59 KQHKKDIAQEAARKK-----RRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALR 113
++ K++I EA + R++ K P+SR EV+Q TE+ + + A A +
Sbjct: 168 EEKKEEIEPEAENPETEAWLRKNRKSPWSRK-----KEVVQGELTEEQK-KYAEDHAKKK 221
Query: 114 EIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPK 154
E K + +TK E A K+ K +K + APK K
Sbjct: 222 EEKGQQGETKGEHYADKSTFHGKEEKDYQGRSWIEAPKDAK 262
>At1g10320 unknown protein
Length = 757
Score = 32.3 bits (72), Expect = 0.12
Identities = 25/80 (31%), Positives = 42/80 (52%), Gaps = 3/80 (3%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRD-AAREAALREIK 116
RKQ +K+IA + + + P + + A +E RR EK E++D E A RE
Sbjct: 47 RKQVRKEIAAKEREEAKAKLNDPAEQERLKA-IEEEDARRREK-ELKDFEESERAWREAM 104
Query: 117 ERVKKTKDEKRAKKAEVQTK 136
E +K ++E+ AK+ E + +
Sbjct: 105 EIKRKKEEEEEAKREEEERR 124
Score = 29.3 bits (64), Expect = 0.98
Identities = 28/96 (29%), Positives = 46/96 (47%), Gaps = 14/96 (14%)
Query: 58 RKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQK---RRTEKPEVRD------AAR 108
+K +K + +E A K+R K + L+ I++ RR EK E++D A R
Sbjct: 43 KKLKRKQVRKEIAAKEREEAKAKLNDPAEQERLKAIEEEDARRREK-ELKDFEESERAWR 101
Query: 109 EAA----LREIKERVKKTKDEKRAKKAEVQTKAQKS 140
EA +E +E K+ ++E+R K E K + S
Sbjct: 102 EAMEIKRKKEEEEEAKREEEERRWKDLEELRKLEAS 137
>At5g62390 KED - like protein
Length = 446
Score = 32.0 bits (71), Expect = 0.15
Identities = 22/76 (28%), Positives = 39/76 (50%), Gaps = 10/76 (13%)
Query: 66 AQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVKKTKDE 125
A EA +KK ++ KK Y+ T EV +R E + + ++ KK K E
Sbjct: 190 AAEAEKKKNKNKKKSYN-----WTTEVKSER-----ENGEVSHTYIIKATTGGEKKKKHE 239
Query: 126 KRAKKAEVQTKAQKSQ 141
++ KK +++TK++K +
Sbjct: 240 EKEKKEKIETKSKKKE 255
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,378,335
Number of Sequences: 26719
Number of extensions: 127705
Number of successful extensions: 1341
Number of sequences better than 10.0: 160
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 100
Number of HSP's that attempted gapping in prelim test: 1134
Number of HSP's gapped (non-prelim): 267
length of query: 162
length of database: 11,318,596
effective HSP length: 91
effective length of query: 71
effective length of database: 8,887,167
effective search space: 630988857
effective search space used: 630988857
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0132.6


