

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.4
(74 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
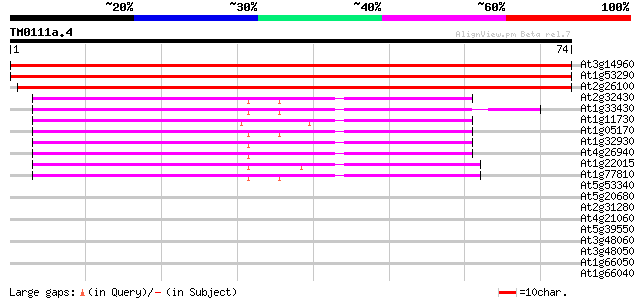
Score E
Sequences producing significant alignments: (bits) Value
At3g14960 galactosyltransferase, putative 137 7e-34
At1g53290 unknown protein (At1g53290) 137 7e-34
At2g26100 unknown protein 112 3e-26
At2g32430 unknown protein 47 2e-06
At1g33430 elicitor response protein, putative 47 2e-06
At1g11730 Avr9 elicitor response-like protein 46 4e-06
At1g05170 putative AVR9 elicitor response protein 45 7e-06
At1g32930 unknown protein 45 9e-06
At4g26940 Avr9 elicitor response like protein 42 4e-05
At1g22015 unknown protein 42 4e-05
At1g77810 similar to Avr9 elicitor response protein emb|CAA06925... 40 2e-04
At5g53340 Avr9 elicitor response protein-like 29 0.49
At5g20680 unknown protein 27 2.4
At2g31280 unknown protein 26 3.2
At4g21060 putative protein 26 4.1
At5g39550 zinc finger -like protein 25 5.4
At3g48060 putative protein 25 5.4
At3g48050 putative protein 25 5.4
At1g66050 hypothetical protein 25 5.4
At1g66040 hypothetical protein 25 5.4
>At3g14960 galactosyltransferase, putative
Length = 343
Score = 137 bits (346), Expect = 7e-34
Identities = 62/74 (83%), Positives = 66/74 (88%)
Query: 1 FRMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELH 60
FRMFSNEDVTIGAWMLAMNVNHEN H LC PEC+ SIAVWDIPKCSGLCNPEKRMLELH
Sbjct: 268 FRMFSNEDVTIGAWMLAMNVNHENLHTLCEPECSPYSIAVWDIPKCSGLCNPEKRMLELH 327
Query: 61 QMESCTQSPTAESD 74
+ESC++SPT SD
Sbjct: 328 MLESCSKSPTLPSD 341
>At1g53290 unknown protein (At1g53290)
Length = 345
Score = 137 bits (346), Expect = 7e-34
Identities = 60/74 (81%), Positives = 68/74 (91%)
Query: 1 FRMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELH 60
FRMF+NEDVTIGAWMLAMNVNHEN+H LC PEC+ +S+AVWDIPKCSGLCNPEKRMLELH
Sbjct: 270 FRMFNNEDVTIGAWMLAMNVNHENHHILCEPECSPSSVAVWDIPKCSGLCNPEKRMLELH 329
Query: 61 QMESCTQSPTAESD 74
+ ESC++SPT SD
Sbjct: 330 KQESCSKSPTLPSD 343
>At2g26100 unknown protein
Length = 333
Score = 112 bits (281), Expect = 3e-26
Identities = 48/73 (65%), Positives = 60/73 (81%)
Query: 2 RMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQ 61
RMF+NEDVTIG+WMLAM+V+HE+N LC P C+ SIAVWDIPKCSGLC+PE R+ ELH+
Sbjct: 257 RMFNNEDVTIGSWMLAMDVHHEDNRALCDPHCSPKSIAVWDIPKCSGLCDPESRLKELHK 316
Query: 62 MESCTQSPTAESD 74
+ C++SPT D
Sbjct: 317 TDMCSKSPTLPPD 329
>At2g32430 unknown protein
Length = 409
Score = 46.6 bits (109), Expect = 2e-06
Identities = 23/67 (34%), Positives = 39/67 (57%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDVT+GAW + ++V H ++ LC P+C + +A +D CSG+C
Sbjct: 330 YANEDVTLGAWFIGLDVTHIDDRRLCCGTPPDCEWKAQAGNICVASFDW-TCSGICRSAD 388
Query: 55 RMLELHQ 61
R+ E+H+
Sbjct: 389 RIKEVHK 395
>At1g33430 elicitor response protein, putative
Length = 395
Score = 46.6 bits (109), Expect = 2e-06
Identities = 25/76 (32%), Positives = 40/76 (51%), Gaps = 12/76 (15%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDV++GAWML + V H + +C P+C + A +D CSG+C
Sbjct: 314 YANEDVSLGAWMLGLEVEHVDERSMCCGTPPDCQWKAQAGNVCAASFDW-SCSGICKSVD 372
Query: 55 RMLELHQMESCTQSPT 70
RM +H+ +C + T
Sbjct: 373 RMARVHR--ACAEGDT 386
>At1g11730 Avr9 elicitor response-like protein
Length = 384
Score = 45.8 bits (107), Expect = 4e-06
Identities = 22/67 (32%), Positives = 40/67 (58%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELC---APECTSTSI------AVWDIPKCSGLCNPEK 54
++NEDV++G+W + +NV H + LC + +C ++ A +D KCSG+C +
Sbjct: 305 YANEDVSLGSWFIGLNVEHVDEKRLCCSTSQDCELKAMMGHVCAASFDW-KCSGICRSAE 363
Query: 55 RMLELHQ 61
RM ++H+
Sbjct: 364 RMADVHE 370
>At1g05170 putative AVR9 elicitor response protein
Length = 404
Score = 45.1 bits (105), Expect = 7e-06
Identities = 22/67 (32%), Positives = 39/67 (57%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDV++GAW + ++V H ++ LC P+C + +A +D CSG+C
Sbjct: 325 YANEDVSLGAWFIGIDVKHIDDRRLCCGTPPDCEWKAQAGNICVASFDW-SCSGICRSAD 383
Query: 55 RMLELHQ 61
R+ E+H+
Sbjct: 384 RIKEVHR 390
>At1g32930 unknown protein
Length = 399
Score = 44.7 bits (104), Expect = 9e-06
Identities = 22/67 (32%), Positives = 37/67 (54%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---------PECTSTSIAVWDIPKCSGLCNPEK 54
++NEDV++G+W + ++V H ++ LC + + A +D CSG+C
Sbjct: 320 YANEDVSLGSWFIGLDVEHIDDRSLCCGTPLDCEWKGQAGNPCAASFDW-SCSGICKSVD 378
Query: 55 RMLELHQ 61
RMLE+HQ
Sbjct: 379 RMLEVHQ 385
>At4g26940 Avr9 elicitor response like protein
Length = 407
Score = 42.4 bits (98), Expect = 4e-05
Identities = 20/66 (30%), Positives = 36/66 (54%), Gaps = 9/66 (13%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA--------PECTSTSIAVWDIPKCSGLCNPEKR 55
+ NEDV++G+W L ++V H ++ LC + + +A +D CSG+C R
Sbjct: 329 YVNEDVSLGSWFLGLDVEHVDDRRLCCGTTDCEWKAQAGNICVASFDW-SCSGICRSADR 387
Query: 56 MLELHQ 61
M ++H+
Sbjct: 388 MKDVHR 393
>At1g22015 unknown protein
Length = 398
Score = 42.4 bits (98), Expect = 4e-05
Identities = 22/68 (32%), Positives = 36/68 (52%), Gaps = 10/68 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECTSTS------IAVWDIPKCSGLCNPEK 54
++NEDVT+G+W + + V ++ C P+C + +A +D KCSG+C
Sbjct: 316 YANEDVTLGSWFIGLEVEQIDDRNFCCGTPPDCEMRAEAGEMCVATFDW-KCSGVCRSVD 374
Query: 55 RMLELHQM 62
RM +H M
Sbjct: 375 RMWMVHVM 382
>At1g77810 similar to Avr9 elicitor response protein emb|CAA06925;
similar to EST gb|AA597316
Length = 414
Score = 40.4 bits (93), Expect = 2e-04
Identities = 19/68 (27%), Positives = 37/68 (53%), Gaps = 10/68 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDV++G+W + + V H ++ C P+C +A ++ CSG+C +
Sbjct: 290 YANEDVSLGSWFIGLEVEHIDDRNFCCGTPPDCRWKAEAGDVCVASFEW-SCSGICKSVE 348
Query: 55 RMLELHQM 62
RM +H++
Sbjct: 349 RMKIVHEV 356
>At5g53340 Avr9 elicitor response protein-like
Length = 338
Score = 28.9 bits (63), Expect = 0.49
Identities = 9/35 (25%), Positives = 22/35 (62%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSI 38
++++DV+ G+W + ++V H + + C +S +I
Sbjct: 300 YAHDDVSTGSWFVGLDVKHVDEGKFCCSAWSSEAI 334
>At5g20680 unknown protein
Length = 551
Score = 26.6 bits (57), Expect = 2.4
Identities = 13/39 (33%), Positives = 16/39 (40%)
Query: 33 CTSTSIAVWDIPKCSGLCNPEKRMLELHQMESCTQSPTA 71
C + I WD S PE L+L E + PTA
Sbjct: 40 CATVVIWTWDRTPTSAFLPPESHYLKLQSEEKVEKLPTA 78
>At2g31280 unknown protein
Length = 720
Score = 26.2 bits (56), Expect = 3.2
Identities = 14/50 (28%), Positives = 23/50 (46%), Gaps = 3/50 (6%)
Query: 25 NHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQMESCTQSPTAESD 74
N+ LC P+ S + P CSG + + +++ + TQ T SD
Sbjct: 169 NNSLCLPKMPSEGLHAEAFPDCSGEVD---KAMDVEESNILTQYKTRRSD 215
>At4g21060 putative protein
Length = 739
Score = 25.8 bits (55), Expect = 4.1
Identities = 12/38 (31%), Positives = 20/38 (52%), Gaps = 5/38 (13%)
Query: 2 RMFSNEDVTIGAWMLAMN-----VNHENNHELCAPECT 34
R+F EDV++G W+ N V + ++ + C CT
Sbjct: 670 RLFKMEDVSMGLWVEQFNASMQPVEYSHSWKFCQYGCT 707
>At5g39550 zinc finger -like protein
Length = 617
Score = 25.4 bits (54), Expect = 5.4
Identities = 9/22 (40%), Positives = 13/22 (58%)
Query: 31 PECTSTSIAVWDIPKCSGLCNP 52
PE ++S W+ P CSG+ P
Sbjct: 44 PESLASSTGEWECPDCSGVVVP 65
>At3g48060 putative protein
Length = 1611
Score = 25.4 bits (54), Expect = 5.4
Identities = 14/48 (29%), Positives = 21/48 (43%)
Query: 22 HENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQMESCTQSP 69
H LC E +S A + KCSG + ++ + Q S + SP
Sbjct: 531 HAKTGNLCGKEDARSSTAGSTLKKCSGGSSRHRKSNNVFQGSSSSASP 578
>At3g48050 putative protein
Length = 1613
Score = 25.4 bits (54), Expect = 5.4
Identities = 14/48 (29%), Positives = 21/48 (43%)
Query: 22 HENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQMESCTQSP 69
H LC E +S A + KCSG + ++ + Q S + SP
Sbjct: 531 HAKTGNLCGKEDARSSTAGSTLKKCSGGSSRHRKSNNVFQGSSSSASP 578
>At1g66050 hypothetical protein
Length = 598
Score = 25.4 bits (54), Expect = 5.4
Identities = 14/48 (29%), Positives = 22/48 (45%), Gaps = 6/48 (12%)
Query: 5 SNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNP 52
S E +T G + +V+ PE ++S W+ P CSG+ P
Sbjct: 24 SEETLTCGTCVTPWHVS------CLLPESLASSTGDWECPDCSGVVVP 65
>At1g66040 hypothetical protein
Length = 622
Score = 25.4 bits (54), Expect = 5.4
Identities = 14/48 (29%), Positives = 22/48 (45%), Gaps = 6/48 (12%)
Query: 5 SNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNP 52
S E +T G + +V+ PE ++S W+ P CSG+ P
Sbjct: 24 SEETLTCGTCVTPWHVS------CLLPESLASSTGDWECPDCSGVVVP 65
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.127 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,820,732
Number of Sequences: 26719
Number of extensions: 59345
Number of successful extensions: 135
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 105
Number of HSP's gapped (non-prelim): 26
length of query: 74
length of database: 11,318,596
effective HSP length: 50
effective length of query: 24
effective length of database: 9,982,646
effective search space: 239583504
effective search space used: 239583504
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0111a.4


