

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.15
(184 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
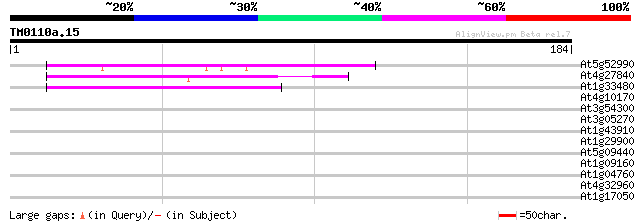
Sequences producing significant alignments: (bits) Value
At5g52990 unknown protein 58 3e-09
At4g27840 unknown protein 55 2e-08
At1g33480 unknown protein 44 6e-05
At4g10170 unknown protein 40 0.001
At3g54300 synaptobrevin -like protein 34 0.039
At3g05270 unknown protein 29 1.2
At1g43910 unknown protein 29 1.2
At1g29900 carbamoyl phosphate synthetase large chain (carB) 29 1.6
At5g09440 unknown protein 28 3.6
At1g09160 putative protein phosphatase 2C 28 3.6
At1g04760 synaptobrevin 7B, putative 28 3.6
At4g32960 unknown protein 27 4.7
At1g17050 prenyl transferase (prephytoene pyrophosphatase dehydr... 27 6.2
>At5g52990 unknown protein
Length = 272
Score = 57.8 bits (138), Expect = 3e-09
Identities = 39/113 (34%), Positives = 62/113 (54%), Gaps = 5/113 (4%)
Query: 13 HSLFSHTVNNRTYTFLI-DPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETL-DSVND-S 69
HS+FSHTV+ +TYTF I D F+YF+I D K ++ LNR+RS++++ + D +D
Sbjct: 46 HSMFSHTVHKKTYTFAIDDDSFVYFSISDESMEKPESFWVLNRLRSAIEDLIKDGGSDVE 105
Query: 70 TAPPPLS--LQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSA 120
T P+S LQ D + EI+ + D +VG + + +DS+
Sbjct: 106 TLINPVSHCLQLKLDPVFAEIVGVVDLELLDMDLVGSPRSVARESRNPSIDSS 158
>At4g27840 unknown protein
Length = 260
Score = 55.1 bits (131), Expect = 2e-08
Identities = 35/102 (34%), Positives = 52/102 (50%), Gaps = 14/102 (13%)
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRS---SLKETLDSVNDS 69
HS+ SHTV+ RTY +ID F YFAI D KS+++ NR++S SL E + +
Sbjct: 46 HSMISHTVHKRTYALIIDGLFSYFAILDEVVAKSESVWLFNRLKSATESLMEDGSTADSL 105
Query: 70 TAPPPLSLQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLK 111
P LQ+ D + EI A +G +H++ L+
Sbjct: 106 DNPTQHCLQSKLDPVFAEI-----------AAIGGNHNKDLE 136
>At1g33480 unknown protein
Length = 508
Score = 43.5 bits (101), Expect = 6e-05
Identities = 23/77 (29%), Positives = 37/77 (47%)
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAP 72
H + T +T+ FL+ F+YFAI D KS L FL ++R LKE + +
Sbjct: 300 HKWYFETRGKKTFGFLMKDDFVYFAIVDDVFKKSSVLDFLEKLRDELKEANKKNSRGSFS 359
Query: 73 PPLSLQTPFDLILQEIL 89
+S D I++ ++
Sbjct: 360 GSISFSNVQDQIVRRLI 376
>At4g10170 unknown protein
Length = 254
Score = 39.7 bits (91), Expect = 0.001
Identities = 18/49 (36%), Positives = 26/49 (52%)
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKE 61
H+ + T+ R + FLI F+YFAI D +S L FL +R K+
Sbjct: 45 HTWYFETIGKRRFGFLIGDGFVYFAIVDEVLKRSSVLKFLEHLRDEFKK 93
>At3g54300 synaptobrevin -like protein
Length = 240
Score = 34.3 bits (77), Expect = 0.039
Identities = 15/61 (24%), Positives = 31/61 (50%), Gaps = 2/61 (3%)
Query: 14 SLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDS--VNDSTA 71
S ++++ + T+ FL+D F++ + D +S FL R++ K+ ++ ND
Sbjct: 44 SKYTYSCDGHTFNFLVDNGFVFLVVADESTGRSVPFVFLERVKEDFKKRYEASIKNDERH 103
Query: 72 P 72
P
Sbjct: 104 P 104
>At3g05270 unknown protein
Length = 603
Score = 29.3 bits (64), Expect = 1.2
Identities = 27/111 (24%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 76 SLQTPFDL-ILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIAN--RTSAKGL 132
S+ T D+ ++ + L ++ P + G H E K+ + + + +TS + +
Sbjct: 284 SMATSVDIGLMDDFLEMEKLAALPHSEPGRKHSESNKELEKSNAHVNQLKHELKTSLRRI 343
Query: 133 AAAAA-TELVELEKDAVEEALVGSAIQIEGFRRVWLDLRSRLSLLHESRSE 182
+ E+VE+EK +E AL GS QIE + ++ +LS + + +E
Sbjct: 344 SELEEKVEMVEVEKLQLEMALNGSKEQIEALQSRLKEIEGKLSEMKKLEAE 394
>At1g43910 unknown protein
Length = 475
Score = 29.3 bits (64), Expect = 1.2
Identities = 18/55 (32%), Positives = 28/55 (50%), Gaps = 1/55 (1%)
Query: 11 STHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDS 65
S + SH N + YT+ D AIF+H HT +TL ++ +L + LD+
Sbjct: 170 SAEKIMSHRENLKIYTYNQDRSKWESAIFEH-HTTFETLAVEPDLKKTLIDDLDA 223
>At1g29900 carbamoyl phosphate synthetase large chain (carB)
Length = 1187
Score = 28.9 bits (63), Expect = 1.6
Identities = 28/112 (25%), Positives = 49/112 (43%), Gaps = 11/112 (9%)
Query: 50 TFLNR----IRSSLKETLDSVNDSTAPPPLSLQTPFDLILQEILHLDDGNRSP--PAVVG 103
TF +R RS K+ S + ST PP L+ ++ +L+ + L D P P +VG
Sbjct: 34 TFFSRSAIYYRSKPKQASSSSSFSTFPPCLNRKSSLTHVLKPVSELADTTTKPFSPEIVG 93
Query: 104 ISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEEALVGS 155
D KK M++ + ++ + + A + L ++ E L+ S
Sbjct: 94 KRTD---LKKIMILGAGPIVIGQACEFDYSGTQACK--ALREEGYEVILINS 140
>At5g09440 unknown protein
Length = 278
Score = 27.7 bits (60), Expect = 3.6
Identities = 14/51 (27%), Positives = 28/51 (54%)
Query: 112 KKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEEALVGSAIQIEGF 162
++ ++VD I++ T+AKG + A+ + E K V +VG + +E +
Sbjct: 50 QRSIIVDFIRSISSVTAAKGPSVASWWKTTEKYKTGVSTLVVGKQLLLENY 100
>At1g09160 putative protein phosphatase 2C
Length = 428
Score = 27.7 bits (60), Expect = 3.6
Identities = 20/83 (24%), Positives = 38/83 (45%), Gaps = 6/83 (7%)
Query: 69 STAPPPLSLQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTS 128
S AP P+ Q PF L H+D N++ + + E ++ + + V+A+R
Sbjct: 304 SLAPAPMKKQNPFTSFLSRKNHMDTNNKNGNKLSAVGVVE-----ELFEEGSAVLADRL- 357
Query: 129 AKGLAAAAATELVELEKDAVEEA 151
K L + T L++ ++E+
Sbjct: 358 GKDLLSNTETGLLKCAVCQIDES 380
>At1g04760 synaptobrevin 7B, putative
Length = 220
Score = 27.7 bits (60), Expect = 3.6
Identities = 13/61 (21%), Positives = 27/61 (43%)
Query: 11 STHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDST 70
S+++ F++ + T+ +L D F Y + + + FL R++ + ST
Sbjct: 41 SSNNKFTYNCDGHTFNYLADNGFTYCVVVIESAGRQIPMAFLERVKEDFNKRYGGGKAST 100
Query: 71 A 71
A
Sbjct: 101 A 101
>At4g32960 unknown protein
Length = 264
Score = 27.3 bits (59), Expect = 4.7
Identities = 19/67 (28%), Positives = 33/67 (48%), Gaps = 1/67 (1%)
Query: 112 KKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEEALVGSAIQIEGFRRVWLDLRS 171
KK+ D+ + +A+ G +++ +L L + A +EA V A Q FR + + RS
Sbjct: 91 KKEFTSDAESAVASLRGLSGNKSSSRADLTLLFRAAAQEAKVSRA-QNRIFRVILIYCRS 149
Query: 172 RLSLLHE 178
+ HE
Sbjct: 150 SMRPTHE 156
>At1g17050 prenyl transferase (prephytoene pyrophosphatase
dehydrogenase) like protein
Length = 417
Score = 26.9 bits (58), Expect = 6.2
Identities = 22/86 (25%), Positives = 41/86 (47%), Gaps = 4/86 (4%)
Query: 74 PLSLQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLA 133
P+SL+T F+++ ++ L+D S +VG + + + + SA R L
Sbjct: 92 PISLETLFEVVADDLQRLNDNLLS---IVGAENPVLISAAEQIF-SAGGKRMRPGLVFLV 147
Query: 134 AAAATELVELEKDAVEEALVGSAIQI 159
+ A EL L++ VE +G I++
Sbjct: 148 SRATAELAGLKELTVEHRRLGEIIEM 173
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,008,382
Number of Sequences: 26719
Number of extensions: 155996
Number of successful extensions: 490
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 480
Number of HSP's gapped (non-prelim): 13
length of query: 184
length of database: 11,318,596
effective HSP length: 93
effective length of query: 91
effective length of database: 8,833,729
effective search space: 803869339
effective search space used: 803869339
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0110a.15


