

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.6
(378 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
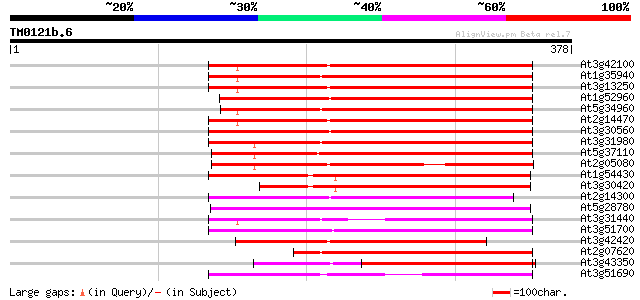
Score E
Sequences producing significant alignments: (bits) Value
At3g42100 putative protein 204 5e-53
At1g35940 hypothetical protein 204 5e-53
At3g13250 hypothetical protein 202 3e-52
At1g52960 hypothetical protein 202 3e-52
At5g34960 putative protein 201 7e-52
At2g14470 pseudogene 200 9e-52
At3g30560 hypothetical protein 191 4e-49
At3g31980 hypothetical protein 191 6e-49
At5g37110 putative helicase 189 3e-48
At2g05080 putative helicase 168 5e-42
At1g54430 hypothetical protein 168 5e-42
At3g30420 hypothetical protein 165 4e-41
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 161 5e-40
At5g28780 putative protein 161 6e-40
At3g31440 hypothetical protein 158 5e-39
At3g51700 unknown protein 157 9e-39
At3g42420 putative protein 150 1e-36
At2g07620 putative helicase 149 2e-36
At3g43350 putative protein 142 2e-34
At3g51690 putative protein 125 5e-29
>At3g42100 putative protein
Length = 1752
Score = 204 bits (520), Expect = 5e-53
Identities = 107/220 (48%), Positives = 146/220 (65%), Gaps = 3/220 (1%)
Query: 135 IFIEPCKDPLLEIVNFSY--PKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGE 192
+ I+ +P+ I Y P L + FFQ RAILAPT E V IN +ML + E
Sbjct: 1503 LLIKEAGNPIEAISKEIYGDPSELHMINDPKFFQRRAILAPTNEDVNTINQYMLEHLKSE 1562
Query: 193 EIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTA 252
E YL+ D+ +D DS N +T +FLN + +G+P+H ++LKVG P+ML+RN+D
Sbjct: 1563 ERIYLSADSIDPTDSDSLANPV-ITPDFLNSIQLTGMPHHALRLKVGAPVMLLRNLDPKG 1621
Query: 253 GLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILC 312
GLCNGTR+ + L K ++ A V+ R+ +G+ IP ++LTPS+ +PFK +RRQFP+ +
Sbjct: 1622 GLCNGTRLQITQLAKQVVQAKVITRDRIGDIVLIPLINLTPSDTKLPFKMRRRQFPLSVA 1681
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
FAMTINKSQGQS VGLYLP+PVF+H QLYVALSRV S+
Sbjct: 1682 FAMTINKSQGQSLEQVGLYLPKPVFSHGQLYVALSRVTSK 1721
>At1g35940 hypothetical protein
Length = 1678
Score = 204 bits (520), Expect = 5e-53
Identities = 107/220 (48%), Positives = 143/220 (64%), Gaps = 3/220 (1%)
Query: 135 IFIEPCKDPLLEIVNFSY--PKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGE 192
+ I P+ I N Y PK+L + FFQ RAILAP E V IN ++L + E
Sbjct: 1429 LLITNADKPIETITNEIYGDPKILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLDAE 1488
Query: 193 EIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTA 252
E YL+ D+ +D DS +N +T +FLN K G+PNH + LKVG P+ML+RN+D
Sbjct: 1489 ERIYLSADSIDPTDSDS-LNNPVITPDFLNSIKLPGLPNHSLCLKVGAPVMLLRNLDPKG 1547
Query: 253 GLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILC 312
GLCNGTR+ + L I+ A V+ + +G IP ++LTP++ +PFK +RRQFP+ +
Sbjct: 1548 GLCNGTRLQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVA 1607
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
FAMTINKSQGQS H+GLYLP+PVF+H QLYVALSRV S+
Sbjct: 1608 FAMTINKSQGQSLEHIGLYLPKPVFSHGQLYVALSRVTSK 1647
>At3g13250 hypothetical protein
Length = 1419
Score = 202 bits (513), Expect = 3e-52
Identities = 106/220 (48%), Positives = 144/220 (65%), Gaps = 3/220 (1%)
Query: 135 IFIEPCKDPLLEIVNFSY--PKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGE 192
+ I+ +P+ I Y P L + FFQ RAILAP E V IN +ML + E
Sbjct: 1171 LLIQEADNPIEAISREIYGDPTKLHEISDPKFFQRRAILAPKNEDVNTINQYMLEHLDSE 1230
Query: 193 EIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTA 252
E YL+ D+ SD DS N +T +FLN K SG+P+H ++LKVG P+ML+RN+D
Sbjct: 1231 ERIYLSADSIDPSDSDSLKNPV-ITPDFLNSIKVSGMPHHSLRLKVGAPVMLLRNLDPKG 1289
Query: 253 GLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILC 312
GLCNGTR+ + L +I+ A V+ + +G+ +IP +++TPS+ +PFK +RRQFP+ +
Sbjct: 1290 GLCNGTRLQITQLCSHIVEAKVITGDRIGQIVYIPLINITPSDTKLPFKMRRRQFPLSVA 1349
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
F MTINKSQGQS VGLYLP+PVF+H QLYVALSRV S+
Sbjct: 1350 FVMTINKSQGQSLEQVGLYLPKPVFSHGQLYVALSRVTSK 1389
>At1g52960 hypothetical protein
Length = 924
Score = 202 bits (513), Expect = 3e-52
Identities = 104/211 (49%), Positives = 140/211 (66%), Gaps = 1/211 (0%)
Query: 142 DPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEIEYLNCDT 201
DP+ I+ Y + FFQ RAIL PT E V IN M++M+ GEE YL+ D+
Sbjct: 709 DPVESIIEAVYGNTFMEEKDPKFFQGRAILCPTNEDVNSINEHMMSMLDGEERIYLSSDS 768
Query: 202 PCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNGTRMI 261
+D S N + +++FLN + SG+PNH ++LKVG P+ML+RN+D GLCNGTR+
Sbjct: 769 IDPADTSSANNDAY-SADFLNSVRVSGLPNHCLRLKVGCPVMLLRNMDPNKGLCNGTRLQ 827
Query: 262 VNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQ 321
V + +I A + N +G+ IPRM +TPS+ +PFK +RRQFP+ + FAMTINKSQ
Sbjct: 828 VTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRLPFKMRRRQFPLSVAFAMTINKSQ 887
Query: 322 GQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
GQ+ VGLYLPRPVF+H QLYVA+SRV S+
Sbjct: 888 GQTLESVGLYLPRPVFSHGQLYVAISRVTSK 918
>At5g34960 putative protein
Length = 1033
Score = 201 bits (510), Expect = 7e-52
Identities = 104/212 (49%), Positives = 141/212 (66%), Gaps = 3/212 (1%)
Query: 143 PLLEIVNFSY--PKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEIEYLNCD 200
P+ I N Y PK+L + FFQ RAILAPT E V IN ++L + EE YL+ D
Sbjct: 792 PIESITNEIYGDPKILHEITDPKFFQGRAILAPTNEDVNTINEYLLEQLHAEERIYLSAD 851
Query: 201 TPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNGTRM 260
+ +D +S +N +T +FLN K +G+PNH ++LKV P+ML+RN+D GLCNGTR+
Sbjct: 852 SIDPTDSNS-LNNPVITPDFLNSIKLAGLPNHSLRLKVSAPVMLLRNLDPKGGLCNGTRL 910
Query: 261 IVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKS 320
+ L I+ A V+ + +G IP ++LTP++ +PFK +RRQFP+ + FAMTIN S
Sbjct: 911 QITQLCTQIVEAKVITGDIIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINTS 970
Query: 321 QGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
QGQS HVGLYLP+ VF+H QLYVALSRV S+
Sbjct: 971 QGQSLEHVGLYLPKAVFSHGQLYVALSRVTSK 1002
>At2g14470 pseudogene
Length = 1265
Score = 200 bits (509), Expect = 9e-52
Identities = 105/220 (47%), Positives = 141/220 (63%), Gaps = 3/220 (1%)
Query: 135 IFIEPCKDPLLEIVNFSY--PKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGE 192
+ I P+ I N Y PK+L + FFQ RAILA E V IN ++L + E
Sbjct: 1033 LLITNADKPIESITNEIYGDPKILHEITDPKFFQGRAILASKNEDVNTINEYLLDQLHAE 1092
Query: 193 EIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTA 252
E YL+ D+ +D DS N +T +FLN K G+PNH ++LKVG P++L+RN+D
Sbjct: 1093 ERIYLSADSIDPTDSDSLSNPV-ITPDFLNSIKLPGLPNHSLRLKVGAPVLLLRNLDPKG 1151
Query: 253 GLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILC 312
GLCNGTR+ + L I+ A V+ + +G IP ++LTP+N +PFK +RRQFP+ +
Sbjct: 1152 GLCNGTRLQITQLCTQIVEAKVITGDRIGHIILIPTVNLTPTNTKLPFKMRRRQFPLSVA 1211
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
F MTINKS+GQS HVGLYLP+PVF+H QLYVALSRV S+
Sbjct: 1212 FVMTINKSEGQSLEHVGLYLPKPVFSHGQLYVALSRVTSK 1251
>At3g30560 hypothetical protein
Length = 1473
Score = 191 bits (486), Expect = 4e-49
Identities = 103/218 (47%), Positives = 139/218 (63%), Gaps = 1/218 (0%)
Query: 135 IFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEI 194
I I +P+ I+ Y FFQ+RAIL PT + V IN+ ML+ + GEE
Sbjct: 1227 ILIPEGDNPIESIIKAVYGTSFDEERDPKFFQDRAILCPTNDDVNSINDHMLSKLTGEEK 1286
Query: 195 EYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGL 254
Y + D+ SD + N + T +FLN K SG+PNH + LKVG P+ML+RN+D GL
Sbjct: 1287 IYRSSDSIDPSDTRADKNPVY-TPDFLNKIKISGLPNHLLWLKVGCPVMLLRNLDSHGGL 1345
Query: 255 CNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFA 314
NGTR+ + L ++ +L +G+ IPRM LTPS+ +PFK +RRQFP+ + FA
Sbjct: 1346 MNGTRLQIVRLGDKLVQGRILTGTRVGKLVIIPRMPLTPSDRRLPFKMKRRQFPLSVAFA 1405
Query: 315 MTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
MTINKSQGQS +VG+YLP+PVF+H QLYVA+SRVKS+
Sbjct: 1406 MTINKSQGQSLGNVGIYLPKPVFSHGQLYVAMSRVKSK 1443
>At3g31980 hypothetical protein
Length = 1099
Score = 191 bits (485), Expect = 6e-49
Identities = 101/220 (45%), Positives = 139/220 (62%), Gaps = 3/220 (1%)
Query: 135 IFIEPCKDPLLEIVNFSYPKLLFNLEKNS--FFQERAILAPTLESVEEINNFMLAMILGE 192
+ I+ DP+ I Y L N FFQ+RAIL P V IN+ ML + GE
Sbjct: 849 LLIKDANDPIEAITKAVYGDLDLLQPNNDPKFFQQRAILCPRNTDVNTINDIMLDKLNGE 908
Query: 193 EIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTA 252
+ YL+ D+ D S +N +T +FLN K SG+PNH + LK+G P+ML+RNID
Sbjct: 909 LVTYLSADSIDPQDAAS-LNNPVLTPDFLNSIKLSGLPNHNLTLKIGTPVMLLRNIDPKG 967
Query: 253 GLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILC 312
GLCNGTR+ V + +I+ A V+ + +G+ I + ++PS+ +PF+ +RRQFP+ +
Sbjct: 968 GLCNGTRLQVTQMGNHILEARVITGDRVGDKVIIIKSQISPSDTKLPFRMRRRQFPIAVA 1027
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
FAMTINKSQGQS VG+YLP+PVF+H QLYVALSRV S+
Sbjct: 1028 FAMTINKSQGQSLKEVGIYLPKPVFSHGQLYVALSRVTSK 1067
>At5g37110 putative helicase
Length = 1307
Score = 189 bits (479), Expect = 3e-48
Identities = 100/218 (45%), Positives = 139/218 (62%), Gaps = 3/218 (1%)
Query: 137 IEPCKDPLLEIVNFSYPKLLFNLEKNS--FFQERAILAPTLESVEEINNFMLAMILGEEI 194
I KDP+ I Y + E+N FFQE+AIL PT E V +IN ML + GEE
Sbjct: 1071 ITKAKDPIEAICTEIYGDITKIHEQNDPIFFQEKAILCPTNEDVNQINETMLDNLQGEEF 1130
Query: 195 EYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGL 254
+L+ D+ +D G N +T +FLN K S +PNH+++LK+G P+ML+RNID GL
Sbjct: 1131 TFLSSDSLDPADI-GGKNNPALTPDFLNSVKVSRLPNHKLRLKIGCPVMLLRNIDPIGGL 1189
Query: 255 CNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFA 314
NGTR+ + + +I+ A++L + G IPR+ L PS+ +PF+ +R Q P+ +CFA
Sbjct: 1190 MNGTRLRITQMGPFILQAMILTGDRAGHLVLIPRLKLAPSDTKLPFRMRRTQLPLAVCFA 1249
Query: 315 MTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
MTINKSQGQS VG++L RP F+H QLYVA+SRV S+
Sbjct: 1250 MTINKSQGQSLKRVGIFLLRPCFSHGQLYVAISRVTSK 1287
>At2g05080 putative helicase
Length = 1219
Score = 168 bits (425), Expect = 5e-42
Identities = 94/219 (42%), Positives = 135/219 (60%), Gaps = 17/219 (7%)
Query: 137 IEPCKDPLLEIVNFSYPKLLFNLEKNS--FFQERAILAPTLESVEEINNFMLAMILGEEI 194
I KDP+ I Y + E+ FFQERAIL PT E V +IN ML + GEE+
Sbjct: 966 ITKAKDPIQAICTEIYGDITKIHEQKDPVFFQERAILCPTNEDVNQINETMLDNLQGEEL 1025
Query: 195 EYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGL 254
+L+ D+ +D S N +T EFLN+ K G+ NH+++LK+G P+ML+RNID GL
Sbjct: 1026 TFLSSDSLDTADIGSRNNPV-LTPEFLNNVKVLGLSNHKLRLKIGSPVMLLRNIDPIGGL 1084
Query: 255 CNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFA 314
NGTR+ + ++ +I+ A++L + ++ +PF+ +R Q P+ +CFA
Sbjct: 1085 MNGTRLQIMQMSPFILQAMILTGDR--------------ADTKLPFRMRRTQLPLAVCFA 1130
Query: 315 MTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSRN 353
MTINKSQGQS VG++LPRP F+H QLYVA+SRV S++
Sbjct: 1131 MTINKSQGQSLKRVGIFLPRPCFSHSQLYVAISRVTSKS 1169
>At1g54430 hypothetical protein
Length = 1639
Score = 168 bits (425), Expect = 5e-42
Identities = 93/222 (41%), Positives = 136/222 (60%), Gaps = 8/222 (3%)
Query: 135 IFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEI 194
+ + ++PL + +P + + A+L P E+V+EIN+++L+ + G
Sbjct: 1383 LLLPETENPLEVLCRSVFPDFTNTFQDLENLKGTAVLTPRNETVDEINDYLLSKVPGLAK 1442
Query: 195 EYLNCDTPCKSDEDSGVNAEWVTS----EFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQ 250
EY + D+ D D + E E+LN + G+P H++ LKVGVPIML+RN++Q
Sbjct: 1443 EYFSADS---IDRDEALTEEGFEMSYPMEYLNSLEFPGLPAHRLCLKVGVPIMLLRNLNQ 1499
Query: 251 TAGLCNGTRMIVNALTKYIIVAIVLNRNTMG-ETTFIPRMSLTPSNADIPFKFQRRQFPV 309
GLCNGTR+IV L ++ A +L+ T + IPR+ L+P ++ PF +RRQFPV
Sbjct: 1500 KEGLCNGTRLIVTHLGDKVLKAEILSDTTKERKKVLIPRIILSPQDSKHPFTLRRRQFPV 1559
Query: 310 ILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKS 351
+C+AMTINKSQGQ+ V LYLP+PVF+H QLYVALSRV S
Sbjct: 1560 RMCYAMTINKSQGQTLNRVALYLPKPVFSHGQLYVALSRVTS 1601
>At3g30420 hypothetical protein
Length = 837
Score = 165 bits (417), Expect = 4e-41
Identities = 88/188 (46%), Positives = 125/188 (65%), Gaps = 8/188 (4%)
Query: 169 AILAPTLESVEEINNFMLAMILGEEIEYLNCDTPCKSDEDSGVNAEWVTS----EFLNDY 224
A+L P E+V+EIN+++L+ + G EY + D+ D+D + E E+LN
Sbjct: 615 AVLTPRNETVDEINDYLLSKVPGLAKEYFSADS---IDQDEALTEEGFEMSYPMEYLNSL 671
Query: 225 KCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMG-ET 283
+ G+P H++ LKVGVPIML+RN++Q GLCNGTR+ V L ++ A +L+ T +
Sbjct: 672 EFPGLPAHRLCLKVGVPIMLLRNLNQKEGLCNGTRLTVTHLGDKVLKAEILSDTTKKRKK 731
Query: 284 TFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLY 343
IPR+ L+P ++ PF +RRQFPV +C+AMT+NKSQGQ+ V LYLP+PVF+H QLY
Sbjct: 732 VLIPRIILSPQDSKHPFTLRRRQFPVRMCYAMTVNKSQGQTLNRVALYLPKPVFSHGQLY 791
Query: 344 VALSRVKS 351
VALSRV S
Sbjct: 792 VALSRVTS 799
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 161 bits (408), Expect = 5e-40
Identities = 88/205 (42%), Positives = 123/205 (59%), Gaps = 1/205 (0%)
Query: 135 IFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEI 194
I I +P+ I+ Y FFQ +AIL PT + V IN+ ML+ + GEE
Sbjct: 1026 ILIPEGDNPIESIIKAVYGTTFAQKRDPKFFQHKAILCPTNDDVNSINDHMLSKLTGEER 1085
Query: 195 EYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGL 254
Y + ++ SD + N + T +FLN K SG+ NH ++LKVG P+ML+RN GL
Sbjct: 1086 IYRSSNSIDPSDTRADKNPIY-TPDFLNKIKISGLANHLLRLKVGCPVMLLRNFYPHGGL 1144
Query: 255 CNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFA 314
NGTR+ + L ++ +L +G+ IPRMSLTPS+ +PFK +RR FP+ + FA
Sbjct: 1145 MNGTRLQIVRLGDKLVQGRILTGTRVGKLVIIPRMSLTPSDRRLPFKMKRRHFPLSVAFA 1204
Query: 315 MTINKSQGQSSTHVGLYLPRPVFTH 339
MTINKSQGQS +VG+YLP+ VF+H
Sbjct: 1205 MTINKSQGQSLGNVGMYLPKAVFSH 1229
>At5g28780 putative protein
Length = 337
Score = 161 bits (407), Expect = 6e-40
Identities = 89/216 (41%), Positives = 129/216 (59%)
Query: 136 FIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEIE 195
F++ + L ++ +Y + + + ER IL P E V+EIN +ML+ + G+ E
Sbjct: 86 FLKHNGNRLQQVTKGAYVQFSVSQPNFQYLTERGILTPHNEYVDEINAYMLSQVGGDSKE 145
Query: 196 YLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGLC 255
YL+ + K+D ++LN + +P H+I LK GVPIM +RN +Q GLC
Sbjct: 146 YLSSYSIGKADTIGADYEALYHVKYLNSLEFPSLPKHKISLKKGVPIMQMRNFNQKEGLC 205
Query: 256 NGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAM 315
NGTR+IV L + +I A ++ G+ IPR L+P ++ PF +R+QFP+ +C+AM
Sbjct: 206 NGTRLIVTNLGEQVIEAQIVTGTHAGKMVSIPRFILSPPQSEHPFTLRRQQFPMRVCYAM 265
Query: 316 TINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKS 351
TI K+QGQS LYLP PVF+HVQLYVALSRV S
Sbjct: 266 TIIKNQGQSLKSDVLYLPNPVFSHVQLYVALSRVTS 301
>At3g31440 hypothetical protein
Length = 536
Score = 158 bits (399), Expect = 5e-39
Identities = 94/220 (42%), Positives = 124/220 (55%), Gaps = 27/220 (12%)
Query: 135 IFIEPCKDPLLEIVNFSY--PKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGE 192
+ I P+ I N Y PK+L + FFQ RAILAP E V IN ++L + E
Sbjct: 311 LLITNADKPIEWITNEIYGDPKILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLHAE 370
Query: 193 EIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTA 252
E YL+ D+ +D DS +N +T +FLN K G
Sbjct: 371 ERIYLSADSIDPTDSDS-LNNPVITPDFLNSIKLPG------------------------ 405
Query: 253 GLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILC 312
GLCNG R+ + L I+ A V+ + +G IP ++LTP++ +PFK +RRQFP+ +
Sbjct: 406 GLCNGARLQITQLFTEIVEAKVITGDRIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVA 465
Query: 313 FAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
FAMTINKSQGQS HVGLYLP+PVF+H QLYVALSRV S+
Sbjct: 466 FAMTINKSQGQSLEHVGLYLPKPVFSHGQLYVALSRVTSK 505
>At3g51700 unknown protein
Length = 344
Score = 157 bits (397), Expect = 9e-39
Identities = 86/218 (39%), Positives = 127/218 (57%), Gaps = 1/218 (0%)
Query: 135 IFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEI 194
+ I CKDP+ IV+ Y + F+QERAIL T + +EIN++ML+ + GEE
Sbjct: 88 LLITECKDPIKTIVDEVYGESFTESYNPDFYQERAILCHTNDVADEINDYMLSQLQGEET 147
Query: 195 EYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGL 254
+ DT + + EFLN K G P+ +++LKVG P+ML+R++ L
Sbjct: 148 KCYGADTIYPTHASPNDKMLYPL-EFLNSIKIPGFPDFKLRLKVGAPVMLLRDLAPYGWL 206
Query: 255 CNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFA 314
GTR+ + + +++ A+++ N GE IPR+ A P K +RRQFPV L FA
Sbjct: 207 RKGTRLQITRVETFVLEAMIITGNNHGEKVLIPRIPSDLREAKFPIKMRRRQFPVKLAFA 266
Query: 315 MTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
MTI++SQ Q+ + VG+YLPR + H Q YVA+S+VKSR
Sbjct: 267 MTIDESQRQTLSKVGIYLPRQLLFHGQRYVAISKVKSR 304
>At3g42420 putative protein
Length = 1018
Score = 150 bits (379), Expect = 1e-36
Identities = 77/169 (45%), Positives = 111/169 (65%), Gaps = 1/169 (0%)
Query: 153 PKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEIEYLNCDTPCKSDEDSGVN 212
P +L E FF +R IL+PT + V IN +ML + GEE +L+ D+ SD DS N
Sbjct: 851 PAMLKVNEDQKFFLKRVILSPTNDDVHTINQYMLNNMEGEERIFLSADSIDPSDSDSLKN 910
Query: 213 AEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVA 272
+T + LN K SG+P+H+++LK+G PI+L+RN+D GLCNGTR+ + +T ++ A
Sbjct: 911 PV-ITPDLLNSIKLSGLPHHELRLKIGAPIILLRNLDPKGGLCNGTRLQITQMTIQVLQA 969
Query: 273 IVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQ 321
V+ + G+ IP +++TPSN +PF+ +RRQFPV L FAMTINKSQ
Sbjct: 970 KVIIGDRSGDIVLIPLINITPSNMKLPFRMRRRQFPVSLAFAMTINKSQ 1018
>At2g07620 putative helicase
Length = 1241
Score = 149 bits (376), Expect = 2e-36
Identities = 76/161 (47%), Positives = 105/161 (65%), Gaps = 1/161 (0%)
Query: 192 EEIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQT 251
E I YL+ D+ D S +N T FLN K SG+ NH + LK+G P+ML++NID
Sbjct: 1050 ESITYLSADSIDPQDPAS-LNNPVFTPYFLNSIKLSGLSNHNLTLKIGTPVMLLKNIDPK 1108
Query: 252 AGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVIL 311
GLCNGTR+ V + +I+ A V+ + + + I + ++PS+ +PF+ +RRQFP+ +
Sbjct: 1109 GGLCNGTRLQVTQMGNHILEARVITGDRVRDKVIIIKAQISPSDTKLPFRMRRRQFPIAV 1168
Query: 312 CFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
FAM I KSQGQS V +YLPRPVF+H QLYVALSRV S+
Sbjct: 1169 AFAMRIKKSQGQSLKEVEIYLPRPVFSHGQLYVALSRVTSK 1209
>At3g43350 putative protein
Length = 830
Score = 142 bits (359), Expect = 2e-34
Identities = 84/190 (44%), Positives = 109/190 (57%), Gaps = 36/190 (18%)
Query: 165 FQERAILAPTLESVEEINNFMLAMILGEEIEYLNCDTPCKSDEDSGVNAEWVTSEFLNDY 224
+Q RAIL PT E V IN M+ M+ GEE YL+ D+ +D S NA ++ ++FLN+
Sbjct: 441 YQGRAILCPTNEDVNSINEHMMRMLDGEERIYLSSDSIDPADISSANNAAYL-ADFLNNV 499
Query: 225 KCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETT 284
+ G+PNH ++LKVG P+ML+RN+D GLCNGTR+ V +T II
Sbjct: 500 RVYGLPNHCLRLKVGCPVMLLRNMDPNKGLCNGTRLQVTQMTDTII-------------- 545
Query: 285 FIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYV 344
Q I FAMTINKSQGQ+ VGLYLPRPVF+H QLYV
Sbjct: 546 ---------------------QARFITAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYV 584
Query: 345 ALSRVKSRNV 354
A+SRV S+ +
Sbjct: 585 AISRVTSKTI 594
Score = 134 bits (338), Expect = 6e-32
Identities = 64/117 (54%), Positives = 85/117 (71%)
Query: 238 VGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNAD 297
+G P+ML+RN+D GLCNGTR+ V + +I A + N +G+ IPRM +TPS+
Sbjct: 594 IGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTR 653
Query: 298 IPFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSRNV 354
+PFK +R+QF + + FAMTINKSQGQ+ VGLYLPRPVF+H QLYVA+SRV S+ V
Sbjct: 654 LPFKMRRKQFALSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSKTV 710
Score = 131 bits (329), Expect = 7e-31
Identities = 63/115 (54%), Positives = 83/115 (71%)
Query: 238 VGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNAD 297
VG P+ML+RN+D GLCNGTR+ V + +I A + N +G+ IPRM +TP +
Sbjct: 710 VGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPLDTR 769
Query: 298 IPFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
+PFK +R+QF + + FAMTINKSQGQ+ VGLYLPRPVF+H QLYVA+SRV S+
Sbjct: 770 LPFKMRRKQFALSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSK 824
>At3g51690 putative protein
Length = 374
Score = 125 bits (313), Expect = 5e-29
Identities = 75/219 (34%), Positives = 121/219 (55%), Gaps = 29/219 (13%)
Query: 135 IFIEPCKDPLLEIVNFSYPKLLFNLEKNSFFQERAILAPTLESVEEINNFMLAMILGEEI 194
+ I KDP+ ++ Y + F + AIL + V++IN++ML+++ GEE
Sbjct: 87 LLITESKDPIKTLLKEVYGEYFAKSYNPDFCHDSAILCHRDDDVDQINDYMLSLLPGEEK 146
Query: 195 EYLNCDTPCKSDEDSGVNAEWVTSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGL 254
E L+ D+ S D +V E LN K G+P+ +++LKVG P+ML+R++D + G
Sbjct: 147 ECLSTDSISPSPNDD----MFVPLEVLNSIKVPGLPDFKLRLKVGAPVMLLRDLDPSRG- 201
Query: 255 CNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFA 314
N G+ +IPR++ P+ + P + +R Q+P+ L FA
Sbjct: 202 -----------------------NKHGKKIWIPRIASYPTETNFPLQMRRTQYPLKLAFA 238
Query: 315 MTINKSQGQSSTHVGLYLPRPVFTH-VQLYVALSRVKSR 352
MTI++SQ + + VGLYLPR VF+H Q++VA+S+VKSR
Sbjct: 239 MTIDESQVHTLSKVGLYLPRQVFSHGRQMFVAISKVKSR 277
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.343 0.151 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,281,312
Number of Sequences: 26719
Number of extensions: 276147
Number of successful extensions: 1134
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1072
Number of HSP's gapped (non-prelim): 40
length of query: 378
length of database: 11,318,596
effective HSP length: 101
effective length of query: 277
effective length of database: 8,619,977
effective search space: 2387733629
effective search space used: 2387733629
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0121b.6


