

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.2
(253 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
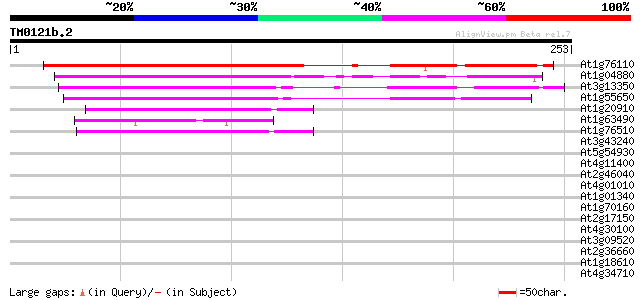
At1g76110 hypothetical protein 207 3e-54
At1g04880 unknown protein 140 7e-34
At3g13350 unknown protein 112 2e-25
At1g55650 unknown protein 108 3e-24
At1g20910 unknown protein 50 1e-06
At1g63490 RB-binding protein -like 46 2e-05
At1g76510 putative DNA-binding protein 43 1e-04
At3g43240 putative protein 38 0.005
At5g54930 unknown protein 34 0.086
At4g11400 putative protein 33 0.11
At2g46040 unknown protein 30 1.2
At4g01010 cyclic nucleotide gated channel (CNGC4) like protein 29 2.1
At1g01340 CaM-regulated potassium ion channel (ACBK1) 29 2.1
At1g70160 unknown protein 29 2.8
At2g17150 unknown protein 28 3.6
At4g30100 putative protein 28 4.7
At3g09520 hypothetical protein 28 4.7
At2g36660 putative poly(A) binding protein 28 4.7
At1g18610 unknown protein 28 4.7
At4g34710 arginine decarboxylase (spe2) 28 6.2
>At1g76110 hypothetical protein
Length = 338
Score = 207 bits (528), Expect = 3e-54
Identities = 117/233 (50%), Positives = 142/233 (60%), Gaps = 43/233 (18%)
Query: 16 KFYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGY 75
K YP PLA HEV+V D+ +F DTLR+FH M +K+MIPVIGG++LDLH LYVEVTRR GY
Sbjct: 25 KEYPEPLALHEVVVKDSSVFWDTLRRFHSIMSTKFMIPVIGGKELDLHVLYVEVTRRGGY 84
Query: 76 EKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVL 135
EKVV EKKWREVG VF FS TTTSAS+VL+KHY NLL+++EQVH F +GP+ P T
Sbjct: 85 EKVVVEKKWREVGGVFRFSATTTSASFVLRKHYLNLLFHYEQVHLFTARGPLLHPIAT-- 142
Query: 136 NSFTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAP---KHANNG 192
HA + S ++ALVEY P ++ N
Sbjct: 143 -------------------FHA--------------NPSTSKEMALVEYTPPSIRYHNTH 169
Query: 193 PESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
P S + G IE KFDCGYLV VKLGSE+L GV++H + P PSS
Sbjct: 170 PPSQGSSSFT---AIGTIEGKFDCGYLVKVKLGSEILNGVLYHSAQ--PGPSS 217
>At1g04880 unknown protein
Length = 448
Score = 140 bits (353), Expect = 7e-34
Identities = 85/222 (38%), Positives = 131/222 (58%), Gaps = 30/222 (13%)
Query: 21 PLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVA 80
P A++E +V D +LF+ +L + H +G+K+M+P+IGGR LDLH L+VEVT R G K++
Sbjct: 21 PEATYEAVVADPRLFMTSLERLHSLLGTKFMVPIIGGRDLDLHKLFVEVTSRGGINKILN 80
Query: 81 EKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTL 140
E++W+EV + F F T T+ASYVL+K+Y++LL N+EQ++FF+ G I PP S
Sbjct: 81 ERRWKEVTATFVFPPTATNASYVLRKYYFSLLNNYEQIYFFRSNGQI-PPDSMQSPSARP 139
Query: 141 CFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAK 200
CF +QG I E +++ + + +P + E+ + SNV
Sbjct: 140 CF-----IQGA---IRPSQELQAL-------TFTPQPKINTAEFL---GGSLAGSNV--- 178
Query: 201 GYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFH--PEETV 240
G I+ KF+ GYLV+V +GSE L+GV++ P+ TV
Sbjct: 179 ------VGVIDGKFESGYLVTVTIGSEQLKGVLYQLLPQNTV 214
>At3g13350 unknown protein
Length = 319
Score = 112 bits (279), Expect = 2e-25
Identities = 73/228 (32%), Positives = 114/228 (49%), Gaps = 49/228 (21%)
Query: 23 ASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEK 82
A ++ +V ++ LF + LR F +P +GG LDLH L++EVT R G E+VV ++
Sbjct: 34 AKYDDLVRNSALFWEKLRAFLGLTSKTLKVPTVGGNTLDLHRLFIEVTSRGGIERVVKDR 93
Query: 83 KWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCF 142
KW+EV F+F TT TSAS+VL+K+Y L+ E V++ ++ P++
Sbjct: 94 KWKEVIGAFSFPTTITSASFVLRKYYLKFLFQLEHVYY--LEKPVS-------------- 137
Query: 143 MEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAKGY 202
SLQ S+ LA P+ + P+ E +G+
Sbjct: 138 ----SLQ---------------------STDEALKSLANESPNPEEGIDEPQVGYEVQGF 172
Query: 203 PRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSSRVVGT 250
I+ KFD GYLV++KLGS+ L+GV++H +T P S + + T
Sbjct: 173 -------IDGKFDSGYLVTMKLGSQELKGVLYHIPQT-PSQSQQTMET 212
>At1g55650 unknown protein
Length = 337
Score = 108 bits (270), Expect = 3e-24
Identities = 67/211 (31%), Positives = 105/211 (49%), Gaps = 48/211 (22%)
Query: 25 HEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKW 84
++ IV + +LF + LR FH K+ IP++GG+ LDLH L+ EVT R G EKV+ +++
Sbjct: 30 YQDIVRNPELFWEMLRDFHESSDKKFKIPIVGGKSLDLHRLFNEVTSRGGLEKVIKDRRC 89
Query: 85 REVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCFME 144
+EV FNF TT T++++VL+K Y +L+ FE +++F Q P+
Sbjct: 90 KEVIDAFNFKTTITNSAFVLRKSYLKMLFEFEHLYYF--QAPL----------------- 130
Query: 145 MDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAKGYPR 204
S+ + + AL K AN +S G
Sbjct: 131 ---------------------------STFWEKEKALKLLIEKSANRDKDSQELKPG--T 161
Query: 205 YGEGRIEEKFDCGYLVSVKLGSEVLRGVIFH 235
G I+ KF+ GYL+S K+GSE L+G+++H
Sbjct: 162 VITGIIDGKFESGYLISTKVGSEKLKGMLYH 192
>At1g20910 unknown protein
Length = 398
Score = 50.1 bits (118), Expect = 1e-06
Identities = 30/103 (29%), Positives = 47/103 (45%), Gaps = 2/103 (1%)
Query: 35 FLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFS 94
FL + F+ ++ P G+ L++ L+ V GYE V K WR+VG FN
Sbjct: 112 FLREVEAFYKESFLEFKPPKFYGQPLNILKLWRAVVNLGGYEVVTTNKLWRQVGESFNPP 171
Query: 95 TTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNS 137
T T+ SY + Y L +E+ + G + P T++ S
Sbjct: 172 KTCTTVSYTFRNFYEKALLEYEKC--LRNNGELNLPGSTLILS 212
>At1g63490 RB-binding protein -like
Length = 1458
Score = 45.8 bits (107), Expect = 2e-05
Identities = 34/95 (35%), Positives = 47/95 (48%), Gaps = 8/95 (8%)
Query: 30 NDAKLFLDTLRQFHFHMGSKYMIPVI-GGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVG 88
N L+ R H+G K V+ G +LDL L+ V R GYEKVV KKW G
Sbjct: 96 NSKTFKLEYNRFLEEHLGKKLKKRVVFEGEELDLCKLFNAVKRFGGYEKVVKGKKW---G 152
Query: 89 SVFNFSTT----TTSASYVLKKHYWNLLYNFEQVH 119
V+ F ++ + A +VL + Y L++FE H
Sbjct: 153 EVYQFMSSGEKISKCAKHVLCQLYKEHLHDFENYH 187
>At1g76510 putative DNA-binding protein
Length = 434
Score = 43.1 bits (100), Expect = 1e-04
Identities = 27/107 (25%), Positives = 47/107 (43%), Gaps = 2/107 (1%)
Query: 31 DAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSV 90
D + F+ + F+ ++ P G+ L+ L+ V + GY+ V K WR+VG
Sbjct: 144 DQEAFIKEVEAFNKENFLEFKAPKFYGQPLNCLKLWRAVIKLGGYDVVTTSKLWRQVGES 203
Query: 91 FNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNS 137
F+ T T+ S+ + Y L +E+ + G + P L S
Sbjct: 204 FHPPKTCTTVSWTFRIFYEKALLEYEK--HLRQNGELNLPGSASLPS 248
>At3g43240 putative protein
Length = 717
Score = 38.1 bits (87), Expect = 0.005
Identities = 34/122 (27%), Positives = 49/122 (39%), Gaps = 24/122 (19%)
Query: 19 PPPLASHE-----VIVNDAKLFLDTLRQFHFHMGSKYMIP----------VIGGRKLDLH 63
P PL H+ + V + FL + QF G ++P V+ ++LDL
Sbjct: 521 PLPLKKHDCGRAHIQVCSEEEFLRDVMQFLLIRGHTRLVPPGGLAEFPDAVLNSKRLDLF 580
Query: 64 TLYVEVTRRSGYEKVVAEKKWREVGSVFN------FSTTTTSASYVLKKHYWNLLYNFEQ 117
LY EV R G+ V W+ G VF+ + T LK+HY L +E
Sbjct: 581 NLYREVVSRGGFH-VGNGINWK--GQVFSKMRNHTLTNRMTGVGNTLKRHYETYLLEYEY 637
Query: 118 VH 119
H
Sbjct: 638 AH 639
>At5g54930 unknown protein
Length = 286
Score = 33.9 bits (76), Expect = 0.086
Identities = 17/36 (47%), Positives = 24/36 (66%), Gaps = 2/36 (5%)
Query: 208 GRIEEKFDCGYLVSVKLGS--EVLRGVIFHPEETVP 241
G IE F+ G+L+SVK+G+ +LRGV+F P P
Sbjct: 68 GVIEATFEAGFLLSVKVGNSDSMLRGVVFKPGHCDP 103
>At4g11400 putative protein
Length = 573
Score = 33.5 bits (75), Expect = 0.11
Identities = 24/80 (30%), Positives = 37/80 (46%), Gaps = 2/80 (2%)
Query: 33 KLFLDTLRQFHFHMGSKYMIP-VIG-GRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSV 90
+LF L F GS +P VIG G+ +DL L+V V R G++ V ++ W V
Sbjct: 28 RLFDQALLVFLEEEGSIKPLPAVIGDGKNVDLFKLFVLVREREGFDTVSRKRLWEVVAEK 87
Query: 91 FNFSTTTTSASYVLKKHYWN 110
F + + ++ Y N
Sbjct: 88 LGFDCSLVPSLILIYLKYLN 107
>At2g46040 unknown protein
Length = 562
Score = 30.0 bits (66), Expect = 1.2
Identities = 12/31 (38%), Positives = 18/31 (57%)
Query: 57 GRKLDLHTLYVEVTRRSGYEKVVAEKKWREV 87
GR +DL L++ VT + G++ V W EV
Sbjct: 76 GRTVDLFNLFLNVTHKGGFDAVSENGSWDEV 106
>At4g01010 cyclic nucleotide gated channel (CNGC4) like protein
Length = 689
Score = 29.3 bits (64), Expect = 2.1
Identities = 20/56 (35%), Positives = 28/56 (49%), Gaps = 11/56 (19%)
Query: 58 RKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLY 113
R L ++ LY EVTR SG +V E W G+ +N S Y+L H + L+
Sbjct: 217 RILRIYPLYTEVTRTSG---IVTETAW--AGAAWNLSL------YMLASHVFGALW 261
>At1g01340 CaM-regulated potassium ion channel (ACBK1)
Length = 706
Score = 29.3 bits (64), Expect = 2.1
Identities = 20/56 (35%), Positives = 28/56 (49%), Gaps = 11/56 (19%)
Query: 58 RKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLY 113
R L ++ LY EVTR SG +V E W G+ +N S Y+L H + L+
Sbjct: 212 RILRMYPLYTEVTRTSG---IVTETAW--AGAAWNLSL------YMLASHVFGALW 256
>At1g70160 unknown protein
Length = 523
Score = 28.9 bits (63), Expect = 2.8
Identities = 20/94 (21%), Positives = 48/94 (50%), Gaps = 10/94 (10%)
Query: 15 GKFYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRS- 73
G YPPPL +H ++++ ++ + +M ++ + +G LDL+ + E RR
Sbjct: 344 GDNYPPPLDAH-LVISVMSMWTRVQPAYAANMWNEALNKRLGTEDLDLYGILEETARRGM 402
Query: 74 GYEKVVA---EKKWREVGSVFNFSTTTTSASYVL 104
+++++ + +W V++ +TT +++L
Sbjct: 403 SFDELLTIPEQDEW-----VYSDGKSTTCVAFIL 431
>At2g17150 unknown protein
Length = 909
Score = 28.5 bits (62), Expect = 3.6
Identities = 19/60 (31%), Positives = 30/60 (49%), Gaps = 1/60 (1%)
Query: 145 MDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAKGYPR 204
MDS+QG +I + + S ++ S + SS P L E P H N ++ + A+ PR
Sbjct: 667 MDSVQGAQGSIQLDSFYTSFPELNSPNMSSNGPSLKSNE-QPSHLNAQTDNGIMAEENPR 725
>At4g30100 putative protein
Length = 1311
Score = 28.1 bits (61), Expect = 4.7
Identities = 22/78 (28%), Positives = 32/78 (40%), Gaps = 5/78 (6%)
Query: 141 CFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGP---ESNV 197
CFMEM+SL +P + V F G S G P ++ P+ + P + +
Sbjct: 1205 CFMEMESLPKDFP-VPKVPSFIPKAPNARGFRSGG-PRTRSIDMHPESRSGTPSEDDKKL 1262
Query: 198 EAKGYPRYGEGRIEEKFD 215
+PR G R E D
Sbjct: 1263 STTTFPRNGNSRRENSVD 1280
>At3g09520 hypothetical protein
Length = 628
Score = 28.1 bits (61), Expect = 4.7
Identities = 28/120 (23%), Positives = 50/120 (41%), Gaps = 16/120 (13%)
Query: 84 WREVGSVFNFSTTTTSASYVLK------KHYWNLLYNFEQVHFFKVQGPITPPAGTVLNS 137
W+ + S+F+F + + S LK + +LL FE K + P G V +
Sbjct: 315 WQAIESIFSFDSISVVRSLALKSLISLSESIRSLLVEFES-GIQKDSSKVVVPGGGV-HP 372
Query: 138 FTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNV 197
T+ M+ SL Y N+ + D+L+GS + L + +++ P S +
Sbjct: 373 LTISVMDHLSLLADYSNV--------LVDILAGSPPPDRSLLPESYFNVSESDDSPSSEL 424
>At2g36660 putative poly(A) binding protein
Length = 609
Score = 28.1 bits (61), Expect = 4.7
Identities = 35/128 (27%), Positives = 54/128 (41%), Gaps = 11/128 (8%)
Query: 132 GTVLNSFTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSS--GKPDLALVEYAP--- 186
G V S+ L + + + YPN AVT + + S + + GK D V Y P
Sbjct: 424 GMVYQSYPLMWKSANMIGSSYPNSEAVTYPPMVANAPSKNRQNRIGKLDRNAVSYVPNVY 483
Query: 187 KHANNGPESNVEAKGY--PRYGEGRIEEKFDCGYLVSVKLGSEV-LRGVIFHP--EETVP 241
+ P S +K YG G+ E K S +G E+ L G + HP E+ P
Sbjct: 484 QSTQMLPLSRDFSKQQHSRTYGRGK-EMKKSIQQRQSETVGMEMQLLGELLHPLVEKLEP 542
Query: 242 PPSSRVVG 249
++++ G
Sbjct: 543 QLANKITG 550
>At1g18610 unknown protein
Length = 574
Score = 28.1 bits (61), Expect = 4.7
Identities = 11/28 (39%), Positives = 17/28 (60%)
Query: 207 EGRIEEKFDCGYLVSVKLGSEVLRGVIF 234
+ RI E + GY + + +VLRGV+F
Sbjct: 408 QARITESYPVGYTMETMIDGKVLRGVLF 435
>At4g34710 arginine decarboxylase (spe2)
Length = 711
Score = 27.7 bits (60), Expect = 6.2
Identities = 21/75 (28%), Positives = 36/75 (48%), Gaps = 8/75 (10%)
Query: 10 SGGDDGKFYPPPLASHEVIVNDAKL----FLDTLRQFHFHMGSKY-MIPVIGGRKLDLHT 64
+ G+ GKF L + +++ KL LD L+ HFH+GS+ ++ +
Sbjct: 260 TSGEKGKF---GLTTTQIVRVVRKLRQSGMLDCLQLLHFHIGSQIPSTSLLSDGVAEAAQ 316
Query: 65 LYVEVTRRSGYEKVV 79
LY E+ R + KV+
Sbjct: 317 LYCELVRLGAHMKVI 331
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,324,212
Number of Sequences: 26719
Number of extensions: 270179
Number of successful extensions: 596
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 579
Number of HSP's gapped (non-prelim): 28
length of query: 253
length of database: 11,318,596
effective HSP length: 97
effective length of query: 156
effective length of database: 8,726,853
effective search space: 1361389068
effective search space used: 1361389068
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0121b.2


