

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0131.12
(78 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
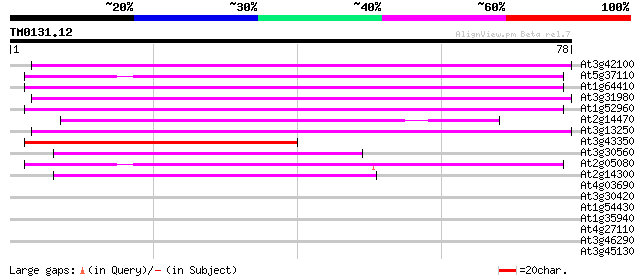
At3g42100 putative protein 54 2e-08
At5g37110 putative helicase 46 3e-06
At1g64410 unknown protein 45 6e-06
At3g31980 hypothetical protein 44 1e-05
At1g52960 hypothetical protein 44 2e-05
At2g14470 pseudogene 43 2e-05
At3g13250 hypothetical protein 43 3e-05
At3g43350 putative protein 41 9e-05
At3g30560 hypothetical protein 39 5e-04
At2g05080 putative helicase 39 5e-04
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 38 8e-04
At4g03690 hypothetical protein 35 0.007
At3g30420 hypothetical protein 34 0.015
At1g54430 hypothetical protein 33 0.020
At1g35940 hypothetical protein 30 0.28
At4g27110 predicted GPI-anchored protein (by homology) 26 3.1
At3g46290 receptor protein kinase -like 25 6.9
At3g45130 oxidosqualene cyclase - like protein 25 9.1
>At3g42100 putative protein
Length = 1752
Score = 53.5 bits (127), Expect = 2e-08
Identities = 27/75 (36%), Positives = 43/75 (57%)
Query: 4 GNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPIL 63
G+ LR FA LL+++ + P+HVW+ TW LLA DI+++ R P + T + L
Sbjct: 1155 GDYLRNFFAMLLLSDSLARPEHVWSETWHLLAEDIENKKREDFKNPDLKLTLAEIRNYTL 1214
Query: 64 LKIDNHLQSNGKSLK 78
+I+ + NG +LK
Sbjct: 1215 QEIEKIMLRNGATLK 1229
>At5g37110 putative helicase
Length = 1307
Score = 46.2 bits (108), Expect = 3e-06
Identities = 27/75 (36%), Positives = 43/75 (57%), Gaps = 2/75 (2%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
SG LR+LF +L + + +P++VW TW+ L+ DIQ+ R +P +I +
Sbjct: 802 SGGYLRQLFVIML--DALISPENVWAATWQHLSEDIQNNKRKYFNRPDLILSDEEKKLYA 859
Query: 63 LLKIDNHLQSNGKSL 77
L +ID+ L+ NG SL
Sbjct: 860 LQEIDHILRRNGTSL 874
>At1g64410 unknown protein
Length = 753
Score = 45.1 bits (105), Expect = 6e-06
Identities = 24/75 (32%), Positives = 40/75 (53%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
S +RRLF +L++ + P+ VW+ TWK+L+ DI+ + R +P + + +
Sbjct: 405 SAKYVRRLFVIMLLSESLTKPEMVWDETWKILSEDIERKKRKEWKRPDLQLSDEERQQYC 464
Query: 63 LLKIDNHLQSNGKSL 77
L +I L NG SL
Sbjct: 465 LQEIARLLTKNGVSL 479
>At3g31980 hypothetical protein
Length = 1099
Score = 43.9 bits (102), Expect = 1e-05
Identities = 22/75 (29%), Positives = 41/75 (54%)
Query: 4 GNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPIL 63
G+ LR LF+ +L+ + P++VW ++L DI+ + R P ++ T + L
Sbjct: 502 GDHLRNLFSMMLLDKCLARPEYVWEKCSRILIEDIETKKRKQYDNPDLVLTDAERRNYAL 561
Query: 64 LKIDNHLQSNGKSLK 78
L+I++ L NG +L+
Sbjct: 562 LEIEDMLLCNGSTLQ 576
>At1g52960 hypothetical protein
Length = 924
Score = 43.5 bits (101), Expect = 2e-05
Identities = 23/75 (30%), Positives = 39/75 (51%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
S +RRLF +L++ + P+ VW+ TW++L+ DI+ R +P + + +
Sbjct: 355 SAKYVRRLFVIMLLSESLTKPEMVWDETWRILSEDIERRKRKEWKRPDLQLSDEERQQYC 414
Query: 63 LLKIDNHLQSNGKSL 77
L +I L NG SL
Sbjct: 415 LQEIARLLTKNGVSL 429
>At2g14470 pseudogene
Length = 1265
Score = 43.1 bits (100), Expect = 2e-05
Identities = 23/61 (37%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 8 RRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKID 67
+ FA LL+++ ++ P HVW+ TW +LA DI + R P I F KP + ID
Sbjct: 717 QEFFAMLLLSDSLSRPAHVWSQTWHILAEDILKKKRDEFKNPEDIDEF---PKPTIDGID 773
Query: 68 N 68
N
Sbjct: 774 N 774
>At3g13250 hypothetical protein
Length = 1419
Score = 42.7 bits (99), Expect = 3e-05
Identities = 22/75 (29%), Positives = 40/75 (53%)
Query: 4 GNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPIL 63
G+ LR F+ +L+++ + P+HVW+ TW LL+ DI + R + T + L
Sbjct: 823 GDYLRNFFSMMLLSDSLARPEHVWSETWHLLSEDILIKKRDEFKNQELTLTEAQIQNYTL 882
Query: 64 LKIDNHLQSNGKSLK 78
+I+ + NG +L+
Sbjct: 883 QEIEKIMLFNGATLE 897
>At3g43350 putative protein
Length = 830
Score = 41.2 bits (95), Expect = 9e-05
Identities = 15/38 (39%), Positives = 26/38 (67%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQH 40
S +RRLF +L++ + P+ VW+ TW++L+ DI+H
Sbjct: 94 SAKYVRRLFVIMLLSESLTKPEMVWDETWRILSKDIEH 131
>At3g30560 hypothetical protein
Length = 1473
Score = 38.9 bits (89), Expect = 5e-04
Identities = 17/43 (39%), Positives = 26/43 (59%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQP 49
+R+LF +LI +++P VW TWK+L+ D Q + R L P
Sbjct: 927 VRKLFVIMLIFESLSSPAVVWEHTWKILSEDFQRKVRDKLKCP 969
>At2g05080 putative helicase
Length = 1219
Score = 38.9 bits (89), Expect = 5e-04
Identities = 27/81 (33%), Positives = 44/81 (53%), Gaps = 8/81 (9%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQP------GMIPTFS 56
SG LR+LF +L + + +P++VW TW+ L+ DIQ+E + +P +I +
Sbjct: 626 SGWYLRQLFVIML--DALISPENVWAATWQHLSEDIQNEKKKYFNRPVTCLFTDLILSDE 683
Query: 57 LSSKPILLKIDNHLQSNGKSL 77
L +ID+ L+ NG SL
Sbjct: 684 EKKVYALQEIDHILRRNGTSL 704
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 38.1 bits (87), Expect = 8e-04
Identities = 16/45 (35%), Positives = 27/45 (59%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGM 51
+R+ F +LI+ +++P VW TWK+L D Q + R L +P +
Sbjct: 734 VRKSFVIMLISESLSSPVVVWEHTWKILFEDFQRKVRDKLERPDL 778
>At4g03690 hypothetical protein
Length = 570
Score = 35.0 bits (79), Expect = 0.007
Identities = 14/38 (36%), Positives = 24/38 (62%)
Query: 9 RLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTL 46
+LF +LI+ +++P VW TWK+L+ D + + R L
Sbjct: 139 KLFVIMLISERLSSPAVVWEHTWKILSEDFKRKLRNQL 176
>At3g30420 hypothetical protein
Length = 837
Score = 33.9 bits (76), Expect = 0.015
Identities = 20/71 (28%), Positives = 37/71 (51%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKI 66
LR LF +LI ++ P +W+ W+ +A+D+ + + L P + K L++I
Sbjct: 235 LRSLFVLILIYCEVSEPLKLWSHCWESMADDVLRKQQRVLNFPQLELKAKELEKYTLIEI 294
Query: 67 DNHLQSNGKSL 77
+ L+ + KSL
Sbjct: 295 ETLLRQHEKSL 305
>At1g54430 hypothetical protein
Length = 1639
Score = 33.5 bits (75), Expect = 0.020
Identities = 20/71 (28%), Positives = 37/71 (51%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKI 66
LR LF +LI ++ P +W+ W+ +A+D+ + + L P + K L++I
Sbjct: 1068 LRSLFVLILIYCEVSEPLKLWSHCWESMADDVFRKQQKVLNFPQLELKAEELEKYTLIEI 1127
Query: 67 DNHLQSNGKSL 77
+ L+ + KSL
Sbjct: 1128 ETLLRQHEKSL 1138
>At1g35940 hypothetical protein
Length = 1678
Score = 29.6 bits (65), Expect = 0.28
Identities = 18/54 (33%), Positives = 26/54 (47%)
Query: 25 HVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKIDNHLQSNGKSLK 78
HV + TW LLA DI R P + T + L +I+ + SNG +L+
Sbjct: 1161 HVRSQTWHLLAEDILKTKRDEFKNPDLTLTETEIKNYTLQEIEKIMLSNGATLE 1214
>At4g27110 predicted GPI-anchored protein (by homology)
Length = 717
Score = 26.2 bits (56), Expect = 3.1
Identities = 10/28 (35%), Positives = 16/28 (56%), Gaps = 1/28 (3%)
Query: 17 TNIMNNPDHVWNITWKLLANDIQHEYRG 44
T+ + DH WN+TW+ + + H RG
Sbjct: 293 TSPLGRLDH-WNLTWEWMRGEFIHSMRG 319
>At3g46290 receptor protein kinase -like
Length = 830
Score = 25.0 bits (53), Expect = 6.9
Identities = 8/25 (32%), Positives = 16/25 (64%)
Query: 8 RRLFASLLITNIMNNPDHVWNITWK 32
R ++ S N +NP+ ++N+TW+
Sbjct: 260 RTVYGSCTEMNSADNPNSIFNVTWE 284
>At3g45130 oxidosqualene cyclase - like protein
Length = 742
Score = 24.6 bits (52), Expect = 9.1
Identities = 9/19 (47%), Positives = 12/19 (62%)
Query: 17 TNIMNNPDHVWNITWKLLA 35
TN+ N H+ N +W LLA
Sbjct: 660 TNLPGNKSHIVNTSWALLA 678
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,819,052
Number of Sequences: 26719
Number of extensions: 60529
Number of successful extensions: 151
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 133
Number of HSP's gapped (non-prelim): 18
length of query: 78
length of database: 11,318,596
effective HSP length: 54
effective length of query: 24
effective length of database: 9,875,770
effective search space: 237018480
effective search space used: 237018480
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0131.12


