

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0122.7
(125 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
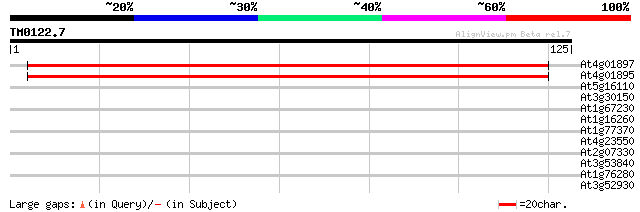
Score E
Sequences producing significant alignments: (bits) Value
At4g01897 unknown protein 158 8e-40
At4g01895 unknown protein 158 8e-40
At5g16110 unknown protein 29 0.69
At3g30150 hypothetical protein 28 0.91
At1g67230 unknown protein 28 0.91
At1g16260 putative wall-associated kinase 28 0.91
At1g77370 unknown protein 28 1.5
At4g23550 WRKY transcription factor 29 (WRKY29) 27 2.0
At2g07330 Mutator-like transposase 27 3.4
At3g53840 protein kinase-like protein 26 4.5
At1g76280 unknown protein 26 5.9
At3g52930 fructose bisphosphate aldolase - like protein 25 7.7
>At4g01897 unknown protein
Length = 122
Score = 158 bits (399), Expect = 8e-40
Identities = 72/116 (62%), Positives = 91/116 (78%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
F+YRIST +EWEE + NGS++G +DKS+ + HLSKLDQV+ TL+ F+++ KE LYLLQ+
Sbjct: 6 FIYRISTEQEWEEFKKNGSSYGAEIDKSTCYYHLSKLDQVQLTLKNFFVDVKEYLYLLQV 65
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSDGRFTCCLL 120
D KKLGDGL+YE VD NSFPHFYGP ++F PL LD V KAEKL+ +G FTC L
Sbjct: 66 DPKKLGDGLIYEAVDEVNSFPHFYGPDKTFVPLPLDSVVKAEKLTFINGNFTCSFL 121
>At4g01895 unknown protein
Length = 122
Score = 158 bits (399), Expect = 8e-40
Identities = 72/116 (62%), Positives = 91/116 (78%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
F+YRIST +EWEE + NGS++G +DKS+ + HLSKLDQV+ TL+ F+++ KE LYLLQ+
Sbjct: 6 FIYRISTEQEWEEFKKNGSSYGAEIDKSTCYYHLSKLDQVQLTLKNFFVDVKEYLYLLQV 65
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSDGRFTCCLL 120
D KKLGDGL+YE VD NSFPHFYGP ++F PL LD V KAEKL+ +G FTC L
Sbjct: 66 DPKKLGDGLIYEAVDEVNSFPHFYGPDKTFVPLPLDSVVKAEKLTFINGNFTCSFL 121
>At5g16110 unknown protein
Length = 178
Score = 28.9 bits (63), Expect = 0.69
Identities = 19/51 (37%), Positives = 33/51 (64%), Gaps = 4/51 (7%)
Query: 61 LLQIDAKKLGDGLVYEMVDGSNSFPHFYG--PSRSFSPLSLDVVTKAEKLS 109
LL+I +K +G + +++ S+S P+F G PSR+ +PL+ D + EKL+
Sbjct: 65 LLEIIRRKEDNGTIGQLL--SSSPPYFPGSPPSRAANPLAQDARFRDEKLN 113
>At3g30150 hypothetical protein
Length = 184
Score = 28.5 bits (62), Expect = 0.91
Identities = 15/50 (30%), Positives = 28/50 (56%)
Query: 31 KSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDG 80
K++AFI + +V++ NC +L L + DAKK+ Y++++G
Sbjct: 22 KNTAFIVPKFVIEVQAYCLSHGFNCSTKLMLFESDAKKIVSCSAYDLLNG 71
>At1g67230 unknown protein
Length = 1132
Score = 28.5 bits (62), Expect = 0.91
Identities = 20/76 (26%), Positives = 33/76 (43%), Gaps = 8/76 (10%)
Query: 8 RISTAKEWEELQANGSTFGGNL-------DKSSAFIHLSKLDQVRSTLERFYMNCKEELY 60
R S KEWEEL + G L +K IHL + ++++ + N + EL
Sbjct: 518 RESFEKEWEELDERKAKIGNELKNITDQKEKLERHIHLEE-ERLKKEKQAANENMERELE 576
Query: 61 LLQIDAKKLGDGLVYE 76
L++ + + YE
Sbjct: 577 TLEVAKASFAETMEYE 592
>At1g16260 putative wall-associated kinase
Length = 720
Score = 28.5 bits (62), Expect = 0.91
Identities = 13/29 (44%), Positives = 17/29 (57%), Gaps = 1/29 (3%)
Query: 73 LVYEMVDGSNSFPHFYGPSRSFSPLSLDV 101
LVYE + N F H + PS F P+S +V
Sbjct: 461 LVYEFIPNRNLFDHLHNPSEDF-PMSWEV 488
>At1g77370 unknown protein
Length = 130
Score = 27.7 bits (60), Expect = 1.5
Identities = 11/42 (26%), Positives = 26/42 (61%), Gaps = 2/42 (4%)
Query: 49 ERFYMNCKEELYLLQIDAKKLGDGLVYEMVD--GSNSFPHFY 88
+R + KEE +++++D ++ GD + YE+++ G + P +
Sbjct: 61 KRIFSQLKEEPFVVELDQREDGDQIQYELLEFVGRRTVPQVF 102
>At4g23550 WRKY transcription factor 29 (WRKY29)
Length = 304
Score = 27.3 bits (59), Expect = 2.0
Identities = 10/26 (38%), Positives = 18/26 (68%)
Query: 64 IDAKKLGDGLVYEMVDGSNSFPHFYG 89
I+ ++L +GL +++ GS +FP F G
Sbjct: 260 IEGRRLSNGLPSDLMSGSGTFPSFTG 285
>At2g07330 Mutator-like transposase
Length = 550
Score = 26.6 bits (57), Expect = 3.4
Identities = 12/44 (27%), Positives = 23/44 (52%)
Query: 34 AFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEM 77
A H+ Q R+ L+ F +NC +E ++ A ++ + YE+
Sbjct: 350 AIRHMFHFKQTRTKLDSFVLNCADEKCDWRVTAHEMPESGYYEI 393
>At3g53840 protein kinase-like protein
Length = 640
Score = 26.2 bits (56), Expect = 4.5
Identities = 24/80 (30%), Positives = 29/80 (36%), Gaps = 16/80 (20%)
Query: 22 GSTFGGNLDKSSAF-IHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGD--------- 71
G F GNLD + + +KL +S Y E L Q+ K L
Sbjct: 367 GEVFKGNLDDGTTVAVKRAKLGNEKS----IYQIVNEVQILCQVSHKNLVKLLGCCIELE 422
Query: 72 --GLVYEMVDGSNSFPHFYG 89
LVYE V F H YG
Sbjct: 423 MPVLVYEFVPNGTLFEHIYG 442
>At1g76280 unknown protein
Length = 812
Score = 25.8 bits (55), Expect = 5.9
Identities = 13/27 (48%), Positives = 16/27 (59%)
Query: 26 GGNLDKSSAFIHLSKLDQVRSTLERFY 52
GG+LDK+S FIH + D S L Y
Sbjct: 137 GGHLDKASEFIHAVREDDRISPLLPIY 163
>At3g52930 fructose bisphosphate aldolase - like protein
Length = 358
Score = 25.4 bits (54), Expect = 7.7
Identities = 21/69 (30%), Positives = 32/69 (45%), Gaps = 9/69 (13%)
Query: 8 RISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEE----LYLLQ 63
++ T K W + +FG L +S+ K + V++ E Y+ CK L +
Sbjct: 283 QLKTKKPW----SLSFSFGRALQQSTLKTWAGKEENVKAAQEALYVRCKANSEATLGTYK 338
Query: 64 IDAKKLGDG 72
DA KLGDG
Sbjct: 339 GDA-KLGDG 346
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,834,982
Number of Sequences: 26719
Number of extensions: 109788
Number of successful extensions: 297
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 288
Number of HSP's gapped (non-prelim): 12
length of query: 125
length of database: 11,318,596
effective HSP length: 87
effective length of query: 38
effective length of database: 8,994,043
effective search space: 341773634
effective search space used: 341773634
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0122.7


