

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.10
(1082 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
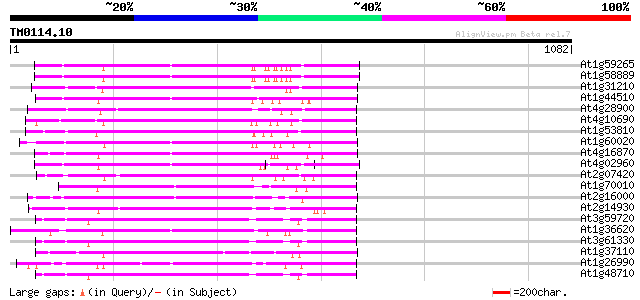
At1g59265 polyprotein, putative 396 e-110
At1g58889 polyprotein, putative 395 e-110
At1g31210 putative reverse transcriptase 368 e-101
At1g44510 polyprotein, putative 366 e-101
At4g28900 putative protein 358 9e-99
At4g10690 retrotransposon like protein 355 8e-98
At1g53810 352 5e-97
At1g60020 hypothetical protein 352 7e-97
At4g16870 retrotransposon like protein 343 3e-94
At4g02960 putative polyprotein of LTR transposon 317 3e-86
At2g07420 putative retroelement pol polyprotein 305 1e-82
At1g70010 hypothetical protein 299 7e-81
At2g16000 putative retroelement pol polyprotein 296 6e-80
At2g14930 pseudogene 291 1e-78
At3g59720 copia-type reverse transcriptase-like protein 275 1e-73
At1g36620 hypothetical protein 273 3e-73
At3g61330 copia-type polyprotein 273 4e-73
At1g37110 271 2e-72
At1g26990 polyprotein, putative 268 9e-72
At1g48710 hypothetical protein 267 2e-71
>At1g59265 polyprotein, putative
Length = 1466
Score = 396 bits (1018), Expect = e-110
Identities = 265/740 (35%), Positives = 364/740 (48%), Gaps = 121/740 (16%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S NL+ + + + V G IPI +G T+L T K +PL
Sbjct: 329 NNWLLDSGATHHITSDFNNLSLHQPYTG-GDDVMVADGSTIPISHTGSTSLST--KSRPL 385
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
L ++L+ P I KNLISV +L N VSV F P F V D TG+PL++ + +LY P
Sbjct: 386 NLHNILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWP 445
Query: 167 VTPSVPFAGLAQ-------SLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVF 218
+ S P + A S WH+RLGHP S L S+ SN ++ + + + C C+
Sbjct: 446 IASSQPVSLFASPSSKATHSSWHARLGHPAPSILNSVISNYSLSVLNPSHKFLSCSDCLI 505
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K ++PF S + P + ++SD+W+SP+LS +RYYV+F+D FT + W +PL KS
Sbjct: 506 NKSNKVPFSQSTINSTRPLEYIYSDVWSSPILSHDNYRYYVIFVDHFTRYTWLYPLKQKS 565
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV E F + + + F + DNG EF + Y + HG+ S PHT NG
Sbjct: 566 QVKETFITFKNLLENRFQTRIGTFYSDNGGEF--VALWEYFSQHGISHLTSPPHTPEHNG 623
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I TLL+HAS+P ++W +A +A YL+N +P L SP Q L+ P
Sbjct: 624 LSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLFGTSP 683
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y LRVFGC CYP + +KL +S CVFLGY L Y C L ++ ISRHV
Sbjct: 684 NYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYISRHVR 743
Query: 459 FDETRFPFA------------------------TMPT-----PPPSTYD---CFSDPIHP 486
FDE FPF+ T+PT P PS D + P P
Sbjct: 744 FDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATPPSSP 803
Query: 487 SIIHQ-----------------WTSPDPS------PTSVVPPAVT--------------- 508
S + +SP+P+ P P T
Sbjct: 804 SAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTSQNNP 863
Query: 509 ----PSQL------PTTSTSSSASP-------PSSPSPPS----PPQPVPPI-------- 539
PSQL P S+SSS SP +SP+PPS PP P+ I
Sbjct: 864 TNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPPPLAQIVNNNNQAP 923
Query: 540 ---RTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALI 596
+M TR+ GI KP ++L+V++ P+ AL D W++AM SE +A I
Sbjct: 924 LNTHSMGTRAKAGIIKPNPKYSLAVSL---AAESEPRTAIQALKDERWRNAMGSEINAQI 980
Query: 597 RNKTWDLV-PRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSP 655
N TWDLV P PS V I+ C WIF K ++G RYKARLV G +Q G+D ETFSP
Sbjct: 981 GNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRPGLDYAETFSP 1040
Query: 656 VVKPATIRTVLTIALSGLGP 675
V+K +IR VL +A+ P
Sbjct: 1041 VIKSTSIRIVLGVAVDRSWP 1060
>At1g58889 polyprotein, putative
Length = 1466
Score = 395 bits (1014), Expect = e-110
Identities = 264/740 (35%), Positives = 363/740 (48%), Gaps = 121/740 (16%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S NL+ + + + V G IPI +G T+L T K +PL
Sbjct: 329 NNWLLDSGATHHITSDFNNLSLHQPYTG-GDDVMVADGSTIPISHTGSTSLST--KSRPL 385
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
L ++L+ P I KNLISV +L N VSV F P F V D TG+PL++ + +LY P
Sbjct: 386 NLHNILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWP 445
Query: 167 VTPSVPFAGLAQ-------SLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVF 218
+ S P + A S WH+RLGHP S L S+ SN ++ + + + C C+
Sbjct: 446 IASSQPVSLFASPSSKATHSSWHARLGHPAPSILNSVISNYSLSVLNPSHKFLSCSDCLI 505
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K ++PF S + P + ++SD+W+SP+LS +RYYV+F+D FT + W +PL KS
Sbjct: 506 NKSNKVPFSQSTINSTRPLEYIYSDVWSSPILSHDNYRYYVIFVDHFTRYTWLYPLKQKS 565
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV E F + + + F + DNG EF + Y + HG+ S PHT NG
Sbjct: 566 QVKETFITFKNLLENRFQTRIGTFYSDNGGEF--VALWEYFSQHGISHLTSPPHTPEHNG 623
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I TLL+HAS+P ++W +A +A YL+N +P L SP Q L+ P
Sbjct: 624 LSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLFGTSP 683
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y LRVFGC CYP + +KL +S CVFLGY L Y C L ++ ISRHV
Sbjct: 684 NYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYISRHVR 743
Query: 459 FDETRFPFA------------------------TMPT-----PPPSTYD---CFSDPIHP 486
FDE FPF+ T+PT P PS D + P P
Sbjct: 744 FDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATPPSSP 803
Query: 487 SIIHQ-----------------WTSPDPS------PTSVVPPAVT--------------- 508
S + +SP+P+ P P T
Sbjct: 804 SAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTSQNNP 863
Query: 509 ----PSQL------PTTSTSSSASP-------PSSPSPPS----PPQPVPPI-------- 539
PSQL P S+SSS SP +SP+PPS PP P+ I
Sbjct: 864 TNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPPPLAQIVNNNNQAP 923
Query: 540 ---RTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALI 596
+M TR+ GI KP ++L+V++ P+ AL D W++AM SE +A I
Sbjct: 924 LNTHSMGTRAKAGIIKPNPKYSLAVSL---AAESEPRTAIQALKDERWRNAMGSEINAQI 980
Query: 597 RNKTWDLV-PRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSP 655
N TWDLV P PS V I+ C WIF K ++G RYKAR V G +Q G+D ETFSP
Sbjct: 981 GNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARFVAKGYNQRPGLDYAETFSP 1040
Query: 656 VVKPATIRTVLTIALSGLGP 675
V+K +IR VL +A+ P
Sbjct: 1041 VIKSTSIRIVLGVAVDRSWP 1060
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 368 bits (944), Expect = e-101
Identities = 233/659 (35%), Positives = 325/659 (48%), Gaps = 40/659 (6%)
Query: 43 TLAPPDST---WYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTL 99
TL D T W+ D+ A++HV SS L S + + + VG G +PI +G TT+
Sbjct: 311 TLRVSDDTGKEWHPDSAATAHVTSSTNGLQSATEYEG-DDAVLVGDGTYLPITHTGSTTI 369
Query: 100 PTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCN 159
+S+ PL + VL P I K+L+SV +L D V FD + D QT +
Sbjct: 370 KSSNGKIPL--NEVLVVPNIQKSLLSVSKLCDDYPCGVYFDANKVCIIDLQTQKVVTTGP 427
Query: 160 SLGDLYPVTPSVPFAGL--------AQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPT 211
LY V + F L + +WH RLGH S ALQ L+++K I +
Sbjct: 428 RRNGLY-VLENQEFVALYSNRQCAATEEVWHHRLGHANSKALQHLQNSKAIQINKSRTSP 486
Query: 212 VCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRYYVLFLDDFTDFLW 270
VCE C GK RLPF+ S++ + P D +H DLW SPV+S+ G +YY +F+DD++ + W
Sbjct: 487 VCEPCQMGKSSRLPFLISDSRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSW 546
Query: 271 TFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSC 330
+PL NKS+ +F S + + +K Q D G EF + + + HG+ R SC
Sbjct: 547 FYPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGIHHRISC 606
Query: 331 PHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPT 390
P+T QNG AERK R + + ++L H+ P FW + A Y++N +P L NLSP
Sbjct: 607 PYTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVLKNLSPY 666
Query: 391 QLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRK 450
+ L+ P Y+ LRVFG CYP + NK PRS CVFLGY ++GY+CF K
Sbjct: 667 EALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGK 726
Query: 451 IIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPS 510
+ ISR+VIF+E+ PF S +S P+ + H S P + V P
Sbjct: 727 VYISRNVIFNESELPF---KEKYQSLVPQYSTPLLQAWQHNKISEISVPAAPVQLFSKPI 783
Query: 511 QLPTTSTSSSASPPSSPSPPS----PPQPVPPI--------------RTMATRSMRGIYK 552
L T + S + P P S + V P+ M TRS GI K
Sbjct: 784 DLNTYAGSQVTEQLTDPEPTSNNEGSDEEVNPVAEEIAANQEQVINSHAMTTRSKAGIQK 843
Query: 553 PRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNI 612
P + L I + PK A+ P W A+ E + + TW LVP D+NI
Sbjct: 844 PNTRYAL---ITSRMNTAEPKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNI 900
Query: 613 IRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
+ W+F K +G ++ KARLV G Q GVD ETFSPVV+ ATIR VL ++ S
Sbjct: 901 LSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTS 959
>At1g44510 polyprotein, putative
Length = 1459
Score = 366 bits (939), Expect = e-101
Identities = 250/719 (34%), Positives = 348/719 (47%), Gaps = 106/719 (14%)
Query: 51 WYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTL 110
W +D+GA+ H+ S L+ + + + + + G G+ I+ +G T LP+ +++ L L
Sbjct: 337 WLLDSGATHHITSDLNALSLHQPYNG-GEYVMIADGTGLTIKQTGSTFLPS--QNRDLAL 393
Query: 111 SHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--PVT 168
VL+ P I KNLISV +L N VSV F P F V D TG L++ + DLY PVT
Sbjct: 394 HKVLYVPDIRKNLISVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDDLYEWPVT 453
Query: 169 P-------SVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNS-PTVCESCVFGK 220
+ P S WHSRLGHP +S L +L S + +S T C C+ K
Sbjct: 454 NPPATALFTSPSPKTTLSSWHSRLGHPSASILNTLLSKFSLPVSVASSNKTSCSDCLINK 513
Query: 221 HVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQV 280
+LPF +S+ + P + + +D+WTSP++S ++YY++ +D +T + W +PL KSQV
Sbjct: 514 SHKLPFATSSIHSSSPLEYIFTDVWTSPIISHDNYKYYLVLVDHYTRYTWLYPLQQKSQV 573
Query: 281 FEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKA 340
F + + F ++ L DNG EF + + ++G+ S PHT NG +
Sbjct: 574 KATFIAFKALVENRFQAKIRTLYSDNGGEF--IALRDFLVSNGISHLTSPPHTPEHNGLS 631
Query: 341 ERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSY 400
ERK R I TLL ASVP +W +A A YL+N +P L SP Q L+ P+Y
Sbjct: 632 ERKHRHIVETGLTLLTQASVPREYWTYAFATAVYLINRMPTPVLCLQSPFQKLFGSSPNY 691
Query: 401 THLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFD 460
LRVFGCLC+P + T NKL+ RS CVFLGY L Y C D+ + ++ SRHV+FD
Sbjct: 692 QRLRVFGCLCFPWLRPYTRNKLEERSKRCVFLGYSLTQTAYLCLDVDNNRLYTSRHVMFD 751
Query: 461 ETRFPFA---------TMPTPPPSTYDCFSDPIHP----SIIHQWTSPDPSPTSVVPP-- 505
E+ +PFA ++ TPP S+ S P + S++ + P SP + PP
Sbjct: 752 ESTYPFAASIREQSQSSLVTPPESSSS--SSPANSGFPCSVLRLQSPPASSPETPSPPQQ 809
Query: 506 ----AVTPSQL--PTTSTSS-------SASPPSSPSPPSPPQ---PVPPIRTMATRSMRG 549
V+P Q PT S S S SP S S P+ P P P ++ G
Sbjct: 810 QNDSPVSPRQTGSPTPSHHSQVRDSTLSPSPSVSNSEPTAPHENGPEPEAQSNPNSPFIG 869
Query: 550 -IYKPRKLFNLSVTI-----DDPTISPLPKN----------------------------- 574
+ P N S +I D T + LP N
Sbjct: 870 PLPNPNPETNPSSSIEQRPVDKSTTTALPPNQTTIAATSNSRSQPPKNNHQMKTRSKNNI 929
Query: 575 --PKL---------------------ALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVN 611
PK AL D W+ AM EFDA RN TWDLVP +
Sbjct: 930 TKPKTKTSLTVALTQPHLSEPNTVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPPNPTQH 989
Query: 612 IIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIAL 670
++ C W+F K NG ++YKARLV G +Q GVD ETFSPV+K TIR VL +A+
Sbjct: 990 LVGCRWVFKLKYLPNGLIDKYKARLVAKGFNQQYGVDYAETFSPVIKATTIRVVLDVAV 1048
>At4g28900 putative protein
Length = 1415
Score = 358 bits (919), Expect = 9e-99
Identities = 243/722 (33%), Positives = 344/722 (46%), Gaps = 107/722 (14%)
Query: 34 DIATAMHTMTLAPPDST-WYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQ 92
D + A M ++ S W D+GA+SH+ +S L S S + VG+ +PI
Sbjct: 275 DFSKAFAAMRVSDQKSNPWVTDSGATSHITNSTSQLQSAQPYSG-EDSVIVGNSDFLPIT 333
Query: 93 GSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTG 152
G L ++ + PL VL P I K+L+SV +LT+D + FD G V D T
Sbjct: 334 HIGSAVLTSNQGNLPLR--DVLVCPNITKSLLSVSKLTSDYPCVIEFDSDGVIVKDKLTK 391
Query: 153 IPLMRCNSLGDLYPVTPSVPFA-------GLAQSLWHSRLGHPGSSALQSLRSNKFITYE 205
L + DLY + A + +WH RLGHP LQ L NK I
Sbjct: 392 QLLTKGTRHNDLYLLENPKFMACYSSRQQATSDEVWHMRLGHPNQDVLQQLLRNKAIVIS 451
Query: 206 HLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRYYVLFLDD 264
S ++C++C GK +LPF SS+ V+ + +H DLW +PV+SS G RYYV+F+D+
Sbjct: 452 K-TSHSLCDACQMGKICKLPFASSDFVSSRLLERVHCDLWGPAPVVSSQGFRYYVIFIDN 510
Query: 265 FTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGL 324
++ F W +PL KS F +F + + + QCD G EF + F ++ A G+
Sbjct: 511 YSRFTWFYPLRLKSDFFSVFLTFQKMVENQCQQKIASFQCDGGGEFISNQFVSHLAECGI 570
Query: 325 IFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSL 384
SCP+T QNG AERK R I + +++ VP W A + +L N++P L
Sbjct: 571 RQLISCPYTPQQNGIAERKHRHITELGSSMMFQGKVPQFLWVEAFYTSNFLCNLLPSSVL 630
Query: 385 SN-LSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKC 443
+ SP ++L + P YT LRVFGC CYP + NK P+S CVF GY ++GYKC
Sbjct: 631 KDQKSPYEVLMGKAPVYTSLRVFGCACYPNLRPYASNKFDPKSLLCVFTGYNEKYKGYKC 690
Query: 444 FDLSHRKIIISRHVIFDETRFPFATMPTPPPSTYDCFSDPI---HPSIIHQWTSPDPSPT 500
F KI I+RHV+FDE++F F+ D +SD + + +++ W S + P
Sbjct: 691 FHPPTGKIYINRHVLFDESKFLFS----------DIYSDKVSGTNSTLVSAWQS-NFLPK 739
Query: 501 SVVPPAVTPSQLPTTSTSSSASP-----------------------------PSSP-SPP 530
S+ PA TP L ++T++S S PSSP +
Sbjct: 740 SI--PA-TPEVLDISNTAASFSDEQGEFSGAVGGGGCGCTADLDSVPIGNSLPSSPVTQQ 796
Query: 531 SPPQPVPPIRT-------------------------------------------MATRSM 547
+ PQP PI + M TRS
Sbjct: 797 NSPQPETPISSAGSGNDAEDSELSENSENSESSVFSEATTETEAADNTNDQSHPMITRSK 856
Query: 548 RGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRP 607
GI+KP + + + P+PK K AL DP W AM E+D+ TWDLVP
Sbjct: 857 SGIFKPNPKYAMFTVKSN---YPVPKTVKTALKDPGWTDAMGEEYDSFEETHTWDLVPPD 913
Query: 608 SDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLT 667
S + + C W+F K KA+G +R KARLV G Q GVD ET+SPVV+ AT+RT+L
Sbjct: 914 SFITPLGCRWVFKTKLKADGTLDRLKARLVAKGYEQEEGVDYMETYSPVVRTATVRTILH 973
Query: 668 IA 669
+A
Sbjct: 974 VA 975
>At4g10690 retrotransposon like protein
Length = 1515
Score = 355 bits (911), Expect = 8e-98
Identities = 242/726 (33%), Positives = 345/726 (47%), Gaps = 95/726 (13%)
Query: 31 VPTDIATAMHTMTLAPPDST----WYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSG 86
+P D+ A M ++ + W D+ A++H+ ++ L + S + + VG+G
Sbjct: 300 LPEDLPNAFAAMRVSDQNQASSHEWLPDSAATAHITNTTDGLQNSQTYSG-DDSVIVGNG 358
Query: 87 QGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSV 146
+PI G T+P + L L VL P I K+L+SV +LT D S FD +
Sbjct: 359 DFLPITHIG--TIPLNISQGTLPLEDVLVCPGITKSLLSVSKLTDDYPCSFTFDSDSVVI 416
Query: 147 HDFQTGIPLMRCNSLGDLYPVTPSVPFAGLAQS--------LWHSRLGHPGSSALQSLRS 198
D +T L + N LY V VPF + +WH RLGHP LQ L
Sbjct: 417 KDKRTQQLLTQGNKHKGLY-VLKDVPFQTYYSTRQQSSDDEVWHQRLGHPNKEVLQHLIK 475
Query: 199 NKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRY 257
K I +S +CE+C GK RLPFV+S V+ P + +H DLW +PV S+ G +Y
Sbjct: 476 TKAIVVNKTSS-NMCEACQMGKVCRLPFVASEFVSSRPLERIHCDLWGPAPVTSAQGFQY 534
Query: 258 YVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHA 317
YV+F+D+++ F W +PL KS F +F + + H + QCD G EF + F A
Sbjct: 535 YVIFIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKIAMFQCDGGGEFVSYKFVA 594
Query: 318 YCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLN 377
+ A+ G+ SCPHT QNG AER+ R + + +L+ H+ VP W A + +L N
Sbjct: 595 HLASCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHSKVPHKLWVEAFFTSNFLSN 654
Query: 378 IIPRKSLS-NLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPL 436
++P +LS N SP ++L+ P YT LRVFG CYP + NK P+S CVFLGY
Sbjct: 655 LLPSSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYAKNKFDPKSLLCVFLGYNN 714
Query: 437 HHRGYKCFDLSHRKIIISRHVIFDETRFP-------FATMPTPP---------------- 473
++GY+C K+ I RHV+FDE +FP F T+ P
Sbjct: 715 KYKGYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTISGSPLFTAWQKGFSSTALSR 774
Query: 474 ------------PSTYDCFSDP--IHPSIIHQWTSPDPSPTS----VVPPA-VTPSQLPT 514
PS S P P+I T+PD + VVPP+ +T + LPT
Sbjct: 775 ETPSTNVEDIIFPSATVSSSVPTGCAPNIAETATAPDVDVAAAHDMVVPPSPITSTSLPT 834
Query: 515 -----------------TSTSSSASPPSSPSPPSPPQPVPPIRT--------------MA 543
T+ SS+ +P S PP+++ M
Sbjct: 835 QPEESTSDQNHYSTDSETAISSAMTPQSINVSLFEDSDFPPLQSVISSTTAAPETSHPMI 894
Query: 544 TRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDL 603
TR+ GI KP + L + P PK+ K AL D W +AM E + TWDL
Sbjct: 895 TRAKSGITKPNPKYAL---FSVKSNYPEPKSVKEALKDEGWTNAMGEEMGTMHETDTWDL 951
Query: 604 VPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIR 663
VP ++ C W+F K ++G +R KARLV G Q GVD ET+SPVV+ AT+R
Sbjct: 952 VPPEMVDRLLGCKWVFKTKLNSDGSLDRLKARLVARGYEQEEGVDYVETYSPVVRSATVR 1011
Query: 664 TVLTIA 669
++L +A
Sbjct: 1012 SILHVA 1017
>At1g53810
Length = 1522
Score = 352 bits (904), Expect = 5e-97
Identities = 236/714 (33%), Positives = 328/714 (45%), Gaps = 81/714 (11%)
Query: 31 VPTDIATAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIP 90
+P +AT T W D+ AS+HV ++ ++ S H + + V G +P
Sbjct: 306 LPMALATMRITDVTDHHGHEWIPDSAASAHVTNNR-HVLQQSQPYHGSDSIMVADGNFLP 364
Query: 91 IQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQ 150
I +G ++ +S PL VL P IVK+L+SV +LT+D SV FD ++D
Sbjct: 365 ITHTGSGSIASSSGKIPL--KEVLVCPDIVKSLLSVSKLTSDYPCSVEFDADSVRINDKA 422
Query: 151 TGIPLMRCNSLGDLYP-------VTPSVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFIT 203
T L+ + LY V S + +WH RLGH + L L S+K I
Sbjct: 423 TKKLLVMGRNRDGLYSLEEPKLQVLYSTRQNSASSEVWHRRLGHANAEVLHQLASSKSII 482
Query: 204 YEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRYYVLFL 262
+ TVCE+C GK RLPF+ S P + +H DLW SP S G RYYV+F+
Sbjct: 483 IINKVVKTVCEACHLGKSTRLPFMLSTFNASRPLERIHCDLWGPSPTSSVQGFRYYVVFI 542
Query: 263 DDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAH 322
D ++ F W +PL KS F F + H +K QCD G EF ++ F + H
Sbjct: 543 DHYSRFTWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFLKHLQDH 602
Query: 323 GLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRK 382
G+ SCP+T QNG AERK R I + +++ + +P +W + A +++N++P
Sbjct: 603 GIQQNMSCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVINLLPTS 662
Query: 383 SL-SNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGY 441
SL +N SP Q LY + P Y+ LRVFGC CYP + K PRS CVFLGY ++GY
Sbjct: 663 SLDNNESPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRSLKCVFLGYNEKYKGY 722
Query: 442 KCFDLSHRKIIISRHVIFDETRFPFATM------------------------PT------ 471
+C +I ISRHV+FDE PF ++ PT
Sbjct: 723 RCLYPPTGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVTPTQPDQSR 782
Query: 472 ------PPPSTYDCFSDPIHPSI------IHQWTSPDPSPTSVV---------------- 503
P P T D + P + TS D SVV
Sbjct: 783 YPVSSIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISVVSGSPERTTGLDSASIG 842
Query: 504 ----PPAVTPSQLPTTSTSSSASPPSSPSPPSPPQ----PVPPIRTMATRSMRGIYKPRK 555
P S +S ++SP SP +P Q PV M TR GI KP K
Sbjct: 843 DSYHSPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTNEHAMVTRGKEGISKPNK 902
Query: 556 LFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRC 615
+ V + P PK AL P W +AMQ E +TW LVP ++N++
Sbjct: 903 RY---VLLTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVLGS 959
Query: 616 MWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
MW+F K A+G ++ KARLV G Q G+D ET+SPVV+ T+R +L +A
Sbjct: 960 MWVFRTKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVA 1013
>At1g60020 hypothetical protein
Length = 1194
Score = 352 bits (903), Expect = 7e-97
Identities = 238/747 (31%), Positives = 352/747 (46%), Gaps = 115/747 (15%)
Query: 20 RPQAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSYSNLSHLNQ 79
+P+A+AA SP + W +D+GA+ H+ S NL+ + + +
Sbjct: 272 QPRANAAIASPYNAN---------------NWLLDSGATHHITSDLNNLSLHQPYTG-GE 315
Query: 80 KLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCF 139
+ + G G+ I +G + T + L L+ VL+ P I KNLISV ++ N VSV F
Sbjct: 316 DVTIADGSGLSISHTGSALISTPSRS--LALTDVLYVPNIHKNLISVYRMCNANKVSVEF 373
Query: 140 DPYGFSVHDFQTGIPLMRCNSLGDLY--PVTPSVPFAGLAQSL-------WHSRLGHPGS 190
P F V D +TG+ L++ + +LY PV P P + + WHSRLGHP
Sbjct: 374 FPAHFQVKDLKTGVQLLQGRTKDELYEWPVNPPKPSSHFTTTTPKTDLTSWHSRLGHPSL 433
Query: 191 SALQSLRSNKFITYEH-LNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPV 249
S L+ + S + + L C C+ K +LPF ++ + P + L+ DLWTSP+
Sbjct: 434 STLKVVVSQFSLPVSNSLQKQFNCSDCLLNKTHKLPFHTNTITSTQPLEYLYIDLWTSPI 493
Query: 250 LSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPE 309
+S +YY++ +D +T + W +P+ KS V ++F + + F + L DNG E
Sbjct: 494 VSIDNFKYYLVIVDHYTRYSWFYPIKQKSHVKDVFMTFKALVANKFQRKIIHLYSDNGGE 553
Query: 310 FDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHAL 369
F + ++ +++G+ + PHT NG +ERK R I TLL AS+P S+W +A
Sbjct: 554 F--IALRSFLSSNGITHLTTPPHTPEHNGISERKHRHIVETGLTLLGQASMPKSYWSYAF 611
Query: 370 QMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPC 429
+A YL+N + + +SP + L+ + P+Y LRVFGCLC+P + T +KL R +PC
Sbjct: 612 TIAIYLINRMSSDVIGGISPYKRLFGQAPNYLKLRVFGCLCFPWLRPYTTHKLDDRPAPC 671
Query: 430 VFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPF--------------------ATM 469
VFLGY Y C + + ++ SRHV F E +PF T+
Sbjct: 672 VFLGYSQTQSAYLCLNRTTGRVYTSRHVQFVENTYPFTKPTLDPFTNLEESNNHSITTTV 731
Query: 470 PTPP----PSTYDCFSDPIHPSIIHQWTSPDP-SPTSVVPPAVT---------------- 508
P+PP PS DP P SP P SP S+ P +T
Sbjct: 732 PSPPFVQLPSVPPPTRDPHQPPPSQPAPSPSPLSPPSMSSPVMTSSPQFSSNRDSTTLHG 791
Query: 509 -----------------PSQLPTTSTSSS-------------ASPPSSP-SPPSPPQPVP 537
P PTTS S S +PP+SP S S P P+P
Sbjct: 792 DYSHVDYGLSSPSNPPGPITSPTTSKSPSEPTSSPSHSNQPNKTPPNSPSSSSSSPTPIP 851
Query: 538 PIRTMATRS------MRGIYKPRKLFNLSVTIDDPTISPLPKNPKL-------ALSDPNW 584
++ S + + R N++ I T++ PK AL DPNW
Sbjct: 852 SPSPQSSNSPPPPPQNQHSMRTRAKNNITKPIKKLTLAATPKGKSKIPTTVAEALRDPNW 911
Query: 585 KSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQI 644
++AM EF+A +RN T+DLVP N + WIF K +G RYKAR + G Q
Sbjct: 912 RNAMSEEFNAGLRNSTYDLVPPKPHQNFVGTRWIFTIKYNPDGSINRYKARFLAKGFHQQ 971
Query: 645 AGVDCDETFSPVVKPATIRTVLTIALS 671
G+D TFSPV+K T++TVL IA+S
Sbjct: 972 HGLDYSNTFSPVIKSTTVQTVLDIAVS 998
>At4g16870 retrotransposon like protein
Length = 1474
Score = 343 bits (880), Expect = 3e-94
Identities = 241/740 (32%), Positives = 344/740 (45%), Gaps = 125/740 (16%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S L + + + + G + I +G T LP++ + L
Sbjct: 331 NNWLLDSGATHHITSDLNALALHQPYN--GDDVMIADGTSLKITKTGSTFLPSNARD--L 386
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
TL+ VL+ P I KNL+SV +L N VSV F P F V D TG L++ + +LY P
Sbjct: 387 TLNKVLYVPDIQKNLVSVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDELYEWP 446
Query: 167 VTP-------SVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYE-HLNSPTVCESCVF 218
VT + P S WHSRLGHP SS L +L S + ++ C C
Sbjct: 447 VTNPKATALFTTPSPKTTLSSWHSRLGHPSSSILNTLISKFSLPVSVSASNKLACSDCFI 506
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K +LPF S+ + P + + SD+W SP+LS ++YY++ +D T + W +PL KS
Sbjct: 507 NKSHKLPFSISSIKSTSPLEYIFSDVWMSPILSPDNYKYYLVLVDHHTRYTWLYPLQQKS 566
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV F + + F ++ L DNG EF + + ++G+ S PHT NG
Sbjct: 567 QVKSTFIAFKALVENRFQAKIRTLYSDNGGEF--IALREFLVSNGISHLTSPPHTPEHNG 624
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I TLL ASVP +W +A A YL+N +P LS SP Q L+ P
Sbjct: 625 LSERKHRHIVETGLTLLTQASVPREYWPYAFAAAVYLINRMPTPVLSMESPFQKLFGSKP 684
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y LRVFGCLC+P + T NKL+ RS CVFLGY L Y CFD+ H+++ SRHV+
Sbjct: 685 NYERLRVFGCLCFPWLRPYTHNKLEERSRRCVFLGYSLTQTAYLCFDVEHKRLYTSRHVV 744
Query: 459 FDETRFPFATMPTP---PPSTYDCFSDPIHPSIIHQWT---SPDPSPTSVV----PPAVT 508
FDE FPF+ + + P T++ S P+ I+ + S SP +V+ PP T
Sbjct: 745 FDEASFPFSNLTSQNSLPTVTFEQSSSPLVTPILSSSSVLPSCLSSPCTVLHQQQPPVTT 804
Query: 509 P----SQLPTT----------------------------STSSSASPPSSPSP--PSPPQ 534
P S PTT S+SS S P++P+ P P
Sbjct: 805 PNSPHSSQPTTSPAPLSPHRSTTMDFQVPQVRSSSPLLSSSSSLNSEPTAPNENGPEPEA 864
Query: 535 PVPPIRTMATRSMRGIYKPRKLFNLSVTID-DPTISPLPK-------------------- 573
PPI ++ + P N + T + +PT +P PK
Sbjct: 865 QSPPIGPLSNPTHEAFIGPLPNPNRNPTNEIEPTPAPHPKPVKPTTTTTTPNRTTVSDAS 924
Query: 574 ------------------------NPKLALSD--PNWKSAMQSEFDALIRNKTW------ 601
N K +L+ PN + + +++K W
Sbjct: 925 HQPTAPQQNQHNMKTRAKNNIKKPNTKFSLTATLPNRSPSEPTNVTQALKDKKWRFAMSD 984
Query: 602 -----------DLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCD 650
DLVP S + ++ C W+F K NG ++YKARLV G +Q GVD
Sbjct: 985 EFDAQQRNHTWDLVPHESQL-LVGCKWVFKLKYLPNGAIDKYKARLVAKGFNQQYGVDYA 1043
Query: 651 ETFSPVVKPATIRTVLTIAL 670
ETFSPV+K TIR VL +A+
Sbjct: 1044 ETFSPVIKSTTIRLVLDVAV 1063
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 317 bits (811), Expect = 3e-86
Identities = 202/579 (34%), Positives = 291/579 (49%), Gaps = 45/579 (7%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S NL+ + + + + G IPI +G +LPTS + L
Sbjct: 308 NNWLLDSGATHHITSDFNNLSFHQPYTG-GDDVMIADGSTIPITHTGSASLPTS--SRSL 364
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
L+ VL+ P I KNLISV +L N VSV F P F V D TG+PL++ + +LY P
Sbjct: 365 DLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWP 424
Query: 167 VTPS-------VPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVF 218
+ S P + S WHSRLGHP + L S+ SN + + + + C C
Sbjct: 425 IASSQAVSMFASPCSKATHSSWHSRLGHPSLAILNSVISNHSLPVLNPSHKLLSCSDCFI 484
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K ++PF +S + P + ++SD+W+SP+LS +RYYV+F+D FT + W +PL KS
Sbjct: 485 NKSHKVPFSNSTITSSKPLEYIYSDVWSSPILSIDNYRYYVIFVDHFTRYTWLYPLKQKS 544
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV + F + F + L DNG EF Y + HG+ S PHT NG
Sbjct: 545 QVKDTFIIFKSLVENRFQTRIGTLYSDNGGEF--VVLRDYLSQHGISHFTSPPHTPEHNG 602
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I M TLL+HASVP ++W +A +A YL+N +P L SP Q L+ + P
Sbjct: 603 LSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPP 662
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y L+VFGC CYP + +KL+ +S C F+GY L Y C + ++ SRHV
Sbjct: 663 NYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQ 722
Query: 459 FDETRFPFATMPTPPPSTYDCFSD-----------PIHPSII--------HQWTSPDP-- 497
FDE FPF+T ++ + SD P P ++ H TSP P
Sbjct: 723 FDERCFPFSTTNFGVSTSQEQRSDSAPNWPSHTTLPTTPLVLPAPPCLGPHLDTSPRPPS 782
Query: 498 SPTSVVPPAVTPSQLPTTSTSS-SASPPSSPSPPSPPQPVPPIRTMATRSMRGIY----- 551
SP+ + V+ S LP++S SS S+S P++PS P P +T + S I
Sbjct: 783 SPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSNSNSPILNNPNP 842
Query: 552 ---KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSA 587
P S P SP P ++S+PN S+
Sbjct: 843 NSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSS 881
Score = 123 bits (308), Expect = 6e-28
Identities = 75/189 (39%), Positives = 107/189 (55%), Gaps = 11/189 (5%)
Query: 494 SPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPP----QPVPPIRT--MATRSM 547
SP SP + P+ + S+ + S+SS+++PP P P+PP P+ T MATR+
Sbjct: 859 SPISSP-HIPTPSTSISEPNSPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAK 917
Query: 548 RGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRP 607
GI KP + ++ + ++ + P+ A+ D W+ AM SE +A I N TWDLVP P
Sbjct: 918 DGIRKPNQKYSYATSL---AANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPP 974
Query: 608 S-DVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVL 666
V I+ C WIF K ++G RYKARLV G +Q G+D ETFSPV+K +IR VL
Sbjct: 975 PPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVL 1034
Query: 667 TIALSGLGP 675
+A+ P
Sbjct: 1035 GVAVDRSWP 1043
>At2g07420 putative retroelement pol polyprotein
Length = 1664
Score = 305 bits (780), Expect = 1e-82
Identities = 209/692 (30%), Positives = 325/692 (46%), Gaps = 94/692 (13%)
Query: 53 MDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSH 112
+D+GAS H+ S + SN+ + + +G IP++G GD L S
Sbjct: 315 IDSGASHHMISDSKLI---SNIEPALGNVVIANGDRIPVKGVGDLDLFDKS-------SK 364
Query: 113 VLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVT---P 169
+ P NL+SV++ TTD N F P D +T L + + LY + P
Sbjct: 365 AFYMPTFTSNLLSVKKATTDLNCYAIFGPNEVHFQDIETSRVLGQGVTKDGLYVLEDTKP 424
Query: 170 SVPFAGLAQSL--------WHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKH 221
SVP + S+ WH+RLGHP S AL+ L + + CE+C+ GKH
Sbjct: 425 SVPLSSHFSSILGNANSESWHARLGHPHSRALKLLLPSTSFKNDE------CEACILGKH 478
Query: 222 VRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVF 281
+ F S+ + FD++HSD+WTSP LS H+Y+V F+D+ + F W L +K +V
Sbjct: 479 CKSVFPKSSTIYEKCFDLIHSDVWTSPCLSRENHKYFVTFIDEKSKFTWFTLLPSKDRVL 538
Query: 282 EMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAE 341
E FT+ + H+ +K L+ DN E+ + +F + HG+I + SCP+T QNG AE
Sbjct: 539 EAFTNFQTYVTNHYDAKIKILRSDNRGEYTSHAFKQHLNKHGIIHQTSCPYTPQQNGVAE 598
Query: 342 RKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYT 401
RK R + + R ++ H +VP FW + A YL+N P K L + SP ++L + P
Sbjct: 599 RKNRHLMEVRRVMMFHTNVPKHFWIDGVVSACYLINQTPTKILLDSSPFEVLNKVKPFIN 658
Query: 402 HLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDE 461
HLRVFGC+C+ L+ NKLQP+S+ +F+GY ++ +GYKC+ L RK++ISR V F E
Sbjct: 659 HLRVFGCVCFVLISGEQRNKLQPKSTKGMFIGYSINQKGYKCYVLETRKVLISRDVKFLE 718
Query: 462 TRFPF-----------ATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPS-PTSVVPPAVTP 509
++ + P+ + + + S I T+P S P ++ P
Sbjct: 719 SKSYYDKKNWEDIQDLTDSPSDRATNLRIILERLGVSNIQTQTTPRTSNPETITQPENME 778
Query: 510 SQ-------------------LPTTSTSSSASPPSS-------------PSPPSPPQPVP 537
+ L T +S +S S +P
Sbjct: 779 EEEEEEEEEEEKQGKEQELITLEETESSKVQEKDTSLLNDDNGHTNNQEEDSNSREEPRI 838
Query: 538 PIRTMATRSMRGIYKPRKLFNLSVTIDDP-----TISPLPKNPKLALS--DPNW------ 584
P R+ + R Y + F+ ++ P T++ LP+ ++ D +W
Sbjct: 839 PRRSEHLKDKRVYYNNQVYFD--NVVEHPIQVVCTLAHLPEEHQVFFGKVDQHWIPQTYE 896
Query: 585 --------KSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARL 636
+ A+ +E A+ N TWD P ++ W+F K K++G ERYKARL
Sbjct: 897 EAITHQVWRDAIAAEKQAMENNHTWDEDELPRGKKVVTSKWVFAIKYKSDGEIERYKARL 956
Query: 637 VGDGRSQIAGVDCDETFSPVVKPATIRTVLTI 668
V G +Q G D +TF+PV K T+R VL++
Sbjct: 957 VARGFTQTYGEDYLDTFAPVAKLHTVRVVLSL 988
>At1g70010 hypothetical protein
Length = 1315
Score = 299 bits (765), Expect = 7e-81
Identities = 199/615 (32%), Positives = 287/615 (46%), Gaps = 61/615 (9%)
Query: 94 SGDTTLPTS---HKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQ 150
S T P S H + L L+ VL PQ NL+SV LT + FD + D
Sbjct: 306 SSSTVCPISGSVHLGRHLILNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDAT 365
Query: 151 TGIPLMRCNSLGDLYPVT----------PSVPFAGL-AQSLWHSRLGHPGSSALQSLRSN 199
+ + + +LY V S+ A + + LWH RLGHP LQ + S
Sbjct: 366 RELMVGMGKQVANLYIVDLDSLSHPGTDSSITVASVTSHDLWHKRLGHPSVQKLQPMSSL 425
Query: 200 KFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTS-PVLSSAGHRYY 258
+ N+ C C K LPFVS NN + PFD++H D W V + G+RY+
Sbjct: 426 LSFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRYF 485
Query: 259 VLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAY 318
+ +DD++ W + L NKS V + + + F +K ++ DN PE + T F+
Sbjct: 486 LTIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELNFTQFYH- 544
Query: 319 CAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNI 378
+ G++ SCP T QN ERK + I N+ R+L + +P S+W + A YL+N
Sbjct: 545 --SKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINR 602
Query: 379 IPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHH 438
+P L + P ++L + P+Y H++VFGCLCY +K PR+ C F+GYP
Sbjct: 603 LPAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSGF 662
Query: 439 RGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPS 498
+GYK DL II+SRHV+F E FPF S Q PD +
Sbjct: 663 KGYKLLDLETHSIIVSRHVVFHEELFPFLGSDL---------------SQEEQNFFPDLN 707
Query: 499 PTSVVPPAVTPSQLPTTSTSSSASPPSSPSP-PSPPQPVPPIRTMATRSMRGIY------ 551
PT PP S + SS+S PS P+ P P ++T ++ + Y
Sbjct: 708 PT---PPMQRQSSDHVNPSDSSSSVEILPSANPTNNVPEPSVQTSHRKAKKPAYLQDYYC 764
Query: 552 ---------KPRKLFNLSVTIDDPTISPL--------PKNPKLALSDPNWKSAMQSEFDA 594
+ RK + I+DP ++ L P N A W+ AM +EFD
Sbjct: 765 HSVVSSTPHEIRKFLSYD-RINDPYLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDF 823
Query: 595 LIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFS 654
L TW++ P+D I C WIF K ++G ERYKARLV G +Q G+D +ETFS
Sbjct: 824 LEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFS 883
Query: 655 PVVKPATIRTVLTIA 669
PV K +++ +L +A
Sbjct: 884 PVAKLNSVKLLLGVA 898
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 296 bits (757), Expect = 6e-80
Identities = 209/653 (32%), Positives = 306/653 (46%), Gaps = 48/653 (7%)
Query: 35 IATAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSY--SNLSHLNQKLYVGSGQGIPIQ 92
+ A HT++ A TW +D+GA+ HV+ +S S LS +N + +G + I
Sbjct: 419 LTVARHTLSSA----TWVIDSGATHHVSHDRSLFSSLDTSVLSAVN----LPTGPTVKIS 470
Query: 93 GSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTG 152
G G L + + L +VL P+ NLIS+ LT D V FD + D G
Sbjct: 471 GVGTLKL-----NDDILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKG 525
Query: 153 IPLMRCNSLGDLYPVT---PSVPFAGLAQ-SLWHSRLGHPGSSALQSLRSNKFITYEHLN 208
L + + +LY + S+ + S+WH RLGH L ++ + T
Sbjct: 526 RMLGQGRRVANLYLLDVGDQSISVNAVVDISMWHRRLGHASLQRLDAISDSLGTTRHKNK 585
Query: 209 SPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTS-PVLSSAGHRYYVLFLDDFTD 267
C C K +L F +SN V FD+LH D+W V + G++Y++ +DD +
Sbjct: 586 GSDFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHSR 645
Query: 268 FLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFR 327
W + L KS+V +F + Q+ + VK ++ DN PE TSF+A G++
Sbjct: 646 ATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPELKFTSFYA---EKGIVSF 702
Query: 328 FSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNL 387
SCP T QN ERK + I N+ R L+ + VP S W + A +L+N P + L N
Sbjct: 703 HSCPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNK 762
Query: 388 SPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLS 447
+P ++L P Y LR FGCLCY +K QPRS C+FLGYP ++GYK DL
Sbjct: 763 TPYEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLE 822
Query: 448 HRKIIISRHVIFDETRFPFATMPTPPPSTYDCFSD--PIHPSIIHQWT-SPD--PSPTSV 502
+ ISR+V F E FP A P S+ F+ P+ II T SP PS S
Sbjct: 823 SNTVFISRNVQFHEEVFPLAKNP-GSESSLKLFTPMVPVSSGIISDTTHSPSSLPSQISD 881
Query: 503 VPPAVTPSQLPTTSTSSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVT 562
+PP + S+ PP+ + TM + I +S +
Sbjct: 882 LPPQI--------SSQRVRKPPAHLNDYH-------CNTMQSDHKYPISSTISYSKISPS 926
Query: 563 ----IDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWI 618
I++ T P+P N A W A+ +E A+ + TW++ P + C W+
Sbjct: 927 HMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGCKWV 986
Query: 619 FHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
F K A+G ERYKARLV G +Q G+D +TFSPV K TI+ +L ++ S
Sbjct: 987 FTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSAS 1039
>At2g14930 pseudogene
Length = 1149
Score = 291 bits (746), Expect = 1e-78
Identities = 213/676 (31%), Positives = 308/676 (45%), Gaps = 68/676 (10%)
Query: 37 TAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGD 96
+A+H +T DS W D+ A++H+ ++ L N + G +PI G
Sbjct: 301 SALH-ITDVSDDSGWVPDSAATAHITNNSSRLQQMQPYLG-NDTVMASDGNFLPITHIGS 358
Query: 97 TTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLM 156
LP++ + PL VL P I K+L+SV +LT D S FD G V D T L
Sbjct: 359 ANLPSTSGNLPL--KDVLVCPNIAKSLLSVSKLTKDYPCSFTFDADGVLVKDKATCKVLT 416
Query: 157 RCNSLGD-LYPVTP-------SVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLN 208
+ +S + LY + S +WH RLGHP LQ L + K I
Sbjct: 417 KGSSTSEGLYKLENPKFQMFYSTRQVKATDEVWHMRLGHPNPQVLQLLANKKAIQINKST 476
Query: 209 SPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRYYVLFLDDFTD 267
S +CESC GK RLPF++S+ + P + +H DLW +PV S G +YYV+F+D+ +
Sbjct: 477 SK-MCESCRLGKSSRLPFIASDFIASRPLERVHCDLWGPAPVSSIQGFQYYVIFIDNRSR 535
Query: 268 FLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFR 327
F W +PL +KS +F + + Q D G EF + F + G+
Sbjct: 536 FCWFYPLKHKSDFCSLFMKFQSFVENLLQTKIGTFQSDGGGEFTSNRFLQHLQESGIQHY 595
Query: 328 FSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSL-SN 386
SCPHT QNG AERK R + TL+ + P FW A A +L N++P +L S+
Sbjct: 596 ISCPHTPQQNGLAERKHRQLTERGLTLMFQSKAPQRFWVEAFFTANFLSNLLPTSALDSS 655
Query: 387 LSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDL 446
+P Q+L+ + P Y+ LR FGC C+P + + NK PRS C+FLGY ++GY+CF
Sbjct: 656 TTPYQVLFGKAPDYSALRTFGCACFPTLRAYARNKFDPRSLKCIFLGYTEKYKGYRCFFP 715
Query: 447 SHRKIIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPA 506
++ +SRHV+FDE+ FPF TY P + W PS +S +
Sbjct: 716 PTNRVYLSRHVLFDESSFPFI-------DTYTSLQHPSPTPMFDAWLKSFPSSSSPL--- 765
Query: 507 VTPSQLPTTSTSSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDP 566
T +S AS P + + P ++ + I P +S D
Sbjct: 766 ---ENDQTAGFNSGASVPVITAQQTQP-------ILSLKDGPNILLPEGEITVSSNNQDI 815
Query: 567 TISPLPKNPKLAL-SDPNWKS----AMQSE--------FDA-LIRNKTWDLVPR------ 606
P+ P L S+ N KS +M SE FD I N PR
Sbjct: 816 EDEPICVTPLQTLSSEDNAKSSETLSMGSEECSECTASFDLDPIGNNALSSSPRHDQLTS 875
Query: 607 -----------PSDVNIIRCMWIFHHKTKAN--GCFERYKARLVGDGRSQIAGVDCDETF 653
P + + + + + K N G ++YKARLV G Q G+D ET+
Sbjct: 876 SIPRAATESTHPMTTRLKKGIIKLNQRVKLNVDGSLDKYKARLVAQGFKQEEGIDYLETY 935
Query: 654 SPVVKPATIRTVLTIA 669
SPVV+ AT+R VL ++
Sbjct: 936 SPVVRSATVRAVLHLS 951
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 275 bits (702), Expect = 1e-73
Identities = 186/637 (29%), Positives = 305/637 (47%), Gaps = 54/637 (8%)
Query: 51 WYMDTGASSHVASSPGNLTSYSNLSH-LNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLT 109
WY+D+GAS+H+ G + ++ L + + +G + ++G G+ + +
Sbjct: 335 WYLDSGASNHMC---GRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQF- 390
Query: 110 LSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQ----TGIPLM--RCNSLGD 163
+S+V + P + N++S+ QL + + S+ D + T +P+ R L
Sbjct: 391 ISNVYYIPSMKTNILSLGQLL-EKGYDIRLKDNNLSIRDKESNLITKVPMSKNRMFVLNI 449
Query: 164 LYPVTPSVPFAGLAQS-LWHSRLGHPGSSALQSLRSNKFIT-YEHLNSPT-VCESCVFGK 220
+ + +S LWH R GH L+ L + + +N P VCE C+ G
Sbjct: 450 RNDIAQCLKMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGN 509
Query: 221 HVRLPFVS-SNNVTVMPFDILHSDLWTSPVLSSAGH-RYYVLFLDDFTDFLWTFPLSNKS 278
++ F S++ P +++H+D+ S G Y++LF+DDF+ W + L KS
Sbjct: 510 QFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKS 569
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
+VFE+F + +K ++ D+G EF + F YC +G+ + + P + QNG
Sbjct: 570 EVFEIFKKFKAHVEKESGLVIKTMRSDSGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNG 629
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
AERK RTI M R++L +P W A+ A YLLN P KS+S +P + R P
Sbjct: 630 VAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKP 689
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+HLRVFG + + VP NKL +S +F+GY + +GYK ++ +K IISR+++
Sbjct: 690 GVSHLRVFGSIAHAHVPDEKRNKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIV 749
Query: 459 FDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
FDE +D S+ + + P PT PP+ P+ PT+ TS
Sbjct: 750 FDE------------EGEWDWNSNEEDYNFFPHFEEDKPEPTREEPPSEEPTTPPTSPTS 797
Query: 519 SSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRK------LFNLSVTIDDPTISPLP 572
S SS RT RS++ +Y+ + LF L + P
Sbjct: 798 SQIEESSSE------------RTPRFRSIQELYEVTENQENLTLFCLFAECE-------P 838
Query: 573 KNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERY 632
+ + A+ W++AM E ++ +N TW+L P+ I W++ K + G ERY
Sbjct: 839 MDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERY 898
Query: 633 KARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
KARLV G SQ AG+D DE F+PV + T+R ++++A
Sbjct: 899 KARLVAKGYSQRAGIDYDEIFAPVARLETVRLIISLA 935
>At1g36620 hypothetical protein
Length = 1152
Score = 273 bits (699), Expect = 3e-73
Identities = 208/725 (28%), Positives = 327/725 (44%), Gaps = 82/725 (11%)
Query: 2 SQWTRPSAPPRQ-PGILGPRPQAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSH 60
+Q + PS+ + I G +A +A + + D AT+ ++ + +D+GAS H
Sbjct: 333 AQTSHPSSSASEFSDIPGVSKEAWSAIRNLLKQDTATSSEKLSGKTNCVDFLIDSGASHH 392
Query: 61 VASSPGNLTSYSNLSHL-----NQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLH 115
+ LT + H N K + + +G I G+ + L+HVL
Sbjct: 393 MTGFLDLLTEIYEIPHSVVVLPNAKHTIATKKGTLILGAN------------MKLTHVLF 440
Query: 116 TPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVTPSVPFAG 175
P + LISV +L + + F + D + + + +Y + + A
Sbjct: 441 VPDLSCTLISVARLLRELHCFAIFTDKVCVIQDRTSKMLIGVGTESNGVYHLQRAEVVAT 500
Query: 176 LA--------QSLWHSRLGHPGSSALQSLRSNKFITYEHLNSP--TVCESCVFGKHVRLP 225
A ++LWH RLGHP S L S+ + ++ +S T+C+ CV K R
Sbjct: 501 SANVVKWKTNKALWHMRLGHPSSKVLSSVLPS-LEDFDSCSSDLKTICDVCVRAKQTRAS 559
Query: 226 FVSSNNVTVMPFDILHSDLWTS-PVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMF 284
F S N F +H D+W SS G Y++ +DD + +W + KS+V +
Sbjct: 560 FSESFNKAEECFSFIHYDVWGPYKHASSCGAHYFLTIVDDHSRAVWIHLMLAKSEVASLL 619
Query: 285 TSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKI 344
F+ VK ++ +NG EF S +Y A G++ + SC +T QNG+ ERK
Sbjct: 620 QQFIAMASRQFNKQVKTVRSNNGTEF--MSLKSYFAERGIVHQISCVYTHQQNGRVERKH 677
Query: 345 RTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLR 404
R I N+ R+LL A +P SFW ++ A YL+N P L +P ++LY + PSY LR
Sbjct: 678 RHILNVARSLLFQAELPISFWEESVLTAAYLINRTPTPILDGKTPYKILYSQPPSYASLR 737
Query: 405 VFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRF 464
VFG LC+ + ++K Q R C+F+GYP +G++ +D+ + +SR V+F E F
Sbjct: 738 VFGSLCFARKHTGRLDKFQERGRKCIFVGYPHGQKGWRIYDIESQIFFVSRDVVFQEDIF 797
Query: 465 PFATMPTPPPSTYDCFSDP--IHPSIIHQWTSP--------DPSPTSVVPPAVTPSQLPT 514
PFA D FS P + PS I + D T+ +P + + P
Sbjct: 798 PFADKKNK-----DTFSSPAAVIPSPILPYDDEFLDIYQIGDVPATNPLPAIIDVNDSPP 852
Query: 515 TSTSSSASPPSSPSPP------------------------SPPQPVPP-----IRTMATR 545
+S +A+P ++ PP Q + I MA
Sbjct: 853 SSPIITATPAAASPPPLRRGLRQRQENVRLKDYQTYSAQCESTQTLSDNIGTCIYPMANY 912
Query: 546 SMRGIYKP-RKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLV 604
I+ P + F ++++ DP P+ A+ + W++A+ E DAL TWD+
Sbjct: 913 VSGEIFSPSNQHFLAAISMVDP-----PQTYNQAIREKEWRNAVFFEVDALEDQGTWDIT 967
Query: 605 PRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRT 664
P V I W+F K +NG ERYKARLV G Q G+D +TF+PVVK T+R
Sbjct: 968 KLPQGVKAIGSKWVFRIKYNSNGTVERYKARLVALGNHQKEGIDFTKTFAPVVKMQTVRL 1027
Query: 665 VLTIA 669
+L +A
Sbjct: 1028 LLDVA 1032
>At3g61330 copia-type polyprotein
Length = 1352
Score = 273 bits (698), Expect = 4e-73
Identities = 184/637 (28%), Positives = 304/637 (46%), Gaps = 54/637 (8%)
Query: 51 WYMDTGASSHVASSPGNLTSYSNLSH-LNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLT 109
WY+D+GAS+H+ G + ++ L + + +G + ++G G+ + +
Sbjct: 335 WYLDSGASNHMC---GRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQF- 390
Query: 110 LSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQ----TGIPLM--RCNSLGD 163
+S+V + P + N++S+ QL + + S+ D + T +P+ R L
Sbjct: 391 ISNVYYIPSMKTNILSLGQLL-EKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNI 449
Query: 164 LYPVTPSVPFAGLAQS-LWHSRLGHPGSSALQSLRSNKFIT-YEHLNSPT-VCESCVFGK 220
+ + +S LWH R GH L+ L + + +N P VCE C+ GK
Sbjct: 450 RNDIAQCLKMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGK 509
Query: 221 HVRLPFVS-SNNVTVMPFDILHSDLWTSPVLSSAGH-RYYVLFLDDFTDFLWTFPLSNKS 278
++ F S++ P +++H+D+ S G Y++LF+DDF+ W + L KS
Sbjct: 510 QFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKS 569
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
+VFE+F + +K ++ D G EF + F YC +G+ + + P + QNG
Sbjct: 570 EVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNG 629
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
ERK RTI M R++L +P W A+ A YLLN P KS+S +P + R P
Sbjct: 630 VVERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKP 689
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+HLRVFG + + VP +KL +S +F+GY + +GYK ++ +K IISR+++
Sbjct: 690 GVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIV 749
Query: 459 FDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
FDE +D S+ + + +P PT PP+ P+ PT+ TS
Sbjct: 750 FDE------------EGEWDWNSNEEDYNFFPHFEEDEPEPTREEPPSEEPTTPPTSPTS 797
Query: 519 SSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRK------LFNLSVTIDDPTISPLP 572
S SS RT RS++ +Y+ + LF L + P
Sbjct: 798 SQIEESSSE------------RTPRFRSIQELYEVTENQENLTLFCLFAECE-------P 838
Query: 573 KNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERY 632
+ + A+ W++AM E ++ +N TW+L P+ I W++ K + G ERY
Sbjct: 839 MDFQKAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERY 898
Query: 633 KARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
KARLV G SQ G+D DE F+PV + T+R ++++A
Sbjct: 899 KARLVAKGYSQRVGIDYDEVFAPVARLETVRLIISLA 935
>At1g37110
Length = 1356
Score = 271 bits (692), Expect = 2e-72
Identities = 191/645 (29%), Positives = 295/645 (45%), Gaps = 36/645 (5%)
Query: 51 WYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTL 110
W +D+G +SH+ S S+ N + +G + QG G + T H L
Sbjct: 308 WILDSGCTSHMTSRRDWFISFQEKG--NTTILLGDDHSVESQGQGTIRIDT-HGGTIKIL 364
Query: 111 SHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVTPS 170
+V + P + +NLIS L + + + V F+ +R + LY + S
Sbjct: 365 ENVKYVPHLRRNLISTGTL---DKLGYRHEGGEGKVRYFKNNKTALRGSLSNGLYVLDGS 421
Query: 171 VPFAGLAQS--------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHV 222
+ L + LWHSRLGH + L+ L I + +N CE CV GK
Sbjct: 422 TVMSELCNAETDKVKTALWHSRLGHMSMNNLKVLAGKGLIDRKEINELEFCEHCVMGKSK 481
Query: 223 RLPFVSSNNVTVMPFDILHSDLWTSPVL--SSAGHRYYVLFLDDFTDFLWTFPLSNKSQV 280
++ F + + +H+DLW SP + S +G +Y++ +DD T +W + L +K +
Sbjct: 482 KVSFNVGKHTSEDALSYVHADLWGSPNVTPSISGKQYFLSIIDDKTRKVWLYFLKSKDET 541
Query: 281 FEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKA 340
F+ F + + VKCL+ DNG EF N+ F +YC HG+ +C +T QNG A
Sbjct: 542 FDKFCEWKSLVENQVNKKVKCLRTDNGLEFCNSRFDSYCKEHGIERHRTCTYTPQQNGVA 601
Query: 341 ERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSY 400
ER RTI +R LL + V FW A A YL+N P ++++ P ++ R P Y
Sbjct: 602 ERMNRTIMEKVRCLLNKSGVEEVFWAEAAATAAYLINRSPASAINHNVPEEMWLNRKPGY 661
Query: 401 THLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFD 460
HLR FG + Y KL+PR+ FLGYP +GYK + L K +ISR+V+F
Sbjct: 662 KHLRKFGSIAY---VHQDQGKLKPRALKGFFLGYPAGTKGYKVWLLEEEKCVISRNVVFQ 718
Query: 461 ETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSS 520
E+ + + T + S + Q + S + V + S+ P T S
Sbjct: 719 ES-VVYRDLKVKEDDTDNLNQKETTSSEVEQNKFAEASGSGGVIQLQSDSE-PITEGEQS 776
Query: 521 ASPPSSPSPPSPPQPVPPIRTMAT------RSMRGIYKPRKL-----FNLSVTIDDPTIS 569
+ Q P + T R R I P + ++ + + I
Sbjct: 777 SDSEEEVEYSEKTQETPKRTGLTTYKLARDRVRRNINPPTRFTEESSVTFALVVVENCIV 836
Query: 570 PLPKNPKLALSDPN---WKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKAN 626
P++ + A+ + W A E D+L++N TWDLV +P D II C W+F K+
Sbjct: 837 QEPQSYQEAMESQDCEKWDMATHDEMDSLMKNGTWDLVDKPKDRKIIGCRWLFKLKSGIP 896
Query: 627 GCF-ERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIAL 670
G R+KARLV G +Q GVD E F+PVVK +IR ++++ +
Sbjct: 897 GVEPTRFKARLVAKGYTQREGVDYQEIFAPVVKHVSIRILMSLVV 941
>At1g26990 polyprotein, putative
Length = 1436
Score = 268 bits (686), Expect = 9e-72
Identities = 203/706 (28%), Positives = 310/706 (43%), Gaps = 72/706 (10%)
Query: 14 PGILGPRPQAHAASTSPVPTDI-----------ATAMHTMTLAPPDST------WYMDTG 56
P I +A A+S++PVP+ +T +T + P T W +D+G
Sbjct: 367 PSISQITDKAIASSSNPVPSISQITGTFFSLYDSTYYEMLTSSIPIETELSLRAWVIDSG 426
Query: 57 ASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHT 116
AS HV +Y L +L +G + I+G+G L + L+L +VL
Sbjct: 427 ASHHVTHERNLYHTYKALDRTFVRL--PNGHTVKIEGTGFIQLTDA-----LSLHNVLFI 479
Query: 117 PQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVT-------- 168
P+ NL+SV LT V F + + L + + +G+LY +
Sbjct: 480 PEFKFNLLSVSVLTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVDV 539
Query: 169 PSVPFAGLAQS------LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVFGKH 221
S P + S +WH RLGHP + + +L + + +N + C C K
Sbjct: 540 SSFPGKSVCSSVKNESEMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLSKQ 599
Query: 222 VRLPFVSSNNVTVMPFDILHSDLW---TSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
LPF S N++ F+++H D W + P + S +RY++ +DDF+ W + L KS
Sbjct: 600 KHLPFKSVNHIREKAFELVHIDTWGPFSVPTVDS--YRYFLTIVDDFSRATWIYLLKQKS 657
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
V +F S + T + V ++ DN E F+ A G+ CP T QN
Sbjct: 658 DVLTVFPSFLKMVETQYHTKVCSVRSDNAHEL---KFNELFAKEGIKADHPCPETPEQNF 714
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
ERK + + N+ R L+ + +P +W + A +L+N + ++N +P + L + P
Sbjct: 715 VVERKHQHLLNVARALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKGKP 774
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
Y+ L+ FGCLCY + K PR+ C+FLGYP+ ++GYK D+ + ISRHVI
Sbjct: 775 DYSSLKAFGCLCYCSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRHVI 834
Query: 459 FDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
F E FPFA+ S + P I P P+ +P + S P
Sbjct: 835 FYEDIFPFAS------SNITDAAKDFFPHIY----LPAPNNDEHLPLVQSSSDAPHNHDE 884
Query: 519 SSAS--PPSSPSP------PSPPQPVPPIRTMATRSMRGIYKPRKLFNLS-------VTI 563
SS+ PS P PS Q T + Y + S I
Sbjct: 885 SSSMIFVPSEPKSTRQRKLPSHLQDFHCYNNTPTTTKTSPYPLTNYISYSYLSEPFGAFI 944
Query: 564 DDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKT 623
+ T + LP+ A D W AM E A +R TW + P+ + C WI K
Sbjct: 945 NIITATKLPQKYSEARLDKVWNDAMGKEISAFVRTGTWSICDLPAGKVAVGCKWIITIKF 1004
Query: 624 KANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
A+G ER+KARLV G +Q G+D TFSPV K T++ +L++A
Sbjct: 1005 LADGSIERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLA 1050
>At1g48710 hypothetical protein
Length = 1352
Score = 267 bits (683), Expect = 2e-71
Identities = 183/637 (28%), Positives = 303/637 (46%), Gaps = 54/637 (8%)
Query: 51 WYMDTGASSHVASSPGNLTSYSNLSH-LNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLT 109
WY+D+GAS+H+ G + ++ L + + +G + ++G G+ + +
Sbjct: 335 WYLDSGASNHMC---GRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQF- 390
Query: 110 LSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQ----TGIPLM--RCNSLGD 163
+S+V + P + N++S+ QL + + S+ D + T +P+ R L
Sbjct: 391 ISNVYYIPSMKTNILSLGQLL-EKGYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNI 449
Query: 164 LYPVTPSVPFAGLAQS-LWHSRLGHPGSSALQSLRSNKFIT-YEHLNSPT-VCESCVFGK 220
+ + +S LWH R GH L+ L + + +N P VCE C+ GK
Sbjct: 450 RNDIAQCLKMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGK 509
Query: 221 HVRLPFVS-SNNVTVMPFDILHSDLWTSPVLSSAGH-RYYVLFLDDFTDFLWTFPLSNKS 278
++ F S++ +++H+D+ S G Y++LF+DDF+ W + L KS
Sbjct: 510 QFKMSFPKESSSRAQKSLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKS 569
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
+VFE+F + +K ++ D G EF + F YC +G+ + + P + QNG
Sbjct: 570 EVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNG 629
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
AERK RTI M R++L +P W A+ A YLLN P KS+S +P + R
Sbjct: 630 VAERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKS 689
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+HLRVFG + + VP +KL +S +F+GY + +GYK ++ +K IISR+++
Sbjct: 690 GVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIV 749
Query: 459 FDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
FDE +D S+ + + +P PT PP+ P+ PT+ TS
Sbjct: 750 FDE------------EGEWDWNSNEEDYNFFPHFEEDEPEPTREEPPSEEPTTPPTSPTS 797
Query: 519 SSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRK------LFNLSVTIDDPTISPLP 572
S SS RT RS++ +Y+ + LF L + P
Sbjct: 798 SQIEESSSE------------RTPRFRSIQELYEVTENQENLTLFCLFAECE-------P 838
Query: 573 KNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERY 632
+ + A+ W++AM E ++ +N TW+L P+ I W++ K + G ERY
Sbjct: 839 MDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKTIGVKWVYKAKKNSKGEVERY 898
Query: 633 KARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
KARLV G Q AG+D DE F+PV + T+R ++++A
Sbjct: 899 KARLVAKGYIQRAGIDYDEVFAPVARLETVRLIISLA 935
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.329 0.140 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,110,603
Number of Sequences: 26719
Number of extensions: 1162476
Number of successful extensions: 14607
Number of sequences better than 10.0: 418
Number of HSP's better than 10.0 without gapping: 262
Number of HSP's successfully gapped in prelim test: 166
Number of HSP's that attempted gapping in prelim test: 6299
Number of HSP's gapped (non-prelim): 2713
length of query: 1082
length of database: 11,318,596
effective HSP length: 110
effective length of query: 972
effective length of database: 8,379,506
effective search space: 8144879832
effective search space used: 8144879832
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0114.10


