

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0112.2
(150 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
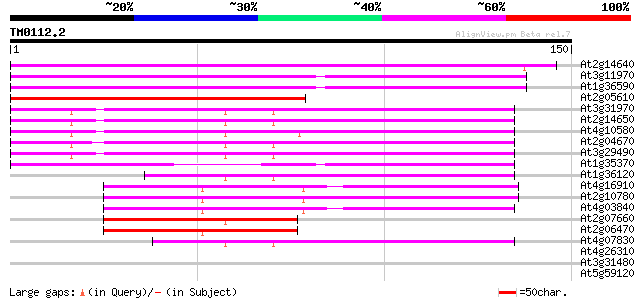
At2g14640 putative retroelement pol polyprotein 106 5e-24
At3g11970 hypothetical protein 102 6e-23
At1g36590 hypothetical protein 102 6e-23
At2g05610 putative retroelement pol polyprotein 79 7e-16
At3g31970 hypothetical protein 68 2e-12
At2g14650 putative retroelement pol polyprotein 63 7e-11
At4g10580 putative reverse-transcriptase -like protein 62 9e-11
At2g04670 putative retroelement pol polyprotein 62 9e-11
At3g29490 hypothetical protein 57 3e-09
At1g35370 hypothetical protein 54 2e-08
At1g36120 putative reverse transcriptase gb|AAD22339.1 54 3e-08
At4g16910 retrotransposon like protein 51 2e-07
At2g10780 pseudogene 51 3e-07
At4g03840 putative transposon protein 48 2e-06
At2g07660 putative retroelement pol polyprotein 45 1e-05
At2g06470 putative retroelement pol polyprotein 45 1e-05
At4g07830 putative reverse transcriptase 42 2e-04
At4g26310 unknown protein 32 0.17
At3g31480 hypothetical protein 31 0.29
At5g59120 cucumisin precursor - like 30 0.64
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 106 bits (264), Expect = 5e-24
Identities = 51/148 (34%), Positives = 87/148 (58%), Gaps = 2/148 (1%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
F+ +WV ++++ R+ ++ R +L+ +YGP+ + + G VAYRL LPEG R+HPVF
Sbjct: 798 FQVGDWVLLRIQPYRQKTLFRRSSQKLSHRFYGPFQVASKHGEVAYRLTLPEGTRIHPVF 857
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H SLLK VG+ ++ L L + P +V+ R+ ++ + + + +QW+G
Sbjct: 858 HVSLLKPWVGDGEPDMGQLPPLRNNGELKLQPTAVLEVRWRSQDKKRVADLLVQWEGLHI 917
Query: 121 DEPTWKDTLNIRSQFP--VFNLEDKVDL 146
++ TW++ + + FP V NLEDKV L
Sbjct: 918 EDATWEEYDQLAASFPEFVLNLEDKVRL 945
>At3g11970 hypothetical protein
Length = 1499
Score = 102 bits (255), Expect = 6e-23
Identities = 53/138 (38%), Positives = 80/138 (57%), Gaps = 2/138 (1%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
FE ++V++KL+ R+ SVV R +L+ Y+GPY II R G VAY+L LP +VHPVF
Sbjct: 1362 FEIGDYVYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPSYSQVHPVF 1421
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H S LK VGN S + L + ++V P V+ + RQ + +V ++W +P
Sbjct: 1422 HVSQLKVLVGNVSTTVHLPSVM--QDVFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPL 1479
Query: 121 DEPTWKDTLNIRSQFPVF 138
+E TW+ +++ FP F
Sbjct: 1480 EEATWEFLFDLQKTFPEF 1497
>At1g36590 hypothetical protein
Length = 1499
Score = 102 bits (255), Expect = 6e-23
Identities = 53/138 (38%), Positives = 80/138 (57%), Gaps = 2/138 (1%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
FE ++V++KL+ R+ SVV R +L+ Y+GPY II R G VAY+L LP +VHPVF
Sbjct: 1362 FEIGDYVYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPSYSQVHPVF 1421
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H S LK VGN S + L + ++V P V+ + RQ + +V ++W +P
Sbjct: 1422 HVSQLKVLVGNVSTTVHLPSVM--QDVFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPL 1479
Query: 121 DEPTWKDTLNIRSQFPVF 138
+E TW+ +++ FP F
Sbjct: 1480 EEATWEFLFDLQKTFPEF 1497
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 79.3 bits (194), Expect = 7e-16
Identities = 38/79 (48%), Positives = 51/79 (64%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
FE ++VF+KL+ R+ SVV R +L+ Y+GPY +I R G VAY+LQLP +VHPVF
Sbjct: 690 FEVGDYVFVKLQPYRQQSVVMRSTQKLSPKYFGPYKVIDRCGEVAYKLQLPANSQVHPVF 749
Query: 61 HASLLKEAVGNNSVELQLL 79
H S L+ VG + LL
Sbjct: 750 HVSQLRVLVGTVTTSTHLL 768
>At3g31970 hypothetical protein
Length = 1329
Score = 67.8 bits (164), Expect = 2e-12
Identities = 40/138 (28%), Positives = 72/138 (51%), Gaps = 5/138 (3%)
Query: 1 FEGFEWVFIKLRALR-ENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRV-HP 58
FE + V++K+ LR N ++ +LT Y GP+ I++R+G VAYRL+LP+ R H
Sbjct: 1184 FEVGDRVYLKMAMLRGPNRSISET--KLTPRYMGPFRIVERVGPVAYRLELPDVMRAFHK 1241
Query: 59 VFHASLLKEAV-GNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQG 117
VFH S+L++ + ++ V ++L+ L P ++ R + +P + + W
Sbjct: 1242 VFHVSMLRKCLHKDDEVLAKILEDLQPNMTLEARPVRILERRIKELRRKKIPLIKVLWNC 1301
Query: 118 KPADEPTWKDTLNIRSQF 135
E TW+ +++ F
Sbjct: 1302 DGVTEETWEPEARMKASF 1319
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 62.8 bits (151), Expect = 7e-11
Identities = 38/138 (27%), Positives = 72/138 (51%), Gaps = 5/138 (3%)
Query: 1 FEGFEWVFIKLRALR-ENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRV-HP 58
FE + V++K+ LR N ++ +L+ Y GP+ I++R+G VAYRL+LP+ R H
Sbjct: 1186 FEVGDRVYLKMAMLRGPNRSISET--KLSPRYMGPFRIVERVGPVAYRLELPDVMRAFHK 1243
Query: 59 VFHASLLKEAV-GNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQG 117
VFH S+L++ + ++ V ++ + L P V+ R + +P + + W
Sbjct: 1244 VFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLWDC 1303
Query: 118 KPADEPTWKDTLNIRSQF 135
+ TW+ ++++F
Sbjct: 1304 DGVTKETWEPEARMKARF 1321
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 62.4 bits (150), Expect = 9e-11
Identities = 38/138 (27%), Positives = 71/138 (50%), Gaps = 5/138 (3%)
Query: 1 FEGFEWVFIKLRALR-ENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRV-HP 58
FE + V++K+ LR N ++ +L+ Y GP+ I++R+ VAYRL+LP+ R H
Sbjct: 1095 FEVGDRVYLKMAMLRGPNRSISET--KLSPRYMGPFKIVERVEPVAYRLELPDVMRAFHK 1152
Query: 59 VFHASLLKEAVGNNSVEL-QLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQG 117
VFH S+L++ + + L ++ + L P V+ R ++ +P + + W
Sbjct: 1153 VFHVSMLRKCLHKDDEALAKIPEDLQPNMTLEARPVRVLERRIKELRQKKIPLIKVLWDC 1212
Query: 118 KPADEPTWKDTLNIRSQF 135
E TW+ ++++F
Sbjct: 1213 DGVTEETWEPEARMKARF 1230
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 62.4 bits (150), Expect = 9e-11
Identities = 38/139 (27%), Positives = 72/139 (51%), Gaps = 7/139 (5%)
Query: 1 FEGFEWVFIKLRALR--ENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRV-H 57
FE + V++K+ LR S++ +L+ Y GP+ I++R+G VAYRL+LP+ R H
Sbjct: 1266 FEVGDRVYLKMAMLRGPNRSILET---KLSPRYMGPFRIVERVGPVAYRLELPDVMRAFH 1322
Query: 58 PVFHASLLKEAV-GNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQ 116
VFH +L++ + ++ V +++ + L P V+ R + +P + + W
Sbjct: 1323 KVFHVLMLRKCLHKDDEVLVKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLWD 1382
Query: 117 GKPADEPTWKDTLNIRSQF 135
E TW+ ++++F
Sbjct: 1383 CDGVTEETWEPEARMKARF 1401
>At3g29490 hypothetical protein
Length = 438
Score = 57.4 bits (137), Expect = 3e-09
Identities = 37/138 (26%), Positives = 69/138 (49%), Gaps = 5/138 (3%)
Query: 1 FEGFEWVFIKLRALR-ENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRV-HP 58
FE + V++K+ LR N ++ +L+ Y GP+ I++R+G VAY L+LP+ R H
Sbjct: 196 FEVGDRVYLKMAMLRGPNRSISET--KLSLRYMGPFRIVERVGPVAYMLELPDVMRAFHK 253
Query: 59 VFHASLLKEAV-GNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQG 117
VFH S+L++ + ++ V ++ + L V+ R Q + + + W
Sbjct: 254 VFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARQVRVLERRIKELQRKKISLIKVLWDC 313
Query: 118 KPADEPTWKDTLNIRSQF 135
E TW+ ++++F
Sbjct: 314 DGVTEETWQPEARMKARF 331
>At1g35370 hypothetical protein
Length = 1447
Score = 54.3 bits (129), Expect = 2e-08
Identities = 36/135 (26%), Positives = 61/135 (44%), Gaps = 25/135 (18%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
F+ ++V++KL+ R+ SVV R +L+ Y+GPY II++ G V
Sbjct: 1333 FDIGDFVYVKLQPYRQQSVVLRVNQKLSPKYFGPYKIIEKCGEV---------------- 1376
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
VGN + QL L ++ P ++ + RQ V ++W G+P
Sbjct: 1377 -------MVGNVTTSTQLPSVL--PDIFEKAPEYILERKLVKRQGRAATMVLVKWIGEPV 1427
Query: 121 DEPTWKDTLNIRSQF 135
+E TWK + + +F
Sbjct: 1428 EEATWKFLFDRQQKF 1442
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 53.9 bits (128), Expect = 3e-08
Identities = 29/101 (28%), Positives = 52/101 (50%), Gaps = 2/101 (1%)
Query: 37 IIQRIGAVAYRLQLPEGGRV-HPVFHASLLKEAV-GNNSVELQLLDHLTGEEVASVHPFS 94
I++R+G VAYRL+LP+ R H VFH S+L++ + ++ V ++ + L P
Sbjct: 1125 IVERVGPVAYRLELPDVMRAFHNVFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARPVR 1184
Query: 95 VITSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQF 135
V+ R + +P + + W E TW+ I+++F
Sbjct: 1185 VLERRIKEVRRKKIPMIKVLWDCDGVTEETWEPEARIKARF 1225
>At4g16910 retrotransposon like protein
Length = 687
Score = 51.2 bits (121), Expect = 2e-07
Identities = 32/117 (27%), Positives = 57/117 (48%), Gaps = 10/117 (8%)
Query: 26 QLTAPYYGPYPIIQRIGAVAYRLQL-PEGGRVHPVFHASLLKEAVGNNSVELQ-----LL 79
+L Y GPY +I+R+GAVAY+L L P+ H VFH S L++ + ++ L
Sbjct: 551 KLRPRYVGPYKVIERVGAVAYKLDLPPKLDAFHNVFHVSQLRKCLSEQEESMEDVPPGLK 610
Query: 80 DHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQFP 136
+++T E P ++ + ++ + I W +E TW+ +++ FP
Sbjct: 611 ENMTVE----AWPVRIMDQMKKGTRGKSMDLLKILWNCGGREEYTWETETKMKANFP 663
>At2g10780 pseudogene
Length = 1611
Score = 50.8 bits (120), Expect = 3e-07
Identities = 29/113 (25%), Positives = 54/113 (47%), Gaps = 2/113 (1%)
Query: 26 QLTAPYYGPYPIIQRIGAVAYRLQL-PEGGRVHPVFHASLLKEAVGNNSVELQ-LLDHLT 83
+L+ Y GPY +I+R+GAVAY+L L P+ H VFH S L++ + + ++ + L
Sbjct: 1467 KLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVEDIPPGLK 1526
Query: 84 GEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQFP 136
P ++ + + + W + +E TW+ +++ FP
Sbjct: 1527 ENMTVEAWPVRIMDRMTKGTRGKARDLLKVLWNCRGREEYTWETENKMKANFP 1579
>At4g03840 putative transposon protein
Length = 973
Score = 48.1 bits (113), Expect = 2e-06
Identities = 30/116 (25%), Positives = 57/116 (48%), Gaps = 10/116 (8%)
Query: 26 QLTAPYYGPYPIIQRIGAVAYRLQL-PEGGRVHPVFHASLLKEAVGNNSVELQ-----LL 79
+L+ Y GPY +I+R+GAVAY+L L P+ H VFH S L++ + N ++ L
Sbjct: 829 KLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSNQEESVEDVPPGLK 888
Query: 80 DHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQF 135
+++T E P ++ + + + + W ++ TW+ +++ F
Sbjct: 889 ENMTVE----AWPVQIMDRMTKGTRGKSRDLLKVLWNCGGREQYTWETENKMKANF 940
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 45.4 bits (106), Expect = 1e-05
Identities = 22/53 (41%), Positives = 34/53 (63%), Gaps = 1/53 (1%)
Query: 26 QLTAPYYGPYPIIQRIGAVAYRLQLPEGGRV-HPVFHASLLKEAVGNNSVELQ 77
+L+ Y GPY +I+R+GAVAY+L LP V H VFH S L++ + + ++
Sbjct: 859 KLSPRYVGPYKVIERVGAVAYKLDLPPKLNVFHNVFHVSQLRKYLSDQEESVE 911
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 45.4 bits (106), Expect = 1e-05
Identities = 21/53 (39%), Positives = 34/53 (63%), Gaps = 1/53 (1%)
Query: 26 QLTAPYYGPYPIIQRIGAVAYRLQL-PEGGRVHPVFHASLLKEAVGNNSVELQ 77
+L+ Y GPY +I+R+GAVAY+L L P+ H VFH S L++ + + ++
Sbjct: 826 KLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVE 878
>At4g07830 putative reverse transcriptase
Length = 611
Score = 41.6 bits (96), Expect = 2e-04
Identities = 25/99 (25%), Positives = 48/99 (48%), Gaps = 2/99 (2%)
Query: 39 QRIGAVAYRLQLPEGGRV-HPVFHASLLKEAV-GNNSVELQLLDHLTGEEVASVHPFSVI 96
QR+G VA+RL+L + R H VFH S+L++ + ++ V ++ + L P V+
Sbjct: 467 QRVGPVAFRLELSDVMRAFHKVFHVSMLRKCLHKDDEVLAKIPEDLQPNMTLEARPVRVL 526
Query: 97 TSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQF 135
R + +P + + E TW+ ++++F
Sbjct: 527 ERRIKELRRKKIPLIKVLRNCDGVTEETWEPEARLKARF 565
>At4g26310 unknown protein
Length = 258
Score = 31.6 bits (70), Expect = 0.17
Identities = 20/70 (28%), Positives = 33/70 (46%), Gaps = 6/70 (8%)
Query: 24 CPQLTAPYYGPYPIIQRIGAVAYRLQLPEG------GRVHPVFHASLLKEAVGNNSVELQ 77
C + T P + P+ +QR G +QL G GR V A ++ G S++++
Sbjct: 52 CCRETPPLHSPWSALQRRGVKVNAIQLRAGNVIERTGRTFRVVEAEHKQQGRGGASIQVE 111
Query: 78 LLDHLTGEEV 87
L D TG ++
Sbjct: 112 LRDVDTGNKL 121
>At3g31480 hypothetical protein
Length = 338
Score = 30.8 bits (68), Expect = 0.29
Identities = 16/47 (34%), Positives = 28/47 (59%), Gaps = 1/47 (2%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYR 47
FE + V++K+ L+ + +L+ Y GP+ I++R+G VAYR
Sbjct: 163 FEVGDRVYLKMDMLQSPKRFILET-KLSPKYMGPFRIVERVGPVAYR 208
>At5g59120 cucumisin precursor - like
Length = 732
Score = 29.6 bits (65), Expect = 0.64
Identities = 30/116 (25%), Positives = 51/116 (43%), Gaps = 16/116 (13%)
Query: 33 GPYPIIQRIGAVA--YRLQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDH---LTGEEV 87
G I++ +GAV YR P+ +HP+ A LL E + L+ D + +
Sbjct: 395 GGLKIVESVGAVGLIYRTPKPDVAFIHPLPAAGLLTEDFESLVSYLESTDSPQAIVLKTE 454
Query: 88 ASVHPFSVITSRFTTRQESTL-----------PQVWIQWQGKPADEPTWKDTLNIR 132
A + S + + F++R +T+ P V I PA EP+ DT +++
Sbjct: 455 AIFNRTSPVIASFSSRGPNTIAVDILKPDITAPGVEILAAYSPAGEPSQDDTRHVK 510
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,542,174
Number of Sequences: 26719
Number of extensions: 137805
Number of successful extensions: 312
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 283
Number of HSP's gapped (non-prelim): 25
length of query: 150
length of database: 11,318,596
effective HSP length: 90
effective length of query: 60
effective length of database: 8,913,886
effective search space: 534833160
effective search space used: 534833160
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0112.2


