

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.16
(1427 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
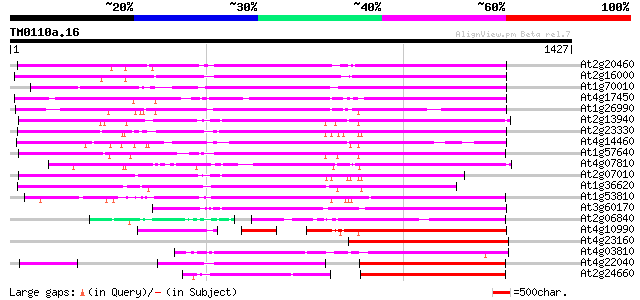
Score E
Sequences producing significant alignments: (bits) Value
At2g20460 putative retroelement pol polyprotein 946 0.0
At2g16000 putative retroelement pol polyprotein 937 0.0
At1g70010 hypothetical protein 909 0.0
At4g17450 retrotransposon like protein 907 0.0
At1g26990 polyprotein, putative 840 0.0
At2g13940 putative retroelement pol polyprotein 814 0.0
At2g23330 putative retroelement pol polyprotein 791 0.0
At4g14460 retrovirus-related like polyprotein 776 0.0
At1g57640 749 0.0
At4g07810 putative polyprotein 742 0.0
At2g07010 putative retroelement pol polyprotein 686 0.0
At1g36620 hypothetical protein 659 0.0
At1g53810 558 e-159
At3g60170 putative protein 473 e-133
At2g06840 putative retroelement pol polyprotein 438 e-122
At4g10990 putative retrotransposon polyprotein 435 e-121
At4g23160 putative protein 418 e-116
At4g03810 putative retrotransposon protein 387 e-107
At4g22040 LTR retrotransposon like protein 369 e-102
At2g24660 putative retroelement pol polyprotein 365 e-101
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 946 bits (2445), Expect = 0.0
Identities = 516/1284 (40%), Positives = 755/1284 (58%), Gaps = 90/1284 (7%)
Query: 21 SSNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDPL 80
+ +PFFLH +D+PGL ++S L+ Y W+ +M I+L KNKLGF++G PRP + DP
Sbjct: 62 TQSPFFLHSADHPGLSIISHRLDETTYGDWSVAMRISLDAKNKLGFVDGSLPRPLESDPN 121
Query: 81 LQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLL 140
+ W R + +V SW+LN VS I S++ A +IW+DL DRF+ N R Y L +++
Sbjct: 122 FRLWSRCNSMVKSWLLNSVSPQIYRSILRLNDATDIWRDLFDRFNLTNLPRTYNLTQEIQ 181
Query: 141 NLQQESLTVTQFYTKHKAIWDELQDYLP-QRSCVCGENRVLLDYFQQEQIMHFLMSLSDS 199
+L+Q +++++++YT K +WD+L C CG+ L ++ +IM FL L++S
Sbjct: 182 DLRQGTMSLSEYYTLLKTLWDQLDSTEALDDPCTCGKAVRLYQKAEKAKIMKFLAGLNES 241
Query: 200 YNQIKSHVLLMNPLPPMNRVFSMVLQEEKQREIAARSADLNSAFAAQATNSGKSGKK--- 256
Y ++ ++ LP + V+ ++ Q+ Q+ A + ++ ++S + +
Sbjct: 242 YAIVRRQIIAKKALPSLAEVYHILDQDNSQKGFFNVVAPPAAFQVSEVSHSPITSPEIMY 301
Query: 257 -----DRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSS----------AVHHVDSS 301
++ RP CS+C ++GH +RC+KKHG+PPG K KSS A +
Sbjct: 302 VQSGPNKGRPTCSFCNRVGHIAERCYKKHGFPPGFTPKGKSSDKPPKPQAVAAQVTLSPD 361
Query: 302 EVSGSSVPQDPPITSSQYQQLLSLLTAQLA---TSPMPSSSTSEPLQIPSPTDLKGFVFS 358
+++G + Q Q L++L ++QL SP +SS E S G +FS
Sbjct: 362 KMTGQLETLAGNFSPDQIQNLIALFSSQLQPQIVSPQTASSQHEASSSQSVAP-SGILFS 420
Query: 359 AS----------SDFSHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKARV 408
S S S + W++DSGA+ HV F++ + ++LP R+
Sbjct: 421 PSTYCFIGILAVSHNSLSSDTWVIDSGATHHVSHDRKLFQTLDTSIVSFVNLPTGPNVRI 480
Query: 409 THIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQE-NKLKTIG 467
+ +GTV + +IL NV ++P F +NL+S+S+L + + F IQ+ K T+G
Sbjct: 481 SGVGTVLINKDIILQNVLFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKGLTLG 540
Query: 468 RGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPS 527
G LY L S P +N++ + SV WH RLGHPS
Sbjct: 541 EGKRIGNLYVLDTQS---------PAISVNAVVDVSV--------------WHKRLGHPS 577
Query: 528 YNRLSLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFH 587
++RL LS + + C VC LAKQK+L F S N+ N +F+L+H DVWGPF
Sbjct: 578 FSRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLHIDVWGPFS 637
Query: 588 VPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAK 647
V T G+++FLTIVDDHSR TW +L+K KS+ AF +V+ Q+ +K VR+DNAK
Sbjct: 638 VETVEGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAK 697
Query: 648 ELQLTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECIST 707
EL T F + +G + SC E P+QNSVVER+HQ +LNVAR+L+FQS+M L +WG+C+ T
Sbjct: 698 ELAFTEFYKAKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLT 757
Query: 708 ATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVL 767
A LIN P+ L NK+P +L + P Y L+ FGCLC+SST RHKF PR+ CV
Sbjct: 758 AVFLINRTPSALLSNKTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRHKFLPRSRACVF 817
Query: 768 VGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQDPFLVPLP*VGLSVP 827
+GY G+KGYKL +L + HISR+V F+E +FP +++ + P+
Sbjct: 818 LGYPFGFKGYKLLDLESNVVHISRNVEFHEELFPLASSQQSATTASDVFTPMD------- 870
Query: 828 PYDVPQPESVPVPSADHIPATPLVSEIVAPSPDAIVPPLAVRRSTRVRHPPGYLADYDC- 886
P+ S + I + L S ++PS ++ RR T+ P +L DY C
Sbjct: 871 ----------PLSSGNSI-TSHLPSPQISPSTQ-----ISKRRITKF---PAHLQDYHCY 911
Query: 887 -PQQTTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAME 945
+ HP+S+ SY +++PS+ +Y ++ P +H+A EW A+ E+ AME
Sbjct: 912 FVNKDDSHPISSSLSYSQISPSHMLYINNISKIPIPQSYHEAKDSKEWCGAIDQEIGAME 971
Query: 946 ANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPV 1005
TW++ SLPPGK +G KWV+ +K H DGS++R KAR+VAKGYTQ+ G+DY +TFSPV
Sbjct: 972 RTDTWEITSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETFSPV 1031
Query: 1006 AKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGY-DVKG----ENLTC 1060
AK+ TV++LL ++A KW L QLDI+NAFLNGDL E +YMK+P GY D+KG N+ C
Sbjct: 1032 AKMATVKLLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVC 1091
Query: 1061 RLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVS 1120
RLKKSIYGLKQASRQWF KFS+ L GF+ + D++LF + G F+VLLVYV+DI+++
Sbjct: 1092 RLKKSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHTLFVRCIGSEFIVLLVYVDDIVIA 1151
Query: 1121 GPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFTD 1180
S+ + LK++FKL++LG LK+FLGLE+A ++ GISLSQR YAL LL D
Sbjct: 1152 STTEQAAQSLTEALKASFKLRELGPLKYFLGLEVARTSEGISLSQRKYALELLTSADMLD 1211
Query: 1181 CRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPT 1240
C+P+S+PM PN++LS + G L D YRRL+G+L+YLTI+RPDI F +NKL QF S P
Sbjct: 1212 CKPSSIPMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPR 1271
Query: 1241 TTHLDALHHLLRYLKTTPGQGFFF 1264
T HL A++ +L+Y+K T GQG F+
Sbjct: 1272 TAHLAAVYKVLQYIKGTVGQGLFY 1295
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 937 bits (2422), Expect = 0.0
Identities = 509/1286 (39%), Positives = 748/1286 (57%), Gaps = 81/1286 (6%)
Query: 13 TSFRIMENSSNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFP 72
TS + + +PFFLH +D+PGL ++S L+ NY W+ +M+I+L KNK GFI+G
Sbjct: 50 TSSESGDPTQSPFFLHSADHPGLNIISHRLDETNYGDWSVAMLISLDAKNKTGFIDGTLS 109
Query: 73 RPADDDPLLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRI 132
RP + D + W R + +V SW+LN VS I S++ A +IW+DL RF+ N R
Sbjct: 110 RPLESDLNFRLWSRCNSMVKSWLLNSVSPQIYRSILRMNDASDIWRDLNSRFNVTNLPRT 169
Query: 133 YQLKKDLLNLQQESLTVTQFYTKHKAIWDELQDYLP-QRSCVCGENRVLLDYFQQEQIMH 191
Y L +++ + +Q +L+++++YT+ K +WD+L C CG+ L +Q +I+
Sbjct: 170 YNLTQEIQDFRQGTLSLSEYYTRLKTLWDQLDSTEALDEPCTCGKAMRLQQKAEQAKIVK 229
Query: 192 FLMSLSDSYNQIKSHVLLMNPLPPMNRVFSMVLQEEKQREI--------AARSADLNSAF 243
FL L++SY ++ ++ LP + V+ ++ Q+ Q+ A + +++ +
Sbjct: 230 FLAGLNESYAIVRRQIIAKKALPSLGEVYHILDQDNSQQSFSNVVAPPAAFQVSEITQSP 289
Query: 244 AAQATNSGKSGKKDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSS---------A 294
+ T ++ RP+CS+ ++GH +RC+KKHG+PPG K K+ A
Sbjct: 290 SMDPTVCYVQNGPNKGRPICSFYNRVGHIAERCYKKHGFPPGFTPKGKAGEKLQKPKPLA 349
Query: 295 VHHVDSSEVSGSSVPQDPPITSSQYQQLLSLLTAQLATSP-----MPSSSTSEPLQI--- 346
+ +SSEV+ S ++ Q QQ +++ ++QL +P S+S S+ L I
Sbjct: 350 ANVAESSEVNTSLESMVGNLSKEQLQQFIAMFSSQLQNTPPSTYATASTSQSDNLGICFS 409
Query: 347 PSPTDLKGFVFSASSDFSHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKA 406
PS G + A S T W++DSGA+ HV S F S + ++LP
Sbjct: 410 PSTYSFIGILTVARHTLSSAT--WVIDSGATHHVSHDRSLFSSLDTSVLSAVNLPTGPTV 467
Query: 407 RVTHIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQEN-KLKT 465
+++ +GT+KL + ++L NV ++P F +NL+S+S+L ++ + F + IQ+ K +
Sbjct: 468 KISGVGTLKLNDDILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRM 527
Query: 466 IGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGH 525
+G+G LY L ++ + +S +WH RLGH
Sbjct: 528 LGQGRRVANLYLLDVGDQSISVNAVVDIS-----------------------MWHRRLGH 564
Query: 526 PSYNRLSLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGP 585
S RL +S ++ + + C VC LAKQ++L F + N FDL+H DVWGP
Sbjct: 565 ASLQRLDAISDSLGTTRHKNKGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGP 624
Query: 586 FHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDN 645
F V T G+++FLTIVDDHSR TW +L+K KSE AF + V+ Q+ +K VR+DN
Sbjct: 625 FSVETVEGYKYFLTIVDDHSRATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDN 684
Query: 646 AKELQLTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECI 705
A EL+ T+F ++G + SC E P+QNSVVER+HQ +LNVAR+L+FQS +PL WG+C+
Sbjct: 685 APELKFTSFYAEKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCV 744
Query: 706 STATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPC 765
TA LIN P+ L NK+P +L P Y+ LR FGCLC+SST P RHKF PR+ C
Sbjct: 745 LTAVFLINRTPSQLLMNKTPYEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRAC 804
Query: 766 VLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQDPFLVPLP*VGLS 825
+ +GY GYKGYKL +L + T ISR+V F+E +FP P S P+ V
Sbjct: 805 LFLGYPSGYKGYKLMDLESNTVFISRNVQFHEEVFPLAK-NPGSESSLKLFTPMVPVSSG 863
Query: 826 VPPYDVPQPESVPVPSADHIPATPLVSEIVAPSPDAIVPPLAVRRSTRVRHPPGYLADYD 885
+ P S+P +D +PP S RVR PP +L DY
Sbjct: 864 IISDTTHSPSSLPSQISD-------------------LPPQI--SSQRVRKPPAHLNDYH 902
Query: 886 CPQQTTPH--PLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAA 943
C + H P+S+ SY K++PS+ Y + PT + +A EW +A+ E+ A
Sbjct: 903 CNTMQSDHKYPISSTISYSKISPSHMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGA 962
Query: 944 MEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFS 1003
ME TW++ +LP GK +G KWV+ LK DG+++R+KARLVAKGYTQ+ G+DY DTFS
Sbjct: 963 MEKTNTWEITTLPKGKKAVGCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFS 1022
Query: 1004 PVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGY-DVKG----ENL 1058
PVAK+TT+++LL ++A KW L QLD++NAFLNG+L EE++MKIP+GY + KG N+
Sbjct: 1023 PVAKMTTIKLLLKVSASKKWFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNV 1082
Query: 1059 TCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDII 1118
RLK+SIYGLKQASRQWF KFSS L GF+ ++ D++LF K FV++LVYV+DI+
Sbjct: 1083 VLRLKRSIYGLKQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKMYDGEFVIVLVYVDDIV 1142
Query: 1119 VSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGF 1178
++ + + + L FKL+DLG LK+FLGLE+A + GIS+ QR YAL LL TG
Sbjct: 1143 IASTSEAAAAQLTEELDQRFKLRDLGDLKYFLGLEVARTTAGISICQRKYALELLQSTGM 1202
Query: 1179 TDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSK 1238
C+P S+PM PNLK+ D G + D QYRR++G+L+YLTI+RPDI F +NKL QF S
Sbjct: 1203 LACKPVSVPMIPNLKMRKDDGDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSA 1262
Query: 1239 PTTTHLDALHHLLRYLKTTPGQGFFF 1264
P TTHL A + +L+Y+K T GQG F+
Sbjct: 1263 PRTTHLTAAYRVLQYIKGTVGQGLFY 1288
>At1g70010 hypothetical protein
Length = 1315
Score = 909 bits (2348), Expect = 0.0
Identities = 511/1238 (41%), Positives = 729/1238 (58%), Gaps = 116/1238 (9%)
Query: 54 MMIALSVKNKLGFINGDFPRPADDDPLLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSA 113
M ++ KNKLGF++G P+P DDDP + W R + +V SW+LN VSK+I +S++ +A
Sbjct: 1 MTTSIEAKNKLGFVDGSIPKPDDDDPYCKIWRRCNSMVKSWLLNSVSKEIYTSILYFPTA 60
Query: 114 KEIWQDLQDRFHQPNGTRIYQLKKDLLNLQQESLTVTQFYTKHKAIWDELQDYLPQRSCV 173
IW+DL RFH+ + R+Y+L++ + +L+Q +L ++ ++T+ + +W+EL V
Sbjct: 61 AAIWKDLYTRFHKSSLPRLYKLRQQIHSLRQGNLDLSSYHTRTQTLWEELTSLQAVPRTV 120
Query: 174 CGENRVLLDYFQQEQIMHFLMSLSDSYNQIKSHVLLMNPLPPMNRVFSMVLQEEKQRE-- 231
LL + +++ FLM L+D Y+ ++S +L+ LP ++ VF+M+ Q+E QR
Sbjct: 121 ----EDLLIERETNRVIDFLMGLNDCYDTVRSQILMKKTLPSLSEVFNMIDQDETQRSAR 176
Query: 232 IAARSADLNSAFAA--QATNSGKSGK--KDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLN 287
I+ +S F Q++ S +G + ++RP+CSYC + GH D C+KKHGYP
Sbjct: 177 ISTTPGMTSSVFPVSNQSSQSALNGDTYQKKERPVCSYCSRPGHVEDTCYKKHGYPTSFK 236
Query: 288 FKNKS-----SAVHHVDSSEVSGSSVPQDPPITSSQYQQLLSLLTAQLATSPMPSSSTSE 342
K K SA + S EV ++ +T+SQ QQL+S L+++L P S+ +
Sbjct: 237 SKQKFVKPSISANAAIGSEEVVNNTSVSTGDLTTSQIQQLVSFLSSKL----QPPSTPVQ 292
Query: 343 PLQIPSPTDLKGFVFSASSDFSHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPD 402
P ++ S S SSD S ++V C +
Sbjct: 293 P-EVHS--------ISVSSDPSSSSTV--------CPIS--------------------- 314
Query: 403 NTKARVTHIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQE-N 461
G+V LG LIL++V ++P F NLLSVS+L ++ I F + V+Q+
Sbjct: 315 ---------GSVHLGRHLILNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDAT 365
Query: 462 KLKTIGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHA 521
+ +G G LY + L+SL + S S + LWH
Sbjct: 366 RELMVGMGKQVANLYIVD----------------LDSLSHPGTDSSITVASVTSHDLWHK 409
Query: 522 RLGHPSYNRLSLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCD 581
RLGHPS +L +SS + + + C VC ++KQK LPFVS N+ ++ FDLIH D
Sbjct: 410 RLGHPSVQKLQPMSSLLSFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDLIHID 469
Query: 582 VWGPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMV 641
WGPF V TH G+R+FLTIVDD+SR TW +L+++KS+ I F MV+ QF IK V
Sbjct: 470 TWGPFSVQTHDGYRYFLTIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGV 529
Query: 642 RTDNAKELQLTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFW 701
R+DNA EL T F +G + SC E PQQNSVVER+HQ +LNVARSL FQSH+P+ +W
Sbjct: 530 RSDNAPELNFTQFYHSKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYW 589
Query: 702 GECISTATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPR 761
G+CI TA +LIN +P P L +K P +L + P+YDH++VFGCLC++ST P RHKFSPR
Sbjct: 590 GDCILTAVYLINRLPAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPR 649
Query: 762 AVPCVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQDPFLVPLP* 821
A C +GY G+KGYKL +L T + +SR VVF+E +FPF + Q+ F
Sbjct: 650 AKACAFIGYPSGFKGYKLLDLETHSIIVSRHVVFHEELFPFLGSDLSQEEQNFF------ 703
Query: 822 VGLSVPPYDVPQPESVPVP-----SADHI-PATPLVSEIVAPS--PDAIVPPLAVRRSTR 873
P+ P P S+DH+ P+ S + PS P VP +V+ S R
Sbjct: 704 ------------PDLNPTPPMQRQSSDHVNPSDSSSSVEILPSANPTNNVPEPSVQTSHR 751
Query: 874 VRHPPGYLADYDCPQ--QTTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYP 931
P YL DY C +TPH + + SYD++ Y + + EP+ + +A +
Sbjct: 752 KAKKPAYLQDYYCHSVVSSTPHEIRKFLSYDRINDPYLTFLACLDKTKEPSNYTEAEKLQ 811
Query: 932 EWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYT 991
W+ AM E +E TW+V SLP K IG +W++K+K + DGS++R+KARLVA+GYT
Sbjct: 812 VWRDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYT 871
Query: 992 QQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGY 1051
Q+ GIDY +TFSPVAK+ +V++LL +AA +K L QLDI+NAFLNGDL EE+YM++PQGY
Sbjct: 872 QKEGIDYNETFSPVAKLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGY 931
Query: 1052 -----DVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDT 1106
D N CRLKKS+YGLKQASRQW+ KFSS L GF SY D++ F K S
Sbjct: 932 ASRQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKISDGI 991
Query: 1107 FVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQR 1166
F+ +LVY++DII++ N +D +K +KS FKL+DLG+LK+FLGLEI S GI +SQR
Sbjct: 992 FLCVLVYIDDIIIASNNDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVRSDKGIHISQR 1051
Query: 1167 AYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIA 1226
YAL LL +TG C+P+S+PMDP++ + D+G + + YRRLIGRL+YL I+RPDI
Sbjct: 1052 KYALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDIT 1111
Query: 1227 FTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQGFFF 1264
F +NKL+QF P HL A++ +L+Y+K T GQG F+
Sbjct: 1112 FAVNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQGLFY 1149
>At4g17450 retrotransposon like protein
Length = 1433
Score = 907 bits (2344), Expect = 0.0
Identities = 515/1299 (39%), Positives = 740/1299 (56%), Gaps = 134/1299 (10%)
Query: 12 NTSFRIMENSSNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDF 71
+ S +N+ +P+FLH SD+PGL +VS L+G NYN+W+ +M ++L KNKL F++G
Sbjct: 55 SASIESYDNAHSPYFLHSSDHPGLNIVSHILDGTNYNNWSIAMRMSLDAKNKLSFVDGSL 114
Query: 72 PRPADDDPLLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTR 131
PRP D + + W R + +V +W+LN V+ E+W DL RF N R
Sbjct: 115 PRPDVSDRMFKIWSRCNSMVKTWLLNVVT--------------EMWNDLFSRFRVSNLPR 160
Query: 132 IYQLKKDLLNLQQESLTVTQFYTKHKAIWDELQD--YLPQRSCVCGENRVLLDYFQQEQI 189
YQL++ + L+Q +L ++ +YTK K +W++L + L R C C + LL+ + +I
Sbjct: 161 KYQLEQSIHTLKQGNLDLSTYYTKKKTLWEQLANTRVLTVRKCNCEHVKELLEEAETSRI 220
Query: 190 MHFLMSLSDSYNQIKSHVLLMNPLPPMNRVFSMVLQEEKQREIAARS-ADLNSAFAAQAT 248
+ FLM L+D++ I+ +L M P P + +++M+ Q+E QR + + ++ +AF QA+
Sbjct: 221 IQFLMGLNDNFAHIRGQILNMKPRPGLTEIYNMLDQDESQRLVGNPTLSNPTAAFQVQAS 280
Query: 249 ----NSGKSGKKDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSSAVHHVDSSEVS 304
+ + +P CSYC KLGH VD+C+KKHGYPPG + K + + +
Sbjct: 281 PIIDSQVNMAQGSYKKPKCSYCNKLGHLVDKCYKKHGYPPGSKW-TKGQTIGSTNLASTQ 339
Query: 305 GSSVPQDP--------PITSSQYQQLLSLLTAQL---ATSPMP---SSSTSEPLQIPSPT 350
V + P ++ Q Q ++S L+ +L + SPMP S+S S +P +
Sbjct: 340 LQPVNETPNEKTDSYEEFSTDQIQTMISYLSTKLHIASASPMPTTSSASISASPSVPMIS 399
Query: 351 DLKGFVFSASSDFSHKTSV-------------WILDSGASCHVCFHLSSFESYHSVRSHT 397
+ G S S+ + + W++DSGA+ HV + + ++ S+ +
Sbjct: 400 QISGTFLSLFSNAYYDMLISSVSQEPAVSPRGWVIDSGATHHVTHNRDLYLNFRSLENTF 459
Query: 398 ISLPDNTKARVTHIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFV 457
+ LP++ ++ IG ++L +++ LHNV Y+P F NL+S
Sbjct: 460 VRLPNDCTVKIAGIGFIQLSDAISLHNVLYIPEFKFNLIS-------------------- 499
Query: 458 IQENKLKTIGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAA 517
+ K IGRG LY L + NN T L S+C + S C+ +
Sbjct: 500 -ELTKELMIGRGSQVGNLYVLDFNENN----HTVSLKGTTSMCPEF---SVCSSVVVDSV 551
Query: 518 LWHARLGHPSYNRLSLLSSTIHCKIPSSINEN-----VCPVCPLAKQKRLPFVSENHFAN 572
WH RLGHP+Y+++ LLS ++ K+ E+ VC VC L+KQK L F S + +
Sbjct: 552 TWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHLSFQSRQNMCS 611
Query: 573 HSFDLIHCDVWGPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQT 632
+FDL+H D WGPF VPT+ TW +L+K+KS+ AF MV T
Sbjct: 612 AAFDLVHIDTWGPFSVPTNDA--------------TWIYLLKNKSDVLHVFPAFINMVHT 657
Query: 633 QFGKVIKMVRTDNAKELQLTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLF 692
Q+ +K VR+DNA EL+ T G + SC E P+QNSVVER+HQ +LNVAR+LLF
Sbjct: 658 QYQTKLKSVRSDNAHELKFTDLFAAHGIVAYHSCPETPEQNSVVERKHQHILNVARALLF 717
Query: 693 QSHMPLVFWGECISTATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLP 752
QS++PL FWG+C+ TA LIN +PTP L+NKSP L P+Y+ L+ FGCLC+SST P
Sbjct: 718 QSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYESLKTFGCLCYSSTSP 777
Query: 753 STRHKFSPRAVPCVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQ 812
RHKF PRA CV +GY GYKGYKL ++ T ISR V+F+E IFPF ++ +
Sbjct: 778 KQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHEDIFPFISSTIKDDIK 837
Query: 813 DPFLVPLP*VGLSVPPYDVPQPESVPVPSADHIPATPLVSEIVAPSPDAIVP--PLAVRR 870
D F P + D+P ++ + + H + S A+VP PL
Sbjct: 838 DFF----PLLQFPARTDDLPLEQTSIIDTHPHQDVS---------SSKALVPFDPL---- 880
Query: 871 STRVRHPPGYLADYDCPQQTTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQY 930
S R + PP +L D+ C Y+ T + + + + P + +A +
Sbjct: 881 SKRQKKPPKHLQDFHC--------------YNNTTEPFHAFINNITNAVIPQRYSEAKDF 926
Query: 931 PEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGY 990
W AM+ E+ AM TW VVSLPP K IG KWV+ +K + DGSI+R+KARLVAKGY
Sbjct: 927 KAWCDAMKEEIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGY 986
Query: 991 TQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQG 1050
TQ+ G+DY +TFSPVAK+T+VRM+L LAA KW + QLDI+NAFLNGDL EE+YMKIP G
Sbjct: 987 TQEEGLDYEETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPG 1046
Query: 1051 Y-DVKGENL----TCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGD 1105
Y D+ GE L CRL KSIYGLKQASRQW+ K S+ L GFQ S D++LF K +
Sbjct: 1047 YADLVGEALPPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKYANG 1106
Query: 1106 TFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQ 1165
+ +LVYV+DI++ + + LKS FKL+DLG K+FLG+EIA S GIS+ Q
Sbjct: 1107 VLMGVLVYVDDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKGISICQ 1166
Query: 1166 RAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDI 1225
R Y L LL TGF +P+S+P+DP++KL+ + G L DS+ YR+L+G+L+YL I+RPDI
Sbjct: 1167 RKYILELLSTTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDI 1226
Query: 1226 AFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQGFFF 1264
A+ +N L QF PT+ HL A+H +LRYLK T GQG F+
Sbjct: 1227 AYAVNTLCQFSHAPTSVHLSAVHKVLRYLKGTVGQGLFY 1265
>At1g26990 polyprotein, putative
Length = 1436
Score = 840 bits (2170), Expect = 0.0
Identities = 492/1316 (37%), Positives = 730/1316 (55%), Gaps = 160/1316 (12%)
Query: 15 FRIMENSSNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRP 74
F++ ++ NP LH SD+PGL +V+ L+G NYNSW+ +M I+L KNKLGF++G RP
Sbjct: 48 FKVSDSGENPLLLHSSDHPGLSIVAHILDGSNYNSWSIAMRISLDAKNKLGFVDGSLLRP 107
Query: 75 ADDDPLLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQ 134
+ DD + W R + +V N R YQ
Sbjct: 108 SVDDSTFRIWSRCNSMVN-----------------------------------NLPRRYQ 132
Query: 135 LKKDLLNLQQESLTVTQFYTKHKAIWDELQDYLPQ--RSCVCGENRVLLDYFQQEQIMHF 192
L++ ++ LQQ L ++ ++TK K +W++L + + + C C + + LL+ + +++ F
Sbjct: 133 LEQAVMTLQQGKLDLSTYFTKKKTLWEQLANTKSRSVKKCDCDQVKELLEEAETSRVIQF 192
Query: 193 LMSLSDSYNQIKSHVLLMNPLPPMNRVFSMVLQEEKQREI--AARSADLNSAFAAQAT-- 248
LM LSD +N I+S + M P P +N +++M+ Q+E QR + AA+S S A Q
Sbjct: 193 LMGLSDDFNTIRSQIFNMKPRPGLNEIYNMLDQDESQRLVGFAAKSVPSPSPAAFQTQGV 252
Query: 249 ----NSGKSGKKDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSSAVHHVDSSEVS 304
N+ + + +P C++C ++GH+VD+C+K HGYPPG +++ V + +
Sbjct: 253 LNDQNTILLAQGNFKKPKCTHCNRIGHTVDKCYKVHGYPPGHPRAKENTYVGSTNLASTD 312
Query: 305 GSSVPQDPPITSSQY--------QQLLSLLTAQL---------------ATSPMPS---- 337
P ++++ + QQL+S L+ +L +++P+PS
Sbjct: 313 QIETQAPPTMSATGHETMSNDHIQQLISYLSTKLQSPSITSCFDKAIASSSNPVPSISQI 372
Query: 338 -----SSTSEPLQIPSPTDLKGFVFSA--SSDFSHKTS-----------VWILDSGASCH 379
+S+S P +PS + + G FS S+ + TS W++DSGAS H
Sbjct: 373 TDKAIASSSNP--VPSISQITGTFFSLYDSTYYEMLTSSIPIETELSLRAWVIDSGASHH 430
Query: 380 VCFHLSSFESYHSVRSHTISLPDNTKARVTHIGTVKLGNSLILHNVCYVPTFTVNLLSVS 439
V + + +Y ++ + LP+ ++ G ++L ++L LHNV ++P F NLLSVS
Sbjct: 431 VTHERNLYHTYKALDRTFVRLPNGHTVKIEGTGFIQLTDALSLHNVLFIPEFKFNLLSVS 490
Query: 440 ALLENPKYSIHFFHKTFVIQE-NKLKTIGRGDTHNGLYYLHASSNNVFSASTSPLSVLNS 498
L + + + F +IQ K +G+G LY L+ + V +S SV +S
Sbjct: 491 VLTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVDVSSFPGKSVCSS 550
Query: 499 LCNQSVFPSHCNKSTCTAALWHARLGHPSYNRLSLLSSTIHC-KIPSSINENVCPVCPLA 557
+ N+S +WH RLGHPS+ ++ LS + K + + + C VC L+
Sbjct: 551 VKNES-------------EMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLS 597
Query: 558 KQKRLPFVSENHFANHSFDLIHCDVWGPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKS 617
KQK LPF S NH +F+L+H D WGPF VPT +R+FLTIVDD SR TW +L+K KS
Sbjct: 598 KQKHLPFKSVNHIREKAFELVHIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLLKQKS 657
Query: 618 EAPLAIMAFFKMVQTQFGKVIKMVRTDNAKELQLTAFLEKQGTLHQFSCVERPQQNSVVE 677
+ +F KMV+TQ+ + VR+DNA EL+ K+G C E P+QN VVE
Sbjct: 658 DVLTVFPSFLKMVETQYHTKVCSVRSDNAHELKFNELFAKEGIKADHPCPETPEQNFVVE 717
Query: 678 RRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINLIPTPNLHNKSPSTLLYQQDPSYD 737
R+HQ LLNVAR+L+FQS +PL +WG+C+ TA LIN + +P ++N++P L + P Y
Sbjct: 718 RKHQHLLNVARALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKGKPDYS 777
Query: 738 HLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNE 797
L+ FGCLC+ ST P +R KF PRA C+ +GY GYKGYKL ++ T + ISR V+F E
Sbjct: 778 SLKAFGCLCYCSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRHVIFYE 837
Query: 798 SIFPFQNTRPYVYSQDPFLVPLP*VGLSVPPYD--VPQPESVPVPSADHIPATPLVSEIV 855
IFPF ++ ++D F P + L P D +P +S +H ++ S I
Sbjct: 838 DIFPFASSNITDAAKDFF----PHIYLPAPNNDEHLPLVQSSSDAPHNHDESS---SMIF 890
Query: 856 APSPDAIVPPLAVRRSTRVRHPPGYLADYDC------PQQTTPHPLSTYYSYDKLTPSYK 909
PS +STR R P +L D+ C +T+P+PL+ Y SY L+ +
Sbjct: 891 VPSEP---------KSTRQRKLPSHLQDFHCYNNTPTTTKTSPYPLTNYISYSYLSEPFG 941
Query: 910 VYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYK 969
+ + + P + +A W AM E++A TW + LP GK +G KW+
Sbjct: 942 AFINIITATKLPQKYSEARLDKVWNDAMGKEISAFVRTGTWSICDLPAGKVAVGCKWIIT 1001
Query: 970 LKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLD 1029
+K DGSI+RHKARLVAKGYTQQ GID+ +TFSPVAK+ TV++LL+LA KW L QLD
Sbjct: 1002 IKFLADGSIERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLAPKMKWYLHQLD 1061
Query: 1030 INNAFLNGDLFEEVYMKIPQGY-DVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHG 1088
I+NA LNGDL EE+YMK+P GY +++G+ ++ K HG
Sbjct: 1062 ISNALLNGDLEEEIYMKLPPGYSEIQGQEVSPNAK----------------------CHG 1099
Query: 1089 FQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKF 1148
D++LF K F+V+LVYV+DI+++ + L S F+L+DLG+ KF
Sbjct: 1100 ------DHTLFVKAQDGFFLVVLVYVDDILIASTTEAASAELTSQLSSFFQLRDLGEPKF 1153
Query: 1149 FLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQY 1208
FLG+EIA +A GISL QR Y L LL + F+DC+P+S+PM+PN KLS DTG L D QY
Sbjct: 1154 FLGIEIARNADGISLCQRKYVLDLLASSDFSDCKPSSIPMEPNQKLSKDTGTLLEDGKQY 1213
Query: 1209 RRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQGFFF 1264
RR++G+L YL ++RPDI F ++KL+Q+ S PT HL ALH +LRYLK T GQG F+
Sbjct: 1214 RRILGKLQYLCLTRPDINFAVSKLAQYSSAPTDIHLQALHKILRYLKGTIGQGLFY 1269
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 814 bits (2102), Expect = 0.0
Identities = 484/1360 (35%), Positives = 714/1360 (51%), Gaps = 157/1360 (11%)
Query: 23 NPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDPLLQ 82
+P+ L SDNPG ++ S LNG+NYN W M+ AL K K GFING PRP +DP +
Sbjct: 30 SPYTLASSDNPGAVISSVELNGDNYNQWATEMLNALQAKRKTGFINGTIPRPPPNDPNYE 89
Query: 83 SWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLLNL 142
+W + +++ WI + + ++V A +W+DL+ RF N RI+Q++ L +
Sbjct: 90 NWTAVNSMIVGWIRTSIEPKVKATVTFISDAHLLWKDLKQRFSVGNKVRIHQIRAQLSSC 149
Query: 143 QQESLTVTQFYTKHKAIWDELQDYLPQRSCVCGENRV-----LLDYFQQEQIMHFLMSLS 197
+Q+ V ++Y + +W+E Y P C CG R ++E+I F++ L
Sbjct: 150 RQDGQAVIEYYGRLSNLWEEYNIYKPVTVCTCGLCRCGATSEPTKEREEEKIHQFVLGLD 209
Query: 198 DS-YNQIKSHVLLMNPLPPMNRVFSMVLQEEK---------QREIA----ARSADLN--- 240
+S + + + ++ M+PLP + ++S V++EE+ Q+E A AR L+
Sbjct: 210 ESRFGGLCATLINMDPLPSLGEIYSRVIREEQRLASVHVREQKEEAVGFLARREQLDHHS 269
Query: 241 ----SAFAAQATNSGKSGKKDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSSAVH 296
S+ ++ T +S + R CS CG+ GH C++ G+P + +N +
Sbjct: 270 RVDASSSRSEHTGGSRSNSIIKGRVTCSNCGRTGHEKKECWQIVGFPDWWSERNGGRGSN 329
Query: 297 ----------------HVDSSEVSGSSVPQDPPITSSQYQQLLSLLTAQLATSPMPSSST 340
V ++ + S+ P T ++ ++LS L + + S S++
Sbjct: 330 GRGRGGRGSNGGRGQGQVMAAHATSSNSSVFPEFTE-EHMRVLSQLVKEKSNSGSTSNNN 388
Query: 341 SEPLQIPSPTDLKGFVFSASSDFSHKTSVW--ILDSGASCHVCFHLSSFESYHSVRSHTI 398
S+ L S KT + ILDSGAS H+ LSS + V +
Sbjct: 389 SDRL-------------------SGKTKLGDIILDSGASHHMTGTLSSLTNVVPVPPCPV 429
Query: 399 SLPDNTKARVTHIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVI 458
D +KA +G + L N++ L NV +VP+ L+SVS LL+ + F +
Sbjct: 430 GFADGSKAFALSVGVLTLSNTVSLTNVLFVPSLNCTLISVSKLLKQTQCLATFTDTLCFL 489
Query: 459 QENKLKT-IGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAA 517
Q+ KT IG G+ G+YYL +P + H A
Sbjct: 490 QDRSSKTLIGSGEERGGVYYL---------TDVTPAKI------------HTANVDSDQA 528
Query: 518 LWHARLGHPSYNRLSLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDL 577
LWH RLGHPS++ LS L + K S++ + C VC AKQ R F + F L
Sbjct: 529 LWHQRLGHPSFSVLSSLP--LFSKTSSTVTSHSCDVCFRAKQTREVFPESINKTEECFSL 586
Query: 578 IHCDVWGPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKV 637
IHCDVWGP+ VP G +FLTIVDD+SR WT+L+ KSE + F K + QFGK
Sbjct: 587 IHCDVWGPYRVPASCGAVYFLTIVDDYSRAVWTYLLLEKSEVRQVLTNFLKYAEKQFGKT 646
Query: 638 IKMVRTDNAKELQ-LTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHM 696
+KMVR+DN E L+++ + G +HQ SCV PQQN VER+H+ +LNVAR+LLFQ+ +
Sbjct: 647 VKMVRSDNGTEFMCLSSYFRENGIIHQTSCVGTPQQNGRVERKHRHILNVARALLFQASL 706
Query: 697 PLVFWGECISTATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRH 756
P+ FWGE I TA +LIN P+ L ++P +L+ P Y LRVFG C+ + +
Sbjct: 707 PIKFWGESILTAAYLINRTPSSILSGRTPYEVLHGSKPVYSQLRVFGSACYVHRVTRDKD 766
Query: 757 KFSPRAVPCVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPF---------QNTRP 807
KF R+ C+ VGY G KG+K+Y++ F +SRDV+F E +FP+ + P
Sbjct: 767 KFGQRSRSCIFVGYPFGKKGWKVYDIERNEFLVSRDVIFREEVFPYAGVNSSTLASTSLP 826
Query: 808 YVYSQDPFLVPLP*VGLSVP-------------------------------PYDVP--QP 834
V D + +P V S+ P D P P
Sbjct: 827 TVSEDDDWAIPPLEVRGSIDSVETERVVCTTDEVVLDTSVSDSEIPNQEFVPDDTPPSSP 886
Query: 835 ESVPVPSADHIPATPLVSEIVAPSPDAIVPPLAVRRSTRVRHPPGYLADY---------- 884
SV + + P TP+V + +P P V P R+S R HPP L DY
Sbjct: 887 LSVSPSGSPNTPTTPIVVPVASPIP---VSPPKQRKSKRATHPPPKLNDYVLYNAMYTPS 943
Query: 885 -------DCPQQTTP-----HPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPE 932
D Q +T PL+ Y S + S++ Y + + EP +F +A+Q
Sbjct: 944 SIHALPADPSQSSTVPGKSLFPLTDYVSDAAFSSSHRAYLAAITDNVEPKHFKEAVQIKV 1003
Query: 933 WQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQ 992
W AM TE+ A+E N+TWD+V LPPGK IGS+WV+K K + DG+++R+KARLV +G Q
Sbjct: 1004 WNDAMFTEVDALEINKTWDIVDLPPGKVAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQ 1063
Query: 993 QNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGYD 1052
G DY +TF+PV ++TTVR LL A +WE++Q+D++NAFL+GDL EEVYMK+P G+
Sbjct: 1064 VEGEDYKETFAPVVRMTTVRTLLRNVAANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFR 1123
Query: 1053 VKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLV 1112
+ CRL+KS+YGLKQA R WF K S L + GF SY DYSLF+ + + +L+
Sbjct: 1124 HSHPDKVCRLRKSLYGLKQAPRCWFKKLSDSLLRFGFVQSYEDYSLFSYTRNNIELRVLI 1183
Query: 1113 YVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSL 1172
YV+D+++ G + + K L F +KDLGKLK+FLG+E++ GI LSQR YAL +
Sbjct: 1184 YVDDLLICGNDGYMLQKFKDYLSRCFSMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALDV 1243
Query: 1173 LHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKL 1232
+ D+G RP P++ N L+SD GP L+D YRRL+GRLLYL +RP+++++++ L
Sbjct: 1244 IADSGNLGSRPAHTPLEQNHHLASDDGPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVL 1303
Query: 1233 SQFLSKPTTTHLDALHHLLRYLKTTPGQGFFFLGQQPPLS 1272
+QF+ P H DA ++RYLK +PGQG L P L+
Sbjct: 1304 AQFMQNPREAHFDAALRVVRYLKGSPGQG-ILLNADPDLT 1342
>At2g23330 putative retroelement pol polyprotein
Length = 1496
Score = 791 bits (2042), Expect = 0.0
Identities = 486/1364 (35%), Positives = 704/1364 (50%), Gaps = 171/1364 (12%)
Query: 20 NSSNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDP 79
+S+ + ++ SDNPG ++ S L NY W+ + L K KLGFI+G P+PA D P
Sbjct: 17 SSTTAYLINASDNPGALISSVVLKENNYAEWSEELQNFLRAKQKLGFIDGSIPKPAAD-P 75
Query: 80 LLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDL 139
L WI + +++ WI + I S+V A ++W++L+ RF NG R LK ++
Sbjct: 76 ELSLWIAINSMIVGWIRTSIDPTIRSTVGFVSEASQLWENLRRRFSVGNGVRKTLLKDEI 135
Query: 140 LNLQQESLTVTQFYTKHKAIWDELQDYLPQRSCVCGENRVLLDYFQQEQIMHFLMSLSDS 199
Q+ V +Y + +W+ELQ+Y R C C + + +++ FL+ L
Sbjct: 136 AACTQDGQPVLAYYGRLIKLWEELQNYKSGRECKCEAASDIEKEREDDRVHKFLLGLDSR 195
Query: 200 YNQIKSHVLLMNPLPPMNRVFSMVLQEEKQREIAARSADL--NSAFAAQATNSGKSGKKD 257
++ I+S + + PLP + +V+S V++EE+ A+R+ D+ A +S +D
Sbjct: 196 FSSIRSSITDIEPLPDLYQVYSRVVREEQNLN-ASRTKDVVKTEAIGFSVQSSTTPRFRD 254
Query: 258 RDRPLCSYCGKLGHSVDRCFKKHGYPP-----------------GLNFK--------NKS 292
+ C++C + GH V +CF HGYP G N + N+S
Sbjct: 255 KSTLFCTHCNRKGHEVTQCFLVHGYPDWWLEQNPQENQPSTRGRGSNGRGSSSGRGGNRS 314
Query: 293 SAVH-----HVDSSEVSGSSVPQDPPITSSQYQQLLSLLTAQLATSPMPSSSTSEPLQIP 347
SA ++++ + +V D + Q QL+SLL AQ PSSS+
Sbjct: 315 SAPTTRGRGRANNAQAAAPTVSGDG---NDQIAQLISLLQAQ-----RPSSSSERLSGNT 366
Query: 348 SPTDLKGFVFSASSDFSHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKAR 407
TD ++D+GAS H+ S + ++ PD ++
Sbjct: 367 CLTD------------------GVIDTGASHHMTGDCSILVDVFDITPSPVTKPDGKASQ 408
Query: 408 VTHIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFV-IQENKLKT- 465
T GT+ L +S LH+V +VP F L+SVS LL+ SI F TF +Q+ L+T
Sbjct: 409 ATKCGTLLLHDSYKLHDVLFVPDFDCTLISVSKLLKQTS-SIAIFTDTFCFLQDRFLRTL 467
Query: 466 IGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGH 525
IG G+ G+YY ++S ++ + LWH RLGH
Sbjct: 468 IGAGEEREGVYYFTGVLAPRVHKASSDFAI-------------------SGDLWHRRLGH 508
Query: 526 PSYNRLSLLSSTIHCKIPSSINENV--CPVCPLAKQKRLPFVSENHFANHSFDLIHCDVW 583
PS S+L S C S + + C C +KQ R F N+ F LIH DVW
Sbjct: 509 PS---TSVLLSLPECNRSSQGFDKIDSCDTCFRSKQTREVFPISNNKTMECFSLIHGDVW 565
Query: 584 GPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRT 643
GP+ P+ G +FLT+VDD+SR WT+L+ K+E I F M + QFGK +K RT
Sbjct: 566 GPYRTPSTTGAVYFLTLVDDYSRSVWTYLMSSKTEVSQLIKNFCAMSERQFGKQVKAFRT 625
Query: 644 DNAKELQ-LTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWG 702
DN E LT + + G LHQ SCV+ PQQN VER+H+ +LNVAR+ LFQ ++P+ FWG
Sbjct: 626 DNGTEFMCLTPYFQTHGILHQTSCVDTPQQNGRVERKHRHILNVARACLFQGNLPVKFWG 685
Query: 703 ECISTATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRA 762
E I TATHLIN P+ L K+P LL+ + PSYD LR FGCLC++ P + KF+ R+
Sbjct: 686 ESILTATHLINRTPSAVLKGKTPYELLFGERPSYDMLRSFGCLCYAHIRPRNKDKFTSRS 745
Query: 763 VPCVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIF----------PFQNTRPYVYSQ 812
CV +GY G K +++Y+L T SRDV F+E I+ P P + +
Sbjct: 746 RKCVFIGYPHGKKAWRVYDLETGKIFASRDVRFHEDIYPYATATQSNVPLPPPTPPMVND 805
Query: 813 DPFLV-----------------PLP*VGLSVPPYDVPQ-----------------PESVP 838
D FL P + PP + Q P S P
Sbjct: 806 DWFLPISTQVDSTNVDSSSSSSPAQSGSIDQPPRSIDQSPSTSTNPVPEEIGSIVPSSSP 865
Query: 839 VPSADHIPA-------TPLVSEIVAPSPDAIVPPLAVRRSTRVRHPPGYLADY------- 884
S D + T L+S + +P + P + + R + L D+
Sbjct: 866 SRSIDRSTSDLSASDTTELLSTGESSTPSSPGLPELLGKGCREKKKSVLLKDFVTNTTSK 925
Query: 885 --------DCPQQTTP-----------------HPLSTYYSYDKLTPSYKVYSVKVASHY 919
P Q P +PLS + + + ++ + +
Sbjct: 926 KKTASHNIHSPSQVLPSGLPTSLSADSVSGKTLYPLSDFLTNSGYSANHIAFMAAILDSN 985
Query: 920 EPTYFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSID 979
EP +F AI EW +AM E+ A+EAN TWD+ LP GK I SKWVYKLK + DG+++
Sbjct: 986 EPKHFKDAILIKEWCEAMSKEIDALEANHTWDITDLPHGKKAISSKWVYKLKYNSDGTLE 1045
Query: 980 RHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDL 1039
RHKARLV G Q+ G+D+ +TF+PVAK+TTVR +LA+AA WE+ Q+D++NAFL+GDL
Sbjct: 1046 RHKARLVVMGNHQKEGVDFKETFAPVAKLTTVRTILAVAAAKDWEVHQMDVHNAFLHGDL 1105
Query: 1040 FEEVYMKIPQGYDVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLF 1099
EEVYM++P G+ + CRL+KS+YGLKQA R WF+K S+ L GF SY DYSLF
Sbjct: 1106 EEEVYMRLPPGFKCSDPSKVCRLRKSLYGLKQAPRCWFSKLSTALRNIGFTQSYEDYSLF 1165
Query: 1100 TKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSAT 1159
+ +GDT + +LVYV+D+IV+G N ID K L F +KDLGKLK+FLGLE++
Sbjct: 1166 SLKNGDTIIHVLVYVDDLIVAGNNLDAIDRFKSQLHKCFHMKDLGKLKYFLGLEVSRGPD 1225
Query: 1160 GISLSQRAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLT 1219
G LSQR YAL ++ +TG C+P+++P+ N KL+S TGP + QYRRL+GR +YLT
Sbjct: 1226 GFCLSQRKYALDIVKETGLLGCKPSAVPIALNHKLASITGPVFTNPEQYRRLVGRFIYLT 1285
Query: 1220 ISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQGFF 1263
I+RPD+++ ++ LSQF+ P H +A L+RYLK +P QG F
Sbjct: 1286 ITRPDLSYAVHILSQFMQAPLVAHWEAALRLVRYLKGSPAQGIF 1329
>At4g14460 retrovirus-related like polyprotein
Length = 1489
Score = 776 bits (2003), Expect = 0.0
Identities = 489/1403 (34%), Positives = 717/1403 (50%), Gaps = 267/1403 (19%)
Query: 18 MENSSNPFFLHHSDNPGLILVSQPLN-GENYNSWNRSMMIALSVKNKLGFINGDFPRPAD 76
++ NP++LH +D+ GLILVS L +++SW RS+++AL+V+NKLGFING +P +
Sbjct: 27 VDQYENPYYLHSADHAGLILVSDRLTTASDFHSWRRSILMALNVRNKLGFINGTITKPPE 86
Query: 77 DDPLLQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLK 136
D +W R + +V +W++N V K I S++ + + IW +L RF Q + RI+ ++
Sbjct: 87 DHRDFGAWSRCNDIVSTWLMNSVDKKIGQSLLYIATVQGIWNNLLSRFKQDDAPRIFDIE 146
Query: 137 KDLLNLQQESLTVTQFYTKHKAIWDELQDYLPQRSCVCG----ENRVLLDYFQQEQIM-- 190
+ L ++Q S+ ++ +YT +W+E ++Y+ C CG + V ++ QQ +
Sbjct: 147 QKLSKIEQGSMDISTYYTALLTLWEEHRNYVELPVCTCGRCECDAAVKWEHLQQRSRVTK 206
Query: 191 --------------HFLM-----SLSDSYNQIKSHVLLMNPLPPMNRVFSMVLQEEKQ-- 229
H LM ++ +++N + N + P+ RV S+ Q
Sbjct: 207 FLKELNEGFDQTRRHILMLKPIPTIKEAFNMVTQDERQRN-VKPLTRVDSVAFQNTSMIN 265
Query: 230 REIAARSADLNSAFAAQ---ATNSGKSGK------KDRDRPLCSYCGKLGHSVDRCFKKH 280
+ A A N+ Q T+ GK G K P G G++ + H
Sbjct: 266 EDENAYVAAYNTVRPNQKPICTHCGKVGHTIQKCYKVHGYPPGMKTGNTGYTYKPNPQLH 325
Query: 281 GYP---------------PGLNFKNKSSAVHHV--DSSEVSGSSVPQDPPI--------- 314
P P N K++ V V ++ Q P +
Sbjct: 326 VQPRMPMMPQPRMQFPAQPYTNSMQKANVVAQVYAETGAYPSEGYSQAPMMNPYGSYPMP 385
Query: 315 --------------TSSQYQQLLSLLTAQLAT-SPMPSSSTSEPLQ-------------- 345
T Q +Q++S AQ+ P SSS PL
Sbjct: 386 HITHGGNNLSLQDFTPQQIEQMISQFQAQVQVPEPAASSSNPSPLATVSEHGFMALTSTS 445
Query: 346 ---IPSPT--------DLK--GFVFSASSDFSHKTSVWILDSGASCHVCFHLSSFESYHS 392
IP P+ DLK SA F + WI+DSGAS HVC L+ F S
Sbjct: 446 GTIIPFPSTSLKYENNDLKFQNHTLSALQKFL-PSDAWIIDSGASSHVCSDLAMFRELKS 504
Query: 393 VRSHTISLPDNTKARVTHIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFF 452
V GTV + LILHNV +VP F NL+SVS+L++ S HF+
Sbjct: 505 VS-----------------GTVHITQKLILHNVLHVPDFKFNLMSVSSLVKTISCSAHFY 547
Query: 453 HKTFVIQE-NKLKTIGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNK 511
+IQE ++ IGRG ++ LY L + + +++ + S+ N
Sbjct: 548 VDCCLIQELSQGLMIGRGRLYHNLYILETENTSPSTSTPAACLFTGSVLNDG-------- 599
Query: 512 STCTAALWHARLGHPSYNRLSLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFA 571
LWH RLGHPS S + L K KRL ++S N+ A
Sbjct: 600 -----HLWHQRLGHPS--------SVV-----------------LQKLKRLAYISHNNLA 629
Query: 572 NHSFDLIHCDVWGPFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQ 631
++ FDL+H D+WGPF + + G R+FLT+VDD +R TW +++++K + F K+V
Sbjct: 630 SNPFDLVHLDIWGPFSIESIEGFRYFLTVVDDCTRTTWVYMLRNKKDVSSVFPEFIKLVS 689
Query: 632 TQFGKVIKMVRTDNAKELQLTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLL 691
TQF IK +R+DNA EL T +++ G LH FSC PQQNSVVER+HQ +LNVAR+LL
Sbjct: 690 TQFNAKIKAIRSDNAPELGFTEIVKEHGMLHHFSCAYTPQQNSVVERKHQHILNVARALL 749
Query: 692 FQSHMPLVFWGECISTATHLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTL 751
FQS++P+ +W +C++TA LIN +P+P L+NKSP L+ + P Y L+ FGCLCF ST
Sbjct: 750 FQSNIPMQYWSDCVTTAVFLINRLPSPLLNNKSPYELILNKQPDYSLLKNFGCLCFVSTN 809
Query: 752 PSTRHKFSPRAVPCVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYS 811
R KF+PRA CV +GY GYKGYK+ +L + + +SR+VVF E +FPF+ + +
Sbjct: 810 AHERTKFTPRARACVFLGYPSGYKGYKVLDLESHSVTVSRNVVFKEHVFPFKTSELLNKA 869
Query: 812 QDPF---LVPLP*VGLSVPPYDVPQPESVPVPSADHIPATPLVSEIVAPSPDAIVPP--- 865
D F ++PLP V + +S+ + D A S + P +I+PP
Sbjct: 870 VDMFPNSILPLPAPLHFVETMPLIDEDSLIPTTTDSRTADNHASSSSSALP-SIIPPSSN 928
Query: 866 ----------LAVRRSTRVRHPPGYLADYDC----------------------------- 886
+ + RS R P YL++Y C
Sbjct: 929 TETQDIDSNAVPITRSKRTTRAPSYLSEYHCSLVPSISTLPPTDSSIPIHPLPEIFTASS 988
Query: 887 PQQTTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEA 946
P++TTP+P+ST SYDK TP + Y + EP F QA++ +W + EL AME
Sbjct: 989 PKKTTPYPISTVVSYDKYTPLCQSYIFAYNTETEPKTFSQAMKSEKWIRVAVEELQAMEL 1048
Query: 947 NQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVA 1006
N+TW V SLPP K+ +G KWV+ +K +PDG+++R+KARLVA+G+TQQ GID++DTFSPVA
Sbjct: 1049 NKTWSVESLPPDKNVVGCKWVFTIKYNPDGTVERYKARLVAQGFTQQEGIDFLDTFSPVA 1108
Query: 1007 KITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGY-----DVKGENLTCR 1061
K+T+ +M+L LAA+ W L Q+D+++AFL+GDL EE++M +PQGY + N CR
Sbjct: 1109 KLTSAKMMLGLAAITGWTLTQMDVSDAFLHGDLDEEIFMSLPQGYTPPAGTILPPNPVCR 1168
Query: 1062 LKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSG 1121
L KSIYGLKQASRQW+ + FV LVY++DI+++
Sbjct: 1169 LLKSIYGLKQASRQWYKR---------------------------FVAALVYIDDIMIAS 1201
Query: 1122 PNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFTDC 1181
N E++++K LL+S FK+KDLG +FFLGL C
Sbjct: 1202 NNDAEVENLKALLRSEFKIKDLGPARFFLGL--------------------------LGC 1235
Query: 1182 RPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTT 1241
+P+S+PMDP L L D G L + + YR+LIGRLLYLTI+RPDI + +++LSQF+S P+
Sbjct: 1236 KPSSIPMDPTLHLVRDMGTPLPNPTAYRKLIGRLLYLTITRPDITYAVHQLSQFISAPSD 1295
Query: 1242 THLDALHHLLRYLKTTPGQGFFF 1264
HL A H +LRY+K PGQG +
Sbjct: 1296 IHLQAAHKVLRYIKANPGQGLMY 1318
>At1g57640
Length = 1444
Score = 749 bits (1935), Expect = 0.0
Identities = 444/1314 (33%), Positives = 690/1314 (51%), Gaps = 132/1314 (10%)
Query: 23 NPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDPLLQ 82
+P+ L +DN G ++ L NY W AL + K GF++G P+P D P L+
Sbjct: 19 SPYDLTAADNSGAVISHPILKTNNYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLE 78
Query: 83 SWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLLNL 142
W+ + +++SW+ + ++++++ A+++W+ ++ RF NG + ++K DL
Sbjct: 79 DWLTINALLVSWMKMTIDSELLTNISHRDVARDLWEQIRKRFSVSNGPKNQKMKADLATC 138
Query: 143 QQESLTVTQFYTKHKAIWDELQDYLPQRSCVCGENRVLLD-----YFQQEQIMHFLMSLS 197
+QE +TV +Y K IWD + Y P R C CG L Y + + + +L L+
Sbjct: 139 KQEGMTVEGYYGKLNKIWDNINSYRPLRICKCGRCICNLGTDQEKYREDDMVHQYLYGLN 198
Query: 198 DS-YNQIKSHVLLMNPLPPMNRVFSMVLQEE---KQREIAARSADLNSAFAAQATNSGK- 252
++ ++ I+S + PLP + V+++V QEE R D+ +AFA Q +
Sbjct: 199 ETKFHTIRSSLTSRVPLPGLEEVYNIVRQEEDMVNNRSSNEERTDV-TAFAVQMRPRSEV 257
Query: 253 ------SGKKDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSSAVHHVDSSEVSGS 306
+ +K +++ LC++C + GHS + CF GYP + + + + +S G
Sbjct: 258 ISEKFANSEKLQNKKLCTHCNRGGHSPENCFVLIGYPEWWGDRPRGKSNSNGSTSRGRGR 317
Query: 307 ---------------SVPQDPPITSSQYQQLLSLLTAQLATSPMPSSSTSEPLQIPSP-- 349
+V P SS++ + + + A S + +++ +
Sbjct: 318 FGPGFNGGQPRPTYVNVVMTGPFPSSEHVNRVITDSDRDAVSGLTDEQWRGVVKLLNAGR 377
Query: 350 TDLKGFVFSASSDFSHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKARVT 409
+D K S + WILD+GAS H+ +L S+ I L D K
Sbjct: 378 SDNKSNAHETQSGTCSLFTSWILDTGASHHMTGNLELLSDMRSMSPVLIILADGNKRVAV 437
Query: 410 HIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQENKLKTIGRG 469
GTV+LG+ LIL +V F ++E + I G
Sbjct: 438 SEGTVRLGSHLILKSV------------------------------FYVKELESDLISVG 467
Query: 470 DTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYN 529
+ + ++A++ V ++ +P LWH RLGH S
Sbjct: 468 QMMDENHCVNAAA--VHTSVKAPFD-----------------------LWHRRLGHASDK 502
Query: 530 RLSLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFHVP 589
++LL + I ENVC C AKQ R F ++ + SF LIHCDVWGP+ P
Sbjct: 503 IVNLLPREL-LSSGKEILENVCDTCMRAKQTRDTFPLSDNRSMDSFQLIHCDVWGPYRAP 561
Query: 590 THVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKE- 648
++ G R+FLTIVDD+SR W +L+ KSE + F +V+ QF IK+VR+DN E
Sbjct: 562 SYSGARYFLTIVDDYSRGVWVYLMTDKSETQKHLKDFIALVERQFDTEIKIVRSDNGTEF 621
Query: 649 LQLTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTA 708
L + + +G H+ SCV P QN VER+H+ +LN+AR+L FQS++P+ FWGECI +A
Sbjct: 622 LCMREYFLHKGIAHETSCVGTPHQNGRVERKHRHILNIARALRFQSYLPIQFWGECILSA 681
Query: 709 THLINLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLV 768
+LIN P+ L KSP +LY+ P Y HLRVFG LC++ KF+ R+ CV V
Sbjct: 682 AYLINRTPSMLLQGKSPYEMLYKTAPKYSHLRVFGSLCYAHNQNHKGDKFAARSRRCVFV 741
Query: 769 GYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPF-------QNTRPYVYSQDPFLV---- 817
GY G KG++L++L + F +SRDV+F E+ FP+ ++ R V P +
Sbjct: 742 GYPHGQKGWRLFDLEEQKFFVSRDVIFQETEFPYSKMSCNEEDERVLVDCVGPPFIEEAI 801
Query: 818 -PLP*VGLSVPPYDV------------------PQPESVPVPSADHIPATPLVSEIVAPS 858
P +G ++ V E V + S D A+ V P
Sbjct: 802 GPRTIIGRNIGEATVGPNVATGPIIPEINQESSSPSEFVSLSSLDPFLASSTVQTADLPL 861
Query: 859 PDAIVPPLAVRRSTRVRHPPGYLADYDC-----------PQQTTPHPLSTYYSYDKLTPS 907
P+ +RRS+R P L ++ ++ +P+ Y + T S
Sbjct: 862 SSTTPAPIQLRRSSRQTQKPMKLKNFVTNTVSVESISPEASSSSLYPIEKYVDCHRFTSS 921
Query: 908 YKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWV 967
+K + V + EPT +++A+ W++AM E+ ++ NQT+ +V+LPPGK +G+KWV
Sbjct: 922 HKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFSIVNLPPGKRALGNKWV 981
Query: 968 YKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQ 1027
YK+K DG+I+R+KARLV G Q+ G+DY +TF+PVAK++TVR+ L +AA W + Q
Sbjct: 982 YKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTVRLFLGVAAARDWHVHQ 1041
Query: 1028 LDINNAFLNGDLFEEVYMKIPQGYDVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQH 1087
+D++NAFL+GDL EEVYMK+PQG+ + CRL KS+YGLKQA R WF+K SS L Q+
Sbjct: 1042 MDVHNAFLHGDLKEEVYMKLPQGFQCDDPSKVCRLHKSLYGLKQAPRCWFSKLSSALKQY 1101
Query: 1088 GFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLK 1147
GF S DYSLF+ + FV +LVYV+D+I+SG + K L+S F +KDLG LK
Sbjct: 1102 GFTQSLSDYSLFSYNNDGIFVHVLVYVDDLIISGSCPDAVAQFKSYLESCFHMKDLGLLK 1161
Query: 1148 FFLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQ 1207
+FLG+E++ +A G LSQR Y L ++ + G RP++ P++ N KLS T P L+DSS+
Sbjct: 1162 YFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPLEQNHKLSLSTSPLLSDSSR 1221
Query: 1208 YRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQG 1261
YRRL+GRL+YL ++RP+++++++ L+QF+ P H +A ++RYLK+ PGQG
Sbjct: 1222 YRRLVGRLIYLVVTRPELSYSVHTLAQFMQNPRQDHWNAAIRVVRYLKSNPGQG 1275
>At4g07810 putative polyprotein
Length = 1366
Score = 742 bits (1915), Expect = 0.0
Identities = 477/1258 (37%), Positives = 671/1258 (52%), Gaps = 159/1258 (12%)
Query: 100 SKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLLNLQQESLTVTQFYTKHKAI 159
S D+ S ++ A +W L+ R Q N ++IY ++ L L Q SL ++ +YT+
Sbjct: 64 SGDLGSGMVYFYDAHLLWLKLEGRSRQSNLSKIYSVQNQLDRLHQGSLDLSAYYTRLTVT 123
Query: 160 --------WDELQDY--LPQRSC---VCGENRVLLDYFQQEQIMHFLMSLSDSYNQIKSH 206
W+EL+++ LP +C CG N + +++ I+ FLM L++S+ Q +
Sbjct: 124 FIRSQCLSWEELKNFEELPSCTCGKCTCGSNDRWIQLYEKHNIVRFLMRLNESFIQARRQ 183
Query: 207 VLLMNPLPPMNRVFSMVLQEEKQREIAARSADLNSAFAAQATNSGKS--GKKDRDRPLCS 264
+L+M+PLP +++ + Q+++QR + F A T ++ + RPLC+
Sbjct: 184 ILMMDPLPEFTNLYNFISQDDQQRSFNSMPTTEKPVFQASITQQKPKFFNQQGKSRPLCT 243
Query: 265 YCGKLGHSVDRCFKKHGYPPGLN------FKNKSS--------AVHHVDSSEVSGSSVPQ 310
YCG LGH+ RC+K HGYPPG + N S +H +G+S
Sbjct: 244 YCGLLGHTNARCYKLHGYPPGYKVPVGTCYNNDKSRGQPYPHNGIHMSHLITYNGNSYAP 303
Query: 311 DPPITSSQYQQLLSLLTAQLATSPMPSSSTSEPLQIPSPTDLKGFVFSASSDFSHKTSVW 370
++ L + + +PM + + + G V S S +H SV
Sbjct: 304 IVQANNNAPYALYNQAYNGNSYAPMAQNFAGNHIISDGSSMSAGNVTSESPTVNH--SVN 361
Query: 371 ILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKARVTHIGTVKLGNSLILHNVCYVPT 430
+++SG F SS V VT + T G+ + V PT
Sbjct: 362 MMNSGRG----FLGSSSHGREQVNQ-----------MVTQLNTQLQGSP---YQVIRPPT 403
Query: 431 FTVNLLSVSA--LLENPKYSIHFFHKTFVIQENKLKTIGRGDTHN--------GLYYLHA 480
+ N S+SA + P + I F +I +N R H +Y++
Sbjct: 404 VSQNHGSISAQGMSPIPSHYISAFEPCLIIPQNTWSLDTRASCHICCDLSLFCNVYHIDH 463
Query: 481 SS----NNV-FSASTSPLSVLNS-LCNQSVF--PS-HCNKSTCTAALWHARLGHPSYNRL 531
++ NN+ S + + LN L VF PS H N + RLGHPS +R+
Sbjct: 464 TNITLPNNIKISINIAETVKLNDRLILHLVFYVPSFHFNLIS--------RLGHPSMSRV 515
Query: 532 SLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFHVPTH 591
LSS +H IP ++E C +C L+KQK L FVS N F LIH D
Sbjct: 516 QALSSNLH--IPQKLSEFHCKICHLSKQKCLSFVSNNKIYEEPFPLIHID---------- 563
Query: 592 VGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKELQL 651
S+ F K+VQTQFG +K +R+DNA ELQ
Sbjct: 564 -------------------------SDVTTIFPEFLKLVQTQFGCTVKSIRSDNAPELQF 598
Query: 652 TAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHL 711
L G H SC PQQN VVER HQ LLNVARSL FQS++PL +W EC+STA L
Sbjct: 599 KDLLATFGIFHYHSCAYTPQQNYVVERNHQHLLNVARSLYFQSNIPLAYWPECVSTAAFL 658
Query: 712 INLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYK 771
IN PTPNL +KSP +LY++ P Y+ LRVF CLC++ST RHKF+ RA CV +GY+
Sbjct: 659 INRTPTPNLEHKSPYEVLYKKLPDYNSLRVFCCLCYASTHQHERHKFTERATSCVFIGYE 718
Query: 772 QGYKGYKLYNLTTKTFHISRDVVFNESIFPF--QNTRPYVYSQDPFLVPLP*-------- 821
G+KGYK+ +L + T ++R+VVF+E+IFPF +++ V D ++P+
Sbjct: 719 SGFKGYKILDLESNTVSVTRNVVFHETIFPFIDKHSTQNVSFFDDSVLPISEKQKENRFQ 778
Query: 822 -------VGLSVPPYDVPQPESVPVPSADHIPA---TPLVSEIVAPSPDAIVPPLAVRRS 871
+ L V P V +P +VP + A T + ++ D + P R+
Sbjct: 779 IYDYFNVLNLEVCP--VIEPTTVPAHTHTRSLAPLSTTVTNDQFGNDMDNTLMP---RKE 833
Query: 872 TRVRHPPGYLADYDCPQ---------QTTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPT 922
TR P YL+ Y C T H LS++ SYDKL+ Y+++ + + EPT
Sbjct: 834 TRA---PSYLSQYHCSNVLKEPSSSLHGTAHSLSSHLSYDKLSNEYRLFCFAIIAEKEPT 890
Query: 923 YFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHK 982
F +A +W AM EL A+ + T ++ SL GK IG KWV+K+K DG+I+R+K
Sbjct: 891 TFKEAALLQKWLDAMNVELDALVSTSTREICSLHDGKRAIGCKWVFKIKYKSDGTIERYK 950
Query: 983 ARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEE 1042
ARLVA GYTQQ G+DY+DTFSP+AK+T+VR++LALAA++ W + Q+D+ NAFL+GD EE
Sbjct: 951 ARLVANGYTQQEGVDYIDTFSPIAKLTSVRLILALAAIHNWSISQMDVTNAFLHGDFEEE 1010
Query: 1043 VYMKIPQGYDV-KGENL----TCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYS 1097
+YM++PQGY KGE L CRL KS+YGLKQASRQWF KFS VL Q+GF S D +
Sbjct: 1011 IYMQLPQGYTPRKGELLPKRPVCRLVKSLYGLKQASRQWFHKFSGVLIQNGFMQSLFDPT 1070
Query: 1098 LFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYS 1157
LF + DTF+ LLVYV+DI++ + + VKQ+L FKLKDLG+ ++FLGLEIA S
Sbjct: 1071 LFVRVREDTFLALLVYVDDIMLVSNKDSAVIEVKQILAKEFKLKDLGQKRYFLGLEIARS 1130
Query: 1158 ATGISLSQRAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLY 1217
GIS+SQR YAL LL + GF C+P PM+ NLKLS + G L D+S YR+LIGRL+Y
Sbjct: 1131 KEGISISQRKYALELLEEFGFLGCKPVPTPMELNLKLSQEDGALLLDASHYRKLIGRLVY 1190
Query: 1218 LTISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQGFFFLGQQPPLSQLT 1275
LT++RPDI F +NKL+Q++S P HL A +LRYLK PGQG F+ P S LT
Sbjct: 1191 LTVTRPDICFAVNKLNQYMSAPREPHLMAARRILRYLKNDPGQGVFY----PASSTLT 1244
>At2g07010 putative retroelement pol polyprotein
Length = 1413
Score = 686 bits (1770), Expect = 0.0
Identities = 421/1222 (34%), Positives = 625/1222 (50%), Gaps = 119/1222 (9%)
Query: 23 NPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDPLLQ 82
+P+ L SDNPG ++ S L G+NYN W+ M+ AL K K GFING +P D+P +
Sbjct: 25 SPYTLASSDNPGAMISSVMLTGDNYNEWSTKMLNALQAKRKTGFINGSISKPPLDNPDYE 84
Query: 83 SWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLLNL 142
+W + +++ WI + + S+V A ++W +L+ RF N ++Q+K L
Sbjct: 85 NWQAVNSMIVGWIRASIEPKVKSTVTFICDAHQLWSELKQRFSVGNKVHVHQIKTQLAAC 144
Query: 143 QQESLTVTQFYTKHKAIWDELQDYLPQRSC-----VCGENRVLLDYFQQEQIMHFLMSLS 197
+Q+ V +Y + +W+E Q Y P C CG ++E+I F++ L
Sbjct: 145 RQDGQPVIDYYGRLCKLWEEFQIYKPITVCKCGLCTCGATLEPSKEREEEKIHQFVLGLD 204
Query: 198 DS-YNQIKSHVLLMNPLPPMNRVFSMVLQEEKQREIAARSADLNSAFA-----AQATNSG 251
DS + + + ++ M+P P + ++S V++EE++ SA ++ T G
Sbjct: 205 DSRFGGLSATLIAMDPFPSLGEIYSRVVREEQRLASVQIREQQQSAIGFLTRQSEVTADG 264
Query: 252 KSGK---KDRDRP-LCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSSAVHHVDSSEVSGSS 307
++ K RDR LCS+CG+ GH C++ G+P + SS
Sbjct: 265 RTDSSIIKSRDRSVLCSHCGRSGHEKKDCWQIVGFPDWWTERTNGGGRGSSSRGRGGRSS 324
Query: 308 VPQDPPITSSQYQQLLSLLTAQLAT----SPMPSSSTSEPLQIPSPTDLKGFVFSASSDF 363
+ Q +TA AT SP P + + I K S
Sbjct: 325 GSNNSGRGRGQ-------VTAAHATTSNLSPFPEFTPDQLRVITQMIQNKNNGTSDKLSG 377
Query: 364 SHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKARVTHIGTVKLGNSLILH 423
K ILD+GAS H+ LS + ++ S ++ D K +GT KL ++ L
Sbjct: 378 KMKLGDVILDTGASHHMTGQLSLLTNIVTIPSCSVGFADGRKTFAISMGTFKLSETVSLS 437
Query: 424 NVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQENKLKT-IGRGDTHNGLYYLHASS 482
NV YVP +L+SVS L++ K F V+Q+ +T IG G+ +G+YYL
Sbjct: 438 NVLYVPALNCSLISVSKLVKQIKCLALFTDTICVLQDRFSRTLIGTGEERDGVYYLT--- 494
Query: 483 NNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYNRLSLLSSTIHCKI 542
A+T+ + H T ALWH RLGHPS++ LS L +
Sbjct: 495 ----DAATTTV--------------HKVDITTDHALWHQRLGHPSFSVLSSLP--LFSGS 534
Query: 543 PSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFHVPTHVGHRFFLTIVD 602
S++ C VC AKQ R F ++ + F LIHCDVWGP+ VP+ G +FLTIVD
Sbjct: 535 SCSVSSRSCDVCFRAKQTREVFPDSSNKSTDCFSLIHCDVWGPYRVPSSCGAVYFLTIVD 594
Query: 603 DHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKELQ-LTAFLEKQGTL 661
D SR WT+L+ KSE + F + QFGK +K++R+DN E L+++ ++QG +
Sbjct: 595 DFSRSVWTYLLLAKSEVRSVLTNFLAYTEKQFGKSVKIIRSDNGTEFMCLSSYFKEQGIV 654
Query: 662 HQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINLIPTPNLH 721
HQ SCV PQQN VER+H+ +LNV+R+LLFQ+ +P+ FWGE + TA +LIN P+ +
Sbjct: 655 HQTSCVGTPQQNGRVERKHRHILNVSRALLFQASLPIKFWGEAVMTAAYLINRTPSSIHN 714
Query: 722 NKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYKGYKLYN 781
SP LL+ P YD LRVFG C++ + + KF R+ C+ VGY G KG+K+Y+
Sbjct: 715 GLSPYELLHGCKPDYDQLRVFGSACYAHRVTRDKDKFGERSRLCIFVGYPFGQKGWKVYD 774
Query: 782 LTTKTFHISRDVVFNESIFPFQN-----------TRPYVYSQD--PFLV----------- 817
L+T F +SRDVVF E++FP+ T P Y +D PF
Sbjct: 775 LSTNEFIVSRDVVFRENVFPYATNEGDTIYTPPVTCPITYDEDWLPFTTLEDRGSDENYL 834
Query: 818 ---PLP*VGLSVPPYDVPQPESVPVPSADHIPATPLVSEIVAP-------SPDAIVPP-- 865
P+ +S + P+S+P P D + + V+ P SP V P
Sbjct: 835 SDPPVCVTNVSESDTEHDTPQSLPTPVDDPLSPSTSVTPTQTPTNSSSSTSPSTNVSPPQ 894
Query: 866 ---------LAVRRSTRVRHPPGYLADY---------DCPQQTTP--------------H 893
R+ R P L DY + P +P +
Sbjct: 895 QDTTPIIENTPPRQGKRQVQQPARLKDYILYNASCTPNTPHVLSPSTSQSSSSIQGNLQY 954
Query: 894 PLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQTWDVV 953
PL+ Y S + + +KV+ + ++ EP +F + ++ W AM E+ A+E N+TWD+V
Sbjct: 955 PLTDYISDECFSAGHKVFLAAITANDEPKHFKEDVKVKVWNDAMYKEVDALEVNKTWDIV 1014
Query: 954 SLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRM 1013
LP GK IGS+WVYK K + DG+++R+KARLV +G Q G DY +TF+PV K+TTVR
Sbjct: 1015 DLPTGKVAIGSQWVYKTKFNADGTVERYKARLVVQGNNQIEGEDYTETFAPVVKMTTVRT 1074
Query: 1014 LLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGYDVKGENLTCRLKKSIYGLKQAS 1073
LL L A +WE++Q+D++NAFL+GDL EEVYMK+P G+ + CRL+KS+YGLKQA
Sbjct: 1075 LLRLVAANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRHSHPDKVCRLRKSLYGLKQAP 1134
Query: 1074 RQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQL 1133
R WF K S L + GF Y DYS F+ + +LVYV+D+I+ G + + K+
Sbjct: 1135 RCWFKKLSDALKRFGFIQGYEDYSFFSYSCKGIELRVLVYVDDLIICGNDEYMVQKFKEY 1194
Query: 1134 LKSAFKLKDLGKLKFFLGLEIA 1155
L F +KDLGKLK+FLG+E++
Sbjct: 1195 LGRCFSMKDLGKLKYFLGIEVS 1216
>At1g36620 hypothetical protein
Length = 1152
Score = 659 bits (1701), Expect = 0.0
Identities = 410/1165 (35%), Positives = 607/1165 (51%), Gaps = 82/1165 (7%)
Query: 21 SSNPFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDPL 80
S++P++LH SD+P +L LNGENY W + L K KLGFI+G +P+ D P
Sbjct: 21 SASPYYLHPSDHPHHVLTPMLLNGENYERWAKLTRNNLQAKQKLGFIDGTLTKPSSDSPD 80
Query: 81 LQSWIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLL 140
W++ + +++ W+ + + S+ +A+ +W+ L+ R+ N +R++QLK D++
Sbjct: 81 YPRWLQTNSMLVGWLYASLDPQVQKSISVVDNARVMWESLRTRYSVGNASRVHQLKYDIV 140
Query: 141 NLQQESLTVTQFYTKHKAIWDELQDYLPQRSCVCGE----NRVLLDYFQQEQIMH-FLMS 195
+Q+ T ++ K K +WD+L DY P +C C +RV + + +H FLM
Sbjct: 141 ACRQDGQTAANYFGKLKVMWDDLDDYEPLLTCCCNRPSCTHRVRQSQRRDHERIHQFLMG 200
Query: 196 LSDS-YNQIKSHVL---LMNPLPPMNRVFSMVLQEEKQREIAARSADLNSA--FAAQA-T 248
L + + ++++L + ++ ++S ++ EE+ I + A FA Q
Sbjct: 201 LDAAKFGTSRTNILGRLSRDDNISLDSIYSEIIAEERHLTITRSKEERVDAVGFAVQTGV 260
Query: 249 NSGKSGKKDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSSAVHHVDSSEVSGSSV 308
N+ S + + C++CG+ HS D CFK HG P K + D+S G
Sbjct: 261 NAIASVTRVNNMGPCTHCGRSNHSADTCFKLHGVPEWYTEK-------YGDTSSGRGRGR 313
Query: 309 PQDPPITSSQYQQLLSLLTAQLATSPMPSSSTSEPLQIPSPTD---------LKGFVFSA 359
P + AQ + PSSS SE IP + LK ++
Sbjct: 314 SSTPRGRGRGHGNSYKANNAQTSH---PSSSASEFSDIPGVSKEAWSAIRNLLKQDTATS 370
Query: 360 SSDFSHKTSV--WILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKARVTHIGTVKLG 417
S S KT+ +++DSGAS H+ L + + + LP+ T GT+ LG
Sbjct: 371 SEKLSGKTNCVDFLIDSGASHHMTGFLDLLTEIYEIPHSVVVLPNAKHTIATKKGTLILG 430
Query: 418 NSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQENKLKT-IGRGDTHNGLY 476
++ L +V +VP + L+SV+ LL F K VIQ+ K IG G NG+Y
Sbjct: 431 ANMKLTHVLFVPDLSCTLISVARLLRELHCFAIFTDKVCVIQDRTSKMLIGVGTESNGVY 490
Query: 477 YLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYNRLS-LLS 535
+L + SA+ K ALWH RLGHPS LS +L
Sbjct: 491 HLQRAEVVATSANVV-------------------KWKTNKALWHMRLGHPSSKVLSSVLP 531
Query: 536 STIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFHVPTHVGHR 595
S SS + +C VC AKQ R F + A F IH DVWGP+ + G
Sbjct: 532 SLEDFDSCSSDLKTICDVCVRAKQTRASFSESFNKAEECFSFIHYDVWGPYKHASSCGAH 591
Query: 596 FFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKE-LQLTAF 654
+FLTIVDDHSR W L+ KSE + F M QF K +K VR++N E + L ++
Sbjct: 592 YFLTIVDDHSRAVWIHLMLAKSEVASLLQQFIAMASRQFNKQVKTVRSNNGTEFMSLKSY 651
Query: 655 LEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINL 714
++G +HQ SCV QQN VER+H+ +LNVARSLLFQ+ +P+ FW E + TA +LIN
Sbjct: 652 FAERGIVHQISCVYTHQQNGRVERKHRHILNVARSLLFQAELPISFWEESVLTAAYLINR 711
Query: 715 IPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGY 774
PTP L K+P +LY Q PSY LRVFG LCF+ KF R C+ VGY G
Sbjct: 712 TPTPILDGKTPYKILYSQPPSYASLRVFGSLCFARKHTGRLDKFQERGRKCIFVGYPHGQ 771
Query: 775 KGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQDPFLVPLP*VGLSVPPYDVP-- 832
KG+++Y++ ++ F +SRDVVF E IFPF + + ++D F P + + PYD
Sbjct: 772 KGWRIYDIESQIFFVSRDVVFQEDIFPFADKK----NKDTFSSPAAVIPSPILPYDDEFL 827
Query: 833 -------QPESVPVPSADHIPATPLVSEIVAPSPDAIVPPLAVRRSTRVRHPPGYLADYD 885
P + P+P+ + +P S I+ +P A PP +RR R R L DY
Sbjct: 828 DIYQIGDVPATNPLPAIIDVNDSPPSSPIITATPAAASPP-PLRRGLRQRQENVRLKDYQ 886
Query: 886 -----CPQQTTP--------HPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPE 932
C T +P++ Y S + +PS + + ++ P ++QAI+ E
Sbjct: 887 TYSAQCESTQTLSDNIGTCIYPMANYVSGEIFSPSNQHFLAAISMVDPPQTYNQAIREKE 946
Query: 933 WQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQ 992
W+ A+ E+ A+E TWD+ LP G IGSKWV+++K + +G+++R+KARLVA G Q
Sbjct: 947 WRNAVFFEVDALEDQGTWDITKLPQGVKAIGSKWVFRIKYNSNGTVERYKARLVALGNHQ 1006
Query: 993 QNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGYD 1052
+ GID+ TF+PV K+ TVR+LL +AA WEL Q+D++NAFL+GDL E++YMK P G+
Sbjct: 1007 KEGIDFTKTFAPVVKMQTVRLLLDVAAAKDWELHQMDVHNAFLHGDLKEDIYMKPPPGFK 1066
Query: 1053 VKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLV 1112
+L C+LKKSIYGLKQA R WF K S+ L + GF S DYSLFT G + ++V
Sbjct: 1067 TTDPSLVCKLKKSIYGLKQAPRCWFEKLSTSLLKFGFTQSKKDYSLFTSIRGSKVLHVIV 1126
Query: 1113 YVNDIIVSGPNATEIDSVKQLLKSA 1137
YV+D+++ G E ++ K L S+
Sbjct: 1127 YVDDVVICGKAVRENNTSKLALGSS 1151
>At1g53810
Length = 1522
Score = 558 bits (1439), Expect = e-159
Identities = 401/1333 (30%), Positives = 626/1333 (46%), Gaps = 199/1333 (14%)
Query: 38 VSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPA-------------DDDPLLQSW 84
V+ LN +NY W LS + LGF+ G PA + +P +W
Sbjct: 15 VTVTLNQQNYILWKSQFESFLSGQGLLGFVTGSISAPAQTRSVTHNNVTSEEPNPEFYTW 74
Query: 85 IRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLLNLQQ 144
+ D VV SW+L ++DI+S V+ C ++ ++W L + F++ + +R+++L++ L L++
Sbjct: 75 HQTDQVVKSWLLGSFAEDILSVVVNCFTSHQVWLTLANHFNRVSSSRLFELQRRLQTLEK 134
Query: 145 ESLTVTQFYTKHKAIWDELQDY---LPQR----SCVCGENRVLLDYFQQEQIMHFLMSLS 197
+ T+ F K I D+L +P++ S + G R + E I + +
Sbjct: 135 KDNTMEVFLKDLKHICDQLASVGSPVPEKMKIFSALNGLGR------EYEPIKTTIENSV 188
Query: 198 DSYNQIKSHVLLMNPLPPMNRVFSMVLQEEKQREIAARSADLNSAF--------AAQATN 249
DS + + +R+ S V + +A +S + +
Sbjct: 189 DSNPSLSLDEVASKLRGYDDRLQSYVTEPTISPHVAFNVTHSDSGYYHNNNRGKGRSNSG 248
Query: 250 SGKSGKKDRDRP------------------LCSYCGKLGHSVDRCFKKHGYPPGLNFKNK 291
SGKS R R +C CGK GH +C+
Sbjct: 249 SGKSSFSTRGRGFHQQISPTSGSQAGNSGLVCQICGKAGHHALKCW-------------- 294
Query: 292 SSAVHHVDSSEVSGSSVPQDPPITSSQYQQLLSLLTAQLATSPMPSSSTSEPLQIPSPTD 351
H D+S Q++ L PM ++ ++I TD
Sbjct: 295 ----HRFDNSY---------------QHEDL-----------PMALAT----MRITDVTD 320
Query: 352 LKGFVFSASSDFSHKTSVWILDSGASCHVCFH---LSSFESYHSVRSHTISLPDNTKARV 408
H WI DS AS HV + L + YH S +I + D +
Sbjct: 321 -------------HHGHEWIPDSAASAHVTNNRHVLQQSQPYHG--SDSIMVADGNFLPI 365
Query: 409 THIGTVKLGNS---LILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQENKLKT 465
TH G+ + +S + L V P +LLSVS L + S+ F + I + K
Sbjct: 366 THTGSGSIASSSGKIPLKEVLVCPDIVKSLLSVSKLTSDYPCSVEFDADSVRINDKATKK 425
Query: 466 I-GRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLG 524
+ G +GLY L L VL S +++ ++ +WH RLG
Sbjct: 426 LLVMGRNRDGLYSLEEPK----------LQVLYST----------RQNSASSEVWHRRLG 465
Query: 525 HPSYNRLSLLSSTIHCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWG 584
H + L L+S+ I + + + VC C L K RLPF+ A+ + IHCD+WG
Sbjct: 466 HANAEVLHQLASSKSIIIINKVVKTVCEACHLGKSTRLPFMLSTFNASRPLERIHCDLWG 525
Query: 585 PFHVPTHVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTD 644
P + G R+++ +D +SRFTW + +K KS+ + F K+V+ Q G IK+ + D
Sbjct: 526 PSPTSSVQGFRYYVVFIDHYSRFTWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCD 585
Query: 645 NAKEL---QLTAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFW 701
E Q L+ G SC PQQN + ER+H+ ++ + S++FQS +PL +W
Sbjct: 586 GGGEFISSQFLKHLQDHGIQQNMSCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYW 645
Query: 702 GECISTATHLINLIPTPNL-HNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSP 760
E TA +INL+PT +L +N+SP LY + P Y LRVFGC C+ + KF P
Sbjct: 646 LESFFTANFVINLLPTSSLDNNESPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDP 705
Query: 761 RAVPCVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQD--PFL-- 816
R++ CV +GY + YKGY+ T +ISR VVF+E+ PF++ +++ QD P L
Sbjct: 706 RSLKCVFLGYNEKYKGYRCLYPPTGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEA 765
Query: 817 --------VPLP*VGLSVPPYDVPQPESVPVPSADHIPAT----PLVSE----------I 854
P P +PQPE+ + +A A P S+ +
Sbjct: 766 WFKSFHHVTPTQPDQSRYPVSSIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISV 825
Query: 855 VAPSP------------DAIVPPLA------VRRSTRVRHPPGYLADYDCPQQ-----TT 891
V+ SP D+ P A RS+ P G QQ T
Sbjct: 826 VSGSPERTTGLDSASIGDSYHSPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTN 885
Query: 892 PHPLST--YYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQT 949
H + T K Y + + KV S EP +A+++P W AM+ E+ + +T
Sbjct: 886 EHAMVTRGKEGISKPNKRYVLLTHKV-SIPEPKTVTEALKHPGWNNAMQEEMGNCKETET 944
Query: 950 WDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKIT 1009
W +V P + +GS WV++ KLH DGS+D+ KARLVAKG+ Q+ GIDY++T+SPV +
Sbjct: 945 WTLVPYSPNMNVLGSMWVFRTKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTP 1004
Query: 1010 TVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGY-DVKGENLTCRLKKSIYG 1068
TVR++L +A + KWEL Q+D+ NAFL+GDL E VYM+ P G+ D + C L KS+YG
Sbjct: 1005 TVRLILHVATVLKWELKQMDVKNAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYG 1064
Query: 1069 LKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEID 1128
LKQ+ R WF +FS+ L + GF S D SLF S + ++LL+YV+D++++G N+ +
Sbjct: 1065 LKQSPRAWFDRFSNFLLEFGFICSLFDPSLFVYSSNNDVILLLLYVDDMVITGNNSQSLT 1124
Query: 1129 SVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPM 1188
+ L F++KD+G++ +FLG++I G+ +SQ+ YA LL +C P P+
Sbjct: 1125 HLLAALNKEFRMKDMGQVHYFLGIQIQTYDGGLFMSQQKYAEDLLITASMANCSPMPTPL 1184
Query: 1189 DPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDALH 1248
L S+ +D + +R L G+L YLT++RPDI F +N + Q + +P+ + + L
Sbjct: 1185 PLQLDRVSNQDEVFSDPTYFRSLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLK 1244
Query: 1249 HLLRYLKTTPGQG 1261
+LRY+K T G
Sbjct: 1245 RILRYIKGTVSMG 1257
>At3g60170 putative protein
Length = 1339
Score = 473 bits (1217), Expect = e-133
Identities = 286/918 (31%), Positives = 467/918 (50%), Gaps = 57/918 (6%)
Query: 364 SHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKARVTHIGTVKL---GNSL 420
+++ VW LDSG S H+ F + T+ L ++T+ V G+VK+ G +
Sbjct: 294 ANRDEVWFLDSGCSNHMTGSKEWFSELEEGFNRTVKLGNDTRMSVVGKGSVKVKVNGVTQ 353
Query: 421 ILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQENKLKTIGRGDTHNGLYYLHA 480
++ V YVP NLLS+ L E + V +K + + N +++L A
Sbjct: 354 VIPEVYYVPELRNNLLSLGQLQERGLAILIRDGTCKVYHPSKGAIMETNMSGNRMFFLLA 413
Query: 481 SSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYNRLSLLSSTIHC 540
S NSLC Q+ +K LWH R GH + L LL+ H
Sbjct: 414 SKPQK-----------NSLCLQT--EEVMDKEN---HLWHCRFGHLNQEGLKLLA---HK 454
Query: 541 KIPSSI-----NENVCPVCPLAKQKRLPFVSENHFANHS-FDLIHCDVWGPFHVPTHVGH 594
K+ + + +C +C KQ R + + + + L+H D+ GP +H G
Sbjct: 455 KMVIGLPILKATKEICAICLTGKQHRESMSKKTSWKSSTQLQLVHSDICGPITPISHSGK 514
Query: 595 RFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKEL---QL 651
R+ L+ +DD +R TW + + KSEA F V+ + G + +RTD E +
Sbjct: 515 RYILSFIDDFTRKTWVYFLHEKSEAFATFKIFKASVEKEIGAFLTCLRTDRGGEFTSNEF 574
Query: 652 TAFLEKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHL 711
F G Q + PQQN V ER+++ ++N RS+L + +P +FW E + H+
Sbjct: 575 GEFCRSHGISRQLTAAFTPQQNGVAERKNRTIMNAVRSMLSERQVPKMFWSEATKWSVHI 634
Query: 712 INLIPTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYK 771
N PT + +P + P ++ RVFGC+ + R K ++ CV +G
Sbjct: 635 QNRSPTAAVEGMTPEEAWSGRKPVVEYFRVFGCIGYVHIPDQKRSKLDDKSKKCVFLGVS 694
Query: 772 QGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQDPFLVPLP*VGLSVPPYDV 831
+ K ++LY+ K IS+DVVF+E + D V V L D
Sbjct: 695 EESKAWRLYDPVMKKIVISKDVVFDED---------KSWDWDQADVEAKEVTLECGDEDD 745
Query: 832 PQP----ESVPVPSADHIPATPLVSEIVAPSPDAIVPPLAVRRSTRVRHPPGYLADYDCP 887
+ E + V S +H+ + VS +P + P + TR R PPG++ADY+
Sbjct: 746 EKNSEVVEPIAVASPNHVGSDNNVSSSPILAPSSPAPSPVAAKVTRERRPPGWMADYETG 805
Query: 888 QQTTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEAN 947
+ +++ + V + + + +P F A++ W++AM E+ ++ N
Sbjct: 806 EG------------EEIEENLSVMLLMMMTEADPIQFDDAVKDKIWREAMEHEIESIVKN 853
Query: 948 QTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAK 1007
TW++ +LP G IG KWVYK KL+ DG +D++KARLVAKGY Q GIDY + F+PVA+
Sbjct: 854 NTWELTTLPKGFTPIGVKWVYKTKLNEDGEVDKYKARLVAKGYAQCYGIDYTEVFAPVAR 913
Query: 1008 ITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGYDVKG-ENLTCRLKKSI 1066
+ TVR +LA+++ + WE+FQLD+ +AFL+G+L EEVY++ P+G+ +G E +L+K++
Sbjct: 914 LDTVRTILAISSQFNWEIFQLDVKSAFLHGELKEEVYVRQPEGFIREGEEEKVYKLRKAL 973
Query: 1067 YGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATE 1126
YGLKQA R W+++ + + F+ +++LFTK +++ +YV+D+I +G +
Sbjct: 974 YGLKQAPRAWYSRIEAYFLKEEFERCPSEHTLFTKTRVGNILIVSLYVDDLIFTGSDKAM 1033
Query: 1127 IDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSL 1186
D K+ + F++ DLGK+K FLG+E+ S GI + QR YA +L G +
Sbjct: 1034 CDEFKKSMMLEFEMSDLGKMKHFLGIEVKQSDGGIFICQRRYAREVLARFGMDESNAVKN 1093
Query: 1187 PMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDA 1246
P+ P KL+ D E D + +++L+G L+YLT++RPD+ + + +S+F+S P +H A
Sbjct: 1094 PIVPGTKLTKDENGEKVDETMFKQLVGSLMYLTVTRPDLMYGVCLISRFMSNPRMSHWLA 1153
Query: 1247 LHHLLRYLKTTPGQGFFF 1264
+LRYLK T G F+
Sbjct: 1154 AKRILRYLKGTVELGIFY 1171
>At2g06840 putative retroelement pol polyprotein
Length = 1102
Score = 438 bits (1126), Expect = e-122
Identities = 253/654 (38%), Positives = 361/654 (54%), Gaps = 72/654 (11%)
Query: 616 KSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKELQL-TAFLEKQGTLHQFSCVERPQQNS 674
K P I F M QF K ++ +R+DN E T++ ++ G LH+ SCV+ PQQN+
Sbjct: 348 KQTLPSLIRNFCAMADRQFRKPVRSIRSDNGTEFMCHTSYFQEHGILHETSCVDTPQQNA 407
Query: 675 VVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINLIPTPNLHNKSPSTLLYQQDP 734
VER+H+ +LNVAR+ LFQ + P P+P L K+P +L+ + P
Sbjct: 408 RVERKHRHILNVARTCLFQGNFPT-----------------PSPVLKGKTPYEVLFGKQP 450
Query: 735 SYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYKGYKLYNLTTKTFHISRDVV 794
SYD LR FGCLC++ P + KF+ R+ C+ +GY + T H S D
Sbjct: 451 SYDMLRTFGCLCYAHIRPRDKDKFASRSRKCIFIGYP--------HETATPNTHDSIDPT 502
Query: 795 FNESIFPFQNTRPYVYSQDPFLVPLP*VGLSVPPYDVPQPESVPVPSADHIPATPLVSEI 854
S +N P P P + P+ P S+ P H + + S
Sbjct: 503 STSSD---ENNTP----------PEPVTPQAEQPHS---PSSISSPHIVHNKGS-VHSRH 545
Query: 855 VAPSPDAIVP--PLAVRRSTRVRHPPGYLADYDCPQ-QTTPHPLSTYYSYDKLTPSYKV- 910
+ D+ P P + + R +HPP YL DY + ++PH S S ++P+
Sbjct: 546 LNEDHDSSSPGLPELLGKGHRPKHPPVYLKDYVAHKVHSSPHTSSPGLSDSNVSPTVSAN 605
Query: 911 ---YSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWV 967
+ + E +F + EW AM+ E+ A+EAN TWDV LP GK I SKWV
Sbjct: 606 HIAFMAAILDSNEQNHFKDDVLIKEWCDAMQKEIEALEANHTWDVTDLPHGKKAISSKWV 665
Query: 968 YKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQ 1027
YKLK + DG+++RHKARLV G Q+ GID+ +TF+PVAK+TTVR+LLA+AA W++FQ
Sbjct: 666 YKLKFNSDGTLERHKARLVVMGNHQKEGIDFKETFAPVAKMTTVRLLLAVAAAKDWDVFQ 725
Query: 1028 LDINNAFLNGDLFEEVYMKIPQGYDVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQH 1087
+D++NAFL+GDL +E S+YGLKQA R WFAK S+ L +
Sbjct: 726 MDVHNAFLHGDLEQE----------------------SLYGLKQAPRCWFAKLSTALRKL 763
Query: 1088 GFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLK 1147
GF SY DYSLF+ T + LVYV+D I+ G N ID K+ L F +KDLGKLK
Sbjct: 764 GFTQSYEDYSLFSLNRDGTVIHFLVYVDDFIIVGNNLKAIDHFKEHLHKCFHMKDLGKLK 823
Query: 1148 FFLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQ 1207
+FLGLE++ A G LSQ+ YAL ++++ G +P+++PM+ + KL S + P + +Q
Sbjct: 824 YFLGLEVSRGADGFCLSQQKYALDIINEAGLLGYKPSAVPMELHHKLGSISSPVFDNPAQ 883
Query: 1208 YRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQG 1261
YRRL+ R +YLTI+RPD+++ ++ LSQF+ P H A L+RYLK +P QG
Sbjct: 884 YRRLVDRFIYLTITRPDLSYAVHILSQFMQTPLEAHWHATLRLVRYLKGSPDQG 937
Score = 92.0 bits (227), Expect = 2e-18
Identities = 112/404 (27%), Positives = 158/404 (38%), Gaps = 96/404 (23%)
Query: 203 IKSHVLLMNPLPPMNRVFSMVLQEEKQREIAARSADLNSA---FAAQATNSGKSGK---- 255
I+S + +PLP NRV+S V++ ++ ++A NS F+ + ++ +
Sbjct: 18 IRSKITDEDPLPSHNRVYSRVIRGQQNLDVARSKETTNSEAINFSVKTPSAPQVAAVYAP 77
Query: 256 KDRDRPLCSYCGKLGHSVDRCFKKHGYPPGLNFKNKSSAVHHVDSSEV------------ 303
K RDR C++C + GH V CF HG+P ++ K + D+ EV
Sbjct: 78 KPRDRS-CTHCHRQGHDVTDCFLVHGFPEWY-YEQKGGSRVSSDNREVVSRLENKPAKRE 135
Query: 304 --------------SGSSVPQDPPITSSQYQQLLSLLTAQLATSPMPSSSTSEPLQIPSP 349
+ + P S Q QL+SLL AQ STSE L
Sbjct: 136 GRSSKGNGRGRGRVNSARAPLSSSNGSDQITQLISLLQAQRP------KSTSERL----- 184
Query: 350 TDLKGFVFSASSDFSHKTSVWILDSGASCHVCFHLSSFESYHSVRSHTISLPDNTKARVT 409
S + T V I+DSGAS H+ S + ++ PD + T
Sbjct: 185 -----------SGNTCLTDV-IIDSGASHHMTGDCSILVDVFDIIPSAVTKPDGKASCAT 232
Query: 410 HIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLENPKYSIHFFHKTFVIQENKLKTIGRG 469
T+ L +S L +V +VP F L+SVS LL+ F +T IG G
Sbjct: 233 KCVTLLLSSSYKLQDVLFVPDFDCTLISVSKLLKQTGP----FSRTL---------IGAG 279
Query: 470 DTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYN 529
+ +YY V AS S + ST + ALWH RLGHPS
Sbjct: 280 EVRERVYYF----TGVLVASAHKTS---------------SDSTSSGALWHRRLGHPS-- 318
Query: 530 RLSLLSSTIHCKIPSSINENV--CPVCPLAKQKRLPFVSENHFA 571
S L S C S + C C AKQ LP + N A
Sbjct: 319 -TSFLLSLPECHQSSKDLGKIDSCDTCSRAKQ-TLPSLIRNFCA 360
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 435 bits (1118), Expect = e-121
Identities = 249/566 (43%), Positives = 349/566 (60%), Gaps = 67/566 (11%)
Query: 756 HKFSPRAVP---CVLVGYKQGYKGYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQ 812
H+FS P V+ Y GYKGYK+ +L + + I+R+VVF+E+ FPF+ ++ S
Sbjct: 318 HQFSCAYTPQQNSVVERYPSGYKGYKVLDLESHSISITRNVVFHETKFPFKTSKFLKESV 377
Query: 813 DPF---LVPLP*VGLSVPPYDVPQPESVPV-----------------PSADHIPATPLVS 852
D F ++PLP P + V ES+P+ SA IP PL S
Sbjct: 378 DMFPNSILPLP-----APLHFV---ESMPLDDDLRADDNNASTSNSASSASSIP--PLPS 427
Query: 853 EIVAPSPDAI---VPPLAVRRSTRVRHPPGYLADYDC----------------------- 886
+ + DA+ + + R R P YL++Y C
Sbjct: 428 TVNTQNTDALDIDTNSVPIARPKRNAKAPAYLSEYHCNSVPFLSSLSPTTSTSIETPSSS 487
Query: 887 --PQQ-TTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAA 943
P++ TTP+P+ST SYDKLTP + Y EP F QA++ +W +A EL A
Sbjct: 488 IPPKKITTPYPMSTAISYDKLTPLFHSYICAYNVETEPKAFTQAMKSEKWTRAANEELHA 547
Query: 944 MEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFS 1003
+E N+TW V SL GK+ +G KWV+ +K +PDGSI+R+KARLVA+G+TQQ GIDYM+TFS
Sbjct: 548 LEQNKTWIVESLTEGKNVVGCKWVFTIKYNPDGSIERYKARLVAQGFTQQEGIDYMETFS 607
Query: 1004 PVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGY-DVKGENL---- 1058
PVAK +V++LL LAA W L Q+D++NAFL+G+L EE+YM +PQGY G +L
Sbjct: 608 PVAKFGSVKLLLGLAAATGWSLTQMDVSNAFLHGELDEEIYMSLPQGYTPPTGISLPSKP 667
Query: 1059 TCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDII 1118
CRL KS+YGLKQASRQW+ + SSV F S D ++F K S + +V+LVYV+D++
Sbjct: 668 VCRLLKSLYGLKQASRQWYKRLSSVFLGANFIQSPADNTMFVKVSCTSIIVVLVYVDDLM 727
Query: 1119 VSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGF 1178
++ +++ ++++K+LL+S FK+KDLG +FFLGLEIA S+ GIS+ QR YA +LL D G
Sbjct: 728 IASNDSSAVENLKELLRSEFKIKDLGPARFFLGLEIARSSEGISVCQRKYAQNLLEDVGL 787
Query: 1179 TDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSK 1238
+ C+P+S+PMDPNL L+ + G L +++ YR L+GRLLYL I+RPDI F ++ LSQFLS
Sbjct: 788 SGCKPSSIPMDPNLHLTKEMGTLLPNATSYRELVGRLLYLCITRPDITFAVHTLSQFLSA 847
Query: 1239 PTTTHLDALHHLLRYLKTTPGQGFFF 1264
PT H+ A H +LRYLK PGQG +
Sbjct: 848 PTDIHMQAAHKVLRYLKGNPGQGLMY 873
Score = 106 bits (264), Expect = 1e-22
Identities = 75/205 (36%), Positives = 99/205 (47%), Gaps = 17/205 (8%)
Query: 326 LTAQLATS-PMPSSSTSEPLQIPSPTDLKGFVFSASSDFSHKTSVWILDSGASCHVCFHL 384
L AQ +TS +P STS + + T + S + S + WI+DSGAS HVC L
Sbjct: 56 LMAQTSTSGTIPFPSTSLKYENNNLTFQNHTLSSLQNVLS--SDAWIIDSGASSHVCSDL 113
Query: 385 SSFESYHSVRSHTISLPDNTKARVTHIGTVKLGNSLILHNVCYVPTFTVNLLSVSALLEN 444
+ F V T++LP+ T+ +TH GT+ + ++LILHNV VP F NL+SV L++
Sbjct: 114 TMFRELIHVSGVTVTLPNGTRVAITHTGTICITSTLILHNVLLVPDFKFNLISVCCLVKT 173
Query: 445 PKYSIHFFHKTFVIQE-NKLKTIGRGDTHNGLYYLHASSNNVFSASTSPLSVLNSLCNQS 503
YS HFF IQE + IGRG T+N LY L + FS S S
Sbjct: 174 LSYSAHFFADCCYIQELTRGLMIGRGKTYNNLYILETQRTS-FSPSLPAASSFTGTVQDD 232
Query: 504 VFPSHCNKSTCTAALWHARLGHPSY 528
LWH RLG Y
Sbjct: 233 ------------CLLWHQRLGIRHY 245
Score = 96.7 bits (239), Expect = 9e-20
Identities = 44/88 (50%), Positives = 61/88 (69%)
Query: 591 HVGHRFFLTIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKELQ 650
H + +FLT+VDD +R TW +++K+KSE F K++ TQ+ IK +R+DN KEL
Sbjct: 247 HYRNLYFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKLIFTQYNAKIKAIRSDNVKELA 306
Query: 651 LTAFLEKQGTLHQFSCVERPQQNSVVER 678
T F+++QG +HQFSC PQQNSVVER
Sbjct: 307 FTKFVKEQGMIHQFSCAYTPQQNSVVER 334
>At4g23160 putative protein
Length = 1240
Score = 418 bits (1075), Expect = e-116
Identities = 211/412 (51%), Positives = 281/412 (67%), Gaps = 7/412 (1%)
Query: 863 VPPLAVRRSTRVRHPPGYLADYDCPQ--QTTPHPLSTYYSYDKLTPSYKVYSVKVASHYE 920
VP +V S R P YL DY C T H +S + SY+K++P Y + V +A E
Sbjct: 26 VPEPSVHTSHRRTRKPAYLQDYYCHSVASLTIHDISQFLSYEKVSPLYHSFLVCIAKAKE 85
Query: 921 PTYFHQAIQYPEWQQAMRTELAAMEANQTWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDR 980
P+ +++A ++ W AM E+ AME TW++ +LPP K IG KWVYK+K + DG+I+R
Sbjct: 86 PSTYNEAKEFLVWCGAMDDEIGAMETTHTWEICTLPPNKKPIGCKWVYKIKYNSDGTIER 145
Query: 981 HKARLVAKGYTQQNGIDYMDTFSPVAKITTVRMLLALAAMYKWELFQLDINNAFLNGDLF 1040
+KARLVAKGYTQQ GID+++TFSPV K+T+V+++LA++A+Y + L QLDI+NAFLNGDL
Sbjct: 146 YKARLVAKGYTQQEGIDFIETFSPVCKLTSVKLILAISAIYNFTLHQLDISNAFLNGDLD 205
Query: 1041 EEVYMKIPQGY-----DVKGENLTCRLKKSIYGLKQASRQWFAKFSSVLTQHGFQHSYHD 1095
EE+YMK+P GY D N C LKKSIYGLKQASRQWF KFS L GF S+ D
Sbjct: 206 EEIYMKLPPGYAARQGDSLPPNAVCYLKKSIYGLKQASRQWFLKFSVTLIGFGFVQSHSD 265
Query: 1096 YSLFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQLLKSAFKLKDLGKLKFFLGLEIA 1155
++ F K + F+ +LVYV+DII+ N +D +K LKS FKL+DLG LK+FLGLEIA
Sbjct: 266 HTYFLKITATLFLCVLVYVDDIIICSNNDAAVDELKSQLKSCFKLRDLGPLKYFLGLEIA 325
Query: 1156 YSATGISLSQRAYALSLLHDTGFTDCRPTSLPMDPNLKLSSDTGPELADSSQYRRLIGRL 1215
SA GI++ QR YAL LL +TG C+P+S+PMDP++ S+ +G + D+ YRRLIGRL
Sbjct: 326 RSAAGINICQRKYALDLLDETGLLGCKPSSVPMDPSVTFSAHSGGDFVDAKAYRRLIGRL 385
Query: 1216 LYLTISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLRYLKTTPGQGFFFLGQ 1267
+YL I+R DI+F +NKLSQF P H A+ +L Y+K T GQG F+ Q
Sbjct: 386 MYLQITRLDISFAVNKLSQFSEAPRLAHQQAVMKILHYIKGTVGQGLFYSSQ 437
>At4g03810 putative retrotransposon protein
Length = 964
Score = 387 bits (994), Expect = e-107
Identities = 265/867 (30%), Positives = 418/867 (47%), Gaps = 85/867 (9%)
Query: 420 LILHNVCYVPTFTVNLLSVSAL-LENPKYSIHFFHKTFVIQENKLKTIGRGDTHNGLYYL 478
L L N YVP N++SVS L +E +SI +NK + R D ++Y
Sbjct: 3 LELKNCYYVPAINKNIISVSCLDMEGFHFSI----------KNKCCSFDRDD----MFYG 48
Query: 479 HASSNNVFSASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYNRLSLLSSTI 538
A +N + + N + F S+ T LWH RLGH + + L S
Sbjct: 49 SAPLDNGLHVLNQSMPIYNIRTKK--FKSNDLNPTF---LWHCRLGHINEKHIQKLHSDG 103
Query: 539 HCKIPSSINENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFHVPTHVGHRFFL 598
+ C C L K + PF + A+ LIH DV GP +++F+
Sbjct: 104 LLNSFDYESYETCESCLLGKMTKAPFTGHSERASDLLGLIHTDVCGPMSTSARGNYQYFI 163
Query: 599 TIVDDHSRFTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKELQLTAF---L 655
T DD SR+ + +L+KHKS++ F VQ QFGK IK +R+D E F L
Sbjct: 164 TFTDDFSRYGYVYLMKHKSKSFENFKEFQNEVQNQFGKSIKALRSDRGGEYLSQVFSDHL 223
Query: 656 EKQGTLHQFSCVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINLI 715
+ G + Q + PQ N V ERR++ LL++ RS++ + +P FWG + T+ ++N
Sbjct: 224 RECGIVSQLTPPGTPQWNGVSERRNRTLLDMVRSMMSHTDLPSPFWGYALETSAFMLNRC 283
Query: 716 PTPNLHNKSPSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYK 775
P+ ++ K+P + + P+ L+++GC ++ L K P++ C VGY + K
Sbjct: 284 PSKSV-EKTPYEIWTGKVPNLSFLKIWGCESYAKRL--ITDKLGPKSDKCYFVGYPKETK 340
Query: 776 GYKLYNLTTKTFHISRDVVFNESIFPFQNTRPYVYSQDPFLVPLP*VGLSVPPYDVPQPE 835
GY Y+ T + R+ F E F + T G V +V +P+
Sbjct: 341 GYYFYHPTDNKVFVVRNGAFLEREFLSKGTS----------------GSKVLLEEVREPQ 384
Query: 836 -SVPVPSADHIPATPLVSEIVAPSPDAIVPPLAVRRSTRVRHPPGYLADYDCPQQTTPHP 894
VP +H V E + P+ VRRS R RH P D+
Sbjct: 385 GDVPTSQEEHQLDLRRVVEPILVEPE-------VRRSERSRHEPDRFRDWVMDD------ 431
Query: 895 LSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPE---WQQAMRTELAAMEANQTWD 951
+++ + EPT + +A+ P+ W +A ++E+ +M N+ W
Sbjct: 432 ----------------HALFMIESDEPTSYEEALMGPDSDKWLEAAKSEMESMSQNKVWT 475
Query: 952 VVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTV 1011
+V LP G I KW++K K+ DG+I +KA LVAKGY Q +GIDY +T+SPVA + ++
Sbjct: 476 LVDLPDGVKPIECKWIFKKKIDMDGNIQIYKAGLVAKGYKQVHGIDYDETYSPVAMLKSI 535
Query: 1012 RMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGYDV-KGENLTCRLKKSIYGLK 1070
R+LLA AA Y +E++Q+D+ AFLNG+L E VYM P+G+ V + C+L +SIYGLK
Sbjct: 536 RILLATAAHYDYEIWQMDVKTAFLNGNLEEHVYMTQPEGFTVPEAARKVCKLHRSIYGLK 595
Query: 1071 QASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSV 1130
QASR W +F+ + + F + + ++ K SG L++YV+DI++ G + + SV
Sbjct: 596 QASRSWNLRFNEAIKEFDFIRNEEEPCVYKKTSGSAVAFLVLYVDDILLLGNDIPLLQSV 655
Query: 1131 KQLLKSAFKLKDLGKLKFFLGLEIAYSATG--ISLSQRAYALSLLHDTGFTDCRPTSLPM 1188
K L S F +KD+G+ + LG+ I I LSQ Y +LH D + +PM
Sbjct: 656 KTWLGSCFSMKDMGEAAYILGIRIYRDRLNKIIGLSQDTYIDKVLHRFNMHDSKKGFIPM 715
Query: 1189 DPNLKLSSDTGPELADSSQ------YRRLIGRLLY-LTISRPDIAFTMNKLSQFLSKPTT 1241
+ LS P D + Y IG ++Y + +RPD+A ++ S++ S P
Sbjct: 716 SHGITLSKTQCPSTHDERERMSKIPYASAIGSIMYAMLYTRPDVACALSMTSRYQSDPGE 775
Query: 1242 THLDALHHLLRYLKTTPGQGFFFLGQQ 1268
+H + ++ +YL+ T + + G +
Sbjct: 776 SHWIVVRNIFKYLRRTKDKFLVYGGSE 802
>At4g22040 LTR retrotransposon like protein
Length = 1109
Score = 369 bits (947), Expect = e-102
Identities = 176/373 (47%), Positives = 258/373 (68%), Gaps = 1/373 (0%)
Query: 889 QTTPHPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQ 948
Q T P S S ++ T S+K + V + EPT +++A+ W++AM E+ ++ NQ
Sbjct: 569 QETEFPYSKM-SCNRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQ 627
Query: 949 TWDVVSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKI 1008
T+ +V+LPPGK +G+KWVYK+K DG+I+R+KARLV G Q+ G+DY +TF+PVAK+
Sbjct: 628 TFSIVNLPPGKRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKM 687
Query: 1009 TTVRMLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGYDVKGENLTCRLKKSIYG 1068
+TVR+ L +AA W + Q+D++NAFL+GDL EEVYMK+PQG+ + CRL KS+YG
Sbjct: 688 STVRLFLGVAAARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQCDDPSKVCRLHKSLYG 747
Query: 1069 LKQASRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEID 1128
LKQA R WF+K SS L Q+GF S DYSLF+ + FV +LVYV+D+I+SG +
Sbjct: 748 LKQAPRCWFSKLSSALKQYGFTQSLSDYSLFSYNNDGVFVHVLVYVDDLIISGSCPDAVA 807
Query: 1129 SVKQLLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPM 1188
K L+S F +KDLG LK+FLG+E++ +A G LSQR Y L ++ + G RP++ P+
Sbjct: 808 QFKSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPL 867
Query: 1189 DPNLKLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDALH 1248
+ N KLS T P L+DSS+YRRL+GRL+YL ++RP+++++++ L+QF+ P H +A
Sbjct: 868 EQNHKLSLSTSPLLSDSSRYRRLVGRLIYLAVTRPELSYSVHTLAQFMQNPRQDHWNAAI 927
Query: 1249 HLLRYLKTTPGQG 1261
++RYLK+ PGQG
Sbjct: 928 RVVRYLKSNPGQG 940
Score = 285 bits (728), Expect = 2e-76
Identities = 164/431 (38%), Positives = 230/431 (53%), Gaps = 28/431 (6%)
Query: 376 ASCHVCFHLSSFESYHSVRSHTISLPDNTKARVTHIGTVKLGNSLILHNVCYVPTFTVNL 435
AS H+ +L S+ I L D K GTV+LG+ LIL +V YV +L
Sbjct: 169 ASHHMTGNLELLSGMRSMSPVLIILADGNKRVAVSEGTVRLGSHLILKSVFYVKELESDL 228
Query: 436 LSVSALLENPKYSIHFFHKTFVIQENKLKTI-GRGDTHNGLYYLHASSNN--VFSASTSP 492
+SV +++ + VIQ+ + + G G NG + N V ++ +P
Sbjct: 229 ISVGQMMDENHCVVQLADHFLVIQDRTTRMVTGIGKRENGSFCFRGMENAAAVHTSVKAP 288
Query: 493 LSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYNRLSLLSSTIHCKIPSSINENVCP 552
LWH RLGH S ++LL + I ENVC
Sbjct: 289 FD-----------------------LWHRRLGHASDKIVNLLPREL-LSSGKEILENVCD 324
Query: 553 VCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFHVPTHVGHRFFLTIVDDHSRFTWTFL 612
C AKQ R F ++ + SF LIHCDVWGP+ P++ G R+FLTIVDD+SR W +L
Sbjct: 325 TCMRAKQTRDTFPLRDNRSMDSFQLIHCDVWGPYRTPSYSGARYFLTIVDDYSRGVWVYL 384
Query: 613 IKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKE-LQLTAFLEKQGTLHQFSCVERPQ 671
+ KSE + F +V+ QF IK VR+DN E L + + +G H+ SCV P
Sbjct: 385 MTDKSETQKHLKDFMALVERQFDTEIKTVRSDNGTEFLCMREYFLHKGIAHETSCVGTPH 444
Query: 672 QNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINLIPTPNLHNKSPSTLLYQ 731
QN VER+H+ +LN+AR+L FQS++P+ FWGECI +A +LIN P+ L KSP +LY+
Sbjct: 445 QNGRVERKHRHILNIARALRFQSYLPIQFWGECILSAAYLINRTPSMLLQGKSPYEMLYK 504
Query: 732 QDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYKGYKLYNLTTKTFHISR 791
P Y HLRVFG LC++ KF+ R+ CV VGY G KG++L++L + F +SR
Sbjct: 505 TAPKYSHLRVFGSLCYAHNQNHKGDKFAARSRRCVFVGYPHGQKGWRLFDLEEQKFFVSR 564
Query: 792 DVVFNESIFPF 802
DV+F E+ FP+
Sbjct: 565 DVIFQETEFPY 575
Score = 102 bits (253), Expect = 2e-21
Identities = 45/149 (30%), Positives = 80/149 (53%)
Query: 24 PFFLHHSDNPGLILVSQPLNGENYNSWNRSMMIALSVKNKLGFINGDFPRPADDDPLLQS 83
P+ L +DN G ++ L NY W AL + K GF++G P+P D P L+
Sbjct: 20 PYDLTAADNSGAVISHPILKTNNYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLED 79
Query: 84 WIRNDHVVMSWILNCVSKDIVSSVICCRSAKEIWQDLQDRFHQPNGTRIYQLKKDLLNLQ 143
W+ + +++SW+ + ++++++ A+++W+ ++ RF NG + ++K DL +
Sbjct: 80 WLTINALLVSWMKMTIDSELLTNISHRDVARDLWEQIRKRFSVSNGPKNQKMKADLATCK 139
Query: 144 QESLTVTQFYTKHKAIWDELQDYLPQRSC 172
QE +TV +Y K IWD + Y P R C
Sbjct: 140 QEGMTVEGYYGKLNKIWDNINSYRPLRIC 168
>At2g24660 putative retroelement pol polyprotein
Length = 1156
Score = 365 bits (938), Expect = e-101
Identities = 176/369 (47%), Positives = 250/369 (67%)
Query: 893 HPLSTYYSYDKLTPSYKVYSVKVASHYEPTYFHQAIQYPEWQQAMRTELAAMEANQTWDV 952
+PLS Y S D + +K + + ++ EP +F +A++ W AM E+ A+E N+TWD+
Sbjct: 611 YPLSDYVSDDCFSAGHKAFLAAITANDEPKHFKEAVRIKVWNDAMFKEVDALEINKTWDI 670
Query: 953 VSLPPGKHEIGSKWVYKLKLHPDGSIDRHKARLVAKGYTQQNGIDYMDTFSPVAKITTVR 1012
V LPPGK IGS+WVYK K + DGSI+R+KARLV +G Q G DY +TF+PV K+TTVR
Sbjct: 671 VDLPPGKVAIGSQWVYKTKYNADGSIERYKARLVVQGNKQVEGEDYNETFAPVVKMTTVR 730
Query: 1013 MLLALAAMYKWELFQLDINNAFLNGDLFEEVYMKIPQGYDVKGENLTCRLKKSIYGLKQA 1072
LL L A +WE++Q+D+NNAFL+GDL EEVYMK+P G+ + CRL+KS+YGLKQA
Sbjct: 731 TLLRLVAANQWEVYQMDVNNAFLHGDLDEEVYMKLPPGFRHSHPDKVCRLRKSLYGLKQA 790
Query: 1073 SRQWFAKFSSVLTQHGFQHSYHDYSLFTKGSGDTFVVLLVYVNDIIVSGPNATEIDSVKQ 1132
R WF K S L + GF + DYS F+ + +LVYV+D+++ G + + K+
Sbjct: 791 PRCWFKKLSDALLRFGFVQGHEDYSFFSYTRNGIELRVLVYVDDLLICGNDGYMLQKFKE 850
Query: 1133 LLKSAFKLKDLGKLKFFLGLEIAYSATGISLSQRAYALSLLHDTGFTDCRPTSLPMDPNL 1192
L F +KDLGKLK+FLG+E++ + GI LSQR YAL ++ D+G CRP P++ N
Sbjct: 851 YLGRCFSMKDLGKLKYFLGIEVSRGSEGIFLSQRKYALDIITDSGNLGCRPALTPLEQNH 910
Query: 1193 KLSSDTGPELADSSQYRRLIGRLLYLTISRPDIAFTMNKLSQFLSKPTTTHLDALHHLLR 1252
L++D GP LAD+ YRRL+GRLLYL +RP+++++++ LSQF+ P HL A ++R
Sbjct: 911 HLATDDGPLLADAKPYRRLVGRLLYLLHTRPELSYSVHVLSQFMQTPREAHLAAAMRIVR 970
Query: 1253 YLKTTPGQG 1261
+LK +PGQG
Sbjct: 971 FLKGSPGQG 979
Score = 266 bits (681), Expect = 5e-71
Identities = 154/390 (39%), Positives = 212/390 (53%), Gaps = 38/390 (9%)
Query: 439 SALLENPKYSIHFFHK--TFVIQENKLKT----------IGRGDTHNGLYYLHASSNNVF 486
S+L N + SI H+ T + LK IG G+ +G+YYL +V
Sbjct: 20 SSLNSNKEESISSEHRLVTLTLTHENLKACKLDRFSRTLIGSGEERDGVYYL----TDVA 75
Query: 487 SASTSPLSVLNSLCNQSVFPSHCNKSTCTAALWHARLGHPSYNRLSLLSSTIHCKIPSSI 546
+A H K + ALWH RLGHPS++ LS L + S+
Sbjct: 76 TAKI-----------------HTAKVSSDQALWHQRLGHPSFSVLSSLPVLTSSSL--SV 116
Query: 547 NENVCPVCPLAKQKRLPFVSENHFANHSFDLIHCDVWGPFHVPTHVGHRFFLTIVDDHSR 606
C VC AKQ R F + + F LIHCDVWGP+ VP+ G +FLTIVDD SR
Sbjct: 117 GSRSCDVCFRAKQTREVFPVSTNKSIECFSLIHCDVWGPYRVPSSCGAVYFLTIVDDFSR 176
Query: 607 FTWTFLIKHKSEAPLAIMAFFKMVQTQFGKVIKMVRTDNAKELQ-LTAFLEKQGTLHQFS 665
WT+L+ KSE + F + QFGK +K++R+DN E L ++ + G +HQ S
Sbjct: 177 AVWTYLLLAKSEVRTVLTNFLVYTEKQFGKSVKVLRSDNGTEFMCLASYFREHGIVHQTS 236
Query: 666 CVERPQQNSVVERRHQQLLNVARSLLFQSHMPLVFWGECISTATHLINLIPTPNLHNK-S 724
CV PQQN VER+H+ +LNVAR++LFQ+ +P+ FWGE + TA +LIN PT +LHN S
Sbjct: 237 CVGTPQQNGRVERKHRHILNVARAILFQASLPIQFWGEAVLTAAYLINRTPT-SLHNGLS 295
Query: 725 PSTLLYQQDPSYDHLRVFGCLCFSSTLPSTRHKFSPRAVPCVLVGYKQGYKGYKLYNLTT 784
P +L+ P+Y+HLRVFG C+ + KF R+ CV +GY KG+K++++
Sbjct: 296 PYEILHNSKPNYEHLRVFGSACYVHRASRDKDKFGERSRLCVFIGYPFAQKGWKVFDMEK 355
Query: 785 KTFHISRDVVFNESIFPFQNTRPYVYSQDP 814
K F +SRDVVF E +FP+ T S P
Sbjct: 356 KEFLVSRDVVFREDVFPYAATNTDHVSASP 385
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.136 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,240,408
Number of Sequences: 26719
Number of extensions: 1492447
Number of successful extensions: 4978
Number of sequences better than 10.0: 175
Number of HSP's better than 10.0 without gapping: 120
Number of HSP's successfully gapped in prelim test: 58
Number of HSP's that attempted gapping in prelim test: 3944
Number of HSP's gapped (non-prelim): 511
length of query: 1427
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1315
effective length of database: 8,326,068
effective search space: 10948779420
effective search space used: 10948779420
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0110a.16


