

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0118c.1
(565 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
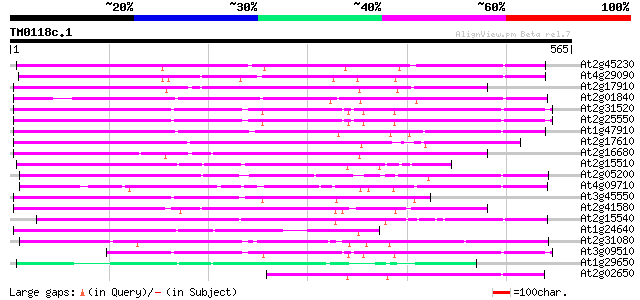
Score E
Sequences producing significant alignments: (bits) Value
At2g45230 putative non-LTR retroelement reverse transcriptase 248 5e-66
At4g29090 putative protein 241 6e-64
At2g17910 putative non-LTR retroelement reverse transcriptase 227 1e-59
At2g01840 putative non-LTR retroelement reverse transcriptase 214 8e-56
At2g31520 putative non-LTR retroelement reverse transcriptase 212 4e-55
At2g25550 putative non-LTR retroelement reverse transcriptase 209 3e-54
At1g47910 reverse transcriptase, putative 209 3e-54
At2g17610 putative non-LTR retroelement reverse transcriptase 199 3e-51
At2g16680 putative non-LTR retroelement reverse transcriptase 194 8e-50
At2g15510 putative non-LTR retroelement reverse transcriptase 190 2e-48
At2g05200 putative non-LTR retroelement reverse transcriptase 187 1e-47
At4g09710 RNA-directed DNA polymerase -like protein 186 4e-47
At3g45550 putative protein 180 2e-45
At2g41580 putative non-LTR retroelement reverse transcriptase 178 8e-45
At2g15540 putative non-LTR retroelement reverse transcriptase 167 2e-41
At1g24640 hypothetical protein 160 2e-39
At2g31080 putative non-LTR retroelement reverse transcriptase 145 6e-35
At3g09510 putative non-LTR reverse transcriptase 137 2e-32
At1g29650 reverse transcriptase, putative 135 5e-32
At2g02650 putative reverse transcriptase 115 8e-26
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 248 bits (634), Expect = 5e-66
Identities = 164/559 (29%), Positives = 269/559 (47%), Gaps = 36/559 (6%)
Query: 8 INLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGF 67
+ +A+P+Y MSCF + +C QIE +++ F+W R +HW W L RPK GG+GF
Sbjct: 796 VAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWKAWCHLSRPKAVGGLGF 855
Query: 68 RDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSA 127
++ +AFN ALL K WR++T+ S++AK++K+ YF K L A G RPS+ W S+ ++
Sbjct: 856 KEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSDPLNAPLGSRPSFAWKSIYEAQ 915
Query: 128 WIFQEGGLWRVGNGKNIRA*EDKWV----PHGGPLVYRNDVAESLGV--VH-VSDLMVTN 180
+ ++G +GNG+ I D W+ V R+ + +H V DL++ +
Sbjct: 916 VLIKQGIRAVIGNGETINVWTDPWIGAKPAKAAQAVKRSHLVSQYAANSIHVVKDLLLPD 975
Query: 181 GGCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSE 240
G W+ N V L T IL++ ++ D F W +R G YS ++GY + + ++
Sbjct: 976 GRDWNWNLVSLLFPDNTQENILALRPGGKETRDRFTWEYSRSGHYSVKSGYWVMTEIINQ 1035
Query: 241 GEASSSSARVMPPAL---WMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCC 297
++ V+ P+L + ++WK D P+ WR + L V S L R + C
Sbjct: 1036 ---RNNPQEVLQPSLDPIFQQIWKLDVPPKIHHFLWRCVNNCLSVASNLAYRHLAREKSC 1092
Query: 298 PRCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSV-------HDFFLSFLATADTN 350
RC ETV+H LF CP R+ W SPL E + H + +++
Sbjct: 1093 VRCPSHGETVNHLLFKCPFARLTWAISPLPAPPGGEWAESLFRNMHHVLSVHKSQPEESD 1152
Query: 351 IAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPS---------RVLLDDGPRA 401
+ +L+ +W+ RN+L+FK ++ + Q++ +A+ + +V R
Sbjct: 1153 HHALIPWILWRLWKNRNDLVFKGREFTAPQVILKATEDMDAWNNRKEPQPQVTSSTRDRC 1212
Query: 402 VQVLPSSWSRPLSGVCKMNIDASVTSS-KEAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
V+ W P G K N D + + G G V+RN+ G +L L S +
Sbjct: 1213 VK-----WQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLLWLGLRALPSQQSVLE 1267
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDV 520
E +RWA+ ++ +RR+ E+D L + + + LA + D R+LL +F+
Sbjct: 1268 TEVEALRWAVLSLSRFNYRRVIFESDSQYLVSLIQNE-MDIPSLAPRIQDIRNLLRHFEE 1326
Query: 521 FNFTFVRRTSNGAADALAR 539
F F RR N AD AR
Sbjct: 1327 VKFQFTRREGNNVADRTAR 1345
>At4g29090 putative protein
Length = 575
Score = 241 bits (616), Expect = 6e-64
Identities = 168/556 (30%), Positives = 265/556 (47%), Gaps = 32/556 (5%)
Query: 10 LAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRD 69
+A+P+Y M+CFLL +C QI +++ F+W + MHW WD L K GGIGF+D
Sbjct: 1 MALPTYTMACFLLPKTVCKQIISVLADFWWRNKQEAKGMHWKAWDHLSCYKAEGGIGFKD 60
Query: 70 FKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWI 129
+AFN ALL K WR++++ +S++AK++K+ YF K L A G RPS+VW S+ S I
Sbjct: 61 IEAFNLALLGKQMWRMLSRPESLMAKVFKSRYFHKSDPLNAPLGSRPSFVWKSIHASQEI 120
Query: 130 FQEGGLWRVGNGKNIRA*EDKWV---PHGGPL----VYRNDVAESLGVVHVSDLMVTNGG 182
++G VGNG++I KW+ P L V + A ++ VSDL+ +G
Sbjct: 121 LRQGARAVVGNGEDIIIWRHKWLDSKPASAALRMQRVPPQEYASVSSILKVSDLIDESGR 180
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRD-DCFFWPETRDGCYSTRTGY----SFI*KK 237
W ++ +E L +P R ++ P +R D + W T G Y+ ++GY I K+
Sbjct: 181 EWRKDVIEML-FPEVERKLIGELRPGGRRILDSYTWDYTSSGDYTVKSGYWVLTQIINKR 239
Query: 238 FSEGEASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCC 297
S E S S ++ K+WK+ P+ W+ + LPV L R + C
Sbjct: 240 SSPQEVSEPSLN----PIYQKIWKSQTSPKIQHFLWKCLSNSLPVAGALAYRHLSKESAC 295
Query: 298 PRCSEVEETVHHALFSCPLVRMIWFAS----PLNLRFDEEGSVHDFFLSFLATAD---TN 350
RC +ETV+H LF C R+ W S PL + + V+ +++ L +
Sbjct: 296 IRCPSCKETVNHLLFKCTFARLTWAISSIPIPLGGEWADSIYVNLYWVFNLGNGNPQWEK 355
Query: 351 IAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQRASS------LRPSRVLLDDGPRAVQV 404
+ + +L+ +W+ RNEL+F+ ++ + ++L+RA +R P+ +
Sbjct: 356 ASQLVPWLLWRLWKNRNELVFRGREFNAQEVLRRAEDDLEEWRIRTEAESCGTKPQVNRS 415
Query: 405 LPSSWSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEA 463
W P K N DA+ E G G V+RN GEV L + S + AE
Sbjct: 416 SCGRWRPPPHQWVKCNTDATWNRDNERCGIGWVLRNEKGEVKWMGARALPKLKSVLEAEL 475
Query: 464 LGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNF 523
MRWA+ ++ + + E+D L N + L + D + LL F F
Sbjct: 476 EAMRWAVLSLSRFQYNYVIFESDSQVLIEILNND-EIWPSLKPTIQDLQRLLSQFTEVKF 534
Query: 524 TFVRRTSNGAADALAR 539
F+ R N A+ +AR
Sbjct: 535 VFIPREGNTLAERVAR 550
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 227 bits (579), Expect = 1e-59
Identities = 146/491 (29%), Positives = 231/491 (46%), Gaps = 20/491 (4%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ I LA+P YVMSCF L LC ++ ++ F+W R +HWL W L PK GG
Sbjct: 771 LKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGG 830
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
GF+D + FN ALLAK WR++ + S+ ++++++ YF FL A +G RPSY W S+
Sbjct: 831 FGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSIL 890
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGG---PLVYRNDVAESLGVVHVSDLMVTNG 181
+ +G +GNG+ DKW+ G PL R + L V + D N
Sbjct: 891 FGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLNRRRFINVDLKVSQLIDPTSRN- 949
Query: 182 GCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEG 241
W+ N + L +P I+ PL ++D F W + +G YS +TGY F+ K+
Sbjct: 950 --WNLNMLRDL-FPWKDVEIILKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQVHHR 1006
Query: 242 EASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCS 301
+ + +L+ K+W P+ W+A +PV L RG+ + C C
Sbjct: 1007 LYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLMCD 1066
Query: 302 EVEETVHHALFSCPLVRMIWFASPLNLRFDE-EGSVHDFFLSFLATADTN-----IAGIF 355
ET++H LF CPL R +W + L+ E SV+ + N + +
Sbjct: 1067 TENETINHILFECPLARQVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVS 1126
Query: 356 LAVLYSVWQARNELLFKWKDSSIDQLLQRASSLR----PSRVLLDDGPRAVQVLPSSWSR 411
+L+ +W+ RN LLF+ K S L+ +A ++ + + + +++ + W
Sbjct: 1127 PWILWFLWKNRNALLFEGKGSITTTLVDKAYEAYHEWFSAQTHMQNDEKHLKI--TKWCP 1184
Query: 412 PLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWAL 470
PL G K NI + + +GA V+R++ G+VL + V SP A+ WAL
Sbjct: 1185 PLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSAKIRSWEWAL 1244
Query: 471 RLATKLGFRRI 481
T F R+
Sbjct: 1245 ESMTHHHFDRV 1255
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 214 bits (546), Expect = 8e-56
Identities = 158/559 (28%), Positives = 260/559 (46%), Gaps = 47/559 (8%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I I +A+P Y M+CFLL +C++I +I+ F+WG + G
Sbjct: 1155 IKAIAMALPVYSMNCFLLPTLICNEINSLITAFWWGKENE------------------GD 1196
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GF+D FN ALLAK WRI+T +S+LA++YK +Y+P ++L A KG SY W S+Q
Sbjct: 1197 LGFKDLHQFNRALLAKQAWRILTNPQSLLARLYKGLYYPNTTYLRANKGGHASYGWNSIQ 1256
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCW 184
+ + Q+G R+G+G+ + ED W+P P R + + + V+DL N W
Sbjct: 1257 EGKLLLQQGLRVRLGDGQTTKIWEDPWLPTLPPRPARGPILDE--DMKVADLWRENKREW 1314
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*K-KFSEGEA 243
D E + P + S+ L D + W TR+ Y+ R+GY +E E
Sbjct: 1315 DPVIFEGVLNPEDQQLAKSLYLSNYAARDSYKWAYTRNTQYTVRSGYWVATHVNLTEEEI 1374
Query: 244 SSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEV 303
+ +P L ++W+ P+ WR L + L R + +P C RC
Sbjct: 1375 INPLEGDVP--LKQEIWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNA 1432
Query: 304 EETVHHALFSCPLVRMIW----FASPLNLRFDEEGSVHDFFLSFLATADTNIA---GIF- 355
+ET++H +F+C +++W F+ L F + + L + N+ G+
Sbjct: 1433 DETINHIIFTCSYAQVVWRSANFSGSNRLCF-TDNLEENIRLILQGKKNQNLPILNGLMP 1491
Query: 356 LAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRV--LLDDGPRAVQVLPSS----- 408
+++ +W++RNE LF+ D ++ Q+A V +++D + S+
Sbjct: 1492 FWIMWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATEWVETMVNDTAISHNTAQSNDRPLS 1551
Query: 409 ----WSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEA 463
WS P G K N D+ ++ G ++R+ +G VL + +L S + AEA
Sbjct: 1552 RSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSYSALQAEA 1611
Query: 464 LGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSH-LATVVFDCRSLLPNFDVFN 522
LG AL++ G+ +W E D LEL N+ E H L T+++D R + +
Sbjct: 1612 LGFLHALQMVWIRGYCYVWFEGDNLELTNLINKT--EDHHLLETLLYDIRFWMTKLPFSS 1669
Query: 523 FTFVRRTSNGAADALARLA 541
+V R N AAD L + A
Sbjct: 1670 IGYVNRERNLAADKLTKYA 1688
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 212 bits (540), Expect = 4e-55
Identities = 164/567 (28%), Positives = 261/567 (45%), Gaps = 41/567 (7%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ + LA+P Y MSCF L G+ S+IE ++ F+W + R + W+ W L K GG
Sbjct: 949 LKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGG 1008
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD FN ALLAK WR++ S+ A++ KA YF S L+AK + SY W SL
Sbjct: 1009 LGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLL 1068
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGG-- 182
+ ++G +G+G+NIR D V P + E+ + +++L G
Sbjct: 1069 DGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTE--ETYKEMTINNLFERKGSYY 1126
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGE 242
WD +K+ I I L ++ D W G Y+ R+GY + +
Sbjct: 1127 FWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLL-----THD 1181
Query: 243 ASSSSARVMPP----ALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCP 298
S++ + PP L ++W +P+ WRA L L RG+ +P CP
Sbjct: 1182 PSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPICP 1241
Query: 299 RCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDF------FLSFLATADTNIA 352
RC E+++HALF+CP M W+ S +L ++ S +DF L+F+ DT ++
Sbjct: 1242 RCHRENESINHALFTCPFATMAWWLSDSSLIRNQLMS-NDFEENISNILNFV--QDTTMS 1298
Query: 353 GIF----LAVLYSVWQARNELLF-KWKDSSIDQLLQRAS------SLRPSRVLLDDGPRA 401
+ +++ +W+ARN ++F K+++S +L + + S R
Sbjct: 1299 DFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQ 1358
Query: 402 VQVLPSSWSRPLSGVCKMNIDASVTSSK-EAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
+ W P + K N DA K EA G ++RN++G ++ + +L +P+
Sbjct: 1359 IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLE 1418
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEF-SHLATVVFDCRSLLPNFD 519
AE + AL+ G+ ++++E DC L N G F S LA + D F
Sbjct: 1419 AETKALLAALQQTWIRGYTQVFMEGDCQTLINLIN--GISFHSSLANHLEDISFWANKFA 1476
Query: 520 VFNFTFVRRTSNGAADALARLAFHYGC 546
F F+RR N A LA+ YGC
Sbjct: 1477 SIQFGFIRRKGNKLAHVLAK----YGC 1499
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 209 bits (533), Expect = 3e-54
Identities = 163/567 (28%), Positives = 260/567 (45%), Gaps = 41/567 (7%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ + LA+P Y MSCF L G+ S+IE ++ F+W + R + W+ W L K GG
Sbjct: 1175 LKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGG 1234
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD FN ALLAK WR++ S+ A++ KA YF S L+AK + SY W SL
Sbjct: 1235 LGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLL 1294
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGG-- 182
+ ++G +G+G+NIR D V P + E+ + +++L G
Sbjct: 1295 DGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTE--ETYKEMTINNLFERKGSYY 1352
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGE 242
WD +K+ I I L ++ D W G Y+ R+GY + +
Sbjct: 1353 FWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLL-----THD 1407
Query: 243 ASSSSARVMPP----ALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCP 298
S++ + PP L ++W +P+ WRA L L RG+ +P CP
Sbjct: 1408 PSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCP 1467
Query: 299 RCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDF------FLSFLATADTNIA 352
RC E+++HALF+CP M W S +L ++ S +DF L+F+ DT ++
Sbjct: 1468 RCHRENESINHALFTCPFATMAWRLSDSSLIRNQLMS-NDFEENISNILNFV--QDTTMS 1524
Query: 353 GIF----LAVLYSVWQARNELLF-KWKDSSIDQLLQRAS------SLRPSRVLLDDGPRA 401
+ +++ +W+ARN ++F K+++S +L + + S R
Sbjct: 1525 DFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQ 1584
Query: 402 VQVLPSSWSRPLSGVCKMNIDASVTSSK-EAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
+ W P + K N DA K EA G ++RN++G ++ + +L +P+
Sbjct: 1585 IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLE 1644
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEF-SHLATVVFDCRSLLPNFD 519
AE + AL+ G+ ++++E DC L N G F S LA + D F
Sbjct: 1645 AETKALLAALQQTWIRGYTQVFMEGDCQTLINLIN--GISFHSSLANHLEDISFWANKFA 1702
Query: 520 VFNFTFVRRTSNGAADALARLAFHYGC 546
F F+R+ N A LA+ YGC
Sbjct: 1703 SIQFGFIRKKGNKLAHVLAK----YGC 1725
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 209 bits (533), Expect = 3e-54
Identities = 148/554 (26%), Positives = 254/554 (45%), Gaps = 28/554 (5%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I + +P YVMSCF L + S++ +++F+W + R MHW+ WD LC K GG
Sbjct: 577 IKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDGG 636
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFR+ FN+ALLAK WR++T S+ AK++K YF K + L++ K Y PSY W S+
Sbjct: 637 LGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSMI 696
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVP--HGGPLVYRNDVAESLGVVHVSDLMVTNGG 182
+ + +G + RVG+G +I D W+P P Y + + + V L+ +
Sbjct: 697 SARSLVYKGLIKRVGSGASISVWNDPWIPAQFPRPAKYGGSIVDP--SLKVKSLIDSRSN 754
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGE 242
W+ + ++ L P I ++P+ +D W T+ G Y+ ++GY +EG
Sbjct: 755 FWNIDLLKELFDPEDVPLISALPIGNPNMEDTLGWHFTKAGNYTVKSGYHTARLDLNEG- 813
Query: 243 ASSSSARVMPPALWMK--LWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRC 300
+ + P +K +WK P+ W+ +PV L +RG++ + C C
Sbjct: 814 ----TTLIGPDLTTLKAYIWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSC 869
Query: 301 SEVEETVHHALFSCPLVRMIWFASPLNLR---FDEEGSVHDFFLSFLATADTNIAGIFLA 357
EE+++H LF C R IW S + F + F + +
Sbjct: 870 GASEESINHTLFQCHPARQIWALSQIPTAPGIFPSNSIFTNLDHLFWRIPSGVDSAPYPW 929
Query: 358 VLYSVWQARNELLFKWKDSSIDQLL----QRASSLRPSRVLLDDGPRA-------VQVLP 406
+++ +W+ARNE +F+ D ++L + A S + ++V L ++V
Sbjct: 930 IIWYIWKARNEKVFENVDKDPMEILLLAVKEAQSWQEAQVELHSERHGSLSIDSRIRVRD 989
Query: 407 SSWSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALG 465
S SG + ID S +S + +G G ++ GE + LSP+ E
Sbjct: 990 VSQDTTFSGF-RCFIDGSWKASDQFSGTGWFCLSSLGESPTMGAANVRRSLSPLHTEMEA 1048
Query: 466 MRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTF 525
+ WA++ + + TDC +L + E+ + + + +S F F+ +
Sbjct: 1049 LLWAMKCMIGADNQNVAFFTDCSDLVKMVS-SPTEWPAFSVYLEELQSDREEFTNFSLSL 1107
Query: 526 VRRTSNGAADALAR 539
+ R++N AD LAR
Sbjct: 1108 ISRSANVKADKLAR 1121
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 199 bits (507), Expect = 3e-51
Identities = 149/525 (28%), Positives = 235/525 (44%), Gaps = 36/525 (6%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I I AIP+Y MSCFLL L QI + F+W + W+ W L PK GG
Sbjct: 248 IKSIVTAIPTYSMSCFLLPMRLIHQITSAMRWFWWSNTKVKHKIPWVAWSKLNDPKKMGG 307
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+ RD K FN ALLAK WRI+ + S++A+++KA YFPKE L+AK + SY W S+
Sbjct: 308 LAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKYFPKERLLDAKATSQSSYAWKSIL 367
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCW 184
+ G + GNG NI+ +D W+P P + VSDL++ G W
Sbjct: 368 HGTKLISRGLKYIAGNGNNIQLWKDNWLPLNPPRPPVGTCDSIYSQLKVSDLLIE--GRW 425
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEAS 244
+ + + L I +I + +D W T DG YS ++GY + K + AS
Sbjct: 426 NEDLLCKLIHQNDIPHIRAIRPSITGANDAITWIYTHDGNYSVKSGYHLLRKLSQQQHAS 485
Query: 245 SSSAR-VMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEV 303
S V ++ +WK +A P+ WR+ + LP L RR ++ + C RC E
Sbjct: 486 LPSPNEVSAQTVFTNIWKQNAPPKIKHFWWRSAHNALPTAGNLKRRRLITDDTCQRCGEA 545
Query: 304 EETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTNIAG-----IFLAV 358
E V+H LF C + + IW + + L + + F + + N + +F +
Sbjct: 546 SEDVNHLLFQCRVSKEIWEQAHIKLCPGDSLMSNSFNQNLESIQKLNQSARKDVSLFPFI 605
Query: 359 LYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRVLLDDGPRAVQVLPSSWSRPLSGVC- 417
+ +W+ RN+L+F K SI +Q+A L+D W L+ C
Sbjct: 606 GWRIWKMRNDLIFNNKRWSIPDSIQKA--------LIDQ---------QQWKESLN--CN 646
Query: 418 -----KMNIDASVTSSKEAGAGMVMRN-NHGEVLAAATTELGPVLSPVLAEALGMRWALR 471
K + S + G +R + + + + + P +P+ AEA ++ A+
Sbjct: 647 EQQQRKPQLHDSHNNKCYGGHLKTLRTLSKAQFVLRGMSSIPPTSTPLEAEAEALKLAMI 706
Query: 472 LATKLGFRRIWIETDCLELYTFWNR--QGGEFSHLATVVFDCRSL 514
+LG+ I D L+ R E+S +AT + D ++
Sbjct: 707 HLQRLGYEDIIFHGDVAALFQPITRTTHSCEYSPIATFLRDIENI 751
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 194 bits (494), Expect = 8e-50
Identities = 134/489 (27%), Positives = 225/489 (45%), Gaps = 17/489 (3%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ I LA+P Y+M+CF L GLC+++ ++ F+W +HW+ L PK GG
Sbjct: 745 LKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFWWNSMEFSNKIHWIGGKKLTLPKSLGG 804
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
GF+D + FN ALLAK WR+ + KS++++I+K+ YF FL A++G RPSY W S+
Sbjct: 805 FGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNTDFLNARQGTRPSYTWRSIL 864
Query: 125 KSAWIFQEGGLWR-VGNGKNIRA*EDKWVPHG---GPLVYRNDVAESLGVVHVSDLMVTN 180
+ GGL R +GNG+ DKW+ G P+ + + + V H+ D + N
Sbjct: 865 YGRELL-NGGLKRLIGNGEQTNVWIDKWLFDGHSRRPMNLHSLMNIHMKVSHLIDPLTRN 923
Query: 181 GGCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSE 240
W+ K+ L + I+ PL +D + W T +G Y+ ++GY ++ +
Sbjct: 924 ---WNLKKLTELFHEKDVQLIMH-QRPLISSEDSYCWAGTNNGLYTVKSGYERSSRETFK 979
Query: 241 GEASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRC 300
+ L+ K+W + VP+ W+A L V L RG+ C C
Sbjct: 980 NLFKEADVYPSVNPLFDKVWSLETVPKIKVFMWKALKGALAVEDRLRSRGIRTADGCLFC 1039
Query: 301 SEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTN------IAGI 354
E ET++H LF CP R +W S + G+ ++ + N + +
Sbjct: 1040 KEEIETINHLLFQCPFARQVWALSLIQAPATGFGTSIFSNINHVIQNSQNFGIPRHMRTV 1099
Query: 355 FLAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRV-LLDDGPRAVQVLPSSWSRPL 413
+L+ +W+ RN+ LF+ + +++ +A + + V W+ P
Sbjct: 1100 SPWLLWEIWKNRNKTLFQGTGLTSSEIVAKAYEECNLWINAQEKSSGGVSPSEHKWNPPP 1159
Query: 414 SGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRL 472
+G K NI + + K+ AG V+R++ G+VL + V S A+ WAL
Sbjct: 1160 AGELKCNIGVAWSRQKQLAGVSWVLRDSMGQVLLHSRRSYSQVYSLFDAKIKSWDWALES 1219
Query: 473 ATKLGFRRI 481
F ++
Sbjct: 1220 MDHFHFDKV 1228
>At2g15510 putative non-LTR retroelement reverse transcriptase
Length = 1138
Score = 190 bits (483), Expect = 2e-48
Identities = 136/462 (29%), Positives = 216/462 (46%), Gaps = 33/462 (7%)
Query: 8 INLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGF 67
+ +A+ Y MSCF L C + +S F+W R HW+ W+ LC K GG+GF
Sbjct: 572 VAMAMQVYAMSCFKLTKTTCKNLTSAMSDFWWNALEHKRKTHWVSWEKLCLAKESGGLGF 631
Query: 68 RDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSA 127
RD ++FN ALLAK WRI+ + A+ +K+ YF E FLEA G RPSY S+
Sbjct: 632 RDIESFNQALLAKQSWRILQYPSFLFARFFKSRYFDDEEFLEADLGVRPSYACCSILFGR 691
Query: 128 WIFQEGGLWRVGNGKNIRA*EDKWVPHGGP-LVYRNDVAESLGVVHVSDLMVTNGGCWDR 186
+ +G VGNGK++ D W+ P L + + +L + V+DL+ CW R
Sbjct: 692 ELLAKGLRKEVGNGKSLNVWMDPWIFDIAPRLPLQRHFSVNLD-LKVNDLINFEDRCWKR 750
Query: 187 NKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEAS-S 245
+ +E L + PT +++ P+ DD + W + G YS ++GY F +
Sbjct: 751 DLLEELFY-PTDVELITKRNPVVNMDDFWVWLHAKTGEYSVKSGYWL---AFQSNKLDLI 806
Query: 246 SSARVMPPALWMK--LWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEV 303
A ++P +K +W + P+ WR LPV + RRG+ +P C C +
Sbjct: 807 QQANLLPSTNGLKEQVWSTKSSPKIKMFLWRILSAALPVADQIIRRGMSIDPRCQICGDE 866
Query: 304 EETVHHALFSCPLVRMIWFAS--PLNLRFDEEGSVHD-----FFLSFLATADTNIAGIFL 356
E+ +H LF+C + R +W S P + S+ F L + + +
Sbjct: 867 GESTNHVLFTCSMARQVWALSGVPTPEFGFQNASIFANIQFLFELKKMILVPDLVKRSWP 926
Query: 357 AVLYSVWQARNELLF------------KWKDSSIDQLLQRASSLRPSRVLLDDGPRAVQV 404
VL+ +W+ RN+L F K ++ ++ + Q S +R S ++G R +
Sbjct: 927 WVLWRLWKNRNKLFFDGITFCPLNSIVKIQEDTL-EWFQAQSQIRVSE--SEEGQRVIP- 982
Query: 405 LPSSWSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVL 445
SW P G K NI ++ + K+ G V+R+ HG V+
Sbjct: 983 FTHSWEPPPEGWVKCNIGSAWSGKKKVCGGAWVLRDEHGSVI 1024
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 187 bits (475), Expect = 1e-47
Identities = 153/547 (27%), Positives = 243/547 (43%), Gaps = 43/547 (7%)
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
++P Y MSCF L LC +I+ +++RF+W R W+ W L PK GG+GFRD
Sbjct: 694 SMPLYSMSCFKLPSALCRKIQSLLTRFWWDTKPDVRKTSWVAWSKLTNPKNAGGLGFRDI 753
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIF 130
+ N +LLAK WR++ +S+L++I Y SF+E K +PS+ W S+ I
Sbjct: 754 ERCNDSLLAKLGWRLLNSPESLLSRILLGKYCHSSSFMECKLPSQPSHGWRSIIAGREIL 813
Query: 131 QEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCWDRNKVE 190
+EG W + NG+ + D W+ PLV + VS L+ N WD NK+
Sbjct: 814 KEGLGWLITNGEKVSIWNDPWLSISKPLVPIGPALREHQDLRVSALINQNTLQWDWNKIA 873
Query: 191 FLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARV 250
+ P I +P P + D W + G Y++R+GY G AS +S +
Sbjct: 874 VIL-PNYENLIKQLPAPSSRGVDKLAWLPVKSGQYTSRSGY---------GIASVASIPI 923
Query: 251 MPPAL-WM-KLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETVH 308
W LWK +P+ W+A ++ LPV L RR + + C RC E T
Sbjct: 924 PQTQFNWQSNLWKLQTLPKIKHLMWKAAMEALPVGIQLVRRHISPSAACHRCGAPESTT- 982
Query: 309 HALFSCPLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVWQARNE 368
H F C +W +PL GS LS L A++ +
Sbjct: 983 HLFFHCEFAAQVWELAPLQETTVPPGSSMLDALSLLKK----------AIILPPTGVTSA 1032
Query: 369 LLFKWKDSSIDQLLQRASSLRPSRVLLD--DGPRAVQVLPSSWS--RPLSGVCKMN---- 420
LF W I + + ++ +R +LD A + LP + + P+S + +
Sbjct: 1033 ALFPW----ICGIYGKLGTM--TRAILDALAWQSAQRCLPKTRNVVHPISQLPVLRSGYF 1086
Query: 421 --IDAS-VTSSKEAGAGMVMRNNHGEVLAAATTELG--PVLSPVLAEALGMRWALRLATK 475
+DA+ + S AG+G V ++ AT G + S + AEA ++ AL A +
Sbjct: 1087 CFVDAAWIAQSSLAGSGWVFQSATALEKETATYSAGCRRLPSALSAEAWAIKSALLHALQ 1146
Query: 476 LGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTSNGAAD 535
LG + + +D + + + ++ + R+L +F F F+ R++N AD
Sbjct: 1147 LGRTDLMVLSDSKSVVDALT-SNISINEIYGLLMEIRALRVSFHSLCFNFISRSANAIAD 1205
Query: 536 ALARLAF 542
A A+L+F
Sbjct: 1206 ATAKLSF 1212
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 186 bits (471), Expect = 4e-47
Identities = 153/549 (27%), Positives = 239/549 (42%), Gaps = 44/549 (8%)
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
++PSY M CF L LC QI+ +++RF+W R M W+ WD L P GG+GFR+
Sbjct: 740 SMPSYAMMCFKLPASLCKQIQSVLTRFWWDSKPDKRKMAWVSWDKLTLPINEGGLGFREI 799
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYV---WLSLQKSA 127
+ AK WRI+ + S+L+++ Y SF++ PS+ W +
Sbjct: 800 E-------AKLSWRILKEPHSLLSRVLLGKYCNTSSFMDCSAS--PSFASHGWRGILAGR 850
Query: 128 WIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCWDRN 187
+ ++G W +G G +I + W+ P E+ + V DL+ + W+
Sbjct: 851 DLLRKGLGWSIGQGDSINVWTEAWLSPSSPQTPIGPPTETNKDLSVHDLICHDVKSWNVE 910
Query: 188 KV--EFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASS 245
+ + R I LPLQ D W + G Y+T+TGY+ K + AS
Sbjct: 911 AIRKHLPQYEDQIRKITINALPLQ---DSLVWLPVKSGEYTTKTGYAL--AKLNSFPASQ 965
Query: 246 SSARVMPPALWMK-LWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVE 304
W K +WK P+ W+A LPV L RR + C RC + E
Sbjct: 966 LDFN------WQKNIWKIHTSPKVKHFLWKAMKGALPVGEALSRRNIEAEVTCKRCGQTE 1019
Query: 305 ETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTNIA----GIFLAVLY 360
++H L CP + +W +P + F+ + H L A +A G+ A LY
Sbjct: 1020 SSLHLMLL-CPYAKKVWELAP--VLFNPSEATHSSVALLLVDAKRMVALPPTGLGSAPLY 1076
Query: 361 -----SVWQARNELLFKWKDSSIDQLLQRA---SSLRPSRVLLDDGPRAVQVLPSSWSRP 412
+W+ARN L+F S + L+ +A + LL P + PS P
Sbjct: 1077 PWLLWHLWKARNRLIFDNHSCSEEGLVLKAILDARAWMEAQLLIHHPSPISDYPS--PTP 1134
Query: 413 LSGVCKMNIDASVTSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRL 472
V +DA+ T+S G G +++ + + + V S ++AE L + AL
Sbjct: 1135 NLKVTSCFVDAAWTTSGYCGMGWFLQDPYKVKIKENQSSSSFVGSALMAETLAVHLALVD 1194
Query: 473 ATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTSNG 532
A G R++ + +DC EL + N G L ++ D R L +F F F+ R SN
Sbjct: 1195 ALSTGVRQLNVFSDCKELISLLN-SGKSIVELRGLLHDIRELSVSFTHLCFFFIPRLSNV 1253
Query: 533 AADALARLA 541
AD+LA+ A
Sbjct: 1254 VADSLAKSA 1262
>At3g45550 putative protein
Length = 851
Score = 180 bits (457), Expect = 2e-45
Identities = 132/440 (30%), Positives = 199/440 (45%), Gaps = 27/440 (6%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ + LA+P Y MSCF L G+ S+IE ++ F+W + R + W+ W L K GG
Sbjct: 414 LKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRLQYSKKEGG 473
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD FN ALLAK WRI+ S+ A++ KA YF S ++AK + SY W SL
Sbjct: 474 LGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKDNSIIDAKTRSQQSYGWSSLL 533
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNG--G 182
+ ++G + +G+GK IR D V P D E + + +L G
Sbjct: 534 SGIALLRKGTRYVIGDGKTIRLGIDNVVDSHPPRPLLTD--EQHNGLSLDNLFQHRGHSR 591
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGE 242
CWD K++ I I L + + D W G Y+ R+GY S +
Sbjct: 592 CWDNAKLQTFVDQSDHDYIKRIYLSTRSKTDRLIWSYNSTGDYTVRSGY-----WLSTHD 646
Query: 243 ASSSSARVMPP----ALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCP 298
S++ + P L K+W +P+ WR LP L RG+ +P CP
Sbjct: 647 PSNTIPTMAKPHGSVDLKTKIWNLPIMPKLKHFLWRILSKALPTTDRLTTRGMRIDPGCP 706
Query: 299 RCSEVEETVHHALFSCPLVRMIWFAS--PLN----LRFDEEGSVHDFFLSFLATADTNIA 352
RC E+++HALF+CP M W S PL L + E ++ + L T T+
Sbjct: 707 RCRRENESINHALFTCPFATMAWRLSDTPLYRSSILSNNIEDNISNILLLLQNTTITDSQ 766
Query: 353 GIF-LAVLYSVWQARNELLFK--WKDSSIDQLLQRASSLRPSRVLLDDGPRAVQVLP--- 406
+ +L+ +W+ARN ++F + SI + +A + GPR +
Sbjct: 767 KLIPFWLLWRIWKARNNVVFNNLRESPSITVVRAKAETNEWLNATQTQGPRRLPKRTTAA 826
Query: 407 --SSWSRPLSGVCKMNIDAS 424
++W +P K N DAS
Sbjct: 827 GNTTWVKPQMPYIKCNFDAS 846
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 178 bits (451), Expect = 8e-45
Identities = 134/500 (26%), Positives = 219/500 (43%), Gaps = 39/500 (7%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ I +A P Y M+CF L LC+++ ++ F+W + +HW+ L PK GG
Sbjct: 522 LKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFWWNSVQDKKKIHWIGAQKLMLPKFLGG 581
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
GF+D + FN ALLAK R+ T S+L++I K+ Y+ FL A KG RPSY W S+
Sbjct: 582 FGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRYYMNSDFLSATKGTRPSYAWQSIL 641
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGV-----VHVSDLMVT 179
+ G +GNG+N D W+ P ESL + + VS L+
Sbjct: 642 YGRELLVSGLKKIIGNGENTYVWMDNWIFDDKP-----RRPESLQIMVDIQLKVSQLIDP 696
Query: 180 NGGCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFS 239
W+ N + L +P I+ P+ R D F W T G Y+ ++ Y ++
Sbjct: 697 FSRNWNLNMLRDL-FPWKEIQIICQQRPMASRQDSFCWFGTNHGLYTVKSEYDLCSRQVH 755
Query: 240 EGEASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPR 299
+ + + L+ K+W ++ P+ W+ + V L RGV+ C
Sbjct: 756 KQMFKEAEEQPSLNPLFGKIWNLNSAPKIKVFLWKVLKGAVAVEDRLRTRGVLIEDGCSM 815
Query: 300 CSEVEETVHHALFSCPLVRMIWFASPL---NLRFDEE---------GSVHDFFLSFLATA 347
C E ET++H LF CPL R +W +P+ N F + G+ H+ LS
Sbjct: 816 CPEKNETLNHILFQCPLARQVWALTPMQSPNHGFGDSIFTNVNHVIGNCHNTELS----- 870
Query: 348 DTNIAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQRASS-----LRPSRVLLDDGPRAV 402
++ + +++ +W+ RN+ LF+ S ++ +A L+ ++ P
Sbjct: 871 -PHLRYVSPWIIWILWKNRNKRLFEGIGSVSLSIVGKALEDCKEWLKAHELICSKEP--- 926
Query: 403 QVLPSSWSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLA 461
+W PL K NI + + + AG V+RN G VL + + S A
Sbjct: 927 -TKDLTWIPPLMNELKCNIGIAWSKKHQMAGVSWVVRNWKGRVLLHSRCSFSQISSHFDA 985
Query: 462 EALGMRWALRLATKLGFRRI 481
+ G +A+ + F R+
Sbjct: 986 KIKGWNYAVESMDQFKFDRV 1005
>At2g15540 putative non-LTR retroelement reverse transcriptase
Length = 1225
Score = 167 bits (422), Expect = 2e-41
Identities = 137/527 (25%), Positives = 227/527 (42%), Gaps = 19/527 (3%)
Query: 28 SQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDFKAFNTALLAKNWWRIMT 87
S++ +++ F+W +HW+ WD LC P GG+GFR + FN LLAK WR++
Sbjct: 696 SKLSSVVANFWWKTREESNGIHWIAWDKLCTPFSDGGLGFRTLEEFNLVLLAKQLWRLIR 755
Query: 88 KLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIFQEGGLWRVGNGKNIRA* 147
S+L+++ + YF ++ K RPS+ W S+ + + G +G+G R
Sbjct: 756 FPNSLLSRVLRGRYFRYSDPIQIGKANRPSFGWRSIMAAKPLLLSGLRRTIGSGMLTRVW 815
Query: 148 EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCWDRNKVEFLCWPPTARAILSIPLP 207
ED W+P P ++ + ++V+DL+ W +++ L P IL I
Sbjct: 816 EDPWIPSFPPRPAKSILNIRDTHLYVNDLIDPVTKQWKLGRLQELVDPSDIPLILGIRPS 875
Query: 208 LQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPALWMKLWKADAVPQ 267
+ D F W T+ G Y+ ++GY + + S AL ++WK +
Sbjct: 876 RTYKSDDFSWSFTKSGNYTVKSGY-WAARDLSRPTCDLPFQGPSVSALQAQVWKIKTTRK 934
Query: 268 CCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPL- 326
W+ L L R + CPRC EE+++H LF CP R IW SP+
Sbjct: 935 FKHFEWQCLSGCLATNQRLFSRHIGTEKVCPRCGAEEESINHLLFLCPPSRQIWALSPIP 994
Query: 327 --NLRFDEEGSVH--DFFLSFLATAD--TNIAGIFLAVLYSVWQARNELLFKWKDSS--- 377
F + DF LS D +I IF +L+ +W++RN +F+ S
Sbjct: 995 SSEYIFPRNSLFYNFDFLLSRGKEFDIAEDIMEIFPWILWYIWKSRNRFIFENVIESPQV 1054
Query: 378 -IDQLLQRASSLRP--SRVLLDDGPRAVQVLPSSWSRPLSGVCKMNIDASVTSSKEAGAG 434
+D +Q A+ + S+ + + P QV+P++ P VC+ + + + +G G
Sbjct: 1055 ILDFAIQEANVWKQANSKEVATEYP-PPQVVPANLP-PTRNVCQFDASWHLKDTL-SGHG 1111
Query: 435 MVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGFRRIWIETDCLELYTFW 494
V+ + VL LSP+ AE + WA+ LG +D + +
Sbjct: 1112 WVL-VDQDIVLLLGLKSARKSLSPLHAEVDSLLWAMECMISLGVSDCSFASDSADFISLL 1170
Query: 495 NRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTSNGAADALARLA 541
E+ + SL+ F F+ F R N AD L++ A
Sbjct: 1171 ENP-SEWPTFVAELATFSSLVCFFPSFSIKFFSRIYNVRADCLSKKA 1216
>At1g24640 hypothetical protein
Length = 1270
Score = 160 bits (405), Expect = 2e-39
Identities = 105/376 (27%), Positives = 167/376 (43%), Gaps = 34/376 (9%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ + LA+P + MSCF L C +E ++ F+W R +HW W+ LC PK GG
Sbjct: 756 LKSVALAMPVFAMSCFKLPITTCENLESAMASFWWDSCDHSRKIHWQSWERLCLPKDSGG 815
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD ++FN ALLAK WR++ +L+++ K+ YF FL+A RPS+ W S+
Sbjct: 816 LGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDATDFLDAALSQRPSFGWRSIL 875
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGG-PLVYRNDVAESLGVVHVSDLMVTNGGC 183
+ +G RVG+G ++ D W+ G +R ++ + + V L+ G
Sbjct: 876 FGRELLSKGLQKRVGDGASLFVWIDPWIDDNGFRAPWRKNLIYDV-TLKVKALLNPRTGF 934
Query: 184 WDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEA 243
WD + L P I +I P+ + D F W + G +S ++ Y + S+
Sbjct: 935 WDEEVLHDLFLPEDILRIKAIK-PVISQADFFVWKLNKSGDFSVKSAYWLAYQTKSQNLR 993
Query: 244 SSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEV 303
S S + L ++W P+ W+ C E+
Sbjct: 994 SEVSMQPSTLGLKTQVWNLQTDPKIKIFLWKV------------------------CGEL 1029
Query: 304 EETVHHALFSCPLVRMIWFASPLNLRFD--EEGSVHDFFLSFLATADT-----NIAGIFL 356
E+ +H LF CPL R IW S D GS++ L D N+ IF
Sbjct: 1030 GESTNHTLFLCPLSRQIWALSDYPFPPDGFSNGSIYSNINHLLENKDNKEWPINLRKIFP 1089
Query: 357 AVLYSVWQARNELLFK 372
+L+ +W+ RN +F+
Sbjct: 1090 WILWRIWKNRNSFIFE 1105
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 145 bits (366), Expect = 6e-35
Identities = 143/552 (25%), Positives = 231/552 (40%), Gaps = 37/552 (6%)
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
+IP +VMS LL ++R F WG + + H L W +C+PK GGIG R
Sbjct: 662 SIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGLRSA 721
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSA--- 127
+ N AL+AK WR++ +S+ A++ + Y K ++ +P W S +S
Sbjct: 722 RDMNKALVAKVGWRLLQDKESLWARVVRKKY--KVGGVQDTSWLKPQPRWSSTWRSVAVG 779
Query: 128 --WIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCWD 185
+ +G W G+G IR D+W+ + D+ + V+ G W+
Sbjct: 780 LREVVVKGVGWVPGDGCTIRFWLDRWLLQEPLVELGTDMIPEGERIKVAADYWLPGSGWN 839
Query: 186 RNKVEFLCWPPTARAILSIPLPL-QQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEAS 244
+ R +LS+ + + D W T+DG ++ R+ YS + +G+
Sbjct: 840 LEILGLYLPETVKRRLLSVVVQVFLGNGDEISWKGTQDGAFTVRSAYSLL-----QGDVG 894
Query: 245 SSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVE 304
R + + ++WK + W +++ RR + N C C+ E
Sbjct: 895 D---RPNMGSFFNRIWKLITPERVRVFIWLVSQNVIMTNVERVRRHLSENAICSVCNGAE 951
Query: 305 ETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDFF----LSFLATADTNIAGIFLAV-- 358
ET+ H L CP + IW L LR H+FF L +L T + GI+ +
Sbjct: 952 ETILHVLRDCPAMEPIW-RRLLPLR-----RHHEFFSQSLLEWLFTNMDPVKGIWPTLFG 1005
Query: 359 --LYSVWQARNELLFKWKDSSIDQL---LQRASSLRPSRV-LLDDGPRAVQV-LPSSWSR 411
++ W+ R +F + D+L A +R V + + P V+V W
Sbjct: 1006 MGIWWAWKWRCCDVFGERKICRDRLKFIKDMAEEVRRVHVGAVGNRPNGVRVERMIRWQV 1065
Query: 412 PLSGVCKMNID-ASVTSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWAL 470
P G K+ D AS + A AG +RN GE L +G +P LAE G + L
Sbjct: 1066 PSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALNIGSCAAP-LAELWGAYYGL 1124
Query: 471 RLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTS 530
+A GFRR+ ++ DC + F + L+ +V C+ + + V R +
Sbjct: 1125 LIAWDKGFRRVELDLDCKLVVGFLSTGVSNAHPLSFLVRLCQGFFTRDWLVRVSHVYREA 1184
Query: 531 NGAADALARLAF 542
N AD LA AF
Sbjct: 1185 NRLADGLANYAF 1196
>At3g09510 putative non-LTR reverse transcriptase
Length = 484
Score = 137 bits (345), Expect = 2e-32
Identities = 127/474 (26%), Positives = 206/474 (42%), Gaps = 41/474 (8%)
Query: 98 KAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGP 157
KA YF S L+AK + SY W SL + ++G +G+G+NIR D V P
Sbjct: 2 KARYFKDVSILDAKVRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPP 61
Query: 158 LVYRNDVAESLGVVHVSDLMVTNGGC--WDRNKVEFLCWPPTARAILSIPLPLQQRDDCF 215
+ E+ + +++L G WD +K+ I I L ++ D
Sbjct: 62 RPLNTE--ETYKEMTINNLFERKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKI 119
Query: 216 FWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARVMPPA----LWMKLWKADAVPQCCET 271
W G Y+ R+GY + + S++ + PP L ++W +P+
Sbjct: 120 IWNYNTTGEYTVRSGYWLL-----THDPSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHF 174
Query: 272 AWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVRMIWFASPLNLRFD 331
WRA L L RG+ +P CPRC E+++HALF+CP M W S +L +
Sbjct: 175 LWRALSQALATTERLTTRGMRIDPSCPRCHRENESINHALFTCPFATMAWRLSDSSLIRN 234
Query: 332 EEGSVHDF------FLSFLATADTNIAGIF----LAVLYSVWQARNELLF-KWKDSSIDQ 380
+ S +DF L+F+ DT ++ + +++ +W+ARN ++F K+++S
Sbjct: 235 QLMS-NDFEENISNILNFV--QDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKT 291
Query: 381 LLQRAS------SLRPSRVLLDDGPRAVQVLPSSWSRPLSGVCKMNIDASVTSSK-EAGA 433
+L + + S R + W P + K N DA K EA
Sbjct: 292 VLSAKAETHDWLNATQSHKKTPSPTRQIAENKIEWRNPPATYVKCNFDAGFDVQKLEATG 351
Query: 434 GMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGFRRIWIETDCLELYTF 493
G ++RN++G ++ + +L +P+ AE + AL+ G+ ++++E DC L
Sbjct: 352 GWIIRNHYGTPISWGSMKLAHTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINL 411
Query: 494 WNRQGGEF-SHLATVVFDCRSLLPNFDVFNFTFVRRTSNGAADALARLAFHYGC 546
N G F S LA + D F F F+RR N A LA+ YGC
Sbjct: 412 IN--GISFHSSLANHLEDISFWANKFASIQFGFIRRKGNKLAHVLAK----YGC 459
>At1g29650 reverse transcriptase, putative
Length = 1557
Score = 135 bits (341), Expect = 5e-32
Identities = 110/465 (23%), Positives = 181/465 (38%), Gaps = 87/465 (18%)
Query: 8 INLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGF 67
+ +A+P Y +SCF L C + ++ F+W R HW+ W+ +C K GG
Sbjct: 1105 VAMAMPVYAISCFKLTKTTCENLSSAMADFWWNALEHKRKTHWVSWEKMCLSKENGGR-- 1162
Query: 68 RDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSA 127
YFP E FLEA G RPSY W S+
Sbjct: 1163 ---------------------------------YFPDEDFLEADLGARPSYAWRSVLHGR 1189
Query: 128 WIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLV-YRNDVAESLGVVHVSDLMVTNGGCWDR 186
+ +G +GNG +I D W+ GP + + + SL + V DL+ +G CW+R
Sbjct: 1190 DLLTKGLRKEIGNGNSISVWMDNWIYDNGPRIPLQKHFSVSLN-LRVKDLINLDGRCWNR 1248
Query: 187 NKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSS 246
+ ++ L + I P+ DD + W ++ G YS ++GY + + +
Sbjct: 1249 DLLQDLFFQVDIDLICK-KKPVVDMDDFWVWLHSKSGEYSVKSGYWLVFQSNKPELLFKA 1307
Query: 247 SARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEET 306
+ L K+W P+ WR LPV + RRG+ +PCC C + E+
Sbjct: 1308 CRQPSTNGLKEKIWSTSTFPKIKLFLWRILSAALPVADQILRRGMNVDPCCQICGQEGES 1367
Query: 307 VHHALFSCPLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVWQAR 366
++H LF+C + R S + G + ++FL+
Sbjct: 1368 INHVLFTCSVARQ---GSSSGKKVLALGFMEEWFLA------------------------ 1400
Query: 367 NELLFKWKDSSIDQLLQRASSLRPSRVLLDDGPRAVQVLPSSWSRPLSGVCKMNIDASVT 426
+ L + D + RA RP W P G K N+ A +
Sbjct: 1401 -QSLIRLVDPEEN---NRADPCRP-----------------RWDPPPIGWVKCNVGAVWS 1439
Query: 427 SSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWAL 470
K+ G V+R++ G+VL + G + + ++ + WA+
Sbjct: 1440 GKKKLCGGSWVVRDDRGQVLLHSRRAFGNLTTKKDSQITCVLWAI 1484
>At2g02650 putative reverse transcriptase
Length = 365
Score = 115 bits (287), Expect = 8e-26
Identities = 84/300 (28%), Positives = 131/300 (43%), Gaps = 21/300 (7%)
Query: 259 LWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETVHHALFSCPLVR 318
+WK P+ WR L + L R + +P C RC EET+HH +F+CP +
Sbjct: 37 IWKLHVAPKIKHFLWRCVTGALATNTRLRSRNIDADPICQRCCIEEETIHHIMFNCPYTQ 96
Query: 319 MIWFASPLNL--RFDEEGSVHD---FFLSFLATADTNIAGIFLA--VLYSVWQARNELLF 371
+W ++ + + ++ S D + T TN FL +++ +W++RN LF
Sbjct: 97 SVWRSANIIIGNQWGPPSSFEDNLNRLIQLSKTQTTNSLDRFLPFWIMWRLWKSRNVFLF 156
Query: 372 KWKDSSID-----------QLLQRASSLRPSRVLLDDGPRAVQVLPSS-WSRPLSGVCKM 419
+ K S D + L + + V + P SS W+ P G K
Sbjct: 157 QQKCQSPDYEARKGIQDATEWLNANETTENTNVHVATNPIQTSRRDSSQWNPPPEGWVKC 216
Query: 420 NIDASVT-SSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRLATKLGF 478
N D+ T S +G +R +G ++ +L + AEALG AL++ G
Sbjct: 217 NFDSGYTQGSPYTRSGWTIRECNGHIVLCGNAKLQSSTCSLHAEALGFLHALQVIWAHGL 276
Query: 479 RRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTSNGAADALA 538
R +W E+D L T N G + S L T+++D R + + FV R N AADALA
Sbjct: 277 RYVWFESDSKSLVTLIN-NGEDHSLLGTLIYDIRHWMLKLPYCSLEFVNRERNSAADALA 335
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.139 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,348,120
Number of Sequences: 26719
Number of extensions: 570316
Number of successful extensions: 1710
Number of sequences better than 10.0: 126
Number of HSP's better than 10.0 without gapping: 93
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 1410
Number of HSP's gapped (non-prelim): 181
length of query: 565
length of database: 11,318,596
effective HSP length: 104
effective length of query: 461
effective length of database: 8,539,820
effective search space: 3936857020
effective search space used: 3936857020
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0118c.1


