

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0126.7
(542 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
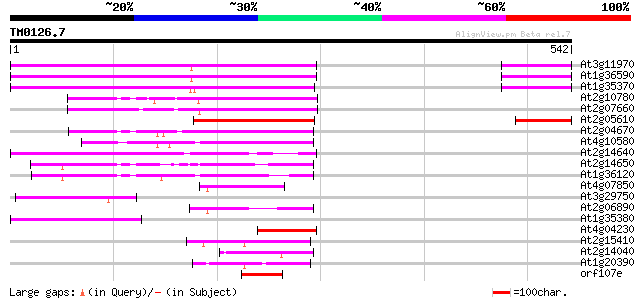
Score E
Sequences producing significant alignments: (bits) Value
At3g11970 hypothetical protein 157 2e-38
At1g36590 hypothetical protein 157 2e-38
At1g35370 hypothetical protein 148 9e-36
At2g10780 pseudogene 110 2e-24
At2g07660 putative retroelement pol polyprotein 107 2e-23
At2g05610 putative retroelement pol polyprotein 101 1e-21
At2g04670 putative retroelement pol polyprotein 100 3e-21
At4g10580 putative reverse-transcriptase -like protein 99 5e-21
At2g14640 putative retroelement pol polyprotein 95 1e-19
At2g14650 putative retroelement pol polyprotein 88 1e-17
At1g36120 putative reverse transcriptase gb|AAD22339.1 68 1e-11
At4g07850 putative polyprotein 59 5e-09
At3g29750 hypothetical protein 57 3e-08
At2g06890 putative retroelement integrase 55 1e-07
At1g35380 hypothetical protein 52 6e-07
At4g04230 putative transposon protein 49 5e-06
At2g15410 putative retroelement pol polyprotein 44 3e-04
At2g14040 putative retroelement pol polyprotein 44 3e-04
At1g20390 hypothetical protein 44 3e-04
orf107e -mitochondrial genome- 42 9e-04
>At3g11970 hypothetical protein
Length = 1499
Score = 157 bits (396), Expect = 2e-38
Identities = 97/309 (31%), Positives = 165/309 (53%), Gaps = 13/309 (4%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
+S+ +++ ++++V G + + +L+D G +HNF+ + V V
Sbjct: 376 ISVNAVSGIAGYKTMRVKGTYDKKIIFILIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVS 435
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ DG K+ +GK D +Q + + L L G+D+VLG+ W +LG I F++L
Sbjct: 436 VADGRKLRVEGKVTDFSWKLQTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFKKLE 495
Query: 120 LRFHVG-EKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQE-ELVAV--- 174
+RF +KVLL G + L K + Q + + Q ++E E EL +
Sbjct: 496 MRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQEDQVQLAMLCVQEVSESTEGELCTINAL 555
Query: 175 ------PEQLQGLMGDYSELFSTIWDLPPSRVQ-DHAIHLQEGAAIPNIRPYRCPHYQKS 227
++ ++ +Y ++F LPP R + +H I L EG+ N RPYR +QK+
Sbjct: 556 TSELGEESVVEEVLNEYPDIFIEPTALPPFREKHNHKIKLLEGSNPVNQRPYRYSIHQKN 615
Query: 228 EIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIID 287
EI+KLV+++L G + S S ++S V+LVKKKDG+WR C+DYR + +T+ + FPI +I+
Sbjct: 616 EIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIE 675
Query: 288 ELLYKLGEA 296
+L+ +LG A
Sbjct: 676 DLMDELGGA 684
Score = 47.8 bits (112), Expect = 2e-05
Identities = 25/67 (37%), Positives = 40/67 (59%)
Query: 476 QSLIFNNFCY*ETGVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLF 535
Q ++ + C G VL+Q PLA+ S + + S+YE+EL+AV+ A+++W L
Sbjct: 895 QFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLL 954
Query: 536 GSHFIIR 542
SHFII+
Sbjct: 955 QSHFIIK 961
>At1g36590 hypothetical protein
Length = 1499
Score = 157 bits (396), Expect = 2e-38
Identities = 97/309 (31%), Positives = 165/309 (53%), Gaps = 13/309 (4%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
+S+ +++ ++++V G + + +L+D G +HNF+ + V V
Sbjct: 376 ISVNAVSGIAGYKTMRVKGTYDKKIIFILIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVS 435
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ DG K+ +GK D +Q + + L L G+D+VLG+ W +LG I F++L
Sbjct: 436 VADGRKLRVEGKVTDFSWKLQTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFKKLE 495
Query: 120 LRFHVG-EKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQE-ELVAV--- 174
+RF +KVLL G + L K + Q + + Q ++E E EL +
Sbjct: 496 MRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQEDQVQLAMLCVQEVSESTEGELCTINAL 555
Query: 175 ------PEQLQGLMGDYSELFSTIWDLPPSRVQ-DHAIHLQEGAAIPNIRPYRCPHYQKS 227
++ ++ +Y ++F LPP R + +H I L EG+ N RPYR +QK+
Sbjct: 556 TSELGEESVVEEVLNEYPDIFIEPTALPPFREKHNHKIKLLEGSNPVNQRPYRYSIHQKN 615
Query: 228 EIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIID 287
EI+KLV+++L G + S S ++S V+LVKKKDG+WR C+DYR + +T+ + FPI +I+
Sbjct: 616 EIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIE 675
Query: 288 ELLYKLGEA 296
+L+ +LG A
Sbjct: 676 DLMDELGGA 684
Score = 47.8 bits (112), Expect = 2e-05
Identities = 25/67 (37%), Positives = 40/67 (59%)
Query: 476 QSLIFNNFCY*ETGVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLF 535
Q ++ + C G VL+Q PLA+ S + + S+YE+EL+AV+ A+++W L
Sbjct: 895 QFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLL 954
Query: 536 GSHFIIR 542
SHFII+
Sbjct: 955 QSHFIIK 961
>At1g35370 hypothetical protein
Length = 1447
Score = 148 bits (373), Expect = 9e-36
Identities = 92/307 (29%), Positives = 167/307 (53%), Gaps = 13/307 (4%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
+S+ +++ + +++ V G + + + +L+D G +HNFI + + V V
Sbjct: 355 ISVNAVSGISGYKTMGVKGTVDKRDLFILIDSGSTHNFIDSTVAAKLGCHVESAGLTKVA 414
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ DG K+ G+ + +Q + + L L G+D+VLG+ W +LG I F++L
Sbjct: 415 VADGRKLNVDGQIKGFTWKLQSTTFQSDILLIPLQGVDMVLGVQWLETLGRISWEFKKLE 474
Query: 120 LRF-HVGEKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVA----- 173
++F + ++V L G + ++ L K Q + V + + +E+ +
Sbjct: 475 MQFFYKNQRVWLHGIITGSVRDIKAHKLQKTQADQIQLAMVCVREVVSDEEQEIGSISAL 534
Query: 174 ---VPEQ--LQGLMGDYSELFSTIWDLPPSRVQ-DHAIHLQEGAAIPNIRPYRCPHYQKS 227
V E+ +Q ++ ++ ++F+ DLPP R + DH I L EGA N RPYR +QK
Sbjct: 535 TSDVVEESVVQNIVEEFPDVFAEPTDLPPFREKHDHKIKLLEGANPVNQRPYRYVVHQKD 594
Query: 228 EIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIID 287
EI+K+V++M+ G + S S F+S V+LVKKKDG+WR C+DY + +T+ ++F I +I+
Sbjct: 595 EIDKIVQDMIKSGTIQVSSSPFASPVVLVKKKDGTWRLCVDYTELNGMTVKDRFLIPLIE 654
Query: 288 ELLYKLG 294
+L+ +LG
Sbjct: 655 DLMDELG 661
Score = 43.1 bits (100), Expect = 4e-04
Identities = 23/67 (34%), Positives = 38/67 (56%)
Query: 476 QSLIFNNFCY*ETGVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLF 535
Q ++ + C VL+Q PLA+ S + + S+YE+EL+A + A+++W L
Sbjct: 866 QFMVETDACGQGIRAVLMQKGHPLAYISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYLL 925
Query: 536 GSHFIIR 542
SHFII+
Sbjct: 926 PSHFIIK 932
>At2g10780 pseudogene
Length = 1611
Score = 110 bits (276), Expect = 2e-24
Identities = 75/248 (30%), Positives = 133/248 (53%), Gaps = 17/248 (6%)
Query: 57 MVELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFE 116
MV G + G R + + + + + + + L DV+LG+ W LG +A+ +
Sbjct: 506 MVRAAGGQAMYPTGLVRGISVVVNGVNMPADLIIVPLKKHDVILGMDW---LGKYKAHID 562
Query: 117 ELSLRFHVGEKVLLRGELGLIR----KNASLSSLVKALQ-QQAEGFWVEFQHLAEVQEEL 171
H G R E G+++ + S S ++ A+Q ++ G E +LA + +
Sbjct: 563 -----CHRGRIQFERDE-GMLKFQGIRTTSGSLVISAIQAERMLGKGCE-AYLATITTKE 615
Query: 172 VAVPEQLQGL--MGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEI 229
V +L+ + + ++S++F+ + +PP R I L+ G + PYR + +++
Sbjct: 616 VGASAELKDIPIVNEFSDVFAAVSGVPPDRSDPFTIELEPGTTPISKAPYRMAPAEMAKL 675
Query: 230 EKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDEL 289
+K ++E+LD G RPS S + + V+ VKKKDGS+R CIDYR +K+T+ NK+P+ IDEL
Sbjct: 676 KKQLEELLDKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDEL 735
Query: 290 LYKLGEAK 297
+ +LG A+
Sbjct: 736 MDQLGGAQ 743
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 107 bits (267), Expect = 2e-23
Identities = 69/244 (28%), Positives = 125/244 (50%), Gaps = 9/244 (3%)
Query: 57 MVELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASL-GHIRANF 115
MV G + R + + + + + + + L DV+LG+ W GHI +
Sbjct: 1 MVRAAGGQAMYPTRLVRGISVVVNGVNMPADLIIVPLKKHDVILGMDWLGKYKGHIDCHR 60
Query: 116 EELSLRFHVGEKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVAVP 175
E + G +L+ + + + S ++A + +G +LA + + V
Sbjct: 61 ERIQFERDEG---MLKFQGIRTTSGSLVISAIQAERMLGKGCEA---YLATITTKEVGAS 114
Query: 176 EQLQGLM--GDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLV 233
+L+ ++ ++S++F+ + +PP R I L+ G + PYR + +E++K +
Sbjct: 115 AELKDILIVNEFSDVFAAVSGVPPDRSDPFTIELEPGTTPISKAPYRMAPAEMAELKKQL 174
Query: 234 KEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
+E+L G RPS S + + V+ VKKKDGS+R CIDYR +K+T+ NK+P+ IDEL+ +L
Sbjct: 175 EELLAKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQL 234
Query: 294 GEAK 297
G A+
Sbjct: 235 GGAQ 238
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 101 bits (251), Expect = 1e-21
Identities = 50/118 (42%), Positives = 80/118 (67%), Gaps = 1/118 (0%)
Query: 178 LQGLMGDYSELFSTIWDLPPSRV-QDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEM 236
++ ++ ++ ++F DLPP R DH I L E + N RPYR +QK+EI+K+V +M
Sbjct: 7 VEKVVTEFPDIFVEPTDLPPFRAPHDHKIELLEDSNPVNQRPYRYVVHQKNEIDKIVDDM 66
Query: 237 LDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLG 294
L G + S S ++S V+LVKKKDG+WR C+DYR + +T+ ++FPI +I++L+ +LG
Sbjct: 67 LASGTIQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELG 124
Score = 47.4 bits (111), Expect = 2e-05
Identities = 23/54 (42%), Positives = 34/54 (62%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL+Q PLAF S + + S+YE+EL+AV+ +++W L SHFII+
Sbjct: 299 GAVLMQEGHPLAFISRQLKGKQLHLSIYEKELLAVIFVVRKWRHYLISSHFIIK 352
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 100 bits (248), Expect = 3e-21
Identities = 72/245 (29%), Positives = 122/245 (49%), Gaps = 21/245 (8%)
Query: 58 VELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEE 117
V++ G + G+ + V + I + + + + DV+LG+ W L H R + +
Sbjct: 365 VKVAGGQFLAVLGRAKGVDIQIAGESMPADLIISPVELYDVILGMDW---LDHYRVHLD- 420
Query: 118 LSLRFHVGEKVLLRGELGLIRKNA--SLSSLV-------KALQQQAEGFWVEFQHLAEVQ 168
H G R E L+ + + SLV K +++ E + V +
Sbjct: 421 ----CHRGRVSFERPEGRLVYQGVRPTSGSLVISAVQAKKMIEKGCEAYLVTIS----MP 472
Query: 169 EELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSE 228
E L V ++ ++ ++F ++ LPPSR I L+ G A + PYR + +E
Sbjct: 473 ESLGQVAVSDIRVIQEFEDVFQSLQGLPPSRSDPFTIELEPGTAPLSKAPYRMAPAEMTE 532
Query: 229 IEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDE 288
++K ++++L G RPS S + + V+ VKKKDGS+R CIDYR + +T+ NK+P+ IDE
Sbjct: 533 LKKQLEDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRGLNWVTVKNKYPLPRIDE 592
Query: 289 LLYKL 293
LL +L
Sbjct: 593 LLDQL 597
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 99.4 bits (246), Expect = 5e-21
Identities = 67/233 (28%), Positives = 118/233 (49%), Gaps = 21/233 (9%)
Query: 70 GKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVL 129
G+ + V + I + + + + DV+LG+ W ++ + L +H G
Sbjct: 351 GRAKGVDIQIAGESMPADLIISPVELYDVILGMDWL--------DYYRVHLDWHRGRVFF 402
Query: 130 LRGELGLIRKNA---SLSSLVKALQQQ------AEGFWVEFQHLAEVQEELVAVPEQLQG 180
R E L+ + S S ++ A+Q + E + V V + V+ +Q
Sbjct: 403 ERPEGRLVYQGVRPISGSLVISAVQAEKMIEKGCEAYLVTISMPESVGQVAVSDIRVVQ- 461
Query: 181 LMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVG 240
++ ++F ++ LPPS+ I L+ G A + PYR + +E++K +K++L G
Sbjct: 462 ---EFQDVFQSLQGLPPSQSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLKDLLGKG 518
Query: 241 IFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
RPS S + + V+ VKKKDGS+R CIDYR +++T+ N++P+ IDELL +L
Sbjct: 519 FIRPSTSPWGAPVLFVKKKDGSFRLCIDYRELNRVTVKNRYPLPRIDELLDQL 571
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 94.7 bits (234), Expect = 1e-19
Identities = 73/299 (24%), Positives = 138/299 (45%), Gaps = 37/299 (12%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVET-PYMVE 59
++L ++ + +++V + + R+I L++ G +HNFI ++ + + T P+ V
Sbjct: 304 ITLYALEGVDTTSTIRVRATIHRNRLIALINSGSTHNFIGEKAVRGMNLKATTTKPFTVR 363
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ +G + C+ + + + + + + L GLD+ +G+ W ++LG N++E +
Sbjct: 364 VVNGMPLVCRSRYEAIPVVMGGVVFPVTLYALPLMGLDLAMGVQWLSTLGPTLCNWKEQT 423
Query: 120 LRFH-VGEKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVAVPE-Q 177
L+FH G++V L G + ++ K + F + H + P+ +
Sbjct: 424 LQFHWAGDEVRLMGIKPTGLRGVEHKTITKKARMGHTIFAITMAHNSSDPN----TPDGE 479
Query: 178 LQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEML 237
++ L+ +++ LF T LP R H I L++G N+RPYR YQK E+ +
Sbjct: 480 IKKLIEEFAGLFETPTQLPSERPIVHRIALKKGTDPVNVRPYRYAFYQKDEMSR------ 533
Query: 238 DVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEA 296
GI RPS S F S V+L +FPI +D++L +L A
Sbjct: 534 -AGIIRPSSSPFLSPVLL-----------------------ERFPIPTVDDMLDELNGA 568
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 88.2 bits (217), Expect = 1e-17
Identities = 77/275 (28%), Positives = 129/275 (46%), Gaps = 38/275 (13%)
Query: 21 LGQLRVIVLVDCGVSHNFISQELMSSAHF--PVVETPYMVELEDGHKVECQGKCRDVQLS 78
+G + VL D G SH FI+ E S + E V++ G V G+ + V +
Sbjct: 308 VGGVEAHVLFDSGASHCFITPESASRGNIRGDPGEQLGAVKVAGGQFVAVLGRTKGVDIQ 367
Query: 79 IQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVLLRGELGLIR 138
I + + + + DV+LG+ W L H R + + H G R E L+
Sbjct: 368 IAGESMPADLIISPVELYDVILGMDW---LDHYRVHLD-----CHRGRVSFERPEGSLVY 419
Query: 139 KNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVAVPEQLQGLMGDYSELFSTIWDLPPS 198
+ +S G V ++ VQ E + + + + + ++ ++F ++ LPPS
Sbjct: 420 QGVRPTS----------GSLV----ISAVQAEKM-IEKGCEAYL-EFEDVFQSLQGLPPS 463
Query: 199 RVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKK 258
R I L+ G A + PYR + +E++K ++++L + V+ VKK
Sbjct: 464 RSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLEDLL------------GAPVLFVKK 511
Query: 259 KDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
KDGS+R CIDYR + +T+ NK+P+ IDELL +L
Sbjct: 512 KDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQL 546
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 68.2 bits (165), Expect = 1e-11
Identities = 68/283 (24%), Positives = 123/283 (43%), Gaps = 41/283 (14%)
Query: 22 GQLRVIVLVDCGVSHNFISQELMSSAHF--PVVETPYMVELEDGHKVECQGKCRDVQLSI 79
G+ R VL D G SH FI+ E S + E V++ G + G+ + V + I
Sbjct: 345 GRCRGHVLFDSGASHCFITPESASRDNIRGEPGEQFGAVKIAGGQFLTVLGRAKGVNIQI 404
Query: 80 QHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVLLRGELGLIRK 139
+ + + + + D +LG+ W L H R + + H G R E L+ +
Sbjct: 405 EEESMPADLIINPVVLYDAILGMDW---LDHYRVHLD-----CHRGRVSFERPEGRLVYQ 456
Query: 140 NASLSS---------LVKALQQQAEGFWVEFQHLAEVQEELVAVPEQLQGLMGDYSELFS 190
+S + K +++ E + V V + V+ +Q ++ ++F
Sbjct: 457 GVRPTSRSLVISEVQVEKMIEKGCEAYLVTISMSESVGQVAVSDIWVVQ----EFEDVFQ 512
Query: 191 TIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFS 250
++ LPPSR I L+ G A + PYR + +E++K ++++L G RPS S++
Sbjct: 513 SLQGLPPSRSDPFTIELELGTAPLSKTPYRMVPAEIAELKKQLEDLLGKGFIRPSTSRWG 572
Query: 251 SLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
+ +++T+ NK+P+ IDELL +L
Sbjct: 573 A------------------PGLNRVTLKNKYPLPRIDELLDQL 597
>At4g07850 putative polyprotein
Length = 1138
Score = 59.3 bits (142), Expect = 5e-09
Identities = 34/84 (40%), Positives = 47/84 (55%), Gaps = 2/84 (2%)
Query: 184 DYSELF--STIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGI 241
DY ++F LPP R +H I GA++PN YR + E+EK V E+++ G
Sbjct: 447 DYQDVFPEENPEGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVETKELEKQVNELMERGH 506
Query: 242 FRPSISQFSSLVILVKKKDGSWRF 265
S+S + V+LV KKDGSWRF
Sbjct: 507 ICESMSPCAVPVLLVPKKDGSWRF 530
Score = 39.3 bits (90), Expect = 0.006
Identities = 19/53 (35%), Positives = 31/53 (57%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
G VL+Q +P+A++S + Y++EL A+VRA+Q W L+ F+I
Sbjct: 637 GAVLMQDKKPIAYFSEKLGGATLNYPTYDKELYALVRALQTWQHYLWPKEFVI 689
>At3g29750 hypothetical protein
Length = 421
Score = 56.6 bits (135), Expect = 3e-08
Identities = 32/120 (26%), Positives = 57/120 (46%), Gaps = 3/120 (2%)
Query: 6 IAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPV-VETPYMVELEDGH 64
+ + T + ++ +G + +V+V +D G + NFI EL S P + V L
Sbjct: 115 VIDLTRNKGMRFYGFILDHKVVVAIDSGATDNFILVELAFSLKLPTSITNQASVLLGQRQ 174
Query: 65 KVECQGKCRDVQLSIQHIPIEQEFFLFNLG--GLDVVLGL*WFASLGHIRANFEELSLRF 122
++ G C ++L +Q + I + F L +L +DV+LG W + LG N++ F
Sbjct: 175 CIQSVGTCLGIRLWVQEVEITENFLLLDLAKTDVDVILGYEWLSKLGETMVNWQNQDFSF 234
>At2g06890 putative retroelement integrase
Length = 1215
Score = 54.7 bits (130), Expect = 1e-07
Identities = 34/122 (27%), Positives = 55/122 (44%), Gaps = 28/122 (22%)
Query: 174 VPEQLQGLMGDYSELF--STIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEK 231
+P ++ L+ DY ++F LPP R +H I GA++PN YR + E+++
Sbjct: 404 LPSEMTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVETKELQR 463
Query: 232 LVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLY 291
+KDGSWR C D RA + +T+ PI +D++L
Sbjct: 464 --------------------------QKDGSWRMCFDCRAINNVTVKYCHPIPRLDDMLD 497
Query: 292 KL 293
+L
Sbjct: 498 EL 499
Score = 37.7 bits (86), Expect = 0.016
Identities = 20/53 (37%), Positives = 31/53 (57%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
G VL+Q + +AF+S + Y++EL A+VRA+QRW L+ F+I
Sbjct: 727 GAVLMQDQKLIAFFSEKLGGATLNYPTYDKELYALVRALQRWQHYLWPKVFVI 779
>At1g35380 hypothetical protein
Length = 508
Score = 52.4 bits (124), Expect = 6e-07
Identities = 29/128 (22%), Positives = 64/128 (49%), Gaps = 1/128 (0%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPY-MVE 59
+S+ +I+ + R+++V G + + +L+D +H+F+ ++ + V
Sbjct: 335 ISVNAISGVSDYRTMRVKGLYEKQVLSILIDPDSTHSFMDSKVAEKLKCVLKPASMAQVS 394
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ DG + K Q ++ ++ + + L G D+VLG W +LG I +F++
Sbjct: 395 VADGRLLGLNAKIDKFQWELRGTQLQADLMVITLRGCDMVLGAQWLETLGPITCDFKKSV 454
Query: 120 LRFHVGEK 127
++FH+G+K
Sbjct: 455 MQFHIGQK 462
>At4g04230 putative transposon protein
Length = 315
Score = 49.3 bits (116), Expect = 5e-06
Identities = 25/57 (43%), Positives = 36/57 (62%)
Query: 240 GIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEA 296
G R S+S + V+LV KKDG+WR C+D RA + ITI + PI + ++L +L A
Sbjct: 128 GYIRESLSPCAVPVLLVPKKDGTWRMCLDCRAINNITIKYRHPIPRLYDMLDELSGA 184
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 43.5 bits (101), Expect = 3e-04
Identities = 31/127 (24%), Positives = 57/127 (44%), Gaps = 8/127 (6%)
Query: 172 VAVPEQLQGLMGDY--SELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQ---- 225
+ +P +LQ + ++ + W + D A+ E P +P + +
Sbjct: 734 IDLPSELQNELVNFLRQNAATFAWSVEDMPGIDSAVTCHELNVDPTYKPLKQKRRKLGPD 793
Query: 226 -KSEIEKLVKEMLDVG-IFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPI 283
++ + VK++LD G I + ++VKKK+G WR CID+ +K + FP+
Sbjct: 794 RTKDVNEEVKKLLDAGSIVEVRYPDWLRNPVVVKKKNGKWRVCIDFTDLNKACPKDSFPL 853
Query: 284 LIIDELL 290
ID L+
Sbjct: 854 PHIDRLV 860
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 43.5 bits (101), Expect = 3e-04
Identities = 35/111 (31%), Positives = 53/111 (47%), Gaps = 21/111 (18%)
Query: 203 HAIHLQEGAAIPNIRPYRCPHYQ-KSEIEKLVKEMLDVGIFRP-SISQFSSLVILVKKKD 260
H IHL EG +I ++ R + + ++K + ++LD GI P S S + S V +V KK
Sbjct: 364 HMIHL-EGESITSVEHQRRLNSNLRDVVKKEIMKLLDAGIIYPISDSTWVSPVHVVPKKG 422
Query: 261 G------------------SWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
G R CIDYR + T + FP+ ID++L +L
Sbjct: 423 GVIVIKNEKNELIPTRTVTGHRMCIDYRKLNSATRKDNFPLSFIDQMLERL 473
>At1g20390 hypothetical protein
Length = 1791
Score = 43.5 bits (101), Expect = 3e-04
Identities = 32/123 (26%), Positives = 57/123 (46%), Gaps = 14/123 (11%)
Query: 177 QLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQ---------KS 227
+L L+ S+ F+ W + + D AI E P +P + +
Sbjct: 738 ELIALLKRNSKTFA--WSIEDMKGIDPAITAHELNVDPTFKPVKQKRRKLGPERARAVNE 795
Query: 228 EIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIID 287
E+EKL+K + + P ++ + ++VKKK+G WR C+DY +K + +P+ ID
Sbjct: 796 EVEKLLKAGQIIEVKYP---EWLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPHID 852
Query: 288 ELL 290
L+
Sbjct: 853 RLV 855
>orf107e -mitochondrial genome-
Length = 107
Score = 42.0 bits (97), Expect = 9e-04
Identities = 18/39 (46%), Positives = 29/39 (74%)
Query: 225 QKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSW 263
+++ ++ + EML+ I +PSIS +SS V+LV+KKDG W
Sbjct: 41 RRTRLKNWLGEMLEARIIQPSISPYSSPVLLVQKKDGGW 79
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.349 0.158 0.543
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,518,239
Number of Sequences: 26719
Number of extensions: 405161
Number of successful extensions: 1585
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 32
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1514
Number of HSP's gapped (non-prelim): 63
length of query: 542
length of database: 11,318,596
effective HSP length: 104
effective length of query: 438
effective length of database: 8,539,820
effective search space: 3740441160
effective search space used: 3740441160
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0126.7


