

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.1
(661 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
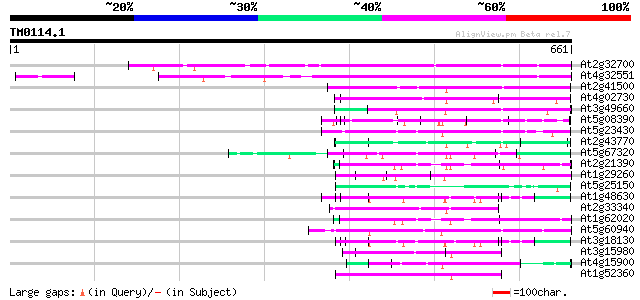
Score E
Sequences producing significant alignments: (bits) Value
At2g32700 unknown protein 341 9e-94
At4g32551 Leunig protein 327 1e-89
At2g41500 putative U4/U6 small nuclear ribonucleoprotein 97 2e-20
At4g02730 putative WD-repeat protein 86 7e-17
At3g49660 putative WD-40 repeat - protein 82 1e-15
At5g08390 katanin p80 subunit - like protein 79 6e-15
At5g23430 unknown protein 79 8e-15
At2g43770 putative splicing factor 75 2e-13
At5g67320 unknown protein 74 3e-13
At2g21390 coatomer alpha subunit 72 1e-12
At1g29260 peroxisomal targeting signal type 2 receptor 72 1e-12
At5g25150 transcription initiation factor IID-associated factor-... 71 2e-12
At1g48630 unknown protein 69 8e-12
At2g33340 putative PRP19-like spliceosomal protein 69 1e-11
At1g62020 hypothetical protein 68 1e-11
At5g60940 cleavage stimulation factor 50K chain 68 2e-11
At3g18130 protein kinase C-receptor/G-protein, putative 68 2e-11
At3g15980 putative coatomer complex subunit 62 8e-10
At4g15900 PRL1 protein 62 1e-09
At1g52360 coatomer complex subunit, putative 62 1e-09
>At2g32700 unknown protein
Length = 787
Score = 341 bits (874), Expect = 9e-94
Identities = 204/541 (37%), Positives = 303/541 (55%), Gaps = 38/541 (7%)
Query: 141 GKQVQKYVFKVVTQFTSIYTNENHKLSG--------SGVTMRVGRDVPRDPPDVVQKTML 192
G QVQK + +QF + H++ + M G PR + + +
Sbjct: 265 GPQVQKSFLQNQSQFQLSPQQQQHQMLAQVQAQGNMTNSPMYGGDMDPRRFTGLPRGNLN 324
Query: 193 PLDGPHQTNNNEALNLVPQPLNGG--------PVNTPNSHQQCQVLKTQNQ-DAALTQAP 243
P DG N+ + P+ PV +S QQ +L Q+Q + + P
Sbjct: 325 PKDGQQNANDGS----IGSPMQSSSSKHISMPPVQQSSSQQQDHLLSQQSQQNNRKRKGP 380
Query: 244 ACT-PNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQSQN 302
+ + P T T N P P T + + I+ H N + M +
Sbjct: 381 SSSGPANSTGTGNTVGPSNSQPSTPSTHTPVDGVA-----IAGNMHHVNSMPKGPMMYGS 435
Query: 303 VEKSTKRMITESTRPVESAQD-CADAKPADENVESFLSFEDEPADNRIEPFSNLKRISAT 361
+ + + + ++ D D ++NVESFLS +D + F LKR S+
Sbjct: 436 --DGIGGLASSANQLLQDDMDQFGDVGALEDNVESFLSQDDGDGGSL---FGTLKRNSSV 490
Query: 362 CSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASAD 421
+ +K FS E+ C+ KS +KV+ C FS DGK++ASAGH+KKVFIWNME Q ++ +
Sbjct: 491 HTET-SKPFSFNEVSCIRKSASKVICCSFSYDGKLLASAGHDKKVFIWNMETLQVESTPE 549
Query: 422 THSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMD 481
H+H+ITDVRF+P ST ATSSFD+++++WDAS+P L ++GH VMS+DFHP++ +
Sbjct: 550 EHAHIITDVRFRPNSTQLATSSFDKTIKIWDASDPGYFLRTISGHAAPVMSIDFHPKKTE 609
Query: 482 LLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDS- 540
LLCS DSN+ IR W++N S C+ +G S QVRFQP+ GQ LA A N+++I D+E ++
Sbjct: 610 LLCSCDSNNDIRFWDINAS-CVRAVKGASTQVRFQPRTGQFLAAASENTVSIFDIENNNK 668
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFH 600
++ KGH +V S+CW +G +ASVSED+ ++WS +S G CI EL + GNKF S +FH
Sbjct: 669 RVNIFKGHSSNVHSVCWSPNGELVASVSEDAVKLWSLSS-GDCIHELSNSGNKFHSVVFH 727
Query: 601 PGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
P Y LL+IGGYQ++E W+ E +K ++A H+ +I+ LA SP ++ASASHD SVK+W
Sbjct: 728 PSYPDLLVIGGYQAIELWNTME-NKCMTVAGHECVISALAQSPSTGVVASASHDKSVKIW 786
Query: 661 K 661
K
Sbjct: 787 K 787
>At4g32551 Leunig protein
Length = 931
Score = 327 bits (838), Expect = 1e-89
Identities = 187/498 (37%), Positives = 265/498 (52%), Gaps = 37/498 (7%)
Query: 176 GRDVPRDPPDVVQKTMLPLDGPHQTNNNEALNLVPQPLNGGPVNTPNSHQQC-----QVL 230
G + P+ P L L P ++N +++ + GG + S QVL
Sbjct: 459 GGNPPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGGGSITMDGSISNSFRGNEQVL 518
Query: 231 KTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGE--H 288
K Q+ + P + T N P + S + +IS H
Sbjct: 519 KNQSGRKRKQPVSSSGPANSSGTA------NTAGPSPSSAPSTPSTHTPGDVISMPNLPH 572
Query: 289 QCNQDQQIQM---QSQNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPA 345
+ + M + S + + R VE D+NVESFLS ED
Sbjct: 573 SGGSSKSMMMFGTEGTGTLTSPSNQLADMDRFVEDGS-------LDDNVESFLSQEDGD- 624
Query: 346 DNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKK 405
+R + T + +KGF+ E+ + S KV CHFSSDGK++ASAGH+KK
Sbjct: 625 ----------QRDAVTRCMDVSKGFTFTEVNSVRASTTKVTCCHFSSDGKMLASAGHDKK 674
Query: 406 VFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTG 465
+W + + + + H+ +ITD+RF P ATSSFD++VR+WDA N SL G
Sbjct: 675 AVLWYTDTMKPKTTLEEHTAMITDIRFSPSQLRLATSSFDKTVRVWDADNKGYSLRTFMG 734
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLAT 525
H V SLDFHP + DL+CS D+++ IR W++N C +GGS Q+RFQP+ G+ LA
Sbjct: 735 HSSMVTSLDFHPIKDDLICSCDNDNEIRYWSINNGSCTRVYKGGSTQIRFQPRVGKYLAA 794
Query: 526 AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWS--AASDGKC 583
+ N + ++DVET + H L+GH + S+CWD SG+FLASVSED ++W+ S+G+C
Sbjct: 795 SSANLVNVLDVETQAIRHSLQGHANPINSVCWDPSGDFLASVSEDMVKVWTLGTGSEGEC 854
Query: 584 IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSP 643
+ EL GNKFQSC+FHP Y SLL+IG YQSLE W+ +E +KT ++ AH+GLI LA S
Sbjct: 855 VHELSCNGNKFQSCVFHPAYPSLLVIGCYQSLELWNMSE-NKTMTLPAHEGLITSLAVST 913
Query: 644 VGELIASASHDNSVKVWK 661
L+ASASHD VK+WK
Sbjct: 914 ATGLVASASHDKLVKLWK 931
Score = 43.1 bits (100), Expect = 5e-04
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H E + M + PQ+
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH---SEVAASYIETQMIKAREQQLQQSQHPQVSQ 101
Query: 68 NEHRPQQFQ 76
+ + QQ Q
Sbjct: 102 QQQQQQQQQ 110
>At2g41500 putative U4/U6 small nuclear ribonucleoprotein
Length = 554
Score = 97.4 bits (241), Expect = 2e-20
Identities = 78/293 (26%), Positives = 126/293 (42%), Gaps = 11/293 (3%)
Query: 375 IGCLHKSINKVLT-CHFSSDGKVMASAGHEKKVFIWNMENF-QYFASADTHSHLITDVRF 432
+ C + ++ LT C FS DGK++A+ +W M A H TDV F
Sbjct: 247 LDCSNFGDDRPLTGCSFSRDGKILATCSLSGVTKLWEMPQVTNTIAVLKDHKERATDVVF 306
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P AT+S DR+ +LW + L GH +++ + FHP L ++ +
Sbjct: 307 SPVDDCLATASADRTAKLWKTDGTL--LQTFEGHLDRLARVAFHPSGK-YLGTTSYDKTW 363
Query: 493 RLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHD 549
RLW++N + G S+ V FQ + + + + D+ T + +GH
Sbjct: 364 RLWDINTGAELLLQEGHSRSVYGIAFQQDGALAASCGLDSLARVWDLRTGRSILVFQGHI 423
Query: 550 KDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLI 608
K V S+ + +G LAS ED+ RIW K + + + N + P L
Sbjct: 424 KPVFSVNFSPNGYHLASGGEDNQCRIWDLRMR-KSLYIIPAHANLVSQVKYEPQEGYFLA 482
Query: 609 IGGYQ-SLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
Y + WS + S S+A H+ +A L + IA+ SHD ++K+W
Sbjct: 483 TASYDMKVNIWSGRDFSLVKSLAGHESKVASLDITADSSCIATVSHDRTIKLW 535
Score = 41.2 bits (95), Expect = 0.002
Identities = 19/62 (30%), Positives = 33/62 (52%)
Query: 393 DGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
+G +A+A ++ KV IW+ +F S H + + S+ AT S DR+++LW
Sbjct: 477 EGYFLATASYDMKVNIWSGRDFSLVKSLAGHESKVASLDITADSSCIATVSHDRTIKLWT 536
Query: 453 AS 454
+S
Sbjct: 537 SS 538
>At4g02730 putative WD-repeat protein
Length = 333
Score = 85.9 bits (211), Expect = 7e-17
Identities = 66/276 (23%), Positives = 117/276 (41%), Gaps = 46/276 (16%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS+DG ++ASA +K + +W+ N+ + HS I+D+ + S ++S D ++R
Sbjct: 51 FSNDGNLLASASVDKTMILWSATNYSLIHRYEGHSSGISDLAWSSDSHYTCSASDDCTLR 110
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
+WDA +P L +L GH V ++F+P +L+ S ++ IR+W V +C+
Sbjct: 111 IWDARSPYECLKVLRGHTNFVFCVNFNP-PSNLIVSGSFDETIRIWEVKTGKCVRM---- 165
Query: 510 SKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSE 569
+K H + S+ ++R G+ + S S
Sbjct: 166 -----------------------------------IKAHSMPISSVHFNRDGSLIVSASH 190
Query: 570 D-SARIWSAASDGKCIGELHSVGNKFQS-CIFHPGYRSLLIIGGYQSLEAWSPTEGSKTW 627
D S +IW A +G C+ L + S F P + +L+ +L+ + G
Sbjct: 191 DGSCKIWD-AKEGTCLKTLIDDKSPAVSFAKFSPNGKFILVATLDSTLKLSNYATGKFLK 249
Query: 628 SIAAHQGLIAGLADS---PVGELIASASHDNSVKVW 660
H + + + G+ I S S DN V +W
Sbjct: 250 VYTGHTNKVFCITSAFSVTNGKYIVSGSEDNCVYLW 285
Score = 46.2 bits (108), Expect = 6e-05
Identities = 40/202 (19%), Positives = 86/202 (41%), Gaps = 10/202 (4%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V +F+ ++ S ++ + IW ++ + HS I+ V F ++ ++
Sbjct: 129 NFVFCVNFNPPSNLIVSGSFDETIRIWEVKTGKCVRMIKAHSMPISSVHFNRDGSLIVSA 188
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S D S ++WDA ++ V F P +L ++ + ++L N +
Sbjct: 189 SHDGSCKIWDAKEGTCLKTLIDDKSPAVSFAKFSPNGKFILVAT-LDSTLKLSNYATGKF 247
Query: 503 MHTTRGGSKQV-----RFQPQYGQ-LLATAIGNSITIIDVETDSHLHDLKGHDKDVLSIC 556
+ G + +V F G+ +++ + N + + D++ + L L+GH V+S+
Sbjct: 248 LKVYTGHTNKVFCITSAFSVTNGKYIVSGSEDNCVYLWDLQARNILQRLEGHTDAVISVS 307
Query: 557 WDRSGNFLASVS---EDSARIW 575
N ++S + + RIW
Sbjct: 308 CHPVQNEISSSGNHLDKTIRIW 329
>At3g49660 putative WD-40 repeat - protein
Length = 317
Score = 81.6 bits (200), Expect = 1e-15
Identities = 67/284 (23%), Positives = 112/284 (38%), Gaps = 49/284 (17%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N + FSSD + + SA +K + +W++E + H++ V F P S M +
Sbjct: 72 NGISDVAFSSDARFIVSASDDKTLKLWDVETGSLIKTLIGHTNYAFCVNFNPQSNMIVSG 131
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
SFD +VR+WD + + L +L H + V ++DF+ R L+ SS + + R+W+ C
Sbjct: 132 SFDETVRIWDVTTG-KCLKVLPAHSDPVTAVDFN-RDGSLIVSSSYDGLCRIWDSGTGHC 189
Query: 503 MHTTRGGSKQ----VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWD 558
+ T VRF P +L + N++ + ++ + L GH
Sbjct: 190 VKTLIDDENPPVSFVRFSPNGKFILVGTLDNTLRLWNISSAKFLKTYTGH---------- 239
Query: 559 RSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQS--LE 616
+ CI SV N G R I+ G + +
Sbjct: 240 -------------------VNAQYCISSAFSVTN---------GKR---IVSGSEDNCVH 268
Query: 617 AWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
W + H + +A P LIAS S D +V++W
Sbjct: 269 MWELNSKKLLQKLEGHTETVMNVACHPTENLIASGSLDKTVRIW 312
Score = 75.1 bits (183), Expect = 1e-13
Identities = 61/252 (24%), Positives = 114/252 (45%), Gaps = 14/252 (5%)
Query: 422 THSHLITDVRFQPGSTMFATSSFDRSVRLWDAS---NPIRS-LGILTGHDEQVMSLDFHP 477
+H+ ++ V+F + A++S D+++R + + +PI + TGH+ + + F
Sbjct: 22 SHNRAVSSVKFSSDGRLLASASADKTIRTYTINTINDPIAEPVQEFTGHENGISDVAFSS 81
Query: 478 RRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ---VRFQPQYGQLLATAIGNSITII 534
++ +SD ++LW+V + T G + V F PQ +++ + ++ I
Sbjct: 82 DARFIVSASDDK-TLKLWDVETGSLIKTLIGHTNYAFCVNFNPQSNMIVSGSFDETVRIW 140
Query: 535 DVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNK 593
DV T L L H V ++ ++R G+ + S S D RIW + + G C+ L N
Sbjct: 141 DVTTGKCLKVLPAHSDPVTAVDFNRDGSLIVSSSYDGLCRIWDSGT-GHCVKTLIDDENP 199
Query: 594 FQSCI-FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH---QGLIAGLADSPVGELIA 649
S + F P + +L+ +L W+ + + H Q I+ G+ I
Sbjct: 200 PVSFVRFSPNGKFILVGTLDNTLRLWNISSAKFLKTYTGHVNAQYCISSAFSVTNGKRIV 259
Query: 650 SASHDNSVKVWK 661
S S DN V +W+
Sbjct: 260 SGSEDNCVHMWE 271
Score = 38.1 bits (87), Expect = 0.016
Identities = 17/74 (22%), Positives = 37/74 (49%), Gaps = 1/74 (1%)
Query: 379 HKSINKVLTCHFS-SDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGST 437
H + ++ FS ++GK + S + V +W + + + + H+ + +V P
Sbjct: 239 HVNAQYCISSAFSVTNGKRIVSGSEDNCVHMWELNSKKLLQKLEGHTETVMNVACHPTEN 298
Query: 438 MFATSSFDRSVRLW 451
+ A+ S D++VR+W
Sbjct: 299 LIASGSLDKTVRIW 312
Score = 37.7 bits (86), Expect = 0.021
Identities = 29/125 (23%), Positives = 50/125 (39%), Gaps = 5/125 (4%)
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIG----ELHSVGNKFQ 595
H L H++ V S+ + G LAS S D + R ++ + I E N
Sbjct: 16 HSQTLTSHNRAVSSVKFSSDGRLLASASADKTIRTYTINTINDPIAEPVQEFTGHENGIS 75
Query: 596 SCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDN 655
F R ++ ++L+ W GS ++ H + +P +I S S D
Sbjct: 76 DVAFSSDARFIVSASDDKTLKLWDVETGSLIKTLIGHTNYAFCVNFNPQSNMIVSGSFDE 135
Query: 656 SVKVW 660
+V++W
Sbjct: 136 TVRIW 140
>At5g08390 katanin p80 subunit - like protein
Length = 823
Score = 79.3 bits (194), Expect = 6e-15
Identities = 66/269 (24%), Positives = 119/269 (43%), Gaps = 15/269 (5%)
Query: 395 KVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDAS 454
+V+ + G + KV +W + S HS I V F + A + +++LWD
Sbjct: 123 RVLVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSVTFDASEGLVAAGAASGTIKLWDLE 182
Query: 455 NPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG---GSK 511
+ + LTGH +S++FHP + S + +++W++ + C+HT +G G
Sbjct: 183 E-AKVVRTLTGHRSNCVSVNFHPFG-EFFASGSLDTNLKIWDIRKKGCIHTYKGHTRGVN 240
Query: 512 QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED- 570
+RF P +++ N + + D+ LH+ K H+ + S+ + LA+ S D
Sbjct: 241 VLRFTPDGRWIVSGGEDNVVKVWDLTAGKLLHEFKSHEGKIQSLDFHPHEFLLATGSADK 300
Query: 571 SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIA 630
+ + W + + IG + + F+P +S+L G +SL+ P G T S
Sbjct: 301 TVKFWDLET-FELIGSGGTETTGVRCLTFNPDGKSVL-CGLQESLKRTEPMSGGATQS-N 357
Query: 631 AHQGLIAGLADSPVGELIASASHDNSVKV 659
+H +G G +DNS KV
Sbjct: 358 SHPEKTSGSGRDQAG------LNDNSSKV 380
Score = 60.5 bits (145), Expect = 3e-09
Identities = 52/208 (25%), Positives = 81/208 (38%), Gaps = 5/208 (2%)
Query: 457 IRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG---GSKQV 513
I S H V L + +L + + + LW + + + + G G V
Sbjct: 99 ISSFREFVAHSAAVNCLKIGRKSSRVLVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSV 158
Query: 514 RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDS-A 572
F G + A A +I + D+E + L GH + +S+ + G F AS S D+
Sbjct: 159 TFDASEGLVAAGAASGTIKLWDLEEAKVVRTLTGHRSNCVSVNFHPFGEFFASGSLDTNL 218
Query: 573 RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH 632
+IW G CI F P R ++ G ++ W T G +H
Sbjct: 219 KIWDIRKKG-CIHTYKGHTRGVNVLRFTPDGRWIVSGGEDNVVKVWDLTAGKLLHEFKSH 277
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVW 660
+G I L P L+A+ S D +VK W
Sbjct: 278 EGKIQSLDFHPHEFLLATGSADKTVKFW 305
Score = 50.1 bits (118), Expect = 4e-06
Identities = 37/154 (24%), Positives = 68/154 (44%), Gaps = 12/154 (7%)
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
+S+G V A A + +W++E + + H V F P FA+ S D ++++
Sbjct: 162 ASEGLVAAGAA-SGTIKLWDLEEAKVVRTLTGHRSNCVSVNFHPFGEFFASGSLDTNLKI 220
Query: 451 WDASNPIRSLGIL---TGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH--- 504
WD IR G + GH V L F P ++ + N V+++W++ + +H
Sbjct: 221 WD----IRKKGCIHTYKGHTRGVNVLRFTPDGRWIVSGGEDN-VVKVWDLTAGKLLHEFK 275
Query: 505 TTRGGSKQVRFQPQYGQLLATAIGNSITIIDVET 538
+ G + + F P L + ++ D+ET
Sbjct: 276 SHEGKIQSLDFHPHEFLLATGSADKTVKFWDLET 309
Score = 50.1 bits (118), Expect = 4e-06
Identities = 42/211 (19%), Positives = 88/211 (40%), Gaps = 13/211 (6%)
Query: 386 LTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFD 445
++ +F G+ AS + + IW++ + H+ + +RF P + D
Sbjct: 198 VSVNFHPFGEFFASGSLDTNLKIWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWIVSGGED 257
Query: 446 RSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM-- 503
V++WD + + L H+ ++ SLDFHP LL + ++ ++ W++ E +
Sbjct: 258 NVVKVWDLTAG-KLLHEFKSHEGKIQSLDFHPHEF-LLATGSADKTVKFWDLETFELIGS 315
Query: 504 -HTTRGGSKQVRFQPQYGQLLATAIGNSITIID------VETDSHLHDLKGHDKDVLSIC 556
T G + + F P G+ + + S+ + +++SH G +D +
Sbjct: 316 GGTETTGVRCLTFNPD-GKSVLCGLQESLKRTEPMSGGATQSNSHPEKTSGSGRDQAGLN 374
Query: 557 WDRSGNFLASVSEDSARIWSAASDGKCIGEL 587
D S + S ++ + K +G+L
Sbjct: 375 -DNSSKVILGKLPGSQKVDPLLKETKSLGKL 404
Score = 47.8 bits (112), Expect = 2e-05
Identities = 30/120 (25%), Positives = 52/120 (43%), Gaps = 4/120 (3%)
Query: 368 KGFSLQEIGCLHK---SINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHS 424
K + +++ GC+H V F+ DG+ + S G + V +W++ + +H
Sbjct: 219 KIWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWIVSGGEDNVVKVWDLTAGKLLHEFKSHE 278
Query: 425 HLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLC 484
I + F P + AT S D++V+ WD +G V L F+P +LC
Sbjct: 279 GKIQSLDFHPHEFLLATGSADKTVKFWDLET-FELIGSGGTETTGVRCLTFNPDGKSVLC 337
>At5g23430 unknown protein
Length = 837
Score = 79.0 bits (193), Expect = 8e-15
Identities = 73/304 (24%), Positives = 135/304 (44%), Gaps = 25/304 (8%)
Query: 368 KGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLI 427
+ + LQE ++N + SS +V+ + G + KV +W + S HS I
Sbjct: 5 RAYKLQEFVAHSAAVNCLKIGRKSS--RVLVTGGEDHKVNLWAIGKPNAILSLYGHSSGI 62
Query: 428 TDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSD 487
V F + A + +++LWD + + LTGH +S+DFHP + S
Sbjct: 63 DSVTFDASEVLVAAGAASGTIKLWDLEE-AKIVRTLTGHRSNCISVDFHPFG-EFFASGS 120
Query: 488 SNDVIRLWNVNQSECMHTTRG---GSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHD 544
+ +++W++ + C+HT +G G +RF P +++ N + + D+ L +
Sbjct: 121 LDTNLKIWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWVVSGGEDNIVKVWDLTAGKLLTE 180
Query: 545 LKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
K H+ + S+ + LA+ S D + + W + + IG + F+P
Sbjct: 181 FKSHEGQIQSLDFHPHEFLLATGSADRTVKFWDLET-FELIGSGGPETAGVRCLSFNPDG 239
Query: 604 RSLLIIGGYQSLE--AWSPTEGSKTWSIAAHQGLIAG---LADSPV--GELIASASHDNS 656
+++L G +SL+ +W P I H G+ G L+D V G+L+ + + +
Sbjct: 240 KTVL-CGLQESLKIFSWEP--------IRCHDGVDVGWSRLSDMNVHEGKLLGCSYNQSC 290
Query: 657 VKVW 660
V VW
Sbjct: 291 VGVW 294
>At2g43770 putative splicing factor
Length = 343
Score = 74.7 bits (182), Expect = 2e-13
Identities = 55/284 (19%), Positives = 111/284 (38%), Gaps = 12/284 (4%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNME-NFQYFASADTHSHLITDVRFQPGSTMFATSS 443
V T F+ G ++AS H++++F+W + + + F H + I D+ + + ++S
Sbjct: 56 VYTMKFNPAGTLIASGSHDREIFLWRVHGDCKNFMVLKGHKNAILDLHWTSDGSQIVSAS 115
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM 503
D++VR WD + + + H V S R L+ S + +LW++ Q +
Sbjct: 116 PDKTVRAWDVETG-KQIKKMAEHSSFVNSCCPTRRGPPLIISGSDDGTAKLWDMRQRGAI 174
Query: 504 HT--TRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
T + V F ++ + N + + D+ L+GH + + G
Sbjct: 175 QTFPDKYQITAVSFSDAADKIFTGGVDNDVKVWDLRKGEATMTLEGHQDTITGMSLSPDG 234
Query: 562 NFLASVSEDSAR-IWSA---ASDGKCI----GELHSVGNKFQSCIFHPGYRSLLIIGGYQ 613
++L + D+ +W A +C+ G H+ C + P + +
Sbjct: 235 SYLLTNGMDNKLCVWDMRPYAPQNRCVKIFEGHQHNFEKNLLKCSWSPDGTKVTAGSSDR 294
Query: 614 SLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSV 657
+ W T + + H G + P +I S S D ++
Sbjct: 295 MVHIWDTTSRRTIYKLPGHTGSVNECVFHPTEPIIGSCSSDKNI 338
Score = 65.5 bits (158), Expect = 9e-11
Identities = 55/242 (22%), Positives = 95/242 (38%), Gaps = 48/242 (19%)
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H + ++F P T+ A+ S DR + LW ++ +L GH ++ L
Sbjct: 52 HPSAVYTMKFNPAGTLIASGSHDREIFLWRVHGDCKNFMVLKGHKNAILDL--------- 102
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHL 542
H T GS+ V P ++ DVET +
Sbjct: 103 ---------------------HWTSDGSQIVSASPD----------KTVRAWDVETGKQI 131
Query: 543 HDLKGHDKDVLSICWDRSGN-FLASVSED-SARIWSAASDGKCIGELHSVGNKFQ-SCIF 599
+ H V S C R G + S S+D +A++W D + G + + +K+Q + +
Sbjct: 132 KKMAEHSSFVNSCCPTRRGPPLIISGSDDGTAKLW----DMRQRGAIQTFPDKYQITAVS 187
Query: 600 HPGYRSLLIIGGYQS-LEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVK 658
+ GG + ++ W +G T ++ HQ I G++ SP G + + DN +
Sbjct: 188 FSDAADKIFTGGVDNDVKVWDLRKGEATMTLEGHQDTITGMSLSPDGSYLLTNGMDNKLC 247
Query: 659 VW 660
VW
Sbjct: 248 VW 249
Score = 62.8 bits (151), Expect = 6e-10
Identities = 53/237 (22%), Positives = 94/237 (39%), Gaps = 27/237 (11%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITD-VRFQPGSTMFAT 441
N +L H++SDG + SA +K V W++E + HS + + G + +
Sbjct: 97 NAILDLHWTSDGSQIVSASPDKTVRAWDVETGKQIKKMAEHSSFVNSCCPTRRGPPLIIS 156
Query: 442 SSFDRSVRLWDASNPIRSLGILTGHDE--QVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
S D + +LWD +R G + + Q+ ++ F + NDV ++W++ +
Sbjct: 157 GSDDGTAKLWD----MRQRGAIQTFPDKYQITAVSFSDAADKIFTGGVDNDV-KVWDLRK 211
Query: 500 SECMHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVET-----------DSHLHDL 545
E T G + P LL + N + + D+ + H H+
Sbjct: 212 GEATMTLEGHQDTITGMSLSPDGSYLLTNGMDNKLCVWDMRPYAPQNRCVKIFEGHQHNF 271
Query: 546 KGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHP 601
+K++L W G + + S D IW S + I +L C+FHP
Sbjct: 272 ---EKNLLKCSWSPDGTKVTAGSSDRMVHIWDTTS-RRTIYKLPGHTGSVNECVFHP 324
Score = 33.1 bits (74), Expect = 0.51
Identities = 16/75 (21%), Positives = 33/75 (43%)
Query: 376 GCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPG 435
G H +L C +S DG + + ++ V IW+ + + H+ + + F P
Sbjct: 266 GHQHNFEKNLLKCSWSPDGTKVTAGSSDRMVHIWDTTSRRTIYKLPGHTGSVNECVFHPT 325
Query: 436 STMFATSSFDRSVRL 450
+ + S D+++ L
Sbjct: 326 EPIIGSCSSDKNIYL 340
Score = 30.4 bits (67), Expect = 3.3
Identities = 11/33 (33%), Positives = 19/33 (57%)
Query: 629 IAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
++ H + + +P G LIAS SHD + +W+
Sbjct: 49 LSGHPSAVYTMKFNPAGTLIASGSHDREIFLWR 81
>At5g67320 unknown protein
Length = 613
Score = 73.9 bits (180), Expect = 3e-13
Identities = 94/445 (21%), Positives = 173/445 (38%), Gaps = 73/445 (16%)
Query: 258 KRKNKHPKIPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQSQNVEKSTKRMITESTRP 317
+++ + KI + + + +KME+ I E +D+ + + +E+ +R E +
Sbjct: 155 EKEREREKIER--EKEREREKMEREIFERE----KDRLKLEKEREIEREREREKIEREKS 208
Query: 318 VESAQDCADAK----PADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQ 373
E AD + D+ + S EP D + P S I +
Sbjct: 209 HEKQLGDADREMVIDQTDKEIAGDGSTGAEPMDIVMTPTSQTSHIPNS------------ 256
Query: 374 EIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFA--------------- 418
++ L ++V C +S ++AS + IW++ + A
Sbjct: 257 DVRILEGHTSEVCACAWSPSASLLASGSGDATARIWSIPEGSFKAVHTGRNINALILKHA 316
Query: 419 --SADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFH 476
++ S +T + + T+ AT S D R+W + + + L+ H + SL ++
Sbjct: 317 KGKSNEKSKDVTTLDWNGEGTLLATGSCDGQARIWTLNGEL--ISTLSKHKGPIFSLKWN 374
Query: 477 PRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDV 536
+ LL S + +W+V E +F+ G L N+++
Sbjct: 375 KKGDYLLTGSVDRTAV-VWDVKAEEWKQ---------QFEFHSGPTLDVDWRNNVSFATS 424
Query: 537 ETDSHLHDLK-----------GHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCI 584
TDS ++ K GH +V + WD +G+ LAS S+DS A+IW+ +
Sbjct: 425 STDSMIYLCKIGETRPAKTFTGHQGEVNCVKWDPTGSLLASCSDDSTAKIWNI-KQSTFV 483
Query: 585 GELHSVGNKFQSCIF--------HPGYRSLLIIGGYQS-LEAWSPTEGSKTWSIAAHQGL 635
+L + + + +P + L + S ++ W G S H+
Sbjct: 484 HDLREHTKEIYTIRWSPTGPGTNNPNKQLTLASASFDSTVKLWDAELGKMLCSFNGHREP 543
Query: 636 IAGLADSPVGELIASASHDNSVKVW 660
+ LA SP GE IAS S D S+ +W
Sbjct: 544 VYSLAFSPNGEYIASGSLDKSIHIW 568
Score = 68.6 bits (166), Expect = 1e-11
Identities = 56/236 (23%), Positives = 103/236 (42%), Gaps = 17/236 (7%)
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
I L K + + ++ G + + ++ +W+++ ++ + HS DV ++
Sbjct: 358 ISTLSKHKGPIFSLKWNKKGDYLLTGSVDRTAVVWDVKAEEWKQQFEFHSGPTLDVDWR- 416
Query: 435 GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
+ FATSS D + L R TGH +V + + P LL S + ++
Sbjct: 417 NNVSFATSSTDSMIYLCKIGET-RPAKTFTGHQGEVNCVKWDPTG-SLLASCSDDSTAKI 474
Query: 495 WNVNQSECMHTTRGGSKQV---RFQP---------QYGQLLATAIGNSITIIDVETDSHL 542
WN+ QS +H R +K++ R+ P + L + + +++ + D E L
Sbjct: 475 WNIKQSTFVHDLREHTKEIYTIRWSPTGPGTNNPNKQLTLASASFDSTVKLWDAELGKML 534
Query: 543 HDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSC 597
GH + V S+ + +G ++AS S D S IWS +GK + G F+ C
Sbjct: 535 CSFNGHREPVYSLAFSPNGEYIASGSLDKSIHIWS-IKEGKIVKTYTGNGGIFEVC 589
Score = 51.6 bits (122), Expect = 1e-06
Identities = 41/188 (21%), Positives = 73/188 (38%), Gaps = 51/188 (27%)
Query: 394 GKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGST---------MFATSSF 444
G ++AS + IWN++ + H+ I +R+ P A++SF
Sbjct: 460 GSLLASCSDDSTAKIWNIKQSTFVHDLREHTKEIYTIRWSPTGPGTNNPNKQLTLASASF 519
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D +V+LWDA + L GH E V SL F P + + S + I +W++ +
Sbjct: 520 DSTVKLWDAELG-KMLCSFNGHREPVYSLAFSPNG-EYIASGSLDKSIHIWSIKE----- 572
Query: 505 TTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFL 564
G+++ T GN + +CW++ GN +
Sbjct: 573 ---------------GKIVKTYTGNG--------------------GIFEVCWNKEGNKI 597
Query: 565 ASVSEDSA 572
A+ D++
Sbjct: 598 AACFADNS 605
>At2g21390 coatomer alpha subunit
Length = 1218
Score = 71.6 bits (174), Expect = 1e-12
Identities = 63/281 (22%), Positives = 122/281 (42%), Gaps = 29/281 (10%)
Query: 389 HFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSV 448
HF + + S G + K+ +WN + + + H I V+F + ++S D+++
Sbjct: 58 HFHNSQPLFVSGGDDYKIKVWNYKTHRCLFTLLGHLDYIRTVQFHHENPWIVSASDDQTI 117
Query: 449 RLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG 508
R+W+ + + +LTGH+ VM FHP+ DL+ S+ + +R+W++ G
Sbjct: 118 RIWNWQSR-TCISVLTGHNHYVMCASFHPKE-DLVVSASLDQTVRVWDI----------G 165
Query: 509 GSKQVRFQP-----QYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNF 563
K+ P ++ Q+ + G I+ + L+GHD+ V + +
Sbjct: 166 ALKKKSASPADDLMRFSQMNSDLFGGVDAIVK-------YVLEGHDRGVNWASFHPTLPL 218
Query: 564 LASVSED-SARIWSAASDGKC--IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
+ S ++D ++W ++ K + L N S +FH ++ +S+ W
Sbjct: 219 IVSGADDRQVKLW-RMNETKAWEVDTLRGHMNNVSSVMFHAKQDIIVSNSEDKSIRVWDA 277
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
T+ + + LA P L+A A HDN + V+K
Sbjct: 278 TKRTGIQTFRREHDRFWILAVHPEINLLA-AGHDNGMIVFK 317
Score = 55.8 bits (133), Expect = 7e-08
Identities = 59/292 (20%), Positives = 113/292 (38%), Gaps = 40/292 (13%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N+V F + ++ H + +W+ D H + V F +F +
Sbjct: 10 NRVKGLSFHPKRPWILASLHSGVIQLWDYRMGTLIDRFDEHEGPVRGVHFHNSQPLFVSG 69
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
D +++W+ R L L GH + + ++ FH ++ +SD + IR+WN C
Sbjct: 70 GDDYKIKVWNYKTH-RCLFTLLGHLDYIRTVQFHHENPWIVSASD-DQTIRIWNWQSRTC 127
Query: 503 MHTTRGGSKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDR 559
+ G + V F P+ +++ ++ ++ + D+ LK
Sbjct: 128 ISVLTGHNHYVMCASFHPKEDLVVSASLDQTVRVWDIGA------LKKKS---------- 171
Query: 560 SGNFLASVSEDSARIWSAASD--------GKCIGELHSVGNKFQSCIFHPGYRSLLIIGG 611
AS ++D R SD K + E H G + S FHP ++
Sbjct: 172 -----ASPADDLMRFSQMNSDLFGGVDAIVKYVLEGHDRGVNWAS--FHPTLPLIVSGAD 224
Query: 612 YQSLEAWSPTEGSKTW---SIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ ++ W E +K W ++ H ++ + ++I S S D S++VW
Sbjct: 225 DRQVKLWRMNE-TKAWEVDTLRGHMNNVSSVMFHAKQDIIVSNSEDKSIRVW 275
Score = 55.1 bits (131), Expect = 1e-07
Identities = 47/223 (21%), Positives = 88/223 (39%), Gaps = 65/223 (29%)
Query: 382 INKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFAT 441
++ + T F + + SA ++ + IWN ++ + H+H + F P + +
Sbjct: 93 LDYIRTVQFHHENPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCASFHPKEDLVVS 152
Query: 442 SSFDRSVRLWD-------ASNPIRSL-------------------GILTGHDEQVMSLDF 475
+S D++VR+WD +++P L +L GHD V F
Sbjct: 153 ASLDQTVRVWDIGALKKKSASPADDLMRFSQMNSDLFGGVDAIVKYVLEGHDRGVNWASF 212
Query: 476 HPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIID 535
HP + L+ S + ++LW +N+++ +
Sbjct: 213 HP-TLPLIVSGADDRQVKLWRMNETKAW-------------------------------E 240
Query: 536 VETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSA 577
V+T L+GH +V S+ + + + S SED S R+W A
Sbjct: 241 VDT------LRGHMNNVSSVMFHAKQDIIVSNSEDKSIRVWDA 277
>At1g29260 peroxisomal targeting signal type 2 receptor
Length = 317
Score = 71.6 bits (174), Expect = 1e-12
Identities = 61/231 (26%), Positives = 108/231 (46%), Gaps = 17/231 (7%)
Query: 445 DRSVRLWDA-----SNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
D SV+++D SNPIRS H +V S+D++P R D +S +D ++LW +++
Sbjct: 82 DGSVKIYDTALPPPSNPIRSF---QEHAREVQSVDYNPTRRDSFLTSSWDDTVKLWAMDR 138
Query: 500 SECMHTTRGGS---KQVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSI 555
+ T + + Q + P++G + A+A G+ ++ I DV + HD ++LS
Sbjct: 139 PASVRTFKEHAYCVYQAVWNPKHGDVFASASGDCTLRIWDVREPGSTMIIPAHDFEILSC 198
Query: 556 CWDRSGNFLASVS--EDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGY- 612
W++ + + + S + + ++W S + L+ G + F P RSL+ Y
Sbjct: 199 DWNKYDDCILATSSVDKTVKVWDVRSYRVPLAVLNGHGYAVRKVKFSPHRRSLIASCSYD 258
Query: 613 QSLEAWS-PTEGSKTWSIAAHQGLIAGLADSPVGE-LIASASHDNSVKVWK 661
S+ W E + H G+ S + E L+AS D V VW+
Sbjct: 259 MSVCLWDYMVEDALVGRYDHHTEFAVGIDMSVLVEGLMASTGWDELVYVWQ 309
Score = 50.8 bits (120), Expect = 2e-06
Identities = 36/111 (32%), Positives = 51/111 (45%), Gaps = 23/111 (20%)
Query: 408 IWNMENFQYFASADTHSHL-ITDVRFQPGSTMF---------------------ATSSFD 445
+WN ++ FASA L I DVR +PGSTM ATSS D
Sbjct: 156 VWNPKHGDVFASASGDCTLRIWDVR-EPGSTMIIPAHDFEILSCDWNKYDDCILATSSVD 214
Query: 446 RSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWN 496
++V++WD + L +L GH V + F P R L+ S + + LW+
Sbjct: 215 KTVKVWDVRSYRVPLAVLNGHGYAVRKVKFSPHRRSLIASCSYDMSVCLWD 265
Score = 46.6 bits (109), Expect = 4e-05
Identities = 27/115 (23%), Positives = 61/115 (52%), Gaps = 3/115 (2%)
Query: 384 KVLTCHFSS-DGKVMASAGHEKKVFIWNMENFQY-FASADTHSHLITDVRFQPGS-TMFA 440
++L+C ++ D ++A++ +K V +W++ +++ A + H + + V+F P ++ A
Sbjct: 194 EILSCDWNKYDDCILATSSVDKTVKVWDVRSYRVPLAVLNGHGYAVRKVKFSPHRRSLIA 253
Query: 441 TSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLW 495
+ S+D SV LWD +G H E + +D L+ S+ ++++ +W
Sbjct: 254 SCSYDMSVCLWDYMVEDALVGRYDHHTEFAVGIDMSVLVEGLMASTGWDELVYVW 308
>At5g25150 transcription initiation factor IID-associated
factor-like protein
Length = 700
Score = 70.9 bits (172), Expect = 2e-12
Identities = 66/282 (23%), Positives = 110/282 (38%), Gaps = 62/282 (21%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + FS G + S+ + + +W+ + H++ + D +F P FA+ S
Sbjct: 421 VYSATFSPPGDFVLSSSADTTIRLWSTKLNANLVCYKGHNYPVWDAQFSPFGHYFASCSH 480
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
DR+ R+W I+ L I+ GH + +D+HP + + + S+ +RLW
Sbjct: 481 DRTARIWSMDR-IQPLRIMAGH---LSDVDWHPN-CNYIATGSSDKTVRLW--------- 526
Query: 505 TTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFL 564
DV+T + GH VLS+ G ++
Sbjct: 527 ------------------------------DVQTGECVRIFIGHRSMVLSLAMSPDGRYM 556
Query: 565 ASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGS 624
AS ED + S +CI L SC++ YR ++ GY W G
Sbjct: 557 ASGDEDGTIMMWDLSTARCITPLMG----HNSCVWSLSYR---VVQGYDI--EW---YGF 604
Query: 625 KTWSIAAHQGLIAGLADSPV------GELIASASHDNSVKVW 660
+W + GL+ + + G L+AS S D +VK+W
Sbjct: 605 LSWVVECCIGLMCWVVNGMTGFGCGEGSLLASGSADCTVKLW 646
Score = 29.6 bits (65), Expect = 5.6
Identities = 21/94 (22%), Positives = 37/94 (39%), Gaps = 21/94 (22%)
Query: 568 SEDSARIWSAASDGKC-IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKT 626
S+ S ++W A G+ G L + + I G RS ++ G
Sbjct: 372 SDSSIKVWDMAKIGQAGSGALQAENDSSDQSIGPNGRRSYTLLLG--------------- 416
Query: 627 WSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
H G + SP G+ + S+S D ++++W
Sbjct: 417 -----HSGPVYSATFSPPGDFVLSSSADTTIRLW 445
>At1g48630 unknown protein
Length = 326
Score = 68.9 bits (167), Expect = 8e-12
Identities = 62/218 (28%), Positives = 103/218 (46%), Gaps = 31/218 (14%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNM--ENFQYFASADTHSHLITDVRFQPGSTM--FA 440
VL+ FS+D + + SA ++ + +WN E + AD H ++ VRF P + +
Sbjct: 108 VLSVAFSTDNRQIVSASRDRTIKLWNTLGECKYTISEADGHKEWVSCVRFSPNTLVPTIV 167
Query: 441 TSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSND-VIRLWNVNQ 499
++S+D++V++W+ N + L GH + ++ P LC+S D VI LW++ +
Sbjct: 168 SASWDKTVKVWNLQN-CKLRNTLAGHSGYLNTVAVSPDGS--LCASGGKDGVILLWDLAE 224
Query: 500 SECMHTTRGGS--KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLK----------- 546
+ +++ GS + F P L A A NSI I D+E+ S + DLK
Sbjct: 225 GKKLYSLEAGSIIHSLCFSPNRYWLCA-ATENSIRIWDLESKSVVEDLKVDLKAEAEKTD 283
Query: 547 -----GHDKDVL---SICWDRSGNFLASVSEDSA-RIW 575
G+ V+ S+ W GN L S D R+W
Sbjct: 284 GSTGIGNKTKVIYCTSLNWSADGNTLFSGYTDGVIRVW 321
Score = 57.8 bits (138), Expect = 2e-08
Identities = 43/195 (22%), Positives = 90/195 (46%), Gaps = 8/195 (4%)
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDG+ S + ++ +W++ + H+ + V F + ++S DR+++L
Sbjct: 72 SSDGQFALSGSWDGELRLWDLATGESTRRFVGHTKDVLSVAFSTDNRQIVSASRDRTIKL 131
Query: 451 WDASNPIR-SLGILTGHDEQVMSLDFHPRRM-DLLCSSDSNDVIRLWNVNQSECMHTTRG 508
W+ + ++ GH E V + F P + + S+ + +++WN+ + +T G
Sbjct: 132 WNTLGECKYTISEADGHKEWVSCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNTLAG 191
Query: 509 GS---KQVRFQPQYGQLLATAIGNSITII-DVETDSHLHDLKGHDKDVLSICWDRSGNFL 564
S V P G L A+ + + ++ D+ L+ L+ + S+C+ + +L
Sbjct: 192 HSGYLNTVAVSPD-GSLCASGGKDGVILLWDLAEGKKLYSLEA-GSIIHSLCFSPNRYWL 249
Query: 565 ASVSEDSARIWSAAS 579
+ +E+S RIW S
Sbjct: 250 CAATENSIRIWDLES 264
Score = 57.4 bits (137), Expect = 3e-08
Identities = 61/265 (23%), Positives = 113/265 (42%), Gaps = 21/265 (7%)
Query: 368 KGFSLQEIGCLHKS-INKVLTCHFSSDGKVMASAGHEKKVFIWNM--ENFQYFASADT-- 422
+G L+ C H + + T +SD V+ ++ +K + +W + E+ Y +
Sbjct: 3 EGLVLKGTMCAHTDMVTAIATPVDNSD--VIVTSSRDKSIILWKLTKEDKSYGVAQRRMT 60
Query: 423 -HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMD 481
HSH + DV + S+D +RLWD + S GH + V+S+ F
Sbjct: 61 GHSHFVQDVVLSSDGQFALSGSWDGELRLWDLATG-ESTRRFVGHTKDVLSVAFSTDNRQ 119
Query: 482 LLCSSDSNDVIRLWNVNQSECMHT--TRGGSKQ----VRFQPQ--YGQLLATAIGNSITI 533
++ S+ + I+LWN EC +T G K+ VRF P +++ + ++ +
Sbjct: 120 IV-SASRDRTIKLWN-TLGECKYTISEADGHKEWVSCVRFSPNTLVPTIVSASWDKTVKV 177
Query: 534 IDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNK 593
+++ + L GH + ++ G+ AS +D + ++GK + L + G+
Sbjct: 178 WNLQNCKLRNTLAGHSGYLNTVAVSPDGSLCASGGKDGVILLWDLAEGKKLYSLEA-GSI 236
Query: 594 FQSCIFHPGYRSLLIIGGYQSLEAW 618
S F P R L S+ W
Sbjct: 237 IHSLCFSPN-RYWLCAATENSIRIW 260
Score = 52.0 bits (123), Expect = 1e-06
Identities = 55/248 (22%), Positives = 96/248 (38%), Gaps = 51/248 (20%)
Query: 423 HSHLITDVRFQ-PGSTMFATSSFDRSVRLWDASNPIRSLGI----LTGHDEQVMSLDFHP 477
H+ ++T + S + TSS D+S+ LW + +S G+ +TGH V
Sbjct: 14 HTDMVTAIATPVDNSDVIVTSSRDKSIILWKLTKEDKSYGVAQRRMTGHSHFVQ------ 67
Query: 478 RRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQL-LATAIGNSITIIDV 536
D++ SSD GQ L+ + + + D+
Sbjct: 68 ---DVVLSSD--------------------------------GQFALSGSWDGELRLWDL 92
Query: 537 ETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQ 595
T GH KDVLS+ + + S S D + ++W+ + K ++
Sbjct: 93 ATGESTRRFVGHTKDVLSVAFSTDNRQIVSASRDRTIKLWNTLGECKYTISEADGHKEWV 152
Query: 596 SCI-FHPGYRSLLIIGGY--QSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASAS 652
SC+ F P I+ ++++ W+ ++A H G + +A SP G L AS
Sbjct: 153 SCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNTLAGHSGYLNTVAVSPDGSLCASGG 212
Query: 653 HDNSVKVW 660
D + +W
Sbjct: 213 KDGVILLW 220
>At2g33340 putative PRP19-like spliceosomal protein
Length = 565
Score = 68.6 bits (166), Expect = 1e-11
Identities = 54/208 (25%), Positives = 99/208 (46%), Gaps = 13/208 (6%)
Query: 378 LHKSINKVLTCHFS--SDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPG 435
LHK+ NK C V+A+ G + +++ + Q ++ HS +T V+F
Sbjct: 217 LHKT-NKPGICSMDILHSKDVIATGGVDATAVLFDRPSGQILSTLTGHSKKVTSVKFVGD 275
Query: 436 STMFATSSFDRSVRLW-DASNPIRSLGI-LTGHDEQVMSLDFHPRRMDLLCSSDSNDVIR 493
S + T+S D++VR+W + + + G L H +V ++ HP + S+ +
Sbjct: 276 SDLVLTASADKTVRIWRNPGDGNYACGYTLNDHSAEVRAVTVHPTNKYFV-SASLDGTWC 334
Query: 494 LWNVNQSECMHTTRGGSKQV-----RFQPQYGQLLATAIGNSITII-DVETDSHLHDLKG 547
++++ C+ SK V F P G +L T S+ I DV++ +++ G
Sbjct: 335 FYDLSSGSCLAQVSDDSKNVDYTAAAFHPD-GLILGTGTSQSVVKIWDVKSQANVAKFDG 393
Query: 548 HDKDVLSICWDRSGNFLASVSEDSARIW 575
H +V +I + +G FLA+ +ED R+W
Sbjct: 394 HTGEVTAISFSENGYFLATAAEDGVRLW 421
>At1g62020 hypothetical protein
Length = 1216
Score = 68.2 bits (165), Expect = 1e-11
Identities = 62/281 (22%), Positives = 121/281 (42%), Gaps = 29/281 (10%)
Query: 389 HFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSV 448
HF + + S G + K+ +WN +N + + H I V+F ++S D+++
Sbjct: 58 HFHNSQPLFVSGGDDYKIKVWNYKNHRCLFTLLGHLDYIRTVQFHHEYPWIVSASDDQTI 117
Query: 449 RLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG 508
R+W+ + + +LTGH+ VM FHP+ DL+ S+ + +R+W++ G
Sbjct: 118 RIWNWQSR-TCVSVLTGHNHYVMCASFHPKE-DLVVSASLDQTVRVWDI----------G 165
Query: 509 GSKQVRFQP-----QYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNF 563
++ P + Q+ + G I+ + L+GHD+ V + +
Sbjct: 166 ALRKKTVSPADDIMRLTQMNSDLFGGVDAIVK-------YVLEGHDRGVNWAAFHPTLPL 218
Query: 564 LASVSED-SARIWSAASDGKC--IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
+ S ++D ++W ++ K + L N S +FH ++ +S+ W
Sbjct: 219 IVSGADDRQVKLW-RMNETKAWEVDTLRGHMNNVSSVMFHAKQDIIVSNSEDKSIRVWDA 277
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
T+ + + LA P L+A A HD+ + V+K
Sbjct: 278 TKRTGLQTFRREHDRFWILAVHPEMNLLA-AGHDSGMIVFK 317
Score = 51.2 bits (121), Expect = 2e-06
Identities = 47/223 (21%), Positives = 87/223 (38%), Gaps = 65/223 (29%)
Query: 382 INKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFAT 441
++ + T F + + SA ++ + IWN ++ + H+H + F P + +
Sbjct: 93 LDYIRTVQFHHEYPWIVSASDDQTIRIWNWQSRTCVSVLTGHNHYVMCASFHPKEDLVVS 152
Query: 442 SSFDRSVRLWD----------ASNPIRSLG----------------ILTGHDEQVMSLDF 475
+S D++VR+WD ++ I L +L GHD V F
Sbjct: 153 ASLDQTVRVWDIGALRKKTVSPADDIMRLTQMNSDLFGGVDAIVKYVLEGHDRGVNWAAF 212
Query: 476 HPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIID 535
HP + L+ S + ++LW +N+++ +
Sbjct: 213 HP-TLPLIVSGADDRQVKLWRMNETKAW-------------------------------E 240
Query: 536 VETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSA 577
V+T L+GH +V S+ + + + S SED S R+W A
Sbjct: 241 VDT------LRGHMNNVSSVMFHAKQDIIVSNSEDKSIRVWDA 277
>At5g60940 cleavage stimulation factor 50K chain
Length = 429
Score = 67.8 bits (164), Expect = 2e-11
Identities = 79/336 (23%), Positives = 137/336 (40%), Gaps = 38/336 (11%)
Query: 353 SNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNME 412
++ K S T ++E+K S HKS+ V FS DG A+ G + + ++ +
Sbjct: 102 NHAKGSSKTIPKHESKTLSE------HKSV--VRCARFSPDGMFFATGGADTSIKLFEVP 153
Query: 413 NFQYFASADT-----------HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLG 461
+ S DT H+ I D+ F P ST+ +S+ D ++ +D S
Sbjct: 154 KVKQMISGDTQARPLIRTFYDHAEPINDLDFHPRSTILISSAKDNCIKFFDFSKTTAKRA 213
Query: 462 ILTGHD-EQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTR-------GGSKQV 513
D V S+ FHP LL +D + + L++VN +C + G QV
Sbjct: 214 FKVFQDTHNVRSISFHPSGEFLLAGTD-HPIPHLYDVNTYQCFLPSNFPDSGVSGAINQV 272
Query: 514 RFQPQYGQLLATAIGNSITIIDVETDSHLHDL-KGHDK-DVLSICWDRSGNFLASVSEDS 571
R+ + + +I + D + + + H K +V S + + F+ S +DS
Sbjct: 273 RYSSTGSIYITASKDGAIRLFDGVSAKCVRSIGNAHGKSEVTSAVFTKDQRFVLSSGKDS 332
Query: 572 -ARIWSAASDGKCIGE-LHSVGNKFQS-CIFHPGYRSLLIIG-GYQSLEAWSPTEGSKT- 626
++W S G+ + E L + K +S IF+ ++ I + W K
Sbjct: 333 TVKLWEIGS-GRMVKEYLGAKRVKLRSQAIFNDTEEFVISIDEASNEVVTWDARTADKVA 391
Query: 627 -WSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
W + H G + SPV + + D S++ WK
Sbjct: 392 KWP-SNHNGAPRWIEHSPVESVFVTCGIDRSIRFWK 426
>At3g18130 protein kinase C-receptor/G-protein, putative
Length = 326
Score = 67.8 bits (164), Expect = 2e-11
Identities = 61/218 (27%), Positives = 103/218 (46%), Gaps = 31/218 (14%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNM--ENFQYFASADTHSHLITDVRFQPGSTM--FA 440
VL+ FS+D + + SA ++ + +WN E + D H ++ VRF P + +
Sbjct: 108 VLSVAFSTDNRQIVSASRDRTIKLWNTLGECKYTISEGDGHKEWVSCVRFSPNTLVPTIV 167
Query: 441 TSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSND-VIRLWNVNQ 499
++S+D++V++W+ N + L GH + ++ P LC+S D VI LW++ +
Sbjct: 168 SASWDKTVKVWNLQN-CKLRNSLVGHSGYLNTVAVSPD--GSLCASGGKDGVILLWDLAE 224
Query: 500 SECMHTTRGGS--KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLK----------- 546
+ +++ GS + F P L A A NSI I D+E+ S + DLK
Sbjct: 225 GKKLYSLEAGSIIHSLCFSPNRYWLCA-ATENSIRIWDLESKSVVEDLKVDLKSEAEKNE 283
Query: 547 -----GHDKDVL---SICWDRSGNFLASVSEDS-ARIW 575
G+ K V+ S+ W G+ L S D R+W
Sbjct: 284 GGVGTGNQKKVIYCTSLNWSADGSTLFSGYTDGVVRVW 321
Score = 55.1 bits (131), Expect = 1e-07
Identities = 51/236 (21%), Positives = 100/236 (41%), Gaps = 18/236 (7%)
Query: 396 VMASAGHEKKVFIWNM-ENFQYFASADT----HSHLITDVRFQPGSTMFATSSFDRSVRL 450
++ +A +K + +W + ++ + + A HSH + DV + S+D +RL
Sbjct: 30 IIVTASRDKSIILWKLTKDDKSYGVAQRRLTGHSHFVEDVVLSSDGQFALSGSWDGELRL 89
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WD + + GH + V+S+ F ++ S+ + I+LWN EC +T G
Sbjct: 90 WDLATG-ETTRRFVGHTKDVLSVAFSTDNRQIV-SASRDRTIKLWN-TLGECKYTISEGD 146
Query: 511 KQ------VRFQPQ--YGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGN 562
VRF P +++ + ++ + +++ + L GH + ++ G+
Sbjct: 147 GHKEWVSCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNSLVGHSGYLNTVAVSPDGS 206
Query: 563 FLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAW 618
AS +D + ++GK + L + G+ S F P R L S+ W
Sbjct: 207 LCASGGKDGVILLWDLAEGKKLYSLEA-GSIIHSLCFSPN-RYWLCAATENSIRIW 260
Score = 54.3 bits (129), Expect = 2e-07
Identities = 42/195 (21%), Positives = 90/195 (45%), Gaps = 8/195 (4%)
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDG+ S + ++ +W++ + H+ + V F + ++S DR+++L
Sbjct: 72 SSDGQFALSGSWDGELRLWDLATGETTRRFVGHTKDVLSVAFSTDNRQIVSASRDRTIKL 131
Query: 451 WDASNPIR-SLGILTGHDEQVMSLDFHPRRM-DLLCSSDSNDVIRLWNVNQSECMHTTRG 508
W+ + ++ GH E V + F P + + S+ + +++WN+ + ++ G
Sbjct: 132 WNTLGECKYTISEGDGHKEWVSCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNSLVG 191
Query: 509 GS---KQVRFQPQYGQLLATAIGNSITII-DVETDSHLHDLKGHDKDVLSICWDRSGNFL 564
S V P G L A+ + + ++ D+ L+ L+ + S+C+ + +L
Sbjct: 192 HSGYLNTVAVSPD-GSLCASGGKDGVILLWDLAEGKKLYSLEA-GSIIHSLCFSPNRYWL 249
Query: 565 ASVSEDSARIWSAAS 579
+ +E+S RIW S
Sbjct: 250 CAATENSIRIWDLES 264
Score = 50.1 bits (118), Expect = 4e-06
Identities = 55/248 (22%), Positives = 95/248 (38%), Gaps = 51/248 (20%)
Query: 423 HSHLITDVRFQ-PGSTMFATSSFDRSVRLWDASNPIRSLGI----LTGHDEQVMSLDFHP 477
H+ ++T + S + T+S D+S+ LW + +S G+ LTGH V
Sbjct: 14 HTDIVTAIATPIDNSDIIVTASRDKSIILWKLTKDDKSYGVAQRRLTGHSHFVE------ 67
Query: 478 RRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQL-LATAIGNSITIIDV 536
D++ SSD GQ L+ + + + D+
Sbjct: 68 ---DVVLSSD--------------------------------GQFALSGSWDGELRLWDL 92
Query: 537 ETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQ 595
T GH KDVLS+ + + S S D + ++W+ + K ++
Sbjct: 93 ATGETTRRFVGHTKDVLSVAFSTDNRQIVSASRDRTIKLWNTLGECKYTISEGDGHKEWV 152
Query: 596 SCI-FHPGYRSLLIIGGY--QSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASAS 652
SC+ F P I+ ++++ W+ S+ H G + +A SP G L AS
Sbjct: 153 SCVRFSPNTLVPTIVSASWDKTVKVWNLQNCKLRNSLVGHSGYLNTVAVSPDGSLCASGG 212
Query: 653 HDNSVKVW 660
D + +W
Sbjct: 213 KDGVILLW 220
>At3g15980 putative coatomer complex subunit
Length = 921
Score = 62.4 bits (150), Expect = 8e-10
Identities = 43/178 (24%), Positives = 75/178 (41%), Gaps = 6/178 (3%)
Query: 408 IWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHD 467
++N + HS I V P +SS D ++LWD N I GH
Sbjct: 86 VYNYNTMDKVKVFEAHSDYIRCVAVHPTLPYVLSSSDDMLIKLWDWENGWACTQIFEGHS 145
Query: 468 EQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQY-----GQL 522
VM + F+P+ + S+ + I++WN+ + T K V + L
Sbjct: 146 HYVMQVVFNPKDTNTFASASLDRTIKIWNLGSPDPNFTLDAHQKGVNCVDYFTGGDKPYL 205
Query: 523 LATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAAS 579
+ + ++ + D +T S + L GH +V ++C+ + + SED + RIW A +
Sbjct: 206 ITGSDDHTAKVWDYQTKSCVQTLDGHTHNVSAVCFHPELPIIITGSEDGTVRIWHATT 263
Score = 43.1 bits (100), Expect = 5e-04
Identities = 28/123 (22%), Positives = 53/123 (42%), Gaps = 4/123 (3%)
Query: 393 DGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPG--STMFATSSFDRSVRL 450
D ASA ++ + IWN+ + + D H + V + G T S D + ++
Sbjct: 157 DTNTFASASLDRTIKIWNLGSPDPNFTLDAHQKGVNCVDYFTGGDKPYLITGSDDHTAKV 216
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WD + L GH V ++ FHP + ++ + + +R+W+ +T G
Sbjct: 217 WDYQTK-SCVQTLDGHTHNVSAVCFHP-ELPIIITGSEDGTVRIWHATTYRLENTLNYGL 274
Query: 511 KQV 513
++V
Sbjct: 275 ERV 277
>At4g15900 PRL1 protein
Length = 486
Score = 62.0 bits (149), Expect = 1e-09
Identities = 45/182 (24%), Positives = 76/182 (41%), Gaps = 5/182 (2%)
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H + V F P + F T S DR++++WD + + L LTGH EQV L R +
Sbjct: 175 HLGWVRSVAFDPSNEWFCTGSADRTIKIWDVATGVLKL-TLTGHIEQVRGLAVSNRHTYM 233
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRG---GSKQVRFQPQYGQLLATAIGNSITIIDVETD 539
+ D V + W++ Q++ + + G G + P LL + + D+ T
Sbjct: 234 FSAGDDKQV-KCWDLEQNKVIRSYHGHLSGVYCLALHPTLDVLLTGGRDSVCRVWDIRTK 292
Query: 540 SHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIF 599
+ L GHD V S+ + + + S D+ + GK + L ++
Sbjct: 293 MQIFALSGHDNTVCSVFTRPTDPQVVTGSHDTTIKFWDLRYGKTMSTLTHHKKSVRAMTL 352
Query: 600 HP 601
HP
Sbjct: 353 HP 354
Score = 48.5 bits (114), Expect = 1e-05
Identities = 54/269 (20%), Positives = 101/269 (37%), Gaps = 42/269 (15%)
Query: 397 MASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNP 456
M SAG +K+V W++E + S H + + P + T D R+WD
Sbjct: 233 MFSAGDDKQVKCWDLEQNKVIRSYHGHLSGVYCLALHPTLDVLLTGGRDSVCRVWDIRTK 292
Query: 457 IRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQ 516
++ + L+GHD V S+ P ++ S + I+ W++ + M T K VR
Sbjct: 293 MQ-IFALSGHDNTVCSVFTRPTDPQVVTGS-HDTTIKFWDLRYGKTMSTLTHHKKSVRAM 350
Query: 517 PQYGQ--LLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARI 574
+ + A+A ++ + H++ K +++ +V+ED +
Sbjct: 351 TLHPKENAFASASADNTKKFSLPKGEFCHNMLSQQKTIIN---------AMAVNEDGVMV 401
Query: 575 WSAASDGKCIGELHSVGNKFQ--SCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH 632
+ + S G+ FQ I PG S+ +
Sbjct: 402 TGGDNGSIWFWDWKS-GHSFQQSETIVQPG-------------------------SLESE 435
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVWK 661
G+ A D+ G + + D ++K+WK
Sbjct: 436 AGIYAACYDN-TGSRLVTCEADKTIKMWK 463
Score = 43.9 bits (102), Expect = 3e-04
Identities = 45/210 (21%), Positives = 72/210 (33%), Gaps = 42/210 (20%)
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
W A P ++ ++ GH V S+ F P + C+ ++ I++W+V T G
Sbjct: 162 WHA--PWKNYRVIQGHLGWVRSVAFDPSN-EWFCTGSADRTIKIWDVATGVLKLTLTGHI 218
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
+QVR A+ N T + G DK V CWD N
Sbjct: 219 EQVR---------GLAVSNRHTYMFSA---------GDDKQVK--CWDLEQN-------- 250
Query: 571 SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIA 630
K I H + HP LL G W + ++++
Sbjct: 251 -----------KVIRSYHGHLSGVYCLALHPTLDVLLTGGRDSVCRVWDIRTKMQIFALS 299
Query: 631 AHQGLIAGLADSPVGELIASASHDNSVKVW 660
H + + P + + SHD ++K W
Sbjct: 300 GHDNTVCSVFTRPTDPQVVTGSHDTTIKFW 329
>At1g52360 coatomer complex subunit, putative
Length = 926
Score = 62.0 bits (149), Expect = 1e-09
Identities = 44/201 (21%), Positives = 84/201 (40%), Gaps = 6/201 (2%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + F + + + + + + ++N + HS I V P +SS
Sbjct: 60 VRSAKFVARKQWVVAGADDMYIRVYNYNTMDKVKVFEAHSDYIRCVAVHPTLPYVLSSSD 119
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D ++LWD I GH VM + F+P+ + S+ + I++WN+ +
Sbjct: 120 DMLIKLWDWEKGWACTQIFEGHSHYVMQVTFNPKDTNTFASASLDRTIKIWNLGSPDPNF 179
Query: 505 TTRGGSKQVRFQPQY-----GQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDR 559
T K V + L+ + ++ + D +T S + L+GH +V ++C+
Sbjct: 180 TLDAHQKGVNCVDYFTGGDKPYLITGSDDHTAKVWDYQTKSCVQTLEGHTHNVSAVCFHP 239
Query: 560 SGNFLASVSED-SARIWSAAS 579
+ + SED + RIW A +
Sbjct: 240 ELPIIITGSEDGTVRIWHATT 260
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.130 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,983,172
Number of Sequences: 26719
Number of extensions: 728223
Number of successful extensions: 3140
Number of sequences better than 10.0: 209
Number of HSP's better than 10.0 without gapping: 113
Number of HSP's successfully gapped in prelim test: 96
Number of HSP's that attempted gapping in prelim test: 2041
Number of HSP's gapped (non-prelim): 748
length of query: 661
length of database: 11,318,596
effective HSP length: 106
effective length of query: 555
effective length of database: 8,486,382
effective search space: 4709942010
effective search space used: 4709942010
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0114.1


