

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.7
(147 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
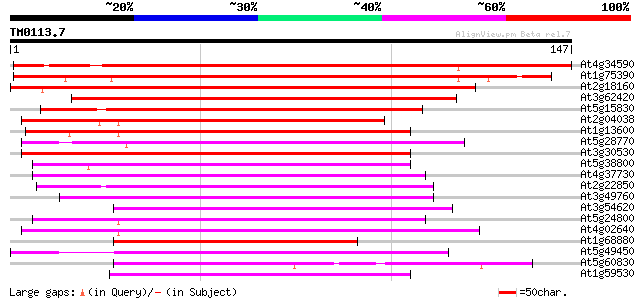
Score E
Sequences producing significant alignments: (bits) Value
At4g34590 bZIP transcription factor ATB2/ Atbzip11 148 1e-36
At1g75390 bZip transcription factor AtbZip44 135 8e-33
At2g18160 G-box binding bZIP transcription factor GBF5 / AtbZip2 127 2e-30
At3g62420 bZip transcription factor AtbZip53 89 6e-19
At5g15830 bZIP transcription factor AtbZip3 84 3e-17
At2g04038 bZip transcription factor AtbZip48 82 1e-16
At1g13600 bZip transcription factor AtbZip58 81 2e-16
At5g28770 bZip transcription factor BZO2H3 / AtbZip63 80 5e-16
At3g30530 bZip transcription factor AtbZip42 78 2e-15
At5g38800 bZIP transcription factor AtbZip43 75 2e-14
At4g37730 bZip transcription factor AtbZip7 70 4e-13
At2g22850 bZip tarnscription factor AtbZip6 66 6e-12
At3g49760 bZIP transcription factor AtbZip5 64 2e-11
At3g54620 bZIP transcription factor-like protein 62 8e-11
At5g24800 bZIP protein BZO2H2 60 5e-10
At4g02640 bZIP protein BZO2H1 60 5e-10
At1g68880 bZip transcription factor AtbZip8 60 5e-10
At5g49450 bZip protein AtbZIP1 57 3e-09
At5g60830 bZip transcription factor AtbZip70 55 1e-08
At1g59530 bZip transcription factor AtbZip4 55 1e-08
>At4g34590 bZIP transcription factor ATB2/ Atbzip11
Length = 159
Score = 148 bits (373), Expect = 1e-36
Identities = 81/161 (50%), Positives = 119/161 (73%), Gaps = 19/161 (11%)
Query: 2 ASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVA 61
+SSSGT+S S++ +SGSEE +LM+QRKRKRM+SNRESARRSRM+KQK LD+L AQV
Sbjct: 3 SSSSGTTS-STIQTSSGSEE---SLMEQRKRKRMLSNRESARRSRMKKQKLLDDLTAQVN 58
Query: 62 QLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATN----- 116
L+ EN +++TS+++TTQ ++ ++AENSVL AQ+ EL+HRL+SLN+II+F++++N
Sbjct: 59 HLKKENTEIVTSVSITTQHYLTVEAENSVLRAQLDELNHRLQSLNDIIEFLDSSNNNNNN 118
Query: 117 ----------GVFAANSFVDPIPSAYYLNHPIMASADILQY 147
G+ + FV+ + +Y +N P+MAS+D L Y
Sbjct: 119 NMGMCSNPLVGLECDDFFVNQMNMSYIMNQPLMASSDALMY 159
>At1g75390 bZip transcription factor AtbZip44
Length = 173
Score = 135 bits (340), Expect = 8e-33
Identities = 80/157 (50%), Positives = 108/157 (67%), Gaps = 17/157 (10%)
Query: 2 ASSSGTSSGSSM--IQNSGSEEELQA--LMDQRKRKRMISNRESARRSRMRKQKHLDELA 57
+S+SG S S + NSGSE +L+ L+D+RKRKR SNRESARRSRMRKQKHLD+L
Sbjct: 9 SSTSGNCSSVSTTGLANSGSESDLRQRDLIDERKRKRKQSNRESARRSRMRKQKHLDDLT 68
Query: 58 AQVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATN- 116
AQV LR EN Q++ + +TTQ ++ I+AEN +L AQ+ EL+HRL+SLNEI+DFV +++
Sbjct: 69 AQVTHLRKENAQIVAGIAVTTQHYVTIEAENDILRAQVLELNHRLQSLNEIVDFVESSSS 128
Query: 117 --------GVFAANSF---VDPIPSAYYLNHPIMASA 142
G+F F ++P+ +Y N PIMASA
Sbjct: 129 GFGMETGQGLFDGGLFDGVMNPMNLGFY-NQPIMASA 164
>At2g18160 G-box binding bZIP transcription factor GBF5 / AtbZip2
Length = 171
Score = 127 bits (319), Expect = 2e-30
Identities = 69/123 (56%), Positives = 90/123 (73%), Gaps = 1/123 (0%)
Query: 1 MASSSGT-SSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQ 59
MASSS T S SS + + + +D+RKRKRM+SNRESARRSRMRKQKH+D+L AQ
Sbjct: 1 MASSSSTYRSSSSSDGGNNNPSDSVVTVDERKRKRMLSNRESARRSRMRKQKHVDDLTAQ 60
Query: 60 VAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVF 119
+ QL ++N+Q+L SL +T+Q +M I AENSVL+AQM ELS RL+SLNEI+D V + F
Sbjct: 61 INQLSNDNRQILNSLTVTSQLYMKIQAENSVLTAQMEELSTRLQSLNEIVDLVQSNGAGF 120
Query: 120 AAN 122
+
Sbjct: 121 GVD 123
>At3g62420 bZip transcription factor AtbZip53
Length = 146
Score = 89.4 bits (220), Expect = 6e-19
Identities = 43/101 (42%), Positives = 71/101 (69%)
Query: 17 SGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNL 76
S ++ + D+RKRKRMISNRESARRSRMRKQK L +L +V L+++N ++ ++
Sbjct: 12 SDNDPRYATVTDERKRKRMISNRESARRSRMRKQKQLGDLINEVTLLKNDNAKITEQVDE 71
Query: 77 TTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNG 117
+++++ ++++N+VL AQ EL+ RL SLN +++ V +G
Sbjct: 72 ASKKYIEMESKNNVLRAQASELTDRLRSLNSVLEMVEEISG 112
>At5g15830 bZIP transcription factor AtbZip3
Length = 186
Score = 84.0 bits (206), Expect = 3e-17
Identities = 46/100 (46%), Positives = 67/100 (67%), Gaps = 2/100 (2%)
Query: 9 SGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQ 68
S +S + +EE ++++RK++RM+SNRESARRSRMRKQ+HLDEL +QVA LRSEN
Sbjct: 55 SNNSTTSDDATEEIF--VINERKQRRMVSNRESARRSRMRKQRHLDELLSQVAWLRSENH 112
Query: 69 QMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEI 108
Q+L LN + ++ ENS L + EL + S+ ++
Sbjct: 113 QLLDKLNQVSDNNDLVIQENSSLKEENLELRQVITSMKKL 152
>At2g04038 bZip transcription factor AtbZip48
Length = 166
Score = 81.6 bits (200), Expect = 1e-16
Identities = 46/99 (46%), Positives = 68/99 (68%), Gaps = 4/99 (4%)
Query: 4 SSGTSSGSSMIQNSGSEEE--LQALM--DQRKRKRMISNRESARRSRMRKQKHLDELAAQ 59
SS ++S MI N+ + +E Q++M D+RK++RM+SNRESARRSRMRKQ+HLDEL +Q
Sbjct: 44 SSSSNSQDLMISNNSTSDEDHHQSIMVLDERKQRRMLSNRESARRSRMRKQRHLDELWSQ 103
Query: 60 VAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGEL 98
V +LR+EN ++ LN ++ + ENS L + +L
Sbjct: 104 VIRLRNENNCLIDKLNRVSETQNCVLKENSKLKEEASDL 142
>At1g13600 bZip transcription factor AtbZip58
Length = 196
Score = 80.9 bits (198), Expect = 2e-16
Identities = 45/105 (42%), Positives = 69/105 (64%), Gaps = 4/105 (3%)
Query: 5 SGTSSGSSMI--QNSGSEEELQALM--DQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
S +S+G ++ NS S+E+ Q M D+RK++RMISNRESARRSRMRKQ+HLDEL +QV
Sbjct: 57 SSSSNGQDLMTSNNSTSDEDHQQSMVIDERKQRRMISNRESARRSRMRKQRHLDELWSQV 116
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESL 105
+LR++N ++ LN ++ + EN+ L + +L + +
Sbjct: 117 IRLRTDNHCLMDKLNRVSESHELALKENAKLKEETSDLRQLISEI 161
>At5g28770 bZip transcription factor BZO2H3 / AtbZip63
Length = 307
Score = 79.7 bits (195), Expect = 5e-16
Identities = 50/121 (41%), Positives = 67/121 (55%), Gaps = 8/121 (6%)
Query: 4 SSGTSSGSSMIQNSGSEEELQALMDQ-----RKRKRMISNRESARRSRMRKQKHLDELAA 58
SS +SGS + SG EEE + ++ KRM+SNRESARRSR RKQ HL EL
Sbjct: 118 SSAITSGSEL---SGDEEEADGETNMNPTNVKRVKRMLSNRESARRSRRRKQAHLSELET 174
Query: 59 QVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGV 118
QV+QLR EN +++ L TQ F EN VL A + L +++ E + + N +
Sbjct: 175 QVSQLRVENSKLMKGLTDVTQTFNDASVENRVLKANIETLRAKVKMAEETVKRLTGFNPM 234
Query: 119 F 119
F
Sbjct: 235 F 235
>At3g30530 bZip transcription factor AtbZip42
Length = 173
Score = 77.8 bits (190), Expect = 2e-15
Identities = 43/102 (42%), Positives = 63/102 (61%)
Query: 4 SSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQL 63
S SS +S + ++ ++++RK++RMISNRESARRSRMRKQ+HLDEL +QV L
Sbjct: 55 SMSLSSNNSTSDEAEEQQTNNNIINERKQRRMISNRESARRSRMRKQRHLDELWSQVMWL 114
Query: 64 RSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESL 105
R EN Q+L LN ++ + EN+ L + EL + +
Sbjct: 115 RIENHQLLDKLNNLSESHDKVLQENAQLKEETFELKQVISDM 156
>At5g38800 bZIP transcription factor AtbZip43
Length = 165
Score = 74.7 bits (182), Expect = 2e-14
Identities = 44/104 (42%), Positives = 62/104 (59%), Gaps = 5/104 (4%)
Query: 7 TSSGSSMIQNSGS-----EEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVA 61
TS S+I ++ S EE + ++++RK+KR ISNRESARRSRMRKQ+ +DEL +QV
Sbjct: 44 TSQFMSLISSNNSTSDEAEENHKEIINERKQKRKISNRESARRSRMRKQRQVDELWSQVM 103
Query: 62 QLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESL 105
LR EN Q+L LN + + EN L + EL + +
Sbjct: 104 WLRDENHQLLRKLNCVLESQEKVIEENVQLKEETTELKQMISDM 147
>At4g37730 bZip transcription factor AtbZip7
Length = 305
Score = 70.1 bits (170), Expect = 4e-13
Identities = 39/103 (37%), Positives = 60/103 (57%)
Query: 7 TSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSE 66
T+ GS ++ + + D+RKRKRM SNRESA+RSRMRKQ H+D L QV +L E
Sbjct: 174 TNFGSGENNHNRKKMIQPEMTDERKRKRMESNRESAKRSRMRKQSHIDNLREQVNRLDLE 233
Query: 67 NQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEII 109
N+++ L L + ++++N+ L + L RL + I+
Sbjct: 234 NRELGNRLRLVLHQLQRVNSDNNRLVTEQEILRLRLSEMRRIL 276
>At2g22850 bZip tarnscription factor AtbZip6
Length = 227
Score = 66.2 bits (160), Expect = 6e-12
Identities = 37/104 (35%), Positives = 61/104 (58%), Gaps = 1/104 (0%)
Query: 8 SSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSEN 67
S+ ++ +N + LQ + D RKRKRM SNRESA+RSRMRKQ+H+D L + +L EN
Sbjct: 108 SNATNHNRNKLNRSVLQ-VTDDRKRKRMESNRESAKRSRMRKQRHIDNLKDEANRLGLEN 166
Query: 68 QQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDF 111
+++ L + ++ +N+ L ++ L R + +I+ F
Sbjct: 167 RELANRLRIVLYNIALMCTDNNQLLSEQEILRRRFLEMRQILIF 210
>At3g49760 bZIP transcription factor AtbZip5
Length = 156
Score = 64.3 bits (155), Expect = 2e-11
Identities = 34/98 (34%), Positives = 56/98 (56%)
Query: 14 IQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTS 73
+ GS+ D+RK+KR +SNRESA+RSR +KQKHL+E++ Q+ QL+ +NQ++
Sbjct: 56 VNQIGSDMSPTDNTDERKKKRKLSNRESAKRSREKKQKHLEEMSIQLNQLKIQNQELKNQ 115
Query: 74 LNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDF 111
L EN L + L +L ++ +++ F
Sbjct: 116 LRYVLYHCQRTKMENDRLLMEHRILHDKLLNIRQVLMF 153
>At3g54620 bZIP transcription factor-like protein
Length = 403
Score = 62.4 bits (150), Expect = 8e-11
Identities = 31/89 (34%), Positives = 52/89 (57%)
Query: 28 DQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAE 87
D ++ +RM+SNRESARRSR RKQ+ ++E QV QLR+E+ ++ L+ ++ +
Sbjct: 229 DVKRARRMLSNRESARRSRRRKQEQMNEFDTQVGQLRAEHSTLINRLSDMNHKYDAAAVD 288
Query: 88 NSVLSAQMGELSHRLESLNEIIDFVNATN 116
N +L A + L +++ E + V N
Sbjct: 289 NRILRADIETLRTKVKMAEETVKRVTGVN 317
>At5g24800 bZIP protein BZO2H2
Length = 277
Score = 59.7 bits (143), Expect = 5e-10
Identities = 36/111 (32%), Positives = 60/111 (53%), Gaps = 8/111 (7%)
Query: 7 TSSGSSMIQNSGSEEELQALM--------DQRKRKRMISNRESARRSRMRKQKHLDELAA 58
T+SGSS + + + E +A D ++ +RM SNRESA+RSR RKQ++L +L
Sbjct: 91 TTSGSSHVNSDDEDAETEAGQSEMTNDPNDLKRIRRMNSNRESAKRSRRRKQEYLVDLET 150
Query: 59 QVAQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEII 109
QV L+ +N + L TQ+F N VL + + L +++ +++
Sbjct: 151 QVDSLKGDNSTLYKQLIDATQQFRSAGTNNRVLKSDVETLRVKVKLAEDLV 201
>At4g02640 bZIP protein BZO2H1
Length = 417
Score = 59.7 bits (143), Expect = 5e-10
Identities = 38/123 (30%), Positives = 61/123 (48%), Gaps = 3/123 (2%)
Query: 4 SSGTSSGSSMIQNSGSEEELQALM---DQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
+SG+S S ++ E E + D +K +RM+SNRESARRSR RKQ+ +L QV
Sbjct: 194 TSGSSREYSDDEDLDEENETTGSLKPEDVKKSRRMLSNRESARRSRRRKQEQTSDLETQV 253
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVFA 120
L+ E+ +L L+ ++ N +L A + L +++ E + V N +
Sbjct: 254 NDLKGEHSSLLKQLSNMNHKYDEAAVGNRILKADIETLRAKVKMAEETVKRVTGMNPMLL 313
Query: 121 ANS 123
S
Sbjct: 314 GRS 316
>At1g68880 bZip transcription factor AtbZip8
Length = 138
Score = 59.7 bits (143), Expect = 5e-10
Identities = 28/64 (43%), Positives = 46/64 (71%)
Query: 28 DQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDAE 87
++RKR+R +SNRESARRSRMRKQ+H++EL + + QL ++N+ ++ L+ + + + E
Sbjct: 45 NERKRRRKVSNRESARRSRMRKQRHMEELWSMLVQLINKNKSLVDELSQARECYEKVIEE 104
Query: 88 NSVL 91
N L
Sbjct: 105 NMKL 108
>At5g49450 bZip protein AtbZIP1
Length = 145
Score = 57.4 bits (137), Expect = 3e-09
Identities = 36/115 (31%), Positives = 60/115 (51%), Gaps = 14/115 (12%)
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
MA++ TSSGS + D++KRKR +SNRESARRSR++KQK +++ ++
Sbjct: 1 MANAEKTSSGSDI--------------DEKKRKRKLSNRESARRSRLKKQKLMEDTIHEI 46
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNAT 115
+ L ++ QR ++ EN+ L ++ LS + L +I + T
Sbjct: 47 SSLERRIKENSERCRAVKQRLDSVETENAGLRSEKIWLSSYVSDLENMIATTSLT 101
>At5g60830 bZip transcription factor AtbZip70
Length = 188
Score = 55.5 bits (132), Expect = 1e-08
Identities = 40/112 (35%), Positives = 61/112 (53%), Gaps = 5/112 (4%)
Query: 28 DQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTS-LNLTTQRFMIIDA 86
++R+ +RM+SNRESARRSRMRK+K ++EL QV QL N + +NL I+
Sbjct: 77 EERRARRMVSNRESARRSRMRKKKQIEELQQQVEQLMMLNHHLSEKVINLLESNHQILQ- 135
Query: 87 ENSVLSAQMGELSHRLESLNEIIDFVNATNGVFAAN-SFVDPIPSAYYLNHP 137
ENS L ++ S L + ++ NA + + N +++ PS N P
Sbjct: 136 ENSQLKEKVS--SFHLLMADVLLPMRNAESNINDRNVNYLRGEPSNRPTNSP 185
>At1g59530 bZip transcription factor AtbZip4
Length = 148
Score = 55.5 bits (132), Expect = 1e-08
Identities = 32/79 (40%), Positives = 45/79 (56%)
Query: 27 MDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIIDA 86
+D +KR+R ISNRESA+RSRM+K+K +EL +V +L NQ++ L I
Sbjct: 47 VDDKKRRRTISNRESAKRSRMKKKKRFEELTEEVNRLNIRNQELKNRLANVVSCGNFISR 106
Query: 87 ENSVLSAQMGELSHRLESL 105
EN+ L + L RL L
Sbjct: 107 ENNRLKTESVCLEIRLLEL 125
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.124 0.319
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,546,859
Number of Sequences: 26719
Number of extensions: 85048
Number of successful extensions: 588
Number of sequences better than 10.0: 142
Number of HSP's better than 10.0 without gapping: 98
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 471
Number of HSP's gapped (non-prelim): 159
length of query: 147
length of database: 11,318,596
effective HSP length: 90
effective length of query: 57
effective length of database: 8,913,886
effective search space: 508091502
effective search space used: 508091502
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0113.7


