

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.7
(402 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
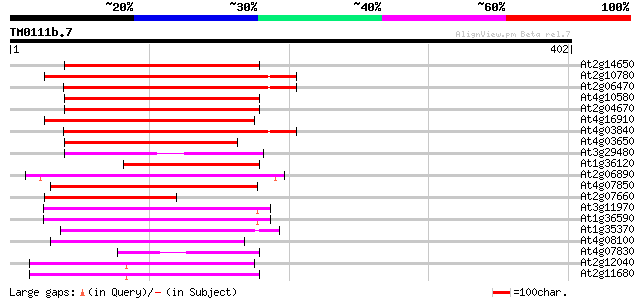
Sequences producing significant alignments: (bits) Value
At2g14650 putative retroelement pol polyprotein 176 2e-44
At2g10780 pseudogene 174 8e-44
At2g06470 putative retroelement pol polyprotein 174 1e-43
At4g10580 putative reverse-transcriptase -like protein 172 3e-43
At2g04670 putative retroelement pol polyprotein 172 4e-43
At4g16910 retrotransposon like protein 171 9e-43
At4g03840 putative transposon protein 169 2e-42
At4g03650 putative reverse transcriptase 148 6e-36
At3g29480 hypothetical protein 125 5e-29
At1g36120 putative reverse transcriptase gb|AAD22339.1 119 2e-27
At2g06890 putative retroelement integrase 115 3e-26
At4g07850 putative polyprotein 115 6e-26
At2g07660 putative retroelement pol polyprotein 108 7e-24
At3g11970 hypothetical protein 105 4e-23
At1g36590 hypothetical protein 105 4e-23
At1g35370 hypothetical protein 95 6e-20
At4g08100 putative polyprotein 95 8e-20
At4g07830 putative reverse transcriptase 91 9e-19
At2g12040 T10J7.2 84 2e-16
At2g11680 putative retroelement pol polyprotein 84 2e-16
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 176 bits (447), Expect = 2e-44
Identities = 84/140 (60%), Positives = 102/140 (72%)
Query: 40 FQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNYL 99
F +K+ LT+ PVL LP EPY VY +AS GLGCVLMQ + +AYAS QL+ HE NY
Sbjct: 740 FVSLKEMLTSTPVLALPEHGEPYMVYTDASGVGLGCVLMQRGKVIAYASRQLRKHEGNYP 799
Query: 100 THDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFT 159
THDLE+A V+FALKIWR YLYG VF++HK LKY+F +LN+ QR WM + DYD
Sbjct: 800 THDLEMAAVIFALKIWRSYLYGGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLE 859
Query: 160 LLYHPGKANVVADALSHQAI 179
+ YHPGKANVVADALSH+ +
Sbjct: 860 IAYHPGKANVVADALSHKRV 879
>At2g10780 pseudogene
Length = 1611
Score = 174 bits (441), Expect = 8e-44
Identities = 89/180 (49%), Positives = 118/180 (65%), Gaps = 1/180 (0%)
Query: 26 EESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMA 85
+++ F+ SF E+K LT APVL LP E EPY VY +AS GLGCVLMQ +A
Sbjct: 919 KDTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSVIA 978
Query: 86 YASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMG 145
YAS QL+ HE+NY THDLE+A VVF LKIWR YLYG+ ++++HK LKY+F +LN+
Sbjct: 979 YASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLYGAKVQIYTDHKSLKYIFTQPELNLR 1038
Query: 146 QRCWMNFIEDYDFTLLYHPGKANVVADALSHQAIHVSSLMVKKLELIETFRDLSLDAKIK 205
QR WM + DY+ + YHPGKAN VADALS + V + +++L+ L ++A K
Sbjct: 1039 QRRWMELVADYNLDIAYHPGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLHVNALSK 1097
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 174 bits (440), Expect = 1e-43
Identities = 90/167 (53%), Positives = 113/167 (66%), Gaps = 1/167 (0%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
SF E+K LT APVL LP E EPY VY +AS GLGCVLMQ +AYAS QL HE+NY
Sbjct: 382 SFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSVIAYASRQLWKHEKNY 441
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDF 158
THDLE+A VVFALKIWR YLYG+ ++++HK LKY+F +LN+ QR WM + DYD
Sbjct: 442 PTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFIQPELNLRQRRWMELVADYDL 501
Query: 159 TLLYHPGKANVVADALSHQAIHVSSLMVKKLELIETFRDLSLDAKIK 205
+ YHPGKAN VADALS + V + +++L+ L ++A K
Sbjct: 502 DIAYHPGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLHVNALSK 547
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 172 bits (436), Expect = 3e-43
Identities = 82/140 (58%), Positives = 100/140 (70%)
Query: 40 FQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNYL 99
F +K+ LT+ PVL LP +PY VY +AS GLGCVLMQH + +AYAS QL HE NY
Sbjct: 765 FVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQHGKVIAYASRQLMKHEGNYP 824
Query: 100 THDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFT 159
THDLE+A V+FALKIWR YLYG VF++HK LKY+F +LN+ QR WM + DYD
Sbjct: 825 THDLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLE 884
Query: 160 LLYHPGKANVVADALSHQAI 179
+ YHPGKANVV DALS + +
Sbjct: 885 IAYHPGKANVVVDALSRKRV 904
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 172 bits (435), Expect = 4e-43
Identities = 82/140 (58%), Positives = 102/140 (72%)
Query: 40 FQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNYL 99
F +K+ LT+ PVL LP +PY VY +AS GLGCVLMQ + +AYAS QL+ HE NY
Sbjct: 791 FVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQRGKVIAYASRQLRKHEGNYP 850
Query: 100 THDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFT 159
THDLE+AVV+FALKIWR YLYG VF++HK LKY+F+ +LN+ Q WM + DYD
Sbjct: 851 THDLEMAVVIFALKIWRSYLYGGKVQVFTDHKSLKYIFNQPELNLRQMRWMELVADYDLE 910
Query: 160 LLYHPGKANVVADALSHQAI 179
+ YHPGKANVVADALS + +
Sbjct: 911 IAYHPGKANVVADALSRKRV 930
>At4g16910 retrotransposon like protein
Length = 687
Score = 171 bits (432), Expect = 9e-43
Identities = 83/150 (55%), Positives = 105/150 (69%)
Query: 26 EESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMA 85
+++ F+ SF E+K LT APVL LP E EPY VY +AS GLGCVL+Q +A
Sbjct: 11 KDTAFNWSEECERSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLIQKGSVIA 70
Query: 86 YASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMG 145
YAS QL+ HE+NY T+DLE+A VVFALKIWR YLYG+ +F++HK LKY+F +LN+
Sbjct: 71 YASRQLRKHEKNYPTNDLEMAAVVFALKIWRSYLYGAKVQIFTDHKSLKYIFTQPELNLR 130
Query: 146 QRCWMNFIEDYDFTLLYHPGKANVVADALS 175
QR W+ + DYD + YHPGKAN V DALS
Sbjct: 131 QRRWIKLVADYDLNIAYHPGKANQVVDALS 160
>At4g03840 putative transposon protein
Length = 973
Score = 169 bits (428), Expect = 2e-42
Identities = 87/167 (52%), Positives = 112/167 (66%), Gaps = 1/167 (0%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
+F E+K LT APVL LP E EPY VY +AS GL CVLMQ +AYAS QL+ HE+NY
Sbjct: 382 NFLELKAMLTNAPVLVLPEEGEPYTVYTDASVVGLECVLMQKGSVIAYASRQLRKHEKNY 441
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDF 158
THDLE+A VVFALKIWR YLYG+ ++++HK LKY+F +LN+ QR WM + DYD
Sbjct: 442 PTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFTQPELNLRQRRWMELVADYDL 501
Query: 159 TLLYHPGKANVVADALSHQAIHVSSLMVKKLELIETFRDLSLDAKIK 205
+ YH GKAN VADALS + V + +++L+ L ++A K
Sbjct: 502 DIAYHAGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLHVNALSK 547
>At4g03650 putative reverse transcriptase
Length = 839
Score = 148 bits (373), Expect = 6e-36
Identities = 70/124 (56%), Positives = 86/124 (68%)
Query: 40 FQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNYL 99
F +K+ LT+ PVL LP EPY VY +AS GLGC LMQ + +AYAS QL+ HE NY
Sbjct: 690 FVSLKEMLTSTPVLALPEHGEPYMVYTDASGVGLGCALMQRGKVIAYASRQLRKHEGNYP 749
Query: 100 THDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFT 159
THDLE+A V+FALKIWR YLYG VF++HK LKY+F +LN+ QR WM + DYD
Sbjct: 750 THDLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLE 809
Query: 160 LLYH 163
+ YH
Sbjct: 810 IAYH 813
>At3g29480 hypothetical protein
Length = 718
Score = 125 bits (313), Expect = 5e-29
Identities = 67/143 (46%), Positives = 87/143 (59%), Gaps = 18/143 (12%)
Query: 40 FQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNYL 99
F +K+ LT+ PVL LP E Y VY +AS GLGCVLMQ + +AYAS QL+ HE NY
Sbjct: 482 FVSLKEILTSTPVLALPEHGESYMVYTDASRVGLGCVLMQRGKVIAYASRQLRKHEGNYP 541
Query: 100 THDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFT 159
THDLE++ VF++HK LKY+ +LN+ QR WM + DYD
Sbjct: 542 THDLEMS------------------AVFTDHKSLKYIITQPELNLRQRRWMELVADYDLE 583
Query: 160 LLYHPGKANVVADALSHQAIHVS 182
+ YH GKA+VVADALS + + V+
Sbjct: 584 IAYHHGKASVVADALSRKRVGVA 606
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 119 bits (299), Expect = 2e-27
Identities = 54/98 (55%), Positives = 69/98 (70%)
Query: 82 EAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KD 141
+ + YAS QL+ HE NY THDLE+A ++FALKIW YLYG VF++HK LK +F +
Sbjct: 723 KVIVYASRQLRKHEGNYPTHDLEMAAIIFALKIWGSYLYGGKVQVFTDHKSLKDIFTQPE 782
Query: 142 LNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQAI 179
LN+ QR WM + DYD + YHPGK NVVADALS + +
Sbjct: 783 LNLRQRRWMELVADYDLEIAYHPGKTNVVADALSRKRV 820
>At2g06890 putative retroelement integrase
Length = 1215
Score = 115 bits (289), Expect = 3e-26
Identities = 71/196 (36%), Positives = 109/196 (55%), Gaps = 10/196 (5%)
Query: 12 RLCKDCRTV----DKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCN 67
R KD T+ + ++ F + +FQ +K +LT APVL L + +E+ C+
Sbjct: 661 RFFKDFSTIVAPLTEVMKKDVGFKWEKAQEEAFQSLKDKLTNAPVLILSEFLKTFEIECD 720
Query: 68 AS*QGLGCVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVF 127
AS G+G VLMQ ++ +A+ S +L NY T+D EL +V AL+ W+HYL+ F +
Sbjct: 721 ASGIGIGAVLMQDQKLIAFFSEKLGGATLNYPTYDKELYALVRALQRWQHYLWPKVFVIH 780
Query: 128 SNHKGLKYLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQAIHVSSLMVK 187
++H+ LK+L + LN W+ FIE + + + Y GK NVVADALS + +S+L VK
Sbjct: 781 TDHESLKHLKGQQKLNKRHARWVEFIETFAYVIKYKKGKDNVVADALSQRYTLLSTLNVK 840
Query: 188 KL------ELIETFRD 197
+ E+ ET D
Sbjct: 841 LMGFEQIKEVYETDHD 856
>At4g07850 putative polyprotein
Length = 1138
Score = 115 bits (287), Expect = 6e-26
Identities = 60/148 (40%), Positives = 91/148 (60%)
Query: 30 FHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASS 89
F + +FQ +K++LT APVL+LP + +E+ C+A G+G VLMQ ++ +AY S
Sbjct: 593 FKWEQAPEDAFQALKEKLTHAPVLSLPDFLKTFEIECDAPGVGIGAVLMQDKKPIAYFSE 652
Query: 90 QLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCW 149
+L NY T+D EL +V AL+ W+HYL+ F + ++H+ LK+L + LN W
Sbjct: 653 KLGGATLNYPTYDKELYALVRALQTWQHYLWPKEFVIHTDHESLKHLKGQQKLNKRHARW 712
Query: 150 MNFIEDYDFTLLYHPGKANVVADALSHQ 177
+ FIE + + + Y GK NVVADALS +
Sbjct: 713 VEFIETFPYVIKYKKGKDNVVADALSRR 740
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 108 bits (269), Expect = 7e-24
Identities = 55/94 (58%), Positives = 65/94 (68%)
Query: 26 EESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMA 85
+++ F+ SF E+K L APVL LP E EPY VY +AS GLGCVLMQ +A
Sbjct: 414 KDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSVIA 473
Query: 86 YASSQLKLHERNYLTHDLELAVVVFALKIWRHYL 119
YAS QL+ HE+NY THDLE+A VVFALKIWR YL
Sbjct: 474 YASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYL 507
>At3g11970 hypothetical protein
Length = 1499
Score = 105 bits (262), Expect = 4e-23
Identities = 58/167 (34%), Positives = 97/167 (57%), Gaps = 4/167 (2%)
Query: 25 SEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAM 84
++ F + +F+++K L APVL+LPL + + V +A QG+G VLMQ +
Sbjct: 859 TKTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEGHPL 918
Query: 85 AYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNM 144
AY S QLK + + ++ EL V+FA++ WRHYL S F + ++ + LKYL + +
Sbjct: 919 AYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTP 978
Query: 145 GQRCWMNFIEDYDFTLLYHPGKANVVADALSH----QAIHVSSLMVK 187
Q+ W+ + ++D+ + Y GK NVVADALS + +H++ +V+
Sbjct: 979 IQQQWLPKLLEFDYEIQYRQGKENVVADALSRVEGSEVLHMAMTVVE 1025
>At1g36590 hypothetical protein
Length = 1499
Score = 105 bits (262), Expect = 4e-23
Identities = 58/167 (34%), Positives = 97/167 (57%), Gaps = 4/167 (2%)
Query: 25 SEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAM 84
++ F + +F+++K L APVL+LPL + + V +A QG+G VLMQ +
Sbjct: 859 TKTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEGHPL 918
Query: 85 AYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNM 144
AY S QLK + + ++ EL V+FA++ WRHYL S F + ++ + LKYL + +
Sbjct: 919 AYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTP 978
Query: 145 GQRCWMNFIEDYDFTLLYHPGKANVVADALSH----QAIHVSSLMVK 187
Q+ W+ + ++D+ + Y GK NVVADALS + +H++ +V+
Sbjct: 979 IQQQWLPKLLEFDYEIQYRQGKENVVADALSRVEGSEVLHMAMTVVE 1025
>At1g35370 hypothetical protein
Length = 1447
Score = 95.1 bits (235), Expect = 6e-20
Identities = 54/157 (34%), Positives = 90/157 (56%), Gaps = 2/157 (1%)
Query: 37 RTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHER 96
+++F +K L APVL LP+ + + V +A QG+ VLMQ +AY S QLK +
Sbjct: 842 QSAFDTLKAVLCNAPVLALPVFDKQFMVETDACGQGIRAVLMQKGHPLAYISRQLKGKQL 901
Query: 97 NYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDY 156
+ ++ EL +FA++ WRHYL S F + ++ + LKYL + + Q+ W+ + ++
Sbjct: 902 HLSIYEKELLAFIFAVRKWRHYLLPSHFIIKTDQRSLKYLLEQRLNTPVQQQWLPKLLEF 961
Query: 157 DFTLLYHPGKANVVADALSHQAIHVSSLMVKKLELIE 193
D+ + Y GK N+VADALS + S ++ L ++E
Sbjct: 962 DYEIQYRQGKENLVADALSR--VEGSEVLHMALSIVE 996
>At4g08100 putative polyprotein
Length = 1054
Score = 94.7 bits (234), Expect = 8e-20
Identities = 49/139 (35%), Positives = 80/139 (57%)
Query: 30 FHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASS 89
F + +FQ +K++LT + V LP + +E+ C+AS G+G VLMQ + +AY S
Sbjct: 706 FKWEDAQENAFQALKEKLTNSSVPILPNFMKSFEIECDASGLGIGAVLMQDHKPIAYFSE 765
Query: 90 QLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCW 149
+L NY T+D EL +V AL+ W+HYL+ F + ++H+ LK+L + LN W
Sbjct: 766 KLGGATLNYPTYDKELYALVRALQTWQHYLWPKKFVIHTDHESLKHLKGQQKLNKRHARW 825
Query: 150 MNFIEDYDFTLLYHPGKAN 168
+ FIE + + + Y +A+
Sbjct: 826 VEFIETFPYVIKYKKREAH 844
>At4g07830 putative reverse transcriptase
Length = 611
Score = 91.3 bits (225), Expect = 9e-19
Identities = 47/102 (46%), Positives = 59/102 (57%), Gaps = 18/102 (17%)
Query: 78 MQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLF 137
MQ + +AY S QLK HE NY THDLE+A V F++HK LKY+F
Sbjct: 1 MQRGKVIAYGSRQLKKHEGNYPTHDLEMAAV------------------FTDHKSLKYIF 42
Query: 138 D*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQAI 179
+LN+ R WM + DYD + YHPGKANVV DALS + +
Sbjct: 43 TQLELNLRLRRWMKLVADYDLEIAYHPGKANVVTDALSRKRV 84
>At2g12040 T10J7.2
Length = 976
Score = 83.6 bits (205), Expect = 2e-16
Identities = 55/165 (33%), Positives = 82/165 (49%), Gaps = 4/165 (2%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + +E F +FQ++K+ L +AP++ P P+EV C+AS +G
Sbjct: 194 KIARPLTSLLCKEVKFEFTQECHDAFQQIKQALISAPIVQPPDWDLPFEVMCDASDFAVG 253
Query: 75 CVLMQHRE----AMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNH 130
VL Q ++ A+ YAS L +RNY T + E VVFA + +R YL G V ++H
Sbjct: 254 AVLGQRKDKKLHAIYYASRTLDDAKRNYATTEKEFLAVVFAFEKFRSYLVGLKVIVHTDH 313
Query: 131 KGLKYLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALS 175
LKYL KD W+ ++++D + G VAD LS
Sbjct: 314 AALKYLMQNKDAKPRLLRWILLLQEFDIEVRDKKGVEKCVADHLS 358
>At2g11680 putative retroelement pol polyprotein
Length = 301
Score = 83.6 bits (205), Expect = 2e-16
Identities = 51/169 (30%), Positives = 87/169 (51%), Gaps = 4/169 (2%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E+ F + T+F+ K+ L +AP++ P P+E+ C+ S +G
Sbjct: 58 KLARPLTRLLCKEAEFIFEEECLTAFKSTKEALVSAPIVQAPNRDYPFEIMCDDSDYAVG 117
Query: 75 CVLMQHRE----AMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNH 130
VL Q + + YAS + + Y T + EL VVFA + +R YL GS TV+++H
Sbjct: 118 AVLGQKIDRKLHVIYYASGTMDDTQVRYATTEKELLAVVFAFEKFRSYLVGSKVTVYTDH 177
Query: 131 KGLKYLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQAI 179
L++++ KD W+ ++++D ++ G N VAD LS I
Sbjct: 178 AALRHIYAKKDTKPRLLRWILLLQEFDMEIVDKKGIENGVADHLSRMRI 226
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.353 0.155 0.544
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,647,793
Number of Sequences: 26719
Number of extensions: 284335
Number of successful extensions: 1308
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 1258
Number of HSP's gapped (non-prelim): 38
length of query: 402
length of database: 11,318,596
effective HSP length: 102
effective length of query: 300
effective length of database: 8,593,258
effective search space: 2577977400
effective search space used: 2577977400
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0111b.7


