

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0126.13
(168 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
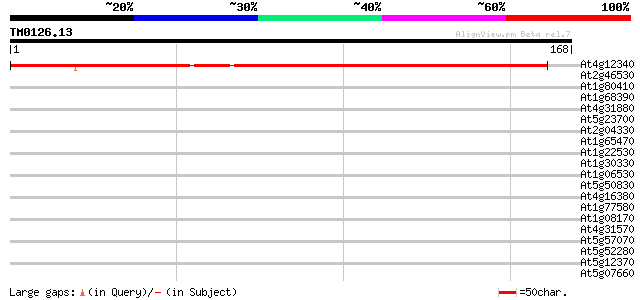
Score E
Sequences producing significant alignments: (bits) Value
At4g12340 putative protein 148 1e-36
At2g46530 ARF1 family auxin responsive transcription factor like... 36 0.009
At1g80410 putative N-terminal acetyltransferase (At1g80410) 33 0.056
At1g68390 hypothetical protein 32 0.12
At4g31880 unknown protein 32 0.16
At5g23700 putative protein 32 0.21
At2g04330 hypothetical protein 32 0.21
At1g65470 hypothetical protein, 3' partial 32 0.21
At1g22530 unknown protein 32 0.21
At1g30330 putative protein 31 0.36
At1g06530 hypothetical protein 31 0.36
At5g50830 unknown protein 30 0.47
At4g16380 unknown protein 30 0.47
At1g77580 unknown protein 30 0.47
At1g08170 hypothetical protein 30 0.61
At4g31570 putative protein 29 1.0
At5g57070 putative protein 29 1.4
At5g52280 hyaluronan mediated motility receptor-like protein 29 1.4
At5g12370 putative protein 29 1.4
At5g07660 SMC-like protein 29 1.4
>At4g12340 putative protein
Length = 170
Score = 148 bits (374), Expect = 1e-36
Identities = 73/162 (45%), Positives = 114/162 (70%), Gaps = 3/162 (1%)
Query: 1 MGEFTIQISNELVNQLVD-DPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQ 59
MG+F+IQIS++L+NQL + + PK++ ++ + KV+ ++++ ++ K N A P+Q
Sbjct: 1 MGDFSIQISSKLINQLAEGNDQPKRRAKKTKPKVSPQSKQKTNQDEEKKPNPVAE-LPMQ 59
Query: 60 SPLFLPAKPPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKL 119
P F P PP A E++ I+SV++ESEKVLE L+ QE+N+++EVT++AKDL +KE+K+
Sbjct: 60 PPFFFPI-PPQGAASTELESIKSVVKESEKVLEKLELQEKNIVREVTERAKDLREKEFKI 118
Query: 120 PNPKPEPCIAERLATLSCYKEHIKDPLKCSAFVTNFAGCLRK 161
P PKP PC ++ A + CYKE+I PLKCS FV +F C R+
Sbjct: 119 PEPKPMPCSSDHEAWMKCYKENIGSPLKCSGFVKSFQDCARR 160
>At2g46530 ARF1 family auxin responsive transcription factor like
protein
Length = 622
Score = 36.2 bits (82), Expect = 0.009
Identities = 29/95 (30%), Positives = 46/95 (47%), Gaps = 10/95 (10%)
Query: 32 KVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQ-------PADPEIDGIRSVL 84
K ET++ +++T +PE + + PL PAKP V +D G SVL
Sbjct: 104 KAEHETDEVYAQITLQPEEDQSEPTSLDPPLVEPAKPTVDSFVKILTASDTSTHGGFSVL 163
Query: 85 RE-SEKVLESLQKQEENMLQEVTQKAKDLHDKEYK 118
R+ + + L SL + QE+ A+DLH E++
Sbjct: 164 RKHATECLPSLDMTQPTPTQELV--ARDLHGYEWR 196
>At1g80410 putative N-terminal acetyltransferase (At1g80410)
Length = 714
Score = 33.5 bits (75), Expect = 0.056
Identities = 23/89 (25%), Positives = 39/89 (42%), Gaps = 15/89 (16%)
Query: 20 PVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQPADPEIDG 79
P KKKI++ ++ A+ ++ +SK + A+ K V+P DP+ G
Sbjct: 591 PAQKKKIKKQKKAEARAKKEAESKSEESTASGASKS----------GKRNVKPVDPDPHG 640
Query: 80 -----IRSVLRESEKVLESLQKQEENMLQ 103
+ + E+ K L LQK N L+
Sbjct: 641 QKLIQVEEPMAEASKYLRLLQKHSPNSLE 669
>At1g68390 hypothetical protein
Length = 408
Score = 32.3 bits (72), Expect = 0.12
Identities = 11/34 (32%), Positives = 22/34 (64%)
Query: 65 PAKPPVQPADPEIDGIRSVLRESEKVLESLQKQE 98
P PP P++PE +G++S + EK++ ++ +E
Sbjct: 84 PPPPPSPPSEPEQNGLKSFIEPPEKLMHDMEDEE 117
>At4g31880 unknown protein
Length = 873
Score = 32.0 bits (71), Expect = 0.16
Identities = 27/118 (22%), Positives = 49/118 (40%), Gaps = 13/118 (11%)
Query: 2 GEFTIQISN-ELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKV----------TGKPEN 50
G+ T +S+ + +L + VPKK + +++ + E KP + + T +P+
Sbjct: 428 GDETANVSSPSMAEELPEQSVPKKTANQKKKESSTEEVKPSASIATEEVSEEPNTSEPQV 487
Query: 51 AAAPGWPVQSPLFLPAKPPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQK 108
G V S KP V P+ + + +KV+ S QE +E +K
Sbjct: 488 TKKSGKKVASS--SKTKPTVPPSKKSTSETKVAKQSEKKVVGSDNAQESTKPKEEKKK 543
>At5g23700 putative protein
Length = 497
Score = 31.6 bits (70), Expect = 0.21
Identities = 17/60 (28%), Positives = 28/60 (46%), Gaps = 7/60 (11%)
Query: 20 PVPKKKIRRNR-------RKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQP 72
P+ RRNR R + ++ +S TG+P + P SPL++P K ++P
Sbjct: 92 PINGSFARRNRSHSPAIGRNITEQVTSVRSSSTGRPSTFSRSSTPNASPLWMPPKASLKP 151
>At2g04330 hypothetical protein
Length = 564
Score = 31.6 bits (70), Expect = 0.21
Identities = 26/114 (22%), Positives = 50/114 (43%), Gaps = 3/114 (2%)
Query: 34 AKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQPADPEIDGIRSVLRESEKVLES 93
A+ET++ + EN G ++ L PV+ A+ + DG+ + E E+ S
Sbjct: 121 AEETQENMQTDEVEDENEKEEGKETETNKELACANPVEEAERQDDGLAVIEEEEER--SS 178
Query: 94 LQKQEENMLQEVTQKA-KDLHDKEYKLPNPKPEPCIAERLATLSCYKEHIKDPL 146
++ N+ + V + +D D++ + P E I E +A + + PL
Sbjct: 179 ASDEDVNVEKSVEDEGDEDERDEDVIVEKPVEERTIDEDIANVDMEEAMAMQPL 232
>At1g65470 hypothetical protein, 3' partial
Length = 618
Score = 31.6 bits (70), Expect = 0.21
Identities = 31/97 (31%), Positives = 46/97 (46%), Gaps = 9/97 (9%)
Query: 56 WPVQSPLFLPAKPPVQPADPEID-GIRSVLR-ESEKVLESL---QKQEENMLQEVTQKAK 110
WP QS + P +P + DPE+D + S E E+ ESL +K E+ L+E KA
Sbjct: 497 WPSQSQVVKPRRPLQK--DPELDYEVDSDEEWEEEEAGESLSDCEKDEDESLEEGCSKAD 554
Query: 111 DLHDKEYKLPNPKPEPCIAERLATLSCYKEHIKDPLK 147
D D E P+ ++E T + + E + D K
Sbjct: 555 DEDDSEDDF--MVPDGYLSEDEVTKAIFNELVIDSYK 589
>At1g22530 unknown protein
Length = 683
Score = 31.6 bits (70), Expect = 0.21
Identities = 26/103 (25%), Positives = 43/103 (41%), Gaps = 23/103 (22%)
Query: 32 KVAKETEKPQSKVTGKPENAAAPGWPV-QSPLFLPAKPPVQPADPEIDGIRSVLRESEKV 90
KV E P+++VT E A G + QS F +E +
Sbjct: 48 KVVSEVAVPETEVTAVKEEEVATGKEILQSESF---------------------KEEGYL 86
Query: 91 LESLQKQEENMLQEVTQKAKD-LHDKEYKLPNPKPEPCIAERL 132
LQ+ E+N L E+ + ++ L+ +E+ P P P P E++
Sbjct: 87 ASELQEAEKNALAELKELVREALNKREFTAPPPPPAPVKEEKV 129
>At1g30330 putative protein
Length = 933
Score = 30.8 bits (68), Expect = 0.36
Identities = 28/92 (30%), Positives = 46/92 (49%), Gaps = 11/92 (11%)
Query: 36 ETEKPQSKVTGKPENAAAPGWP-VQSPLFLPAKPPVQ-------PADPEIDGIRSVLRES 87
ET++ +++T +P NA P + + L +P++ P +D G SV R +
Sbjct: 89 ETDEVYAQMTLQPLNAQEQKDPYLPAELGVPSRQPTNYFCKTLTASDTSTHGGFSVPRRA 148
Query: 88 -EKVLESLQKQEENMLQEVTQKAKDLHDKEYK 118
EKV L ++ QE+ A+DLHD E+K
Sbjct: 149 AEKVFPPLDYSQQPPAQELM--ARDLHDNEWK 178
>At1g06530 hypothetical protein
Length = 323
Score = 30.8 bits (68), Expect = 0.36
Identities = 16/68 (23%), Positives = 34/68 (49%), Gaps = 18/68 (26%)
Query: 70 VQPADPEIDGIRSVLRESEKVLESLQ------------------KQEENMLQEVTQKAKD 111
++ + E+ G+R+V E+EK ++ L+ + EE M +++ K K+
Sbjct: 146 IEELEKEVAGLRTVKEENEKRMKELESKLGALEVKELDEKNKKFRAEEEMREKIDNKEKE 205
Query: 112 LHDKEYKL 119
+HD + K+
Sbjct: 206 VHDLKEKI 213
>At5g50830 unknown protein
Length = 281
Score = 30.4 bits (67), Expect = 0.47
Identities = 24/108 (22%), Positives = 49/108 (45%), Gaps = 7/108 (6%)
Query: 23 KKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLF-----LPAKPPVQPADPEI 77
K+ +R N + ++TE+ + + A AP V+ +P+K V+ E+
Sbjct: 57 KELLRHNSSEDGEDTEENDGSLKASAKVAPAPDHNVEEKKEVIYEKIPSKDDVKEVKEEV 116
Query: 78 DGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPNPKPE 125
+ + E + ++KQEE + V ++ ++ HD + + N K E
Sbjct: 117 --VVNKQEEDNHHQDVVEKQEEENKEVVKKQEEENHDDDVVVINVKKE 162
>At4g16380 unknown protein
Length = 192
Score = 30.4 bits (67), Expect = 0.47
Identities = 21/77 (27%), Positives = 34/77 (43%), Gaps = 3/77 (3%)
Query: 57 PVQSPLFLPAKPPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKE 116
P + P P +PP +P D + + E K LE L++ E+ + +K K+ +
Sbjct: 109 PPKPPQPQPQQPPQKPKDAQPKAPKPKEPEKPKQLEKLKEPEK---PKQPEKPKEPEKTK 165
Query: 117 YKLPNPKPEPCIAERLA 133
P P P P A + A
Sbjct: 166 QPAPAPAPAPAPAAKPA 182
Score = 26.6 bits (57), Expect = 6.8
Identities = 20/56 (35%), Positives = 27/56 (47%), Gaps = 3/56 (5%)
Query: 20 PVPKKKIRRNRRKVAKETEKP-QSKVTGKPENAAAPGWPVQSPLFLP-AKPPVQPA 73
P PK+ + + + KE EKP Q + +PE P P +P P AKP PA
Sbjct: 132 PKPKEPEKPKQLEKLKEPEKPKQPEKPKEPEKTKQPA-PAPAPAPAPAAKPAPAPA 186
>At1g77580 unknown protein
Length = 779
Score = 30.4 bits (67), Expect = 0.47
Identities = 19/76 (25%), Positives = 41/76 (53%), Gaps = 3/76 (3%)
Query: 76 EIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKD--LHDKEYKLPNPKPEPCIAERLA 133
EI+ + S ++E E+ LE L+ ++ + EV ++ +H + ++ + + + E+L
Sbjct: 344 EIEVLTSRIKELEEKLEKLEAEKHELENEVKCNREEAVVHIENSEVLTSRTKE-LEEKLE 402
Query: 134 TLSCYKEHIKDPLKCS 149
L KE +K +KC+
Sbjct: 403 KLEAEKEELKSEVKCN 418
>At1g08170 hypothetical protein
Length = 243
Score = 30.0 bits (66), Expect = 0.61
Identities = 16/62 (25%), Positives = 33/62 (52%), Gaps = 4/62 (6%)
Query: 57 PVQSPLFLPAKPPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKE 116
P ++P+ + +P QP + L++++KV +K++EN ++ +K DL E
Sbjct: 95 PPETPVEVRDEPSPQPPETPASKSEGTLKKTDKV----EKKQENKKKKKKKKRDDLAGDE 150
Query: 117 YK 118
Y+
Sbjct: 151 YR 152
>At4g31570 putative protein
Length = 2712
Score = 29.3 bits (64), Expect = 1.0
Identities = 16/43 (37%), Positives = 22/43 (50%)
Query: 74 DPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKE 116
D EI+ + L E E +E L+ + + QEV QK DL E
Sbjct: 2379 DLEIEALMQALDEEESQMEDLKLRVTELEQEVQQKNLDLQKAE 2421
>At5g57070 putative protein
Length = 607
Score = 28.9 bits (63), Expect = 1.4
Identities = 28/104 (26%), Positives = 41/104 (38%), Gaps = 25/104 (24%)
Query: 23 KKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQPADPEIDGIRS 82
KKK+++++RK E+ VT P+ QS + P+ PP P P
Sbjct: 349 KKKLQKSKRKERIESSPMVEDVTEPPQ--------YQSLIPPPSPPPPPPPPP------P 394
Query: 83 VLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPNPKPEP 126
LR S+ V L K K + K + +P P P P
Sbjct: 395 PLRSSQSVFYGLFK-----------KGVKSNKKIHSVPAPPPPP 427
>At5g52280 hyaluronan mediated motility receptor-like protein
Length = 853
Score = 28.9 bits (63), Expect = 1.4
Identities = 16/53 (30%), Positives = 31/53 (58%), Gaps = 3/53 (5%)
Query: 76 EIDGIRSVLRESEKVLESLQKQEEN---MLQEVTQKAKDLHDKEYKLPNPKPE 125
EID ++ + + + L+S +K+ E +L E+TQ+ + L ++ YK + K E
Sbjct: 425 EIDTLKQQIEDLDWELDSYKKKNEEQEILLDELTQEYESLKEENYKNVSSKLE 477
Score = 28.1 bits (61), Expect = 2.3
Identities = 33/153 (21%), Positives = 64/153 (41%), Gaps = 22/153 (14%)
Query: 20 PVPKKKIRRNRRKVAKETEKPQSKVTGK--PENAAAPGWPVQSPLFLPAKPPVQPADPEI 77
P + RR+ + + +S + + PEN+ G+ + + P++ E+
Sbjct: 218 PATRNGHRRSNTDWSASSTSDESYIESRNSPENSFQRGFSSVTE----SSDPIERLKMEL 273
Query: 78 DGIRSVLRESEKVLESLQKQ---EENMLQEVTQKAKDLHD---------KEYKLPNPKPE 125
+ +R SE +SL+KQ E +QE++++ L ++ +L N + E
Sbjct: 274 EALRRQSELSELEKQSLRKQAIKESKRIQELSKEVSCLKGERDGAMEECEKLRLQNSRDE 333
Query: 126 PCIAERLATL----SCYKEHIKDPLKCSAFVTN 154
RL + S E I+D L C +T+
Sbjct: 334 ADAESRLRCISEDSSNMIEEIRDELSCEKDLTS 366
>At5g12370 putative protein
Length = 863
Score = 28.9 bits (63), Expect = 1.4
Identities = 14/47 (29%), Positives = 25/47 (52%)
Query: 72 PADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYK 118
P PE+DG+ S+ +++ K L L+KQ + L + ++ K K
Sbjct: 84 PFFPEVDGLLSLFKDACKELVDLRKQVDGRLNTLKKEVSTQDSKHRK 130
>At5g07660 SMC-like protein
Length = 1058
Score = 28.9 bits (63), Expect = 1.4
Identities = 17/48 (35%), Positives = 25/48 (51%)
Query: 76 EIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPNPK 123
EI R RE+E LE L+ + ++ TQ KDL KE ++ + K
Sbjct: 654 EIQECRGQKREAEMNLEGLESTMRRLKKQRTQLEKDLTRKELEMQDLK 701
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.132 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,055,615
Number of Sequences: 26719
Number of extensions: 174213
Number of successful extensions: 863
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 39
Number of HSP's that attempted gapping in prelim test: 792
Number of HSP's gapped (non-prelim): 89
length of query: 168
length of database: 11,318,596
effective HSP length: 92
effective length of query: 76
effective length of database: 8,860,448
effective search space: 673394048
effective search space used: 673394048
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0126.13


