

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.13
(359 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
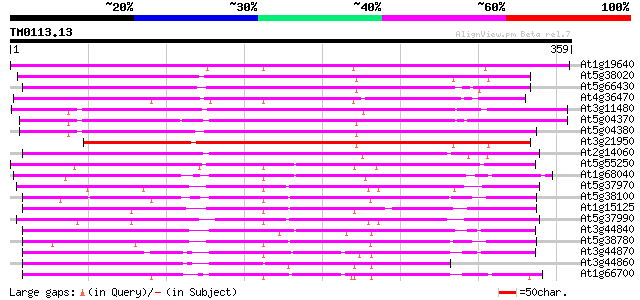
At1g19640 S-adenosyl-L-methionine:jasmonic acid carboxyl methylt... 275 2e-74
At5g38020 SAMT-like protein 271 5e-73
At5g66430 S-adenosyl-L-methionine:salicylic acid carboxyl methyl... 259 2e-69
At4g36470 hypothetical protein 259 2e-69
At3g11480 unknown protein 258 3e-69
At5g04370 S-adenosyl-L-methionine:salicylic acid carboxyl methyl... 249 2e-66
At5g04380 S-adenosyl-L-methionine:salicylic acid carboxyl methyl... 241 6e-64
At3g21950 salicylic acid carboxyl methyltransferase, putative 240 8e-64
At2g14060 hypothetical protein 228 5e-60
At5g55250 S-adenosyl-L-methionine:salicylic acid carboxyl methyl... 181 4e-46
At1g68040 putative S-adenosyl-L-methionine:salicylic acid carbox... 181 4e-46
At5g37970 putative protein 167 8e-42
At5g38100 unknown protein 166 2e-41
At1g15125 putative protein 165 3e-41
At5g37990 putative protein 165 4e-41
At3g44840 proteinkinase AtPP -like protein 161 4e-40
At5g38780 AtPP - like protein 159 2e-39
At3g44870 AtPP -like protein 156 1e-38
At3g44860 AtPP -like protein 152 3e-37
At1g66700 unknown protein 148 4e-36
>At1g19640 S-adenosyl-L-methionine:jasmonic acid carboxyl
methyltransferase (JMT)
Length = 389
Score = 275 bits (704), Expect = 2e-74
Identities = 160/388 (41%), Positives = 226/388 (58%), Gaps = 30/388 (7%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFP-SCLKVADLGC 59
M + +VLHMN G+G+ SYA NS+ Q N++ + ++++++ +L + S + +ADLGC
Sbjct: 1 MEVMRVLHMNKGNGETSYAKNSTAQSNIISLGRRVMDEALKKLMMSNSEISSIGIADLGC 60
Query: 60 SSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQE 119
SSGPN+L+ SNI++T+ + +L+ P + LNDL NDFN SLP+FY R+
Sbjct: 61 SSGPNSLLSISNIVDTIHNLCPDLDRPVPELRVSLNDLPSNDFNYICASLPEFYDRVNNN 120
Query: 120 K-GHKFG-----PCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV----------NY 163
K G FG CF S PGSFYGRLFP S+HF HSS SLHWLS+V
Sbjct: 121 KEGLGFGRGGGESCFVSAVPGSFYGRLFPRRSLHFVHSSSSLHWLSQVPCREAEKEDRTI 180
Query: 164 TKPL-NKGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGR---- 218
T L N G IY++KTSP + HKAY QF+ DF +FL SRS EL+PGG MV++ +GR
Sbjct: 181 TADLENMGKIYISKTSPKSAHKAYALQFQTDFWVFLRSRSEELVPGGRMVLSFLGRRSLD 240
Query: 219 DENNELINAWVVIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQR 278
E W ++ L MA E +IE+ K+DAFN P Y ++EE++ VIE+EGSF I R
Sbjct: 241 PTTEESCYQWELLAQALMSMAKEGIIEEEKIDAFNAPYYAASSEELKMVIEKEGSFSIDR 300
Query: 279 LETIRTDWIKNINVSDDIDEDTRAE--------AVAKYIRAVAESILKSEFGEAIMDELF 330
LE DW + D R++ V+ IRAV E +L+ FGE +MDELF
Sbjct: 301 LEISPIDWEGGSISEESYDLVIRSKPEALASGRRVSNTIRAVVEPMLEPTFGENVMDELF 360
Query: 331 RRFKNKIVKLHGVEKLELANLVMHITKS 358
R+ + + V A +++ + ++
Sbjct: 361 ERYAKIVGEYFYVSSPRYAIVILSLVRA 388
>At5g38020 SAMT-like protein
Length = 368
Score = 271 bits (692), Expect = 5e-73
Identities = 149/343 (43%), Positives = 202/343 (58%), Gaps = 18/343 (5%)
Query: 6 VLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCAT-FPSCLKVADLGCSSGPN 64
VL M GGDG+ SYANNS Q+ + K ++ +++ + T FP C+KVADLGCSSG N
Sbjct: 3 VLSMKGGDGEHSYANNSEGQKRLASDAKPVVVETVKEMIVKTDFPGCIKVADLGCSSGEN 62
Query: 65 ALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKF 124
L+V S I+NT+ Q P LNDL NDFNTTFK +P F+K L+ +
Sbjct: 63 TLLVMSEIVNTIITSYQQKGKNLPEINCCLNDLPDNDFNTTFKLVPAFHKLLKMDVK--- 119
Query: 125 GPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHK 184
G CF SG PGSFY RLFP+ S+HF HSS LHWLSKV NK +YL PP V+K
Sbjct: 120 GKCFISGVPGSFYSRLFPSKSLHFVHSSLCLHWLSKVPDGLEDNKKNVYLRSPCPPNVYK 179
Query: 185 AYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDE----NNELINAWVVIGMVLNDMAS 240
+Y QF+ DF LFL R+ E +P G M +T +GR + + W I L D+ S
Sbjct: 180 SYLTQFKNDFSLFLRLRADETVPNGRMALTFVGRKSLDPLSKDCFQNWSSISDSLLDLVS 239
Query: 241 ENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETI-------RTDWIKNINVS 293
E +++++ +D+FN+P Y P E+R+VIE EGSF I ETI +T + +
Sbjct: 240 EGIVKESDVDSFNLPFYNPDESEVREVIESEGSFKISNFETIFGLLFSYKTGRTEVKDDD 299
Query: 294 DDIDEDTRAEAV---AKYIRAVAESILKSEFGEAIMDELFRRF 333
D++D+ R E + A IR++ E +L + FG+AIMD LF R+
Sbjct: 300 DNLDQSCRFEVIRKRASIIRSITEPMLGAHFGDAIMDRLFERY 342
>At5g66430 S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase-like protein
Length = 354
Score = 259 bits (661), Expect = 2e-69
Identities = 144/339 (42%), Positives = 205/339 (59%), Gaps = 24/339 (7%)
Query: 9 MNGGDGDASYANNSSFQRNVMLTTKHIL-EDSIIRLYCATFPSCLKVADLGCSSGPNALM 67
M+GGDGD SY+ NS Q+ V+ K +L +++ + FP+ +KVADLGC++G N +
Sbjct: 1 MSGGDGDNSYSTNSLLQKKVLSKAKPVLVKNTKGMMINLNFPNYIKVADLGCATGENTFL 60
Query: 68 VASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGPC 127
+ I+NT++ + Q N + P LNDL NDFNTTFK +P F KR++ ++ C
Sbjct: 61 TMAEIVNTINVLCQQCNQKPPEIDCCLNDLPDNDFNTTFKFVPFFNKRVKSKR-----LC 115
Query: 128 FFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAYF 187
F SG PGSFY RLFP S+HF HSSYSLHWLSKV N ++Y+T +SPP +KAY
Sbjct: 116 FVSGVPGSFYSRLFPRKSLHFVHSSYSLHWLSKVPKGLEKNSSSVYITTSSPPNAYKAYL 175
Query: 188 AQFREDFKLFLGSRSCELLPGGAMVITLIGRDE-----NNELINAWVVIGMVLNDMASEN 242
QF+ DFK FL RS E++ G MV+T IGR + + + W ++ L D+ E
Sbjct: 176 NQFQSDFKSFLEMRSEEMVSNGRMVLTFIGRKTLDDPLHRDCCHFWTLLSTSLRDLVYEG 235
Query: 243 LIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDED--- 299
L+ +K+D+FNIP Y P+ EE+ ++I EGSF+I LE + + +S+ DED
Sbjct: 236 LVSASKVDSFNIPFYDPSKEEVMEMIRNEGSFEINDLEIHGFE----LGLSNH-DEDYML 290
Query: 300 -----TRAEAVAKYIRAVAESILKSEFGEAIMDELFRRF 333
+ A IRAV+ES+L ++FG IMD LF++F
Sbjct: 291 HSQISKAGQREANCIRAVSESMLVADFGVDIMDTLFKKF 329
>At4g36470 hypothetical protein
Length = 371
Score = 259 bits (661), Expect = 2e-69
Identities = 147/345 (42%), Positives = 208/345 (59%), Gaps = 25/345 (7%)
Query: 3 IEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSG 62
+E+ +M GGDG SYA NSS Q+ T KHI +++ +LY T P L +ADLGCSSG
Sbjct: 6 MEREFYMTGGDGKTSYARNSSLQKKASDTAKHITLETLQQLYKETRPKSLGIADLGCSSG 65
Query: 63 PNALMVASNIINTVDAVSQNLNLEQPV--FQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
PN L ++ I TV QP+ F FLNDL GNDFN FKSLPDF+ L+++
Sbjct: 66 PNTLSTITDFIKTVQVAHHREIPIQPLPEFSIFLNDLPGNDFNFIFKSLPDFHIELKRDN 125
Query: 121 GHKFGPC---FFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNKGA 171
+ G C F + PGSFYGRLFP +++HF ++S+SLHWLSKV K +NKG
Sbjct: 126 NN--GDCPSVFIAAYPGSFYGRLFPENTIHFVYASHSLHWLSKVPTALYDEQGKSINKGC 183
Query: 172 IYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGR------DENNELI 225
+ + S AV KAY +QF+EDF +FL RS E++ G MV+ ++GR D N
Sbjct: 184 VSICSLSSEAVSKAYCSQFKEDFSIFLRCRSKEMVSAGRMVLIILGREGPDHVDRGNSFF 243
Query: 226 NAWVVIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTD 285
W ++ + D+ ++ E+ KLD++++ Y P+A+EI +++EGSF+++RLE +
Sbjct: 244 --WELLSRSIADLVAQGETEEEKLDSYDMHFYAPSADEIEGEVDKEGSFELERLEMLEVK 301
Query: 286 WIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELF 330
K N DI + +AVAK +RAV ES+L FGE I+D+LF
Sbjct: 302 KDKG-NTEGDI---SYGKAVAKTVRAVQESMLVQHFGEKILDKLF 342
>At3g11480 unknown protein
Length = 379
Score = 258 bits (660), Expect = 3e-69
Identities = 148/365 (40%), Positives = 215/365 (58%), Gaps = 19/365 (5%)
Query: 2 AIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHIL----EDSIIRLYCATFPSCLKVADL 57
A + L M+GGDG SY+ NS Q+ V+ K +L E+ ++ L FP+ +KVA+L
Sbjct: 24 AFVKALCMSGGDGANSYSANSRLQKKVLSMAKPVLVRNTEEMMMNL---DFPTYIKVAEL 80
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQ 117
GCSSG N+ + IINT++ + Q++N P LNDL NDFNTTFK +P F K L
Sbjct: 81 GCSSGQNSFLAIFEIINTINVLCQHVNKNSPEIDCCLNDLPENDFNTTFKFVPFFNKELM 140
Query: 118 QEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKT 177
CF G PGSFY RLF +S+H HSSY+LHWLSKV NKG +Y+T +
Sbjct: 141 ITNKSS---CFVYGAPGSFYSRLFSRNSLHLIHSSYALHWLSKVPEKLENNKGNLYITSS 197
Query: 178 SPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELI-----NAWVVIG 232
SP + +KAY QF++DF +FL RS E++ G MV+T IGR+ N+ + + W ++
Sbjct: 198 SPQSAYKAYLNQFQKDFTMFLRLRSEEIVSNGRMVLTFIGRNTLNDPLYRDCCHFWTLLS 257
Query: 233 MVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINV 292
L D+ E L+ ++KLDAFN+P Y P +E+++VI++EGSF+I LE+ D + +
Sbjct: 258 NSLRDLVFEGLVSESKLDAFNMPFYDPNVQELKEVIQKEGSFEINELESHGFD-LGHYYE 316
Query: 293 SDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLV 352
DD + A IRAV+E +L + FGE I+D LF ++ + + +LV
Sbjct: 317 EDDFEAGRNE---ANGIRAVSEPMLIAHFGEEIIDTLFDKYAYHVTQHANCRNKTTVSLV 373
Query: 353 MHITK 357
+ +TK
Sbjct: 374 VSLTK 378
>At5g04370 S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase-like protein
Length = 396
Score = 249 bits (636), Expect = 2e-66
Identities = 144/360 (40%), Positives = 209/360 (58%), Gaps = 18/360 (5%)
Query: 7 LHMNGGDGDASYANNSSFQRNVMLTTKHIL----EDSIIRLYCATFPSCLKVADLGCSSG 62
L M GGDG SY++NS QR V+ K +L +D +I L FP+ +KVADLGCSSG
Sbjct: 39 LCMRGGDGYNSYSSNSLLQRRVLSKAKPVLVKNTKDLMINL---NFPTYIKVADLGCSSG 95
Query: 63 PNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGH 122
N + S IINT++ Q N P LNDL NDFNTTFK + F+ +
Sbjct: 96 QNTFLAMSEIINTINVFCQQRNQNPPEIDCCLNDLPSNDFNTTFKFI-QFFNGMNITSKE 154
Query: 123 KFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAV 182
+ F G PGSFY RLFP S+HF HSSY LHWLSKV NK ++Y+T +SP +
Sbjct: 155 SY---FVYGVPGSFYSRLFPRRSLHFVHSSYGLHWLSKVPEGLEKNKMSVYITNSSPLST 211
Query: 183 HKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDE-----NNELINAWVVIGMVLND 237
+KAY QF+ DF FL RS E++ G MV+T IGR+ + + + W ++ L D
Sbjct: 212 YKAYLNQFQRDFATFLKLRSEEMVSNGRMVLTFIGRNTIDNPLHRDCCHFWTLLSKSLRD 271
Query: 238 MASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDID 297
+ +E L+ +K+D+F +P Y P +EI++++++EGSF+I+ LET D + + N D+
Sbjct: 272 LVAEGLVSASKVDSFYLPFYDPNEKEIKEMVQKEGSFEIRDLETHGYD-LGHCN-QDESK 329
Query: 298 EDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHITK 357
+ A YIRAV+E +L + FG+AI++ LF +F + + ++V+ +TK
Sbjct: 330 RSKSGQNEANYIRAVSEPLLAAHFGDAIINILFNKFACHVSQHVSCRNKTTVSIVVSLTK 389
>At5g04380 S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase-like protein
Length = 385
Score = 241 bits (614), Expect = 6e-64
Identities = 139/341 (40%), Positives = 193/341 (55%), Gaps = 18/341 (5%)
Query: 7 LHMNGGDGDASYANNSSFQRNVMLTTKHIL----EDSIIRLYCATFPSCLKVADLGCSSG 62
L MNGGD D SY S Q+ V+ T IL E+ + L FP C+KVADLGCSSG
Sbjct: 32 LCMNGGDVDNSYTTKSLLQKRVLSITNPILVKNTEEMLTNL---DFPKCIKVADLGCSSG 88
Query: 63 PNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGH 122
N + S I+NT++ + Q N +P LNDL NDFNTTFK + F K+L
Sbjct: 89 QNTFLAMSEIVNTINVLCQKWNQSRPEIDCCLNDLPTNDFNTTFKFITFFNKKLTSN--- 145
Query: 123 KFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAV 182
G CF SG PGSFY RLFP S+HF +S YS+H+LSKV NK ++Y+T +SP +
Sbjct: 146 --GSCFVSGVPGSFYSRLFPRKSLHFIYSIYSIHFLSKVPDGLEKNKMSVYITSSSPLSE 203
Query: 183 HKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDE-----NNELINAWVVIGMVLND 237
+KAY QF+ DF FL RS E++ G MV+TLIGR+ + + W ++ L D
Sbjct: 204 YKAYLNQFKRDFTTFLRMRSEEMVHNGRMVLTLIGRNTLDNPLYRDCCHCWTLLSNSLRD 263
Query: 238 MASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDID 297
+ E L+ +K+ +F +P Y P EE++++I EGSF I LE D +
Sbjct: 264 LVFEGLLSASKVYSFKMPFYDPNEEEVKEIIRNEGSFQINDLEMHEFDLGHSKEKCSLQS 323
Query: 298 EDTRA-EAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKI 337
+A + A IRAV E++L + FG+ I+D LF ++ + +
Sbjct: 324 HKAKAGQKEASCIRAVTETMLVAHFGDDIIDALFHKYAHHV 364
>At3g21950 salicylic acid carboxyl methyltransferase, putative
Length = 335
Score = 240 bits (613), Expect = 8e-64
Identities = 128/300 (42%), Positives = 182/300 (60%), Gaps = 17/300 (5%)
Query: 48 FPSCLKVADLGCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFK 107
FP C+KVADLGCSSG N +V S I+NT+ Q P LNDL NDFNTTFK
Sbjct: 13 FPGCIKVADLGCSSGENTFLVMSEIVNTIITTYQQNGQNLPEIDCCLNDLPENDFNTTFK 72
Query: 108 SLPDFYKRLQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPL 167
+P F+++L K + G C+ SG PGSFY RLFP+ S+HF HSS+ LHWLSKV
Sbjct: 73 LIPSFHEKL---KMNVKGNCYVSGCPGSFYTRLFPSKSLHFVHSSFCLHWLSKVPDGLEE 129
Query: 168 NKGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDE----NNE 223
NK +YL PP ++++Y+ QF++DF +FL R+ E +P G M +TL+GR + E
Sbjct: 130 NKKNVYLRSPCPPNLYESYWNQFKKDFSMFLRMRAEETMPSGRMALTLVGRKTLDPLSKE 189
Query: 224 LINAWVVIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETI- 282
W ++ L D+ SE +++++ L++FN+P Y P E+++VIE EGSF+I+ ETI
Sbjct: 190 CFKDWSLVSDSLLDLVSEGVVKESDLESFNLPYYSPDESEVKEVIENEGSFEIKNFETIF 249
Query: 283 ------RTDWIKNINVSDDIDEDTRAEAV---AKYIRAVAESILKSEFGEAIMDELFRRF 333
+T + + DD+D R E V A R++ E +L + FGEAI+D LF ++
Sbjct: 250 GLLFSYKTGHSEVKDDDDDVDHSRRFEVVKTRANMTRSIIEPMLVAHFGEAIIDRLFDKY 309
>At2g14060 hypothetical protein
Length = 359
Score = 228 bits (580), Expect = 5e-60
Identities = 127/342 (37%), Positives = 199/342 (58%), Gaps = 16/342 (4%)
Query: 9 MNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCAT-FPSCLKVADLGCSSGPNALM 67
M GG GD SYA NS +QR+V + ++ +++ + FP C+KVADLGCS+G N ++
Sbjct: 1 MKGGTGDHSYATNSHYQRSVFYEIQPLVIENVREMLLKNGFPGCIKVADLGCSTGQNTVL 60
Query: 68 VASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGPC 127
S I T+ Q ++ P +LNDL NDFNTTFK F ++L+ E K+
Sbjct: 61 AMSAIAYTIMESYQQMSKNPPEIDCYLNDLPENDFNTTFKLFHSFQEKLKPEVKGKW--- 117
Query: 128 FFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAYF 187
F SG PGSFY RLFP S+HF HS++S+HWLS++ N +I++ P V+K+Y
Sbjct: 118 FVSGVPGSFYSRLFPRKSLHFVHSAFSIHWLSRIPDGLESNTKSIHIKYPYPSNVYKSYL 177
Query: 188 AQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNEL----INAWVVIGMVLNDMASENL 243
QF+ DF LFL RS E++ G MV+T +GR ++ L W ++ L D+ASE
Sbjct: 178 NQFKIDFSLFLKMRSEEVVHNGHMVLTFVGRKVSDTLSKDCFQVWSLLSDCLLDLASEGF 237
Query: 244 IEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINV---SDDIDEDT 300
+ + + +FN+P Y P EE+R+ I +EGSF+I ++E + D + + +D ++
Sbjct: 238 VNDSMVKSFNMPFYNPNEEEVREFILKEGSFEITKIE--KFDHVVPYKIDREEEDEEQSL 295
Query: 301 RAEA---VAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
+ EA A + R + E +L + FG+AI++ +F ++ + + K
Sbjct: 296 QLEAGIKHASWARCITEPLLVAHFGDAIIEPVFNKYAHYMAK 337
>At5g55250 S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase-like protein
Length = 386
Score = 181 bits (460), Expect = 4e-46
Identities = 116/355 (32%), Positives = 182/355 (50%), Gaps = 24/355 (6%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSI--IRLYCATFPSCLKVADLG 58
M +E++L M GG G SYANNS Q + H+LE+++ + L + P DLG
Sbjct: 13 MKLERLLSMKGGKGQDSYANNSQAQAMHARSMLHLLEETLENVHLNSSASPPPFTAVDLG 72
Query: 59 CSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQ 118
CSSG N + + I+ + ++ P F F +DL NDFNT F+ LP
Sbjct: 73 CSSGANTVHIIDFIVKHISKRFDAAGIDPPEFTAFFSDLPSNDFNTLFQLLPPLVSNTCM 132
Query: 119 EK-----GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPL 167
E+ G++ F +G PGSFY RLFP ++ FFHS++SLHWLS+V +
Sbjct: 133 EECLAADGNR--SYFVAGVPGSFYRRLFPARTIDFFHSAFSLHWLSQVPESVTDRRSAAY 190
Query: 168 NKGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRD--ENNELI 225
N+G +++ AY QF+ D FL +R+ E+ GGAM + +GR + +
Sbjct: 191 NRGRVFIHGAGEKTT-TAYKRQFQADLAEFLRARAAEVKRGGAMFLVCLGRTSVDPTDQG 249
Query: 226 NAWVVIGM----VLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLET 281
A ++ G +D+ E L+ K D FNIP Y P+ ++ ++V++ GSF I +L
Sbjct: 250 GAGLLFGTHFQDAWDDLVREGLVAAEKRDGFNIPVYAPSLQDFKEVVDANGSFAIDKLVV 309
Query: 282 IRTDWIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNK 336
+ +N DD E R A A R+VA ++++ GE + ++LF R +++
Sbjct: 310 YKGGSPLVVNEPDDASEVGR--AFASSCRSVAGVLVEAHIGEELSNKLFSRVESR 362
>At1g68040 putative S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase
Length = 363
Score = 181 bits (460), Expect = 4e-46
Identities = 123/363 (33%), Positives = 184/363 (49%), Gaps = 40/363 (11%)
Query: 3 IEQVLHMNGGDGDASYANNSSFQRNVMLTTKH-----ILEDSIIRLYCATFPSCLKVADL 57
+ L M+GGDG SY+ NS QR K +LE + ++ + ++ADL
Sbjct: 7 VRNSLPMSGGDGPNSYSKNSHLQRKTTSLLKEKIDKLVLEKLNAKTLISSDSNTFRIADL 66
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQ 117
GC++GPN + NII +++ + N +P F F NDL NDFNT F SLP
Sbjct: 67 GCATGPNTFFLVDNIIKSIETSLRKSNSSKPEFLVFFNDLPQNDFNTLFTSLP------- 119
Query: 118 QEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNKGA 171
Q++ + G PGSFYGR+ P SVH + + HWLS V +K NKG
Sbjct: 120 QDRSY-----LAVGVPGSFYGRVLPQSSVHIVVTMGATHWLSSVPKEVLDKSSKAWNKGK 174
Query: 172 IYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELIN----- 226
++ + + V KAY QF D + FL +R+ E++ GG +V+ + G + N
Sbjct: 175 VHYSNAADEVV-KAYRDQFGRDMEKFLEARATEIVSGGLLVVGMCGIPKGMPFSNLADSI 233
Query: 227 AWVVIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETI-RTD 285
+ + VL M SE LI + ++D FNIP Y T EE+ ++ + G F ++ +E + T
Sbjct: 234 MYTSMADVLTQMHSEGLISEEQVDTFNIPIYSATPEEVTVLVVKNGCFTVESMELMDPTA 293
Query: 286 WIKN-INVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVE 344
W+K NV ED R V I+A S+ + FGE ++D++F R K+V L E
Sbjct: 294 WLKRPTNV-----EDVRHWMVC--IKATMGSLFINHFGEHLLDDVFDRLTAKLVGL--TE 344
Query: 345 KLE 347
K+E
Sbjct: 345 KIE 347
>At5g37970 putative protein
Length = 412
Score = 167 bits (423), Expect = 8e-42
Identities = 118/358 (32%), Positives = 178/358 (48%), Gaps = 45/358 (12%)
Query: 5 QVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATF------PSCLKVADLG 58
Q MNGGDG SY +NSS+Q+ + K ++I+ F + L++ D G
Sbjct: 56 QSFPMNGGDGPHSYIHNSSYQKVAIDGVKERTSEAILEKLDLEFLNRNSEENILRIVDFG 115
Query: 59 CSSGPNALMVASNIINTVDAVSQNLN---LEQPV-FQFFLNDLFGNDFNTTFKSLPDFYK 114
CS GPN V NII+TV N + P+ FQ ND NDFNT F++ P F +
Sbjct: 116 CSIGPNTFDVVQNIIDTVKQKRLKENKTYIGAPLEFQVCFNDQPNNDFNTLFRTQPFFSR 175
Query: 115 RLQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLN 168
+ F G PGSF+GR+ P +S+H H+SY+LHWLS V + LN
Sbjct: 176 K----------EYFSVGVPGSFHGRVLPKNSLHIGHTSYTLHWLSNVPQHVCDKKSPALN 225
Query: 169 KGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAW 228
K I V KAY QFR+DF FL +R+ EL+ GG M+++ + W
Sbjct: 226 KSYIQCNNL-VDEVTKAYKIQFRKDFGGFLEARAEELVSGGLMILSGQCLPDGIPKALTW 284
Query: 229 --VVIGMV---LNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIR 283
VVI M+ L D+A + K K++ F++P+Y P E + IE+ +F+++ +E I
Sbjct: 285 QGVVIDMIGDCLMDLAKLGITSKEKIELFSLPTYIPHISEFKANIEQNENFNVETMEEIS 344
Query: 284 --TDWIKNINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
D++ N + + RA+ +I++ FGE +++ELF R ++ K
Sbjct: 345 HPMDYMPLTN-----------DFITSMFRAILNTIIEEHFGEGVVNELFSRLAKRLDK 391
>At5g38100 unknown protein
Length = 359
Score = 166 bits (419), Expect = 2e-41
Identities = 114/348 (32%), Positives = 176/348 (49%), Gaps = 40/348 (11%)
Query: 9 MNGGDGDASYANNSSFQRNVMLT----TKHILEDSIIRLYCATFPSCLKVADLGCSSGPN 64
M+ G SY +NSS+Q+ + + T+ + + + + F + ++AD GCS GPN
Sbjct: 10 MSSGHDQHSYIHNSSYQKAAISSAVEKTRRCIFEKLDLQLSSDFGT-FRIADFGCSIGPN 68
Query: 65 ALMVASNIINTVDAVSQNLNLEQPV----FQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
VA +II+TV + + E + FQ F ND NDFNT F++ P L E+
Sbjct: 69 TFHVAQSIIDTVKSKRLEESTENSLVPLEFQVFFNDQPTNDFNTLFRTQP-----LSPER 123
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNKGAIYL 174
+ F G PGSFYGR+ P +S+H H+SY+ HWLSKV + NK I
Sbjct: 124 EY-----FSVGVPGSFYGRVLPRNSIHIGHTSYTTHWLSKVPDNVCDKKSMAWNKNYIQC 178
Query: 175 TKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV----- 229
V KAY QF +D ++FL +R+ EL+PGG M++ + L W
Sbjct: 179 NNLL-EEVTKAYKVQFIKDMEIFLDARAEELVPGGLMIVIGECLPDGVSLYETWQGYVMD 237
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
IG L DMA + + K+D F++P Y P E++ IE+ GSF I+ +ET + ++
Sbjct: 238 TIGDCLMDMAKSGITSEEKIDLFSLPVYFPQFSELKGEIEKNGSFTIELMET-TSHPLEG 296
Query: 290 INVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKI 337
+++D + RA +I++ FG+ ++DELF R K+
Sbjct: 297 KPLTNDF--------ITSTFRAFLTTIIEKHFGDGVVDELFYRLAKKL 336
>At1g15125 putative protein
Length = 347
Score = 165 bits (418), Expect = 3e-41
Identities = 116/345 (33%), Positives = 172/345 (49%), Gaps = 41/345 (11%)
Query: 9 MNGGDGDASYANNSSFQRNVMLTTKHILEDSI-IRLYCATFP-SCLKVADLGCSSGPNAL 66
MNGGDG +SYA NSS+QR + + +L + I RL S +AD GCSSGPN +
Sbjct: 1 MNGGDGASSYARNSSYQRGAIEAAEALLRNEINARLDITNHSFSSFTIADFGCSSGPNTV 60
Query: 67 MVASNIINTV--DAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKF 124
+ II + S N P FQ F ND+ DFN F LP
Sbjct: 61 IAVDIIIQALYHKFTSSLPNTTTPQFQVFFNDVSHTDFNALFALLPPQR----------- 109
Query: 125 GPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNKGAIYLTKTS 178
P F +G PGSFYG LFP ++ +SS +L WLS + + N+G I+ T S
Sbjct: 110 -PYFVAGVPGSFYGNLFPKAHLNLAYSSCALCWLSDLPSELTDTSSPAYNRGRIHYTGAS 168
Query: 179 PPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRD------ENNELINAWVVIG 232
V +AY +Q+++D KLFL +RS EL G M + + G + + + ++G
Sbjct: 169 -AEVAQAYSSQYKKDIKLFLHARSQELAENGLMALIVPGVPDGFLDCQEASTGSEFDLLG 227
Query: 233 MVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINV 292
L DMA E+ ++++FN+P Y T +E+ +I G I ++ET+ +
Sbjct: 228 SCLMDMAK----EEEEVNSFNLPIYYTTPKELEDIIRSNGELKIDKMETLGS-------- 275
Query: 293 SDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKI 337
D D E+ Y+RAV E ++++ FG I+D+LF R+ K+
Sbjct: 276 MDAQDTMPDLESRVLYLRAVLEGLVRTHFGHQILDDLFDRYALKL 320
>At5g37990 putative protein
Length = 362
Score = 165 bits (417), Expect = 4e-41
Identities = 117/357 (32%), Positives = 178/357 (49%), Gaps = 43/357 (12%)
Query: 5 QVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIR------LYCATFPSCLKVADLG 58
Q MNGGDG SY +NSS+Q+ + K ++I++ L + + L++AD G
Sbjct: 6 QSFPMNGGDGPHSYIHNSSYQKVAIDGAKEKTSEAILKNLDLELLNRNSDENILRIADFG 65
Query: 59 CSSGPNALMVASNIINTV---DAVSQNLNLEQPV-FQFFLNDLFGNDFNTTFKSLPDFYK 114
CS GPN V NII+TV + N + P+ FQ ND NDFNT F++ P K
Sbjct: 66 CSIGPNTFEVVQNIIDTVKQKNLKENNAYIGAPLEFQVCFNDQPNNDFNTLFRTQPISSK 125
Query: 115 RLQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLN 168
+ G PGSF+GR+ P +S+H H +Y+LHWLS V + LN
Sbjct: 126 QAYLSVG----------VPGSFHGRVLPKNSLHIGHITYALHWLSTVPQHVCDKKSPALN 175
Query: 169 KGAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAW 228
K I V +AY QF++D FLG+R+ EL+ GG M+++ + W
Sbjct: 176 KSYIQCNNL-VEEVTEAYRVQFKKDMGDFLGARAEELVSGGLMILSGQCLPDGVPKALTW 234
Query: 229 --VVIGMV---LNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIR 283
VVI M+ L DMA + + K K++ F++P Y P E + IE +F I+ +E
Sbjct: 235 QGVVIDMIGDCLMDMAKQGITTKEKIELFSLPIYIPHISEFKAEIERNENFSIETME--- 291
Query: 284 TDWIKNINVSDDID-EDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
+S +D + + + RA+ +I++ FG+ +++ELF RF K+ K
Sbjct: 292 -------KISHPMDYKPLTNDFITSMFRAILNTIIEEHFGDGVVNELFDRFAKKLNK 341
>At3g44840 proteinkinase AtPP -like protein
Length = 348
Score = 161 bits (408), Expect = 4e-40
Identities = 115/340 (33%), Positives = 174/340 (50%), Gaps = 34/340 (10%)
Query: 9 MNGGDGDASYANNSSFQRNVM-LTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPNALM 67
M GG+G SY +S +Q ++ T+ I E +L + + +AD GCS+GPN
Sbjct: 7 MIGGEGPESYRQHSKYQGGLLEAATEKINEAISTKLNIDLASNLVNIADFGCSTGPNTFR 66
Query: 68 VASNIINTVD-AVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGP 126
II+ V+ Q NLE+ FQ F ND NDFNT FK+LP K
Sbjct: 67 AVQTIIDAVEHKYQQENNLEEIEFQVFFNDSSNNDFNTLFKTLPPARKY----------- 115
Query: 127 CFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV-NYTKPLNKGA----IYLTKTSPPA 181
F +G P SF+GR+ P S+H SSYSLH+LSK+ K + A I+ T S
Sbjct: 116 -FATGVPASFFGRVLPRSSLHVGVSSYSLHFLSKIPKKIKDCDSHAWNKDIHCTGFSKEV 174
Query: 182 VHKAYFAQFREDFKLFLGSRSCELLPGGAMVIT---LIGRDENNELINAWVV--IGMVLN 236
V +AY Q++ D + FL +R+ EL+ GG + + L + +E +N ++ IG LN
Sbjct: 175 V-RAYLDQYKIDMESFLTARAQELVSGGLLFLLGSCLPNGVQMSETLNGMMIDCIGSSLN 233
Query: 237 DMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDI 296
D+A + LI++ KLD F +P Y A EI+Q+IE+ + I+R + I + +++I
Sbjct: 234 DIAKQGLIDQEKLDTFKLPIYVAYAGEIKQIIEDNVYYTIERFDIISQE-------NEEI 286
Query: 297 DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNK 336
D E + + I+ S FG+ +M++ F K K
Sbjct: 287 PLD--PEFLTVSFKVTVGGIVASHFGQHVMEKTFEVVKTK 324
>At5g38780 AtPP - like protein
Length = 361
Score = 159 bits (403), Expect = 2e-39
Identities = 109/349 (31%), Positives = 174/349 (49%), Gaps = 40/349 (11%)
Query: 9 MNGGDGDASYANNSSFQR----NVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPN 64
M+GGD SY +NSS+Q+ V + + +++ L S +AD GCS GPN
Sbjct: 10 MSGGDDQHSYIHNSSYQKAGIDGVQEKARQYILENLDLLNMNPNLSTFTIADFGCSIGPN 69
Query: 65 ALMVASNIINTVDAV----SQNLNLEQPV-FQFFLNDLFGNDFNTTFKSLPDFYKRLQQE 119
NII+ V SQ + P+ FQ + NDL NDFNT F++ P K+
Sbjct: 70 TFHAVQNIIDIVKLKHLKESQEDSRVAPLEFQVYFNDLPNNDFNTLFRTQPPSSKQ---- 125
Query: 120 KGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNKGAIY 173
F G PGSFYGR+ P +S+H ++S++ HWLSKV + NK I+
Sbjct: 126 ------EYFSVGVPGSFYGRVLPRNSIHIGNTSFTTHWLSKVPEEVCDKNSLAWNKNYIH 179
Query: 174 LTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV---- 229
V +AY QF +D +FL +R+ EL+PGG M+ + + W
Sbjct: 180 CNNLI-EEVTEAYKVQFEKDMGVFLKARAEELVPGGLMITLGQCLPDGVAMYETWSGIVK 238
Query: 230 -VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIK 288
IG L DMA+ + + K++ FN+P Y P E++ IE+ F I+ +E + + ++
Sbjct: 239 DTIGDCLQDMATLGVTTEEKIEMFNLPVYFPQVSELKGAIEQNIRFTIEMMEIV-SHPLE 297
Query: 289 NINVSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKI 337
+ +S++ + RA+ ++++ FG +++DELFR+F K+
Sbjct: 298 AVQLSNNF--------ITSMYRAILSTVIERHFGGSVVDELFRQFAKKL 338
>At3g44870 AtPP -like protein
Length = 379
Score = 156 bits (395), Expect = 1e-38
Identities = 114/342 (33%), Positives = 175/342 (50%), Gaps = 41/342 (11%)
Query: 9 MNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCL-KVADLGCSSGPNALM 67
M GG+G SY ++S +Q ++ K + ++I F S L +AD GCSSGPN
Sbjct: 7 MIGGEGPNSYRDHSKYQGALVEAAKEKINEAISTKLDIDFTSNLVNIADFGCSSGPNTFT 66
Query: 68 VASNIINTVD-AVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGP 126
+I+ V+ + N+E FQ F ND NDFNT FK+LP RL
Sbjct: 67 AVQTLIDAVENKYKKESNIE---FQVFFNDSSNNDFNTLFKTLPP--ARLY--------- 112
Query: 127 CFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV-----NYTKPLNKGAIYLTKTSPPA 181
F SG PGSF+GR+ P +S+H S+YSLH++SK+ + P+ I+ + +S
Sbjct: 113 -FASGVPGSFFGRVLPRNSLHLGVSAYSLHFISKIPKEVKDRDSPVWNKDIHCSGSSKE- 170
Query: 182 VHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENN-----ELINAWVV--IGMV 234
V K Y Q++ D FL +R+ EL+ GG ++ L+G N E + ++ IG
Sbjct: 171 VAKLYLGQYKIDVGSFLNARAQELVSGGLLL--LLGSCRPNGVQMFETVEGMMIDFIGAS 228
Query: 235 LNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSD 294
LN++A++ LI++ KLD F +P Y P A+E++Q+IE+ G F I+ E I I+
Sbjct: 229 LNEIANQGLIDQQKLDTFKLPIYAPQADELKQIIEDNGCFTIEVFENI-------IHAKG 281
Query: 295 DIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNK 336
+ D E + + + S FG+ M++ F K K
Sbjct: 282 EYPLD--PEFLTVSFKVTVGGSVASLFGQDGMEKTFELVKEK 321
>At3g44860 AtPP -like protein
Length = 348
Score = 152 bits (384), Expect = 3e-37
Identities = 101/286 (35%), Positives = 153/286 (53%), Gaps = 25/286 (8%)
Query: 9 MNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCL-KVADLGCSSGPNALM 67
M GG+G SY +S +Q +++ K + ++I F S L +AD GCSSGPN
Sbjct: 7 MIGGEGPNSYREHSKYQGALVIAAKEKINEAISTKLDIDFTSNLVNIADFGCSSGPNTFT 66
Query: 68 VASNIINTVD-AVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGP 126
+I+ V+ + N+E FQ F ND NDFNT FK+LP RL
Sbjct: 67 AVQTLIDAVENKYKKESNIEGIEFQVFFNDSSNNDFNTLFKTLPP--ARLY--------- 115
Query: 127 CFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKT-----SPPA 181
F SG PGSF+GR+ P +S+H SSYSLH++SKV + ++ ++ K S
Sbjct: 116 -FASGVPGSFFGRVLPKNSLHVGVSSYSLHFVSKVP-KEIKDRDSLVWNKDIHCSGSSKE 173
Query: 182 VHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRD---ENNELINAWVV--IGMVLN 236
V K Y Q++ D FL +R+ EL+ GG +++ R + E + ++ IG LN
Sbjct: 174 VVKLYLGQYKIDVGSFLTARAQELVSGGLLLLLGSCRPTGVQMFETVEGMMIDFIGSSLN 233
Query: 237 DMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETI 282
++A++ LI++ KLD F +P Y P +E++Q+IE+ F I+ E I
Sbjct: 234 EIANQGLIDQQKLDTFKLPIYAPNVDELKQIIEDNKCFTIEAFEKI 279
>At1g66700 unknown protein
Length = 353
Score = 148 bits (374), Expect = 4e-36
Identities = 116/354 (32%), Positives = 176/354 (48%), Gaps = 45/354 (12%)
Query: 9 MNGGDGDASYANNSSFQRNVMLTTKHILEDSI-IRLYCATFPSCLKVADLGCSSGPNALM 67
M GGDG SY SS+QR ++ TK + +I L + VAD GC+SGPN +
Sbjct: 9 MIGGDGPESYNQQSSYQRALLEATKDKMTKAISANLDLDLISNRFIVADFGCASGPNTFV 68
Query: 68 VASNIINTVDAVSQNLNLEQPV----FQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHK 123
NII+ V+ + + P FQ ND NDFNT F++LP G +
Sbjct: 69 AVQNIIDAVEEKYRRETGQNPADNIEFQVLFNDFSLNDFNTLFQTLPP---------GRR 119
Query: 124 FGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV-----NYTKPLNKGAIYLTKTS 178
+ F +G PGSF+ R+ P +S H SY+ H+ SK+ + PL + T +
Sbjct: 120 Y---FSAGVPGSFFERVLPKESFHIGVMSYAFHFTSKIPKGIMDRDSPLWNKDMQCTGFN 176
Query: 179 PPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIG---RD--ENNELINAWVV--I 231
P AV KAY Q+ D K+ L +R+ EL+PGG M+ L+G RD + +E V+ I
Sbjct: 177 P-AVKKAYLDQYSIDTKILLDARAEELVPGGLML--LLGSCLRDGVKMSETPKGTVMDFI 233
Query: 232 GMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNIN 291
G L+D+A + + E+ K+D F Y EIRQ+IEE G F I+ E I I+
Sbjct: 234 GESLSDLAKQGVTEQEKVDTFRTSIYFAEQGEIRQIIEENGKFTIEAFEDI-------IH 286
Query: 292 VSDDIDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFR----RFKNKIVKLH 341
++ D + A++ +A + + + FG +M + F + + +I +LH
Sbjct: 287 AKNEFPFDPKTLAIS--FKAFYGAFISAHFGVEVMRKAFELVEVKAREQISRLH 338
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,155,178
Number of Sequences: 26719
Number of extensions: 352547
Number of successful extensions: 1021
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 883
Number of HSP's gapped (non-prelim): 31
length of query: 359
length of database: 11,318,596
effective HSP length: 100
effective length of query: 259
effective length of database: 8,646,696
effective search space: 2239494264
effective search space used: 2239494264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0113.13


