

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.13
(1389 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
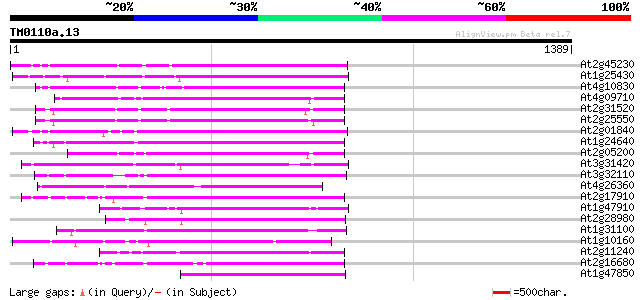
Score E
Sequences producing significant alignments: (bits) Value
At2g45230 putative non-LTR retroelement reverse transcriptase 352 7e-97
At1g25430 hypothetical protein 352 7e-97
At4g10830 putative protein 343 3e-94
At4g09710 RNA-directed DNA polymerase -like protein 340 3e-93
At2g31520 putative non-LTR retroelement reverse transcriptase 335 1e-91
At2g25550 putative non-LTR retroelement reverse transcriptase 335 1e-91
At2g01840 putative non-LTR retroelement reverse transcriptase 323 3e-88
At1g24640 hypothetical protein 323 3e-88
At2g05200 putative non-LTR retroelement reverse transcriptase 323 4e-88
At3g31420 hypothetical protein 322 7e-88
At3g32110 non-LTR reverse transcriptase, putative 320 5e-87
At4g26360 putative protein 313 4e-85
At2g17910 putative non-LTR retroelement reverse transcriptase 313 6e-85
At1g47910 reverse transcriptase, putative 303 3e-82
At2g28980 putative non-LTR retroelement reverse transcriptase 302 1e-81
At1g31100 hypothetical protein 301 1e-81
At1g10160 putative reverse transcriptase 290 3e-78
At2g11240 pseudogene 281 2e-75
At2g16680 putative non-LTR retroelement reverse transcriptase 277 3e-74
At1g47850 hypothetical protein 275 1e-73
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 352 bits (904), Expect = 7e-97
Identities = 243/849 (28%), Positives = 420/849 (48%), Gaps = 32/849 (3%)
Query: 2 RWVSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKMDYVSV 61
R +SWN +G+G R + E PE++ + E+K + ++ V D +V
Sbjct: 2 RILSWNCQGVGNTPTVRHLREIRGLYFPEVIFLCETKKRRNYLENVVGHLGF--FDLHTV 59
Query: 62 DAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDF 121
+ +G +GGL +W S ++ + R I LL + ++ +YG V +ER +
Sbjct: 60 EPIGKSGGLALMWKD-SVQIKVLQSDKRLIDALL--IWQDKEFYLTCIYGEPVQAERGEL 116
Query: 122 FNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESS---FSDFVADSNLIDLPLQ 178
+ L+ L L+ ++ GDFN +++ E+IG ESS F + L ++
Sbjct: 117 WERLTR-LGLSRSGPWMLTGDFNELVDPSEKIGGPARKESSCLEFRQMLNSCGLWEVNHS 175
Query: 179 SDKFTWYSSRAGGLWS-RIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFG 237
+F+WY +R L R+DR + ++ + F + SD P+I +L N+
Sbjct: 176 GYQFSWYGNRNDELVQCRLDRTVANQAWMELFPQAKATYLQKICSDHSPLINNLVGDNWR 235
Query: 238 P-KPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVS 296
F + W+ + F+ L+ ++W +T + ++ +K+ + I +W+ + S
Sbjct: 236 KWAGFKYDKRWVQREGFKDLLCNFWSQQSTK--TNALMMEKIASCRREISKWKRVSKPSS 293
Query: 297 RVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLRE 356
VRI ++ L ++ + + K L+ QEY EE W++K+R+ W+R
Sbjct: 294 AVRIQELQFKLDAATKQIPFDRRELARLKKELS------QEYNNEEQFWQEKSRIMWMRN 347
Query: 357 GDRNTSFFHSISKIRKSKKLISQLVVGG----VSIEDPNMIKDAIFSHFQSFFTSENRLR 412
GDRNT +FH+ +K R+++ I +L+ S ED + +A +F+ F SE+
Sbjct: 348 GDRNTKYFHAATKNRRAQNRIQKLIDEEGREWTSDEDLGRVAEA---YFKKLFASEDVGY 404
Query: 413 PKIRCANLSRL-SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVK 471
NL+ L S +L ++ E+ S + ++ PG DG N ++QFW +
Sbjct: 405 TVEELENLTPLVSDQMNNNLLAPITKEEVQRATFSINPHKCPGPDGMNGFLYQQFWETMG 464
Query: 472 EDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASR 531
+ + + + F +G + +G+N T I LIPK ++DFRPISL YK+I K++A+R
Sbjct: 465 DQITEMVQAFFRSGSIEEGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANR 524
Query: 532 LQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLDFAKDYDNI 588
L+ + P LISE Q AF +GR ISD ILIA+E++ L N K F+ +K D +K YD +
Sbjct: 525 LKKILPSLISETQAAFVKGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRV 584
Query: 589 DWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLF 648
+W FL ++ + F D WI I CV + +L+NG+ G +GLRQGDPLSP+LF
Sbjct: 585 EWPFLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLF 644
Query: 649 NLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDLLCNTLL 707
+C L +L + +G+++ G ++HL FADD++ +C+ ++ + + +
Sbjct: 645 VICTEMLVKMLQSAEQKNQITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIE 704
Query: 708 SFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASF 766
+ SG +VN KS+I G ++ V G +YLG P + +
Sbjct: 705 EYSLASGQRVNYLKSSIYFGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVAT 764
Query: 767 WEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRS 826
+ D+L +K W S F+S G+ L+KA ++P + M+ FK P + ++E ++
Sbjct: 765 LSYLKDRLGKKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAE 824
Query: 827 FLQGGKLHG 835
F K G
Sbjct: 825 FWWKNKKEG 833
>At1g25430 hypothetical protein
Length = 1213
Score = 352 bits (904), Expect = 7e-97
Identities = 272/862 (31%), Positives = 416/862 (47%), Gaps = 48/862 (5%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKMDYVSVD--A 63
WN+RG R + + KP V E+ + + + F NA+ + V+ A
Sbjct: 8 WNIRGFNNVSHRSGFKKWVKANKPIFGGVIETHVKQPKDRK---FINALLPGWSFVENYA 64
Query: 64 VGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFN 123
G + +W+P+ + A K+++ I + S + + VY N + R + +
Sbjct: 65 FSDLGKIWVMWDPSVQVVVVA-KSLQMITCEVLLPGSPSWIIVSVVYAANEVASRKELWI 123
Query: 124 LLSEALSLNNCFCKIMG-------GDFNAILNEGERIG-IQMGVESSFSDF---VADSNL 172
+ +N I+G GDFN +LN E + + V+ + DF + + L
Sbjct: 124 EI-----VNMVVSGIIGDRPWLVLGDFNQVLNPQEHSNPVSLNVDINMRDFRDCLLAAEL 178
Query: 173 IDLPLQSDKFTWYS-SRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRP--IIL 229
DL + + FTW++ S + +IDR LV++ F + SD ++L
Sbjct: 179 SDLRYKGNTFTWWNKSHTTPVAKKIDRILVNDSWNALFPSSLGIFGSLDFSDHVSCGVVL 238
Query: 230 SLGAINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQ 289
+I +PF F+N+ L + +F LV D W + VG S F + +KLK LK IK++
Sbjct: 239 EETSIK-AKRPFKFFNYLLKNLDFLNLVRDNWFTLNVVGSSMFRVSKKLKALKKPIKDFS 297
Query: 290 GYASSVSRVRINTIEADLQVTMDR-LASEEP---GIELRKNRLTILEGLWQEYRK-EEYR 344
S R L DR LA P EL R W EE
Sbjct: 298 RLNYSELEKRTKEAHDFLIGCQDRTLADPTPINASFELEAERK------WHILTAAEESF 351
Query: 345 WRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQS 403
+RQK+R+ W EGD NT +FH ++ R S IS L G G ++ I D S+F S
Sbjct: 352 FRQKSRISWFAEGDGNTKYFHRMADARNSSNSISALYDGNGKLVDSQEGILDLCASYFGS 411
Query: 404 FFTSENRLRPKIRCAN-----LS-RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDG 457
E + P + N LS R S A LE FS +I L S N++ G DG
Sbjct: 412 LLGDE--VDPYLMEQNDMNLLLSYRCSPAQVCELESTFSNEDIRAALFSLPRNKSCGPDG 469
Query: 458 FNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLI 517
F FF W IV +V ++F S+G L+K NAT I LIPK P+ SDFRPIS +
Sbjct: 470 FTAEFFIDSWSIVGAEVTDAIKEFFSSGCLLKQWNATTIVLIPKIVNPTCTSDFRPISCL 529
Query: 518 GSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLN-KKQGGGFL 576
+ YK+I+++L RLQ + +IS Q AF GR +++ +L+A ++V N +
Sbjct: 530 NTLYKVIARLLTDRLQRLLSGVISSAQSAFLPGRSLAENVLLATDLVHGYNWSNISPRGM 589
Query: 577 LKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKG 636
LK+D K +D++ W F+ L+ + +++INWI C+ST T ++ +NG + GFF+ KG
Sbjct: 590 LKVDLKKAFDSVRWEFVIAALRALAIPEKFINWISQCISTPTFTVSINGGNGGFFKSTKG 649
Query: 637 LRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDE 696
LRQGDPLSP+LF L + S LL+ + L+++HL FADD ++F +
Sbjct: 650 LRQGDPLSPYLFVLAMEAFSNLLHSRYESGLIHYHPKASNLSISHLMFADDVMIFFDGGS 709
Query: 697 NQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAP 756
+ +C TL F SGLKVN KS + ++ E+ A YG+ IG LPI YLG P
Sbjct: 710 FSLHGICETLDDFASWSGLKVNKDKSHLYLAGLNQLESNANAA-YGFPIGTLPIRYLGLP 768
Query: 757 LGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKV 816
L R + +EP+L+K+ + R+W +K +S +GR+ L+ + + F M+ F P
Sbjct: 769 LMNRKLRIAEYEPLLEKITARFRSWVNKCLSFAGRIQLISSVIFGSINFWMSTFLLPKGC 828
Query: 817 VGEVEKILRSFLQGGKLHGYHG 838
+ +E + FL G + G
Sbjct: 829 IKRIESLCSRFLWSGNIEQAKG 850
>At4g10830 putative protein
Length = 1294
Score = 343 bits (881), Expect = 3e-94
Identities = 242/781 (30%), Positives = 401/781 (50%), Gaps = 35/781 (4%)
Query: 65 GSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNL 124
G +GGL LW S + + ++ R I + + +++ ++ VYG SER+ +
Sbjct: 424 GHSGGLALLWKD-SVRLSNLYQDDRHIDVHIS--INNINFYLSRVYGHPCQSERHSLWTH 480
Query: 125 LSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLIDLPLQSDK 181
E LS I+ GDFN IL+ E+IG E +F F V+ +L D+ D+
Sbjct: 481 F-ENLSKTRNDPWILIGDFNEILSNNEKIGGPQRDEWTFRGFRNMVSTCDLKDIRSIGDR 539
Query: 182 FTWYSSR-AGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFGP-K 239
F+W R + + +DR ++ + F + SD +P+ LSL +
Sbjct: 540 FSWVGERHSHTVKCCLDRAFINSEGAFLFPFAELEFLEFTGSDHKPLFLSLEKTETRKMR 599
Query: 240 PFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVR 299
PF F L F+ V W A + L +++ + + + + ++ SR+R
Sbjct: 600 PFRFDKRLLEVPHFKTYVKAGWNKA--INGQRKHLPDQVRTCRQAMAKLKHKSNLNSRIR 657
Query: 300 INTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDR 359
IN ++A L M + E R+ I L YR EE W+QK+R +W++EGDR
Sbjct: 658 INQLQAALDKAMSSVNRTE-----RRTISHIQRELTVAYRDEERYWQQKSRNQWMKEGDR 712
Query: 360 NTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKD--AIFSHFQSFFTS--ENRLRPK- 414
NT FFH+ +K R S +++LV + E+ + + I H Q FFT E+ RP
Sbjct: 713 NTEFFHACTKTRFS---VNRLVT--IKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRPVS 767
Query: 415 -IRCANLSRL--SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVK 471
I A + Q N L K S+ EI+ + ++APG DG F+K W IV
Sbjct: 768 IIDFAGFKPIVTEQIND-DLTKDLSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVG 826
Query: 472 EDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASR 531
DV+K + F T + + +N T I +IPK P T+SD+RPI+L YK+ISK L R
Sbjct: 827 PDVIKEVKIFFRTSYMKQSINHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVER 886
Query: 532 LQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLDFAKDYDNI 588
L+ ++S++Q AF GR ++D ++IA+E++ L K+ ++ +K D +K YD +
Sbjct: 887 LKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRV 946
Query: 589 DWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLF 648
+W FL ++ F + WI WI V + S+LVNG G + ++G+RQGDPLSP+LF
Sbjct: 947 EWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLF 1006
Query: 649 NLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDLLCNTLL 707
LC + L+ L+ + + + G+RIG G+ + HLQFADD+L FC+++ L +
Sbjct: 1007 ILCADILNHLIKNRVAEGDIRGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFD 1066
Query: 708 SFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASF 766
+ + SG K+N+ KS I G V R+ + G YLG P ++
Sbjct: 1067 VYEYYSGQKINMSKSMITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDM 1126
Query: 767 WEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRS 826
+ ++++++++ +W++K++S +G+ ++K+ S+P++ M+ FK PL +V E+E +L +
Sbjct: 1127 FNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMN 1186
Query: 827 F 827
F
Sbjct: 1187 F 1187
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 340 bits (873), Expect = 3e-93
Identities = 235/732 (32%), Positives = 364/732 (49%), Gaps = 32/732 (4%)
Query: 112 PNVDSERYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVES---SFSDFVA 168
PN R F++ +S +L ++ GDFN IL+ E+ G + E +F FV+
Sbjct: 51 PNFIDNRSVFWDKIS-SLGAQRSSAWLLTGDFNDILDNSEKQGGPLRWEGFFLAFRSFVS 109
Query: 169 DSNLIDLPLQSDKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPI 227
+ L D+ + +W +R + SR+DR L + F + SD RP+
Sbjct: 110 QNGLWDINHTGNSLSWRGTRYSHFIKSRLDRALGNCSWSELFPMSKCEYLRFEGSDHRPL 169
Query: 228 ILSLGAINFG-PKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIK 286
+ GA KPF F +E LV + W A + K+ + I
Sbjct: 170 VTYFGAPPLKRSKPFRFDRRLREKEEIRALVKEVWELARQDS-----VLYKISRCRQSII 224
Query: 287 EWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWR 346
+W +S S I + L+ L+++ P L + I + L YR+EE W+
Sbjct: 225 KWTKEQNSNSAKAIKKAQQALE---SALSADIPDPSLIGS---ITQELEAAYRQEELFWK 278
Query: 347 QKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQSFF 405
Q +RV+WL GDRN +FH+ ++ R+ +S + G G + I I S+FQ+ F
Sbjct: 279 QWSRVQWLNSGDRNKGYFHATTRTRRMLNNLSVIEDGSGQEFHEEEQIASTISSYFQNIF 338
Query: 406 TSENRLRPKIRCANLSRLSQAN-KMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFK 464
T+ N ++ LS + ++ L K S EI E L S ++APG DGF+ +FF
Sbjct: 339 TTSNNSDLQVVQEALSPIISSHCNEELIKISSLLEIKEALFSISADKAPGPDGFSASFFH 398
Query: 465 QFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLI 524
+W I++ DV + F L LN T ++LIPK + P VSD+RPI+L YK++
Sbjct: 399 AYWDIIEADVSRDIRSFFVDSCLSPRLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIV 458
Query: 525 SKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL---NKKQGGGFLLKLDF 581
+K+L RLQ LIS +Q AF GR I+D +LI +E++ FL K+ +K D
Sbjct: 459 AKILTRRLQPWLSELISLHQSAFVPGRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDM 518
Query: 582 AKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGD 641
+K YD I W FL ++L + F D+WI W+ CV T + S L+NGS G +GLRQGD
Sbjct: 519 SKAYDRIKWNFLQEVLMRLGFHDKWIRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGD 578
Query: 642 PLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQID 700
PLSP+LF LC LS L + + G+R+ G +NHL FADDT+ FC+ +
Sbjct: 579 PLSPYLFILCTEVLSGLCRKAQEKGVMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCG 638
Query: 701 LLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGM-----YGWSIGQLPILYLGA 755
L N L + SG +NL KS I + ++ +R + IG+ YLG
Sbjct: 639 ALSNILKKYELASGQSINLAKSAITFSSKTPQDIKRRVKLSLRIDNEGGIGK----YLGL 694
Query: 756 PLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLK 815
P R+ + ++D++R++ +W+ +F+S +G+ L+KA L S+P + M FK P
Sbjct: 695 PEHFGRRKRDIFSSIVDRIRQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPAS 754
Query: 816 VVGEVEKILRSF 827
+ +++ +L F
Sbjct: 755 LCKQIQSVLTRF 766
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 335 bits (859), Expect = 1e-91
Identities = 235/793 (29%), Positives = 405/793 (50%), Gaps = 59/793 (7%)
Query: 65 GSAGGLISLW-NPASYAMESAVKNVRFIGLLLRALVDSSLLF------IMNVYGPNVDSE 117
G +GGL +W N S ++ S + L+DS + F + VYG SE
Sbjct: 218 GRSGGLALMWKNNVSLSLISQDER----------LIDSHVTFNNKSFYLSCVYGHPTQSE 267
Query: 118 RYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLID 174
R+ + L E +S N ++ GDFN IL+ E+IG M E +F +F V+ ++ D
Sbjct: 268 RHQLWQTL-EHISDNRNAEWLLVGDFNEILSNAEKIGGPMREEWTFRNFRNMVSHCDIED 326
Query: 175 LPLQSDKFTWYSSR-AGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG- 232
+ + D+F+W R + +DR ++ F ++ SD +P+++
Sbjct: 327 MRSKGDRFSWVGERHTHTVKCCLDRVFINSAWTATFPYAETEFLDFTGSDHKPVLVHFNE 386
Query: 233 AINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYA 292
+ K F F N + F+++V W T + + +++ + + + +
Sbjct: 387 SFPRRSKLFRFDNRLIDIPTFKRIVQTSW--RTNRNSRSTPITERISSCRQAMARLKHAS 444
Query: 293 SSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVR 352
+ S RI +++ L M+ + R+ + E L + + EE W+QK+R +
Sbjct: 445 NLNSEQRIKKLQSSLNRAMESTRRVD-----RQLIPQLQESLAKAFSDEEIYWKQKSRNQ 499
Query: 353 WLREGDRNTSFFHSISKIRKSKKLISQLV--VGGVSIEDPNMIKDAIFSHFQSFFT---S 407
W++EGD+NT +FH+ +K R S+ ++ ++ G + D I +H Q FFT S
Sbjct: 500 WMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDKE-----IGNHAQDFFTNIFS 554
Query: 408 ENRLRPK-IRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQ 465
N ++ I A+ S ++ + L K+FS+ EI++ + ++APG DG F+K
Sbjct: 555 TNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKN 614
Query: 466 FWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLIS 525
W IV DV+ + F T + +N T I +IPK P+T+SD+RPI+L YK+IS
Sbjct: 615 CWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVIS 674
Query: 526 KVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGG---FLLKLDFA 582
K L +RL+ ++S++Q AF GR I+D ++IA+EV+ L ++ +K D +
Sbjct: 675 KCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVS 734
Query: 583 KDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDP 642
K YD ++W FL ++ F ++WI WI V + S+L+NGS G+ +G+RQGDP
Sbjct: 735 KAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDP 794
Query: 643 LSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDL 701
LSP+LF LC + LS L+N + GVRIG G A+ HLQFADD+L FC+ +
Sbjct: 795 LSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQA 854
Query: 702 LCNTLLSFLFTSGLKVNLHKSTI-IGCNV------DVRETQRVAGMYGWSIGQLPILYLG 754
L + + + SG K+N+ KS I G V +++ + G YLG
Sbjct: 855 LKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSKLKQILEIPNQGGGG------KYLG 908
Query: 755 APLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPL 814
P ++ +E ++D+++++ TW+++F+S +G+ ++K+ ++P++ M+ FK P
Sbjct: 909 LPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPK 968
Query: 815 KVVGEVEKILRSF 827
+V E+E +L +F
Sbjct: 969 GIVSEIESLLMNF 981
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 335 bits (859), Expect = 1e-91
Identities = 237/792 (29%), Positives = 406/792 (50%), Gaps = 57/792 (7%)
Query: 65 GSAGGLISLW-NPASYAMESAVKNVRFIGLLLRALVDSSLLF------IMNVYGPNVDSE 117
G +GGL +W N S ++ S + L+DS + F + VYG SE
Sbjct: 444 GRSGGLALMWKNNVSLSLISQDER----------LIDSHVTFNNKSFYLSCVYGHPTQSE 493
Query: 118 RYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLID 174
R+ + L E +S N ++ GDFN IL+ E+IG M E +F +F V+ ++ D
Sbjct: 494 RHQLWQTL-EHISDNRNAEWLLVGDFNEILSNAEKIGGPMREEWTFRNFRNMVSHCDIED 552
Query: 175 LPLQSDKFTWYSSR-AGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG- 232
+ + D+F+W R + +DR ++ F ++ SD +P+++
Sbjct: 553 MRSKGDRFSWVGERHTHTVKCCLDRVFINSAWTATFPYAEIEFLDFTGSDHKPVLVHFNE 612
Query: 233 AINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYA 292
+ K F F N + F+++V W T + + +++ + + + +
Sbjct: 613 SFPRRSKLFRFDNRLIDIPTFKRIVQTSW--RTNRNSRSTPITERISSCRQAMARLKHAS 670
Query: 293 SSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVR 352
+ S RI +++ L M+ + R+ + E L + + EE W+QK+R +
Sbjct: 671 NLNSEQRIKKLQSSLNRAMESTRRVD-----RQLIPQLQESLAKAFSDEEIYWKQKSRNQ 725
Query: 353 WLREGDRNTSFFHSISKIRKSKKLISQLV--VGGVSIEDPNMIKDAIFSHFQSFFT---S 407
W++EGD+NT +FH+ +K R S+ ++ ++ G + D I +H Q FFT S
Sbjct: 726 WMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDKE-----IGNHAQDFFTNIFS 780
Query: 408 ENRLRPK-IRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQ 465
N ++ I A+ S ++ + L K+FS+ EI++ + ++APG DG F+K
Sbjct: 781 TNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKN 840
Query: 466 FWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLIS 525
W IV DV+ + F T + +N T I +IPK P+T+SD+RPI+L YK+IS
Sbjct: 841 CWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVIS 900
Query: 526 KVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGG---FLLKLDFA 582
K L +RL+ ++S++Q AF GR I+D ++IA+EV+ L ++ +K D +
Sbjct: 901 KCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVS 960
Query: 583 KDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDP 642
K YD ++W FL ++ F ++WI WI V + S+L+NGS G+ +G+RQGDP
Sbjct: 961 KAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDP 1020
Query: 643 LSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDL 701
LSP+LF LC + LS L+N + GVRIG G A+ HLQFADD+L FC+ +
Sbjct: 1021 LSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQA 1080
Query: 702 LCNTLLSFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPI-----LYLGA 755
L + + + SG K+N+ KS I G V R+ I ++P YLG
Sbjct: 1081 LKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSRLK-----QILEIPNQGGGGKYLGL 1135
Query: 756 PLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLK 815
P ++ +E ++D+++++ TW+++F+S +G+ ++K+ ++P++ M+ FK P
Sbjct: 1136 PEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKG 1195
Query: 816 VVGEVEKILRSF 827
+V E+E +L +F
Sbjct: 1196 IVSEIESLLMNF 1207
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 323 bits (829), Expect = 3e-88
Identities = 233/852 (27%), Positives = 408/852 (47%), Gaps = 46/852 (5%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKM---DYVSVD 62
WN +GLG+ R + E +++ + E+K + + + VKM D +
Sbjct: 368 WNCQGLGQPLTVRRLEEVQRVYFLDMLFLIETKQQDNYTRDL-----GVKMGFEDMCIIS 422
Query: 63 AVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFF 122
G +GGL+ W +++ +VR + L + + ++ +YG + SER+ +
Sbjct: 423 PRGLSGGLVVYWKK-HLSIQVISHDVRLVDLYVE--YKNFNFYLSCIYGHPIPSERHHLW 479
Query: 123 NLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQ---MGVESSFSDFVADSNLIDLPLQS 179
L + +S + +M GDFN ILN E+ G + +G +F++ + N+ DL +
Sbjct: 480 EKL-QRVSAHRSGPWMMCGDFNEILNLNEKKGGRRRSIGSLQNFTNMINCCNMKDLKSKG 538
Query: 180 DKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSL------- 231
+ ++W R + S +DR ++ D F SD P+I+ +
Sbjct: 539 NPYSWVGKRQNETIESCLDRVFINSDWQASFPAFETEFLPIAGSDHAPVIIDIAEEVCTK 598
Query: 232 -GAINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQG 290
G + + F F ++F V W + + +KL + + +W+
Sbjct: 599 RGQFRYDRRHFQF-------EDFVDSVQRGWNRGRSDSHGGYY--EKLHCCRQELAKWKR 649
Query: 291 YASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKAR 350
+ + +I T++ + A+E + L + + L Q YR EE W K+R
Sbjct: 650 RTKTNTAEKIETLKYRVD------AAERDHTLPHQTILRLRQDLNQAYRDEELYWHLKSR 703
Query: 351 VRWLREGDRNTSFFHSISKIRKSKKLISQLV--VGGVSIEDPNMIKDAIFSHFQSFFTSE 408
RW+ GDRNT FF++ +K+RKS+ I + G + D + K A F T++
Sbjct: 704 NRWMLLGDRNTMFFYASTKLRKSRNRIKAITDAQGIENFRDDTIGKVAENYFADLFTTTQ 763
Query: 409 NRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWG 468
+I ++++ L + ++ E+ + + + +RAPG DGF F+ W
Sbjct: 764 TSDWEEIISGIAPKVTEQMNHELLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHLWD 823
Query: 469 IVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVL 528
++ DV F + + +N T I LIPK P +SD+RPISL ++YK+ISK+L
Sbjct: 824 LIGNDVCLMVRHFFESDVMDNQINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKIL 883
Query: 529 ASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKK---QGGGFLLKLDFAKDY 585
RL+ +IS++Q AF G+ ISD +L+A+E++ L + Q G +K D +K Y
Sbjct: 884 IKRLKQCLGDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKAY 943
Query: 586 DNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSP 645
D ++W FL ++ ++ F RW+ WI TCV++ + +L+NGS G +G+RQGDPLSP
Sbjct: 944 DRVEWNFLEKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSP 1003
Query: 646 FLFNLCVNGLSCLLNQLLDDSNFSGVRIGPG-LALNHLQFADDTLLFCENDENQIDLLCN 704
+LF C LS +L + + G++I LA++HL FADD+L FC I+ L
Sbjct: 1004 YLFLFCAEVLSNMLRKAEVNKQIHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQLAL 1063
Query: 705 TLLSFLFTSGLKVNLHKSTII-GCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRR 763
+ SG K+N KS+II G + QR+ + G + YLG P R+
Sbjct: 1064 IFKKYEEASGQKINYAKSSIIFGQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRK 1123
Query: 764 ASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKI 823
+E ++ K++ + W ++S +G+ ++KA ++P++ M F P + E+ +
Sbjct: 1124 VELFEYIVTKVKERTEGWAYNYLSPAGKEIVIKAIAMALPVYSMNCFLLPTLICNEINSL 1183
Query: 824 LRSFLQGGKLHG 835
+ +F G + G
Sbjct: 1184 ITAFWWGKENEG 1195
>At1g24640 hypothetical protein
Length = 1270
Score = 323 bits (829), Expect = 3e-88
Identities = 237/789 (30%), Positives = 398/789 (50%), Gaps = 42/789 (5%)
Query: 60 SVDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLF------IMNVYGPN 113
+V+ VG GGL LW S +++F+ + L+D+ + F + VYG
Sbjct: 21 TVEPVGKCGGLALLWK------SSVQVDLKFVD---KNLMDAQVQFGAVNFCVSCVYGDP 71
Query: 114 VDSERYDFFNLLSE-ALSLNNCFCKIMGGDFNAILNEGERIGIQMGVE---SSFSDFVAD 169
S+R + +S + + +C M GDFN IL+ GE+ G + +F++ +
Sbjct: 72 DRSKRSQAWERISRIGVGRRDKWC--MFGDFNDILHNGEKNGGPRRSDLDCKAFNEMIKG 129
Query: 170 SNLIDLPLQSDKFTWYSSRAGGLW--SRIDRWLVSEDAINKFDGISQSVENWGLSDRRPI 227
+L+++P + FTW + R G W R+DR +++ F +Q+ ++ SD RP+
Sbjct: 130 CDLVEMPAHGNGFTW-AGRRGDHWIQCRLDRAFGNKEWFCFFPVSNQTFLDFRGSDHRPV 188
Query: 228 ILSLGAINFGPK-PFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIK 286
++ L + + F F +L ++ ++ + W S G + V +L+ + +
Sbjct: 189 LIKLMSSQDSYRGQFRFDKRFLFKEDVKEAIIRTW-SRGKHGTNISVAD-RLRACRKSLS 246
Query: 287 EWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILE-GLWQEYRKEEYRW 345
W+ + S +IN +EA L+ E+ + R+++L+ L + YR+EE W
Sbjct: 247 SWKKQNNLNSLDKINQLEAALE-------KEQSLVWPIFQRVSVLKKDLAKAYREEEAYW 299
Query: 346 RQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFS-HFQSF 404
+QK+R +WLR G+RN+ +FH+ K + +K I +L +++ K + + +F +
Sbjct: 300 KQKSRQKWLRSGNRNSKYFHAAVKQNRQRKRIEKLKDVNGNMQTSEAAKGEVAAAYFGNL 359
Query: 405 FTSENRLRPKIRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFF 463
F S N + L R+S+ SL + S EI E + S APG DG + FF
Sbjct: 360 FKSSNPSGFTDWFSGLVPRVSEVMNESLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFF 419
Query: 464 KQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKL 523
+ +W V V + F + G + N T + LIPK P+ + D RPISL YK+
Sbjct: 420 QHYWSTVGNQVTSEVKKFFADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKI 479
Query: 524 ISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLD 580
ISK++A RLQ P ++S+ Q AF R I+D IL+A+E+V L + + F+ +K D
Sbjct: 480 ISKIMAKRLQPWLPEIVSDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEFMAVKSD 539
Query: 581 FAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQG 640
+K YD ++W +L +L + F +W+NWI CVS+ T S+L+N G +++GLRQG
Sbjct: 540 MSKAYDRVEWSYLRSLLLSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQG 599
Query: 641 DPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGP-GLALNHLQFADDTLLFCENDENQI 699
DPLSPFLF LC GL+ LLN+ + G++ G ++HL FADD+L C+ Q
Sbjct: 600 DPLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCKASREQS 659
Query: 700 DLLCNTLLSFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLG 758
+L L + +G +NL+KS+I G VD + + G YLG P
Sbjct: 660 LVLQKILKVYGNATGQTINLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPEC 719
Query: 759 GNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVG 818
+ + + D+L+ K W ++ +S G+ L+K+ ++P+F M+ FK P+
Sbjct: 720 FSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCE 779
Query: 819 EVEKILRSF 827
+E + SF
Sbjct: 780 NLESAMASF 788
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 323 bits (828), Expect = 4e-88
Identities = 214/704 (30%), Positives = 354/704 (49%), Gaps = 39/704 (5%)
Query: 144 NAILNEGERIGIQMGVESSFSDF---VADSNLIDLPLQSDKFTWYSSRAGG-LWSRIDRW 199
N IL+ E+ G + SF DF ++ + L DL + F+W R + R+DR
Sbjct: 36 NEILDNSEKRGGPPRDQGSFIDFRSFISKNGLWDLKYSGNPFSWRGMRYDWFVRQRLDRA 95
Query: 200 LVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFGPKPFLFYNHWLLDKEF-EKLVS 258
+ + + F + SD RP+++ + + +++ L D + L+
Sbjct: 96 MSNNSWLESFPSGRSEYLRFEGSDHRPLVVFVDEARVKRRGQFRFDNRLRDNDVVNALIQ 155
Query: 259 DWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEADLQVTMDRLASEE 318
+ W +A A VL K+ + I W + S I + L+ + L ++
Sbjct: 156 ETWTNAG----DASVLT-KMNQCRREIINWTRLQNLNSAELIEKTQKALE---EALTADP 207
Query: 319 PGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLIS 378
P LE Y+ EE W+Q++RV WL GDRNT +FH++++ R+++ ++
Sbjct: 208 PNPTTIGALTATLE---HAYKLEEQFWKQRSRVLWLHSGDRNTGYFHAVTRNRRTQNRLT 264
Query: 379 QLV-VGGVSIEDPNMIKDAIFSHFQSFFTSENRLRPKIRCANLSRL-SQANKMSLEKQFS 436
+ + GV+ + + I I +FQ FTSE+ + + + SQ + L + +
Sbjct: 265 VMEDINGVAQHEEHQISQIISGYFQQIFTSESDGDFSVVDEAIEPMVSQGDNDFLTRIPN 324
Query: 437 EGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFI 496
+ E+ + + S + ++APG DGF F+ +W I+ DV + F ++ + +N T I
Sbjct: 325 DEEVKDAVFSINASKAPGPDGFTAGFYHSYWHIISTDVGREIRLFFTSKNFPRRMNETHI 384
Query: 497 SLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDC 556
LIPK P V+D+RPI+L YK+++K++ R+Q++ P LISENQ AF GR ISD
Sbjct: 385 RLIPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLISENQSAFVPGRVISDN 444
Query: 557 ILIANEVVDFL---NKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTC 613
+LI +EV+ FL + K+ +K D +K YD ++W FL +L+ F WI+W+ C
Sbjct: 445 VLITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVLQRFGFHSIWIDWVLEC 504
Query: 614 VSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRI 673
V++ + S L+NG+ G +GLRQGDPLSP LF LC LS L + GVR+
Sbjct: 505 VTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSGLCTRAQRLRQLPGVRV 564
Query: 674 G-PGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVR 732
G +NHL FADDT+ F ++D + L L + SG +N HKS++ + R
Sbjct: 565 SINGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFSSKTPR 624
Query: 733 ETQ---------RVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNS 783
+ R G G YLG P R+ + ++DK+R+K +W S
Sbjct: 625 SVKGQVKRILKIRKEGGTG--------KYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWAS 676
Query: 784 KFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSF 827
+F+S +G+ ++KA L S+PL+ M+ FK P + +++ +L F
Sbjct: 677 RFLSQAGKQVMLKAVLASMPLYSMSCFKLPSALCRKIQSLLTRF 720
>At3g31420 hypothetical protein
Length = 1491
Score = 322 bits (826), Expect = 7e-88
Identities = 248/830 (29%), Positives = 401/830 (47%), Gaps = 66/830 (7%)
Query: 30 EIVLVQESKLDSSRVKTVEA---FANAVKMDYVSVDAVGSAGGLISLWNPASYAMESAVK 86
+I+ + E+K R++ V A F N +V+V G +GGL+ WN +S +
Sbjct: 210 DIICLSETKQQDDRIRDVGAELGFCN-----HVTVPPDGLSGGLVVFWN-SSVDIFLCFS 263
Query: 87 NVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNLLSEALSLNNCFCKIMGGDFNAI 146
N + L +++ + ++ VYG S R+ + L + + GDFN I
Sbjct: 264 NSNLVDLHVKS--NEGSFYLSFVYGHPNPSHRHHLWERLERLNTTRQGTAWFIMGDFNEI 321
Query: 147 LNEGERIGIQMGVESSFSDF---VADSNLIDLPLQSDKFTWYSSRAGG-LWSRIDRWLVS 202
L+ E+ G ++ E +F DF V N DL D+F+W R + +DR + +
Sbjct: 322 LSNREKRGGRLRPERTFQDFRNMVRGCNFTDLKSVGDRFSWAGKRGDHHVTCSLDRTMAN 381
Query: 203 EDAINKFDGISQSVENWGLSDRRPIILSLGAINFGPKPFLFYNHWLLDKE-FEKLVSDWW 261
+ F +G SD RP++ ++ A + F Y+ L KE F+++V + W
Sbjct: 382 NEWHTLFPESETVFMEYGESDHRPLVTNISAQKEERRGFFSYDSRLTHKEGFKQVVLNQW 441
Query: 262 CSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEADLQVTMDRLASEEPGI 321
L +KL + I W+ + RI I L DR + G
Sbjct: 442 HRRNGSFEGDSSLNRKLVECRQAISRWKKNNRVNAEERIKIIRHKL----DRAIAT--GT 495
Query: 322 ELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSK-KLIS-Q 379
+ R + + L Q Y EE W+ K+R RWL GDRNT +FHS +K R+ + +L+S Q
Sbjct: 496 ATPRERRQMRQDLNQAYADEEIYWQTKSRSRWLNAGDRNTRYFHSTTKTRRCRNRLLSVQ 555
Query: 380 LVVGGVSIEDPNMIKDAIFSHFQSFFTSENRLRPKIRCANL-----SRLSQANKMSLEKQ 434
G + D N+ K AI ++F + S +R A++ +++ L +
Sbjct: 556 DSDGDICRGDENIAKVAI-NYFDDLYKSTPNT--SLRYADVFQGFQQKITDEINEDLIRP 612
Query: 435 FSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNAT 494
+E EI E + S +R P DGF +F++QFW +K+ V+ F +L + N T
Sbjct: 613 VTELEIEESVFSVAPSRTPDPDGFTADFYQQFWPDIKQKVIDEVTRFFERSELDERHNHT 672
Query: 495 FISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQIS 554
+ LIPK P+T++ FRPI+L +YK+ISK+L +RL+ I+ENQ AF GR I+
Sbjct: 673 NLCLIPKVETPTTIAKFRPIALCNVSYKIISKILVNRLKKHLGGAITENQAAFVPGRLIT 732
Query: 555 DCILIANEVVDFL--NKKQGGGFL-LKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIR 611
+ +IA+EV L K+Q ++ LK D K YD ++W FL + +++M F +WI I
Sbjct: 733 NNAIIAHEVYYALKARKRQANSYMALKTDITKAYDRLEWDFLEETMRQMGFNTKWIERIM 792
Query: 612 TCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGV 671
CV+ S+L+NGS G + E+G+R GDPLSP+LF LC LS ++ Q + G+
Sbjct: 793 ICVTMVRFSVLINGSPHGTIKPERGIRHGDPLSPYLFILCAEVLSHMIKQAEINKKLKGI 852
Query: 672 RIG-PGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVD 730
R+ G ++HL FADD++ F TL N T I
Sbjct: 853 RLSTQGPFISHLLFADDSIFF-------------TL----------ANQRSCTAIKEPDT 889
Query: 731 VRETQRVAGMYG-WSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLS 789
R + + G++ G+ YLG P N ++ + +++K++ K + W+ KF+S
Sbjct: 890 KRRMRHLLGIHNEGGEGK----YLGLPEQFNKKKKELFNYIIEKVKDKTQGWSKKFLSQG 945
Query: 790 GRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSF--LQGGKLHGYH 837
G+ L+KA ++P++ M IFK +V E++ +L F G + G H
Sbjct: 946 GKEVLLKAVALAMPVYSMNIFKLTKEVCEEIDSLLARFWWSSGNETKGMH 995
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 320 bits (819), Expect = 5e-87
Identities = 243/789 (30%), Positives = 377/789 (46%), Gaps = 58/789 (7%)
Query: 61 VDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYD 120
VDAVG +GGL LW + +FI + + + ++ VY S R
Sbjct: 624 VDAVGHSGGLWLLWRTGIGEVSVVDSTDQFIHA--KDVNGKDNVNLVVVYAAPTASRRSG 681
Query: 121 FFNLLSEAL-SLNNCFCKIMGGDFNAILNEGERIGIQMGVES---SFSDFVADSNLIDLP 176
++ L + + S++ ++GGDFN I+ ER G + S +F +++ D +LIDL
Sbjct: 682 LWDRLGDVIRSMDGPV--VIGGDFNTIVRLDERSGGNGRLSSDSLAFGEWINDHSLIDLG 739
Query: 177 LQSDKFTWYSSRAGGLW--SRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG-- 232
+ +KFTW R + R+DR L A K+ S + SD P+ + L
Sbjct: 740 FKGNKFTWKRGREERFFVAKRLDRVLCCAHARLKWQEASVLHLPFLASDHAPLYVQLTPE 799
Query: 233 -AINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGY 291
+ N G +PF F WL F++L+ + W
Sbjct: 800 VSGNRGRRPFRFEAAWLSHPGFKELL---------------------------LTSWNKD 832
Query: 292 ASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARV 351
S+ +++ + DL + D L EE EL K+ +LE +EE W QK+R
Sbjct: 833 ISTPEALKVQEL-LDLHQSDDLLKKEE---ELLKDFDVVLE-------QEEVVWMQKSRE 881
Query: 352 RWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQSFFTSEN- 409
+W GDRNT FFH+ + IR+ + I L G + + ++ +++ ++ ++
Sbjct: 882 KWFVHGDRNTKFFHTSTIIRRRRNQIEMLQDNDGRWLSNAQELETHAIDYYKRLYSLDDL 941
Query: 410 -RLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWG 468
+ ++ + LS+A+ SL K FS E+ ++S +APG DGF F++Q W
Sbjct: 942 DAVVEQLPQEGFTALSEADFSSLTKPFSPLEVEGAIRSMGKYKAPGPDGFQPVFYQQGWE 1001
Query: 469 IVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVL 528
+V E V KF DF S+G + N + LI K P ++ FRPISL +K I+KV+
Sbjct: 1002 VVGESVTKFVMDFFSSGSFPQETNDVLVVLIAKVLKPEKITQFRPISLCNVLFKTITKVM 1061
Query: 529 ASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQG--GGFLLKLDFAKDYD 586
RL+ V LI Q +F GR +D I++ EVV + +K+G G LLKLD K YD
Sbjct: 1062 VGRLKGVINKLIGPAQTSFIPGRLSTDNIVVVQEVVHSMRRKKGVKGWMLLKLDLEKAYD 1121
Query: 587 NIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPF 646
I W L D LK W+ WI CV ++ +L NG T F+ +GLRQGDPLSP+
Sbjct: 1122 RIRWDLLEDTLKAAGLPGTWVQWIMKCVEGPSMRLLWNGEKTDAFKPLRGLRQGDPLSPY 1181
Query: 647 LFNLCVNGLSCLLNQLLDDSNFSGVRIG-PGLALNHLQFADDTLLFCENDENQIDLLCNT 705
LF LC+ L L+ + + ++I G L+H+ FADD +LF E +QI +L
Sbjct: 1182 LFVLCIERLCHLIESSIAAKKWKPIKISQSGPRLSHICFADDLILFAEASIDQIRVLRGV 1241
Query: 706 LLSFLFTSGLKVNLHKSTIIGCNVDVRET-QRVAGMYGWSIGQLPILYLGAPLGGNPRRA 764
L F SG KV+L KS I +R+ +R++ G + YLG P+
Sbjct: 1242 LEKFCGASGQKVSLEKSKIYFSKNVLRDLGKRISEESGMKATKDLGKYLGVPILQKRINK 1301
Query: 765 SFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKIL 824
+ ++ + + W + +S +GRLTL KA L S+ + M+ K P + ++K+
Sbjct: 1302 ETFGEVIKRFSSRLAGWKGRMLSFAGRLTLTKAVLTSILVHTMSTIKLPQSTLDGLDKVS 1361
Query: 825 RSFLQGGKL 833
R+FL G L
Sbjct: 1362 RAFLWGSTL 1370
>At4g26360 putative protein
Length = 1141
Score = 313 bits (802), Expect = 4e-85
Identities = 235/723 (32%), Positives = 355/723 (48%), Gaps = 37/723 (5%)
Query: 68 GGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNLLSE 127
G + LW+P S + K+++ I + + + I VY N D +R + + ++
Sbjct: 61 GKIWVLWDP-SVEVVIVAKSLQMITCEVLFPNSRTWIVISVVYAANEDDKRKELWREITA 119
Query: 128 ALSLNNCFCK--IMGGDFNAILNEGERIG-IQMGVESSFSDF---VADSNLIDLPLQSDK 181
++ F + I+ GDFN +L+ E + + V+ DF + D+ L DL +
Sbjct: 120 LVASPVTFNRPWILLGDFNQVLHPHEHSRHVSLNVDRRIRDFRECLLDAELSDLVYKGSS 179
Query: 182 FTWYS-SRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRP--IILSLGAINFGP 238
FTW++ S+ + +IDR LV+E N F SD ++L L I
Sbjct: 180 FTWWNKSKTRPVAKKIDRILVNESWSNLFPSSFGLFGPPDFSDHASCGVVLELDPIK-AK 238
Query: 239 KPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRV 298
+PF F+N L + EF LV D W S VG S F + +KLK LK IK++ + S +
Sbjct: 239 RPFKFFNFLLKNPEFLNLVWDVWYSTNVVGSSMFRVSKKLKALKKPIKDFSRL--NYSNL 296
Query: 299 RINTIEA-DLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRK-EEYRWRQKARVRWLRE 356
T EA + ++ L + P +E + L + WQ EE +RQ++RV W E
Sbjct: 297 EKRTEEAHETLLSFQNLTLDNPSLENAAHELEA-QRKWQILATAEESFFRQRSRVTWFAE 355
Query: 357 GDRNTSFFHSISKIRKSKKLISQLVV-GGVSIEDPNMIKDAIFSHFQSFFTSENRL---- 411
GD NT +FH ++ RKS I+ LV G I+ I D +F++ + +N
Sbjct: 356 GDGNTRYFHRMADSRKSVNTITTLVDDSGTQIDSQQGIADHCALYFENLLSDDNDPYSLE 415
Query: 412 RPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVK 471
+ + R + LE FS+ +I N+A G DGF
Sbjct: 416 QDDMNLLLTYRCPYSQVADLEAMFSDEDIKAAFFGLPSNKACGPDGF------------- 462
Query: 472 EDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASR 531
V +F +G L+K NAT I LIPK S SDFRPIS + + YK+I+++L R
Sbjct: 463 -PVTAAVREFFISGNLLKQWNATTIVLIPKFPNASCTSDFRPISCMNTLYKVIARLLTDR 521
Query: 532 LQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGG-GFLLKLDFAKDYDNIDW 590
LQ + +IS +Q AF GR +++ +L+A E+V N + +LK+D K +D++ W
Sbjct: 522 LQKLLSCVISPSQSAFLPGRLLAENVLLATEMVHGYNWRNISLRGMLKVDLRKAFDSVRW 581
Query: 591 GFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNL 650
F+ L + ++INWI C+ST T ++ VNG GFF+ KGLRQGDPLSP+LF L
Sbjct: 582 EFIIAALLALGVPTKFINWIHQCISTPTFTVSVNGCCGGFFKSAKGLRQGDPLSPYLFVL 641
Query: 651 CVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFL 710
+ S LLN D L+++HL FADD ++F + + + +C TL F
Sbjct: 642 AMEVFSKLLNSRFDSGYIRYHPKASDLSISHLMFADDVMIFFDGGSSSLHGICETLEDFA 701
Query: 711 FTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPM 770
SGLKVN KS ++ E +A YG+ G LPI YLG PL R + +EP+
Sbjct: 702 SWSGLKVNNDKSHFFCAGLEQAERNSLAA-YGFPQGCLPIRYLGLPLMCRKLRIAEYEPL 760
Query: 771 LDK 773
L+K
Sbjct: 761 LEK 763
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 313 bits (801), Expect = 6e-85
Identities = 245/821 (29%), Positives = 396/821 (47%), Gaps = 46/821 (5%)
Query: 29 PEIVLVQESKLDSSRVKTVEAFANAVKMDYV-SVDAVGSAGGLISLWNP-ASYAMESAVK 86
P+I+ + E+K V V + D++ +V+ G +GGL W A K
Sbjct: 7 PDILFLMETKNSQDFVYKVFCWLG---YDFIHTVEPEGRSGGLAIFWKSHLEIEFLYADK 63
Query: 87 NVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFF-NLLSEALSLNNCFCKIMGGDFNA 145
N+ + + R + + FI VYG V R + +L S L +C I GDFN
Sbjct: 64 NLMDLQVSSR----NKVWFISCVYGLPVTHMRPKLWEHLNSIGLKRAEAWCLI--GDFND 117
Query: 146 ILNEGERIGIQMGVESSFSDF---VADSNLIDLPLQSDKFTWYSSRAGGLW--SRIDRWL 200
I + E++G SSF F + + ++ +L + FTW +R W ++DR
Sbjct: 118 IRSNDEKLGGPRRSPSSFQCFEHMLLNCSMHELGSTGNSFTWGGNR-NDQWVQCKLDRCF 176
Query: 201 VSEDAINKFDGISQ-SVENWGLSDRRPIILSLGAINFGPKPFLFYNHWLLDKEFE----- 254
+ + F Q +E +G SD RP++ + F LF + DK +
Sbjct: 177 GNPAWFSIFPNAHQWFLEKFG-SDHRPVL-----VKFTNDNELFRGQFRYDKRLDDDPYC 230
Query: 255 -KLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEADLQVTMDR 313
+++ W SA + G + L + I W+ + + ++ RI + DL
Sbjct: 231 IEVIHRSWNSAMSQGTHSSFFS--LIECRRAISVWKHSSDTNAQSRIKRLRKDLDAEKSI 288
Query: 314 LASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKS 373
P IE K++L++ Y EE WRQK+R +WL GD+NT FFH+ +
Sbjct: 289 QIPCWPRIEYIKDQLSLA------YGDEELFWRQKSRQKWLAGGDKNTGFFHATVHSERL 342
Query: 374 KKLISQLVVGGVSIEDPNMIKDAIFSHF-QSFFTSENRLRPKIRCANL-SRLSQANKMSL 431
K +S L+ N K I S F ++ FTS L L ++++ +L
Sbjct: 343 KNELSFLLDENDQEFTRNSDKGKIASSFFENLFTSTYILTHNNHLEGLQAKVTSEMNHNL 402
Query: 432 EKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGL 491
++ +E E++ + S + APG DGF FF+Q W +VK ++ F TG L +
Sbjct: 403 IQEVTELEVYNAVFSINKESAPGPDGFTALFFQQHWDLVKHQILTEIFGFFETGVLPQDW 462
Query: 492 NATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGR 551
N T I LIPK P +SD RPISL YK+ISK+L RL+ P ++S Q AF R
Sbjct: 463 NHTHICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQR 522
Query: 552 QISDCILIANEVVDFL---NKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWIN 608
ISD IL+A+E++ L ++ K D +K YD ++W FL ++ + F ++WI+
Sbjct: 523 LISDNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWIS 582
Query: 609 WIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNF 668
WI CV++ + S+L+NG G +G+RQGDPLSP LF LC L +LN+
Sbjct: 583 WIMNCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKI 642
Query: 669 SGVRI-GPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTI-IG 726
+G++ +++NHL FADDTLL C+ + + + L L + SG +NL+KS I G
Sbjct: 643 TGIQFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFG 702
Query: 727 CNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFI 786
NVD++ + G S+ YLG P + + + + +KL+ + W +K +
Sbjct: 703 KNVDIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTL 762
Query: 787 SLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSF 827
S G+ L+K+ ++P+++M+ FK P + ++ ++ F
Sbjct: 763 SQGGKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDF 803
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 303 bits (777), Expect = 3e-82
Identities = 207/639 (32%), Positives = 332/639 (51%), Gaps = 44/639 (6%)
Query: 222 SDRRPIILSLG-AINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKG 280
SD P+I ++ I G + F F W+ + +S W + FV +KL
Sbjct: 4 SDHSPVIATIADKIPRGKQNFRFDKRWIGKDGLLEAISQGWNLDSGFREGQFV--EKLTN 61
Query: 281 LKNRIKEWQGYASSVSRVRINTIEADLQVTM---DRLASEEPGIELRKNRLTILEGLWQE 337
+ I +W+ R I ++A+L V DR E + LR L +
Sbjct: 62 CRRAISKWRKSLIPFGRQTIEDLKAELDVAQRDDDRSREEITELTLR---------LKEA 112
Query: 338 YRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQL--VVGGVSIEDPNMIKD 395
YR EE W QK+R W++ GD N+ FFH+++K R+++ I+ L G SIED + I++
Sbjct: 113 YRDEEQYWYQKSRSLWMKLGDNNSKFFHALTKQRRARNRITGLHDENGIWSIEDDD-IQN 171
Query: 396 AIFSHFQSFFTSENRLRPKIRCANLSRLS-----QANKMSLEKQFSEGEIWEVLKSCDGN 450
S+FQ+ FT+ N P++ L + + N + L +E E+ L
Sbjct: 172 IAVSYFQNLFTTAN---PQVFDEALGEVQVLITDRINDL-LTADATECEVRAALFMIHPE 227
Query: 451 RAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSD 510
+APG DG FF++ W I+K D++ F G K LN T I LIPK P+ +++
Sbjct: 228 KAPGPDGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDKRLNTTNICLIPKTERPTRMTE 287
Query: 511 FRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--N 568
RPISL YK+ISK+L RL+ V P LISE Q AF GR ISD ILIA E+ L N
Sbjct: 288 LRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGRLISDNILIAQEMFHGLRTN 347
Query: 569 KKQGGGFL-LKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSS 627
F+ +K D +K YD ++W F+ +L++M F ++WI+WI C++T +L+NG
Sbjct: 348 SSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWISWIMWCITTVQYKVLINGQP 407
Query: 628 TGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIG-PGLALNHLQFAD 686
G E+GLRQGDPLSP+LF LC L + + + +G+++ P A++HL FAD
Sbjct: 408 KGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLITGIKVATPSPAVSHLLFAD 467
Query: 687 DTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTI-IGCNVD---VRETQRVAGMYG 742
D+L FC+ ++ Q ++ L + SG ++N KS+I G V+ + + + G++
Sbjct: 468 DSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFGHKVEDSIKADIKLILGIH- 526
Query: 743 WSIGQLPILYLGAP--LGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLC 800
++G + YLG P LGG+ + + + D+L+ + W++KF+S G+ ++K+
Sbjct: 527 -NLGGMG-SYLGLPESLGGS--KTKVFSFVRDRLQSRINGWSAKFLSKGGKEVMIKSVAA 582
Query: 801 SVPLFLMAIFKAPLKVVGEVEKILRSF--LQGGKLHGYH 837
++P ++M+ F+ P + ++ + F G G H
Sbjct: 583 TLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMH 621
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 302 bits (773), Expect = 1e-81
Identities = 186/611 (30%), Positives = 317/611 (51%), Gaps = 23/611 (3%)
Query: 237 GPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVL---QQKLKGLKNRIKEWQGYAS 293
G KPF F N +F +V W S+ + S L +KLK LK ++E
Sbjct: 544 GRKPFKFVNVLTKLPQFLPVVESHWASSAPLYVSTSALYRFSKKLKTLKPHLRE------ 597
Query: 294 SVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGL-----WQEYRK-EEYRWRQ 347
+ + ++ + + L ++ ++ TI E L W + EE +Q
Sbjct: 598 -LGKEKLGDLPKRTREAHILLCEKQATTLANPSQETIAEELKAYTDWTHLSELEEGFLKQ 656
Query: 348 KARVRWLREGDRNTSFFHSISKIRKSKKLISQLV-VGGVSIEDPNMIKDAIFSHFQSFFT 406
K+++ W+ GD N S+FH +++RK + I ++ +++ IK F F
Sbjct: 657 KSKLHWMNVGDGNNSYFHKAAQVRKMRNSIREIRGPNAETLQTSEEIKGEAERFFNEFLN 716
Query: 407 SENRLRPKIRCANLSRL-----SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLN 461
++ I +L L S ++ L ++ + EI +VL + N++PG DG+
Sbjct: 717 RQSGDFHGISVEDLRNLMSYRCSVTDQNILTREVTGEEIQKVLFAMPNNKSPGPDGYTSE 776
Query: 462 FFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAY 521
FFK W + D + + F G L KGLNAT ++LIPK + + D+RPIS Y
Sbjct: 777 FFKATWSLTGPDFIAAIQSFFVKGFLPKGLNATILALIPKKDEAIEMKDYRPISCCNVLY 836
Query: 522 KLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVV-DFLNKKQGGGFLLKLD 580
K+ISK+LA+RL+++ P I +NQ AF + R + + +L+A E+V D+ + +K+D
Sbjct: 837 KVISKILANRLKLLLPSFILQNQSAFVKERLLMENVLLATELVKDYHKESVTPRCAMKID 896
Query: 581 FAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQG 640
+K +D++ W FL + L+ +NF + + +WI+ C+STAT S+ VNG GFF +GLRQG
Sbjct: 897 ISKAFDSVQWQFLLNTLEALNFPETFRHWIKLCISTATFSVQVNGELAGFFGSSRGLRQG 956
Query: 641 DPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQID 700
LSP+LF +C+N LS ++++ N + L HL FADD ++F + + I+
Sbjct: 957 CALSPYLFVICMNVLSHMIDEAAVHRNIGYHPKCEKIGLTHLCFADDLMVFVDGHQWSIE 1016
Query: 701 LLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGN 760
+ N F SGL+++L KSTI V + + + ++ GQLP+ YLG PL
Sbjct: 1017 GVINVFKEFAGRSGLQISLEKSTIYLAGVSASDRVQTLSSFPFANGQLPVRYLGLPLLTK 1076
Query: 761 PRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEV 820
+ + P+++ ++ K +W ++ +S +GRL L+ + + S+ F M+ ++ P + E+
Sbjct: 1077 QMTTADYSPLIEAVKTKISSWTARSLSYAGRLALLNSVIVSIANFWMSAYRLPAGCIREI 1136
Query: 821 EKILRSFLQGG 831
EK+ +FL G
Sbjct: 1137 EKLCSAFLWSG 1147
>At1g31100 hypothetical protein
Length = 1090
Score = 301 bits (772), Expect = 1e-81
Identities = 222/740 (30%), Positives = 359/740 (48%), Gaps = 53/740 (7%)
Query: 117 ERYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGER-------IGIQMGVESSFSDFVAD 169
E ++ LLS +LS N IM GDFN +L E + +M V F D + +
Sbjct: 31 ELWEELLLLSVSLS-GNGKPWIMLGDFNQVLCPAEHSQATSLNVNRRMKV---FRDCLFE 86
Query: 170 SNLIDLPLQSDKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPII 228
+ L DL + + FTW++ A + ++DR LV+E ++F SD
Sbjct: 87 AELCDLVFKGNTFTWWNKSATRPVAKKLDRILVNESWCSRFPSAYAVFGEPDFSDHASCG 146
Query: 229 LSLGAINFGPK-PFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKE 287
+ + + K PF FYN L + +F LV + W S VG S F + +KLK LKN I+
Sbjct: 147 VIINPLMHREKRPFRFYNFLLQNPDFISLVGELWYSINVVGSSMFKMSKKLKALKNPIRT 206
Query: 288 W--QGYASSVSRVRI--NTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEY 343
+ + +++ RV+ N + T+ +E+ R ++ + EE
Sbjct: 207 FSMENFSNLEKRVKEAHNLVLYRQNKTLSDPTIPNAALEMEAQRKWLIL-----VKAEES 261
Query: 344 RWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVV-GGVSIEDPNMIKDAIFSHFQ 402
+ Q++RV W+ EGD NTS+FH ++ RK+ I ++ GV I+ IK+ +F
Sbjct: 262 FFCQRSRVTWMGEGDSNTSYFHRMADSRKAVNTIHIIIDDNGVKIDTQLGIKEHCIEYFS 321
Query: 403 SFFTSE----NRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGF 458
+ E ++ R S K L FS +I S N+ G DGF
Sbjct: 322 NLLGGEVGPPMLIQEDFDLLLPFRCSHDQKKELAMSFSRQDIKSAFFSFPSNKTSGPDGF 381
Query: 459 NLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIG 518
+ FFK+ W ++ +V +F ++ L+K NAT + LIPK S ++DFRPIS
Sbjct: 382 PVEFFKETWSVIGTEVTDAVSEFFTSSVLLKQWNATTLVLIPKITNASKMNDFRPISCND 441
Query: 519 ----SAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQ-GG 573
+ YK+I+++L +RLQ + +IS Q AF GR +++ +L+A E+V N++
Sbjct: 442 FGPITLYKVIARLLTNRLQCLLSQVISPFQSAFLPGRFLAENVLLATELVQGYNRQNIDP 501
Query: 574 GFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEM 633
+LK+D K +D+I W F+ LK + DR++ WI C+ST T S+ VNG++ GFF+
Sbjct: 502 RGMLKVDLRKAFDSIRWDFIISALKAIGIPDRFVYWITQCISTPTFSVCVNGNTGGFFKS 561
Query: 634 EKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCE 693
+GLRQG+PLSPFLF L + S LLN L+++HL FADD ++F +
Sbjct: 562 TRGLRQGNPLSPFLFVLAMEVFSSLLNSRFQAGYIHYHPKTSPLSISHLMFADDIMVFFD 621
Query: 694 NDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYL 753
+ + + L F F SGL +N K+ + +D E +A
Sbjct: 622 GGSSSLHGISEALEDFAFWSGLVLNREKTHLYLAGLDRIEASTIA--------------- 666
Query: 754 GAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAP 813
R + + P+L+KL ++ R+W+ K +S +GR+ L+ + + + F ++ F P
Sbjct: 667 ------RKLRIAEYGPLLEKLAKRFRSWSVKCLSFAGRVQLIASVISGIINFWISTFILP 720
Query: 814 LKVVGEVEKILRSFLQGGKL 833
V +E + FL G +
Sbjct: 721 KGCVKRIEALCARFLWSGNI 740
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 290 bits (743), Expect = 3e-78
Identities = 238/823 (28%), Positives = 395/823 (47%), Gaps = 53/823 (6%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFA-NAVKMDYVSVDAV 64
WN+RGL ++R V ++ + E+ + +V A +MD S
Sbjct: 6 WNIRGLNSRNRQRVVRSWIASNNLLVGCFLETHVAQENANSVLASTLPGWRMD--SNYCC 63
Query: 65 GSAGGLISLWNPA-SYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFN 123
G + +W+P+ S + + F + + +L+ S + VYG N + +R +
Sbjct: 64 SELGRIWIVWDPSISVLVFKRTDQIMFCSIKIPSLLQS--FAVAFVYGRNSELDRRSLWE 121
Query: 124 --LLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVES-----SFSDFVADSNLIDLP 176
L+ S + ++ GDFN I E I + + + DS L DLP
Sbjct: 122 DILVLSRTSPLSVTPWLLLGDFNQIAAASEHYSINQSLLNLRGMEDLQCCLRDSQLSDLP 181
Query: 177 LQSDKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAIN 235
+ FTW + + + ++DR L + + F + G SD P I+ + N
Sbjct: 182 SRGVFFTWSNHQQDNPILRKLDRALANGEWFAVFPSALAVFDPPGDSDHAPCIILID--N 239
Query: 236 FGP---KPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYA 292
P K F +++ + +S W T VG F L+Q LK K +
Sbjct: 240 QPPPSKKSFKYFSFLSSHPSYLAALSTAWEENTLVGSHMFSLRQHLKVAKLCCR------ 293
Query: 293 SSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKE--------EYR 344
+++R+R + I+ ++ RL E+ +EL + L RK+ E
Sbjct: 294 -TLNRLRFSNIQQRTAQSLTRL--EDIQVELLTSPSDTLFRREHVARKQWIFFAAALESF 350
Query: 345 WRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQS 403
+RQK+R+RWL EGD NT FFH ++ LI L G +E+ + IK + +++
Sbjct: 351 FRQKSRIRWLHEGDANTRFFHRAVIAHQATNLIKFLRGDDGFRVENVDQIKGMLIAYYSH 410
Query: 404 FF--TSENRLRP----KIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDG 457
SEN + P KI+ R L SE EI +VL S N+APG DG
Sbjct: 411 LLGIPSEN-VTPFSVEKIKGLLPFRCDSFLASQLTTIPSEEEITQVLFSMPRNKAPGPDG 469
Query: 458 FNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLI 517
F + FF + W IVK VV +F +G L +G NAT I+LIPK ++ FRP++
Sbjct: 470 FPVEFFIEAWAIVKSSVVAAIREFFISGNLPRGFNATAITLIPKVTGADRLTQFRPVACC 529
Query: 518 GSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVD-FLNKKQGGGFL 576
+ YK+I+++++ RL++ + NQ F +GR + + +L+A+E+VD F +
Sbjct: 530 TTIYKVITRIISRRLKLFIDQAVQANQVGFIKGRLLCENVLLASELVDNFEADGETTRGC 589
Query: 577 LKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKG 636
L++D +K YDN++W FL +ILK ++ +I+WI C+S+A+ SI NG GFF+ +KG
Sbjct: 590 LQVDISKAYDNVNWEFLINILKALDLPLVFIHWIWVCISSASYSIAFNGELIGFFQGKKG 649
Query: 637 LRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGV-RIGPGL---ALNHLQFADDTLLFC 692
+RQGDP+S LF L ++ +L++ LD +G+ + P + HL FADD L+F
Sbjct: 650 IRQGDPMSSHLFVLVMD----VLSKSLDLGALNGLFNLHPNCLAPIITHLSFADDVLVFS 705
Query: 693 ENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILY 752
+ + I + L F SGL +N K+ ++ + + +A G + G LP+ Y
Sbjct: 706 DGAASSIAGILTILDDFRQGSGLGINREKTELLLDGGNFARNRSLADNLGITHGSLPVRY 765
Query: 753 LGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLM 795
LG PL R ++P++D++ + +W ++ +S +GRL L+
Sbjct: 766 LGVPLMSQKMRRQDYQPLVDRINSRFTSWTARHLSFAGRLQLL 808
>At2g11240 pseudogene
Length = 1044
Score = 281 bits (719), Expect = 2e-75
Identities = 190/616 (30%), Positives = 311/616 (49%), Gaps = 31/616 (5%)
Query: 222 SDRRPIILSLGAINFGPKPFLFYNHWLLDK-EFEKLVSDWWCSATTVGWSAFVLQQKLKG 280
SD RP++ K Y+ L + E LV + W T ++++K+
Sbjct: 57 SDHRPLLTHFDLSKKKKKGVFRYDRRLKNNDEVTALVQEAWNLYDTD-----IVEEKISR 111
Query: 281 LKNRIKEWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRK 340
+ I +W S+ IE + Q + ++S++ EL TI L Y+
Sbjct: 112 CRLEIVKWSRAKQQSSQ---KLIEENRQKLEEAMSSQDHNQELLS---TINTNLLLAYKA 165
Query: 341 EEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVV-GGVSIEDPNMIKDAIFS 399
EE W+Q++R WL GD+N+ +FH+I++ R S + GV + I + I
Sbjct: 166 EEEYWKQRSRQLWLALGDKNSGYFHAITRGRTVINKFSVIEKEDGVPEYEEAGILNVISE 225
Query: 400 HFQSFFTSENRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFN 459
+FQ F++ R + + +A K + + + EI S ++APG DGF+
Sbjct: 226 YFQKLFSANEGARA-------ATIKEAIKPFISPEQNPEEIKSACFSIHADKAPGPDGFS 278
Query: 460 LNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGS 519
+FF+ W V ++V + F S+ L +N T I+LIPK + D+RPI+L
Sbjct: 279 ASFFQSNWMTVGPNIVLEIQSFFSSSTLQPTINKTHITLIPKIQSLKRMVDYRPIALCTV 338
Query: 520 AYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNK---KQGGGFL 576
YK+ISK+L+ RLQ + +ISENQ AF R +D +LI +E + +L ++
Sbjct: 339 FYKIISKLLSRRLQPILQEIISENQSAFVPKRASNDNVLITHEALHYLKSLGAEKRCFMA 398
Query: 577 LKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKG 636
+K + +K YD I+W F+ +++EM F WI+WI C++T + S L+NGS+ G E+G
Sbjct: 399 VKTNMSKAYDRIEWDFIKLVMQEMGFHQTWISWILQCITTVSYSFLLNGSAQGAVTPERG 458
Query: 637 LRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPG-LALNHLQFADDTLLFCEND 695
LRQGDPLSPFLF +C LS L + D + G+R+ G +NHL FADDT+ FC +D
Sbjct: 459 LRQGDPLSPFLFIICSEVLSGLCRKAQLDGSLLGLRVSKGNPRVNHLLFADDTIFFCRSD 518
Query: 696 ENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVD----VRETQRVAGMYGWSIGQLPIL 751
L + SG +N KS I E Q++ G+ +G L
Sbjct: 519 LKSCKTFLCILKKYEEASGQMINKSKSAITFSRKTPDHIKTEAQQILGIQ--LVGGLG-K 575
Query: 752 YLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFK 811
YLG P ++ + ++D++R++ +W+S+F+S +G+ T++K+ L S+P + M+ FK
Sbjct: 576 YLGLPKMFGRKKRDLFNQIVDRIRQRSLSWSSRFLSTAGKTTMLKSVLASMPTYTMSCFK 635
Query: 812 APLKVVGEVEKILRSF 827
+ + ++ L F
Sbjct: 636 LLVSLCKRIQSALTHF 651
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 277 bits (708), Expect = 3e-74
Identities = 232/789 (29%), Positives = 378/789 (47%), Gaps = 53/789 (6%)
Query: 60 SVDAVGSAGGLISLW-NPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSER 118
+++ VG +GGL W N KN+ L L+ F+ VYG V R
Sbjct: 21 TIEPVGKSGGLAIFWKNHLEIDFLFEDKNL----LDLKVSQGKKSWFVSCVYGNPVLHLR 76
Query: 119 YDFFNLLSE-ALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLID 174
Y + LS + N+ +C I GDFN IL+ ++G + SSF F + + ++
Sbjct: 77 YLLLDKLSSIGVQRNSAWCMI--GDFNDILSNDGKLGGPSRLISSFQPFKNMLLNCDMHQ 134
Query: 175 LPLQSDKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQ-SVENWGLSDRRPIILSLG 232
+ + FTW +R + ++DR + + F Q +E G S RP++
Sbjct: 135 MGSSGNSFTWGGTRNDQWIQCKLDRCFGNSEWFTMFSNSHQWFLEKLG-SHHRPVL---- 189
Query: 233 AINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVG-WS----AFVLQQKLKGLKNR--I 285
+NF +F + DK F + D C+A+T+ W + V L+ +K R I
Sbjct: 190 -VNFVNDQEVFRGQFCYDKRFAE---DPQCAASTLSSWIGNGISDVSSSMLRMVKCRKAI 245
Query: 286 KEWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRW 345
W+ + ++ RI + ++L + I + + +L + +R+EE W
Sbjct: 246 SGWKKNSDFNAQNRILRLRSELDEEKSKQYPCWSRISVIQTQLGVA------FREEESFW 299
Query: 346 RQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFS-HFQSF 404
R K++ +WL GDRN+ FF ++ K ++K + LV + N K I S +F++
Sbjct: 300 RLKSKDKWLFGGDRNSKFFQAMVKANRTKNSLRFLVDENGNEHTLNREKGNIASVYFENL 359
Query: 405 FTSENRLRPKIRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFF 463
F S + +R+S+ L + +E EI + S + AP
Sbjct: 360 FMSSYPANSQSALDGFKTRVSEEMNQELTQAVTELEIHSAVFSINVESAPEK-------L 412
Query: 464 KQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKL 523
+ G +++ FFE TG L + N T + LIPK P +SD RPISL YK+
Sbjct: 413 ECCQGSDYIEILGFFE----TGVLPQEWNHTHLYLIPKFTNPQRMSDIRPISLCSVLYKI 468
Query: 524 ISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFLL-KLD 580
ISK+L+ +L+ P ++S +Q AF R ISD ILIA+E+V L N K F++ K D
Sbjct: 469 ISKILSFKLKKHLPSIVSPSQSAFFAERLISDNILIAHEIVHSLRTNDKISKEFMVFKTD 528
Query: 581 FAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQG 640
+K YD ++W FL +IL + F D+WI+WI CV++ T S+L+NG G E+G+RQG
Sbjct: 529 MSKAYDRVEWSFLQEILVALGFNDKWISWIMGCVTSVTYSVLINGQHFGHITPERGIRQG 588
Query: 641 DPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRI-GPGLALNHLQFADDTLLFCENDENQI 699
DP+SPFLF LC L +L Q + SG++ G G ++NHL F DDT L C ++
Sbjct: 589 DPISPFLFVLCTEALIHILQQAENSKKVSGIQFNGSGPSVNHLLFVDDTQLVCRATKSDC 648
Query: 700 DLLCNTLLSFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLG 758
+ + L + SG +N+ KS+I G VD + + G + YLG P
Sbjct: 649 EQMMLCLSQYGHISGQLINVEKSSITFGVKVDEDTKRWIKNRSGIHLEGGTGKYLGLPEN 708
Query: 759 GNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVG 818
+ + + + +KL+ W K +S G+ L+K+ ++P+++M F+ P +
Sbjct: 709 LSGSKQDLFGYIKEKLQSHLSGWYDKTLSQGGKEILLKSIALALPVYIMTCFRLPKGLCT 768
Query: 819 EVEKILRSF 827
++ ++ F
Sbjct: 769 KLTSVMMDF 777
>At1g47850 hypothetical protein
Length = 799
Score = 275 bits (703), Expect = 1e-73
Identities = 145/411 (35%), Positives = 236/411 (57%), Gaps = 1/411 (0%)
Query: 422 RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDF 481
R S ++ L ++ + E +VL + N+ PG DG+ FFK W I +D + + F
Sbjct: 13 RCSATDQDMLTREVTSEENQKVLFAMPSNKFPGPDGYTSEFFKATWSITGQDFIAAIKSF 72
Query: 482 HSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLIS 541
G L KGLNAT ++LIPK + + + D+RPIS YK+ISK++A+RL+V+ P I
Sbjct: 73 FIKGFLPKGLNATILALIPKKDEATLMRDYRPISCCNVIYKVISKIIANRLKVMLPTFIL 132
Query: 542 ENQFAFAQGRQISDCILIANEVV-DFLNKKQGGGFLLKLDFAKDYDNIDWGFLFDILKEM 600
+NQ AF + R + + +L+A E+V D+ +K+D +K +D++ W FL + L+ +
Sbjct: 133 QNQSAFVRERLLIENVLLATELVKDYHKDSISPRCAMKIDISKAFDSVQWQFLLNTLEAL 192
Query: 601 NFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLN 660
NF + + +WI+ C+STAT S+ VNG GFF ++GLRQG LSP+LF +C+N LS +++
Sbjct: 193 NFPENFCHWIKLCISTATFSVQVNGELAGFFGSKRGLRQGCALSPYLFVICMNVLSHMID 252
Query: 661 QLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLH 720
N L+L HL FADD ++F + + ++ + N F SGL ++L
Sbjct: 253 VAAVHRNIGYHPKCKKLSLTHLCFADDLMVFIDGQQRSVEGVINIFKEFAGKSGLHISLE 312
Query: 721 KSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRT 780
KST+ V + + ++ GQLP+ YLG PL + + P+LDK+R K +
Sbjct: 313 KSTLYLAGVSELNRNNILSAFPFASGQLPVRYLGLPLLTKQMTTADYSPLLDKVRSKISS 372
Query: 781 WNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGG 831
W ++ +S +GRL L+ + + S+ F M+ ++ P + E+EK+ +FL G
Sbjct: 373 WTARSLSYAGRLALINSVIVSLSNFWMSAYRLPAGCIKEIEKLCSAFLWSG 423
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.147 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,825,265
Number of Sequences: 26719
Number of extensions: 1262011
Number of successful extensions: 4553
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 4199
Number of HSP's gapped (non-prelim): 130
length of query: 1389
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1277
effective length of database: 8,326,068
effective search space: 10632388836
effective search space used: 10632388836
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0110a.13


