

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.10
(1408 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
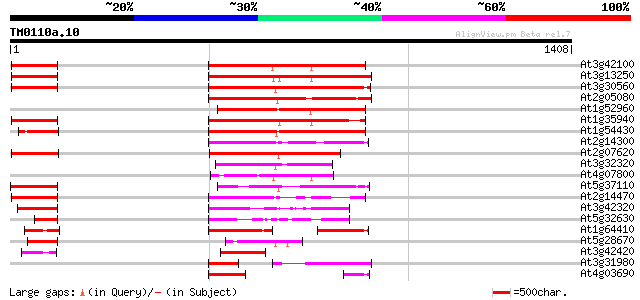
Score E
Sequences producing significant alignments: (bits) Value
At3g42100 putative protein 379 e-105
At3g13250 hypothetical protein 377 e-104
At3g30560 hypothetical protein 371 e-102
At2g05080 putative helicase 365 e-100
At1g52960 hypothetical protein 358 1e-98
At1g35940 hypothetical protein 351 2e-96
At1g54430 hypothetical protein 346 6e-95
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 323 3e-88
At2g07620 putative helicase 300 4e-81
At3g32320 hypothetical protein 243 4e-64
At4g07800 hypothetical protein 234 2e-61
At5g37110 putative helicase 233 6e-61
At2g14470 pseudogene 230 4e-60
At3g42320 putative protein 203 5e-52
At5g32630 putative protein 172 1e-42
At1g64410 unknown protein 149 1e-35
At5g28670 putative protein 125 2e-28
At3g42420 putative protein 121 3e-27
At3g31980 hypothetical protein 117 6e-26
At4g03690 hypothetical protein 107 6e-23
>At3g42100 putative protein
Length = 1752
Score = 379 bits (974), Expect = e-105
Identities = 189/412 (45%), Positives = 262/412 (62%), Gaps = 18/412 (4%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
M+T+EFQKRGLPHAHILL++ A+ KL + ID +I AE+PD P L+E + N M+HG
Sbjct: 778 MHTVEFQKRGLPHAHILLFMDAKSKLPTADDIDKIISAEIPDKDKEPELYEVIKNSMIHG 837
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM G+C+K +PKK+ D+T DGYP+YRRR T + G DNG+V
Sbjct: 838 PCGAANMNSPCMVEGKCSKQYPKKHQDITKVGKDGYPIYRRRMTEDYIEKGGFKCDNGYV 897
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRV------------TLSMSSGR 666
VPYN KL ++YQAHIN+E+CN+S SIKYLFKYINKG DRV T + +SG
Sbjct: 898 VPYNKKLSLRYQAHINVEWCNQSGSIKYLFKYINKGADRVVFIVEPVNQDKTTENATSGE 957
Query: 667 TTQDKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKEN 726
+ DEIK +FDCRY+S EAVWRI+++ + + VQ+L++H KQ V K +
Sbjct: 958 PPNSTEKKKDEIKDWFDCRYVSASEAVWRIYKFPLQDRSTAVQRLSFHDEGKQPVYAKPD 1017
Query: 727 EAIEDVLRRNIVKNSMFLAWMDANCKYVHG------RQLTYVEFPELFVYDPKSKSWHPR 780
IEDVL R ++SMF+AW+ N G R+L Y + P F +D K+K W R
Sbjct: 1018 ADIEDVLERISNEDSMFMAWLTLNKNNDVGKNGKRARELLYSQIPAYFTWDGKNKQWVKR 1077
Query: 781 KQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLK 840
+G S+GR+++V YYLR+LLN+ +G Y+D+++ N V+ +F+ AC A G+L
Sbjct: 1078 IRGFSLGRINYVCRKMEVEYYLRVLLNIVKGPMSYDDIKTFNGVVYPSFKEACFARGILD 1137
Query: 841 DDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
DD+ +IDG+ + + G Y R F LL +S+ P HVW+ETWH+LA+ I
Sbjct: 1138 DDQVYIDGLHEASQFCFGDYLRNFFAMLLLSDSLARPEHVWSETWHLLAEDI 1189
Score = 132 bits (331), Expect = 2e-30
Identities = 62/116 (53%), Positives = 80/116 (68%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K Q++ SP F++CC KG + LP L D P L++NLL D SR+F +NIR YN +FA TS
Sbjct: 340 KKQQNKSPTFTLCCGKGNVKLPLLKDSPALINNLLTGDDALSRNFRENIRIYNMIFAMTS 399
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
+GG+VD + G+GP F + G NYH IGSL P GD K++QLYI DT+NEV NR
Sbjct: 400 LGGRVDNSMPKGKGPNMFRLQGGNYHLIGSLKPNPGDYAKYSQLYIVDTENEVDNR 455
>At3g13250 hypothetical protein
Length = 1419
Score = 377 bits (969), Expect = e-104
Identities = 189/427 (44%), Positives = 269/427 (62%), Gaps = 18/427 (4%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHILL+++A KL + ID +I AE+P+ P L+E + N M+HG
Sbjct: 446 MYTVEFQKRGLPHAHILLFMAANSKLPTADDIDKIISAEIPNKDKEPELYEVIKNSMMHG 505
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM +G+C+K +PKK+ ++T +DGYP+YRRR T + GV DN +V
Sbjct: 506 PCGSANTSSPCMVDGQCSKLYPKKHQEITKVGADGYPIYRRRLTDDYIEKGGVKCDNRYV 565
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTL------SMSSGRTTQDKP 672
VPYN KL ++YQAHIN+E+CN++ SIKYLFKYINKG DRV +S TT
Sbjct: 566 VPYNKKLSLRYQAHINVEWCNQNGSIKYLFKYINKGPDRVVFIVEPIKEATSSDTTAPVV 625
Query: 673 VED------DEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKEN 726
D DEIK +FDCRY+S EA+WRIF++ I H+ PVQKL++H KQ F
Sbjct: 626 ESDTTEKKKDEIKDWFDCRYVSASEAIWRIFKFPIQHRSTPVQKLSFHDKGKQPAYFDAK 685
Query: 727 EAIEDVLRRNIVKNSMFLAWMDANCKYVHG------RQLTYVEFPELFVYDPKSKSWHPR 780
+ DVL R ++S FLAW+ N K G R Y E P F +D ++K + R
Sbjct: 686 AKMADVLERVSNEDSQFLAWLTLNRKNAVGKNGKRARDCLYAEIPAYFTWDGENKQFKKR 745
Query: 781 KQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLK 840
+G S+GR+++V + YYLR+LLN+ RG Y+D+++VN V+ +++ AC A G+L
Sbjct: 746 TRGFSLGRINYVSRKMEDEYYLRVLLNIVRGPQSYDDIKTVNGVVYPSYKLACFARGILD 805
Query: 841 DDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKH 900
DD+ +I+G+++ + G Y R F LL +S+ P HVW+ETWH+L++ I + +
Sbjct: 806 DDQVYINGLIEASQFCFGDYLRNFFSMMLLSDSLARPEHVWSETWHLLSEDILIKKRDEF 865
Query: 901 NNRGMCL 907
N+ + L
Sbjct: 866 KNQELTL 872
Score = 133 bits (334), Expect = 8e-31
Identities = 61/116 (52%), Positives = 80/116 (68%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K S +P FS+CC +G + LP+L + PEL+ LL+ D SRH+ IR YN +FA TS
Sbjct: 9 KKSNSENPKFSLCCGQGSVKLPFLKESPELIKKLLKGNDALSRHYRQFIRIYNMIFAMTS 68
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
+GGKVD + GRGP F + G NYH+IGSL P +GD K++QLYI DT+NEV+NR
Sbjct: 69 LGGKVDKSMPKGRGPAMFRLQGGNYHQIGSLKPKDGDYAKYSQLYIVDTENEVENR 124
>At3g30560 hypothetical protein
Length = 1473
Score = 371 bits (952), Expect = e-102
Identities = 182/419 (43%), Positives = 261/419 (61%), Gaps = 16/419 (3%)
Query: 500 YTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGP 559
YTIEFQKRGLPHAHI++W+ YK +D +I AE+PD + P L++ VS M+HGP
Sbjct: 554 YTIEFQKRGLPHAHIIVWMDPRYKFPIADHVDKIIFAEIPDKENDPKLYQAVSECMIHGP 613
Query: 560 CGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVV 619
C L SPCM+NG+C+K++PK + + T D++GYP+YRRR+TG + DN +VV
Sbjct: 614 CRLVNPNSPCMENGKCSKYYPKNHVENTFLDNEGYPIYRRRDTGRFIKKNKYPCDNRYVV 673
Query: 620 PYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGR-----------TT 668
PYN LL KY+AHIN+E+CN+S S+KYLFKY+NKG DRVT+S+ R T
Sbjct: 674 PYNDFLLRKYRAHINVEWCNQSVSVKYLFKYVNKGPDRVTVSLEPHRKEVVSEENNVGET 733
Query: 669 QDKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEA 728
+ P E ++++ YFDCRY+S CEA+WRI Y IH++ V KLT+H KQ + KE E
Sbjct: 734 NNDPQEQNQVEDYFDCRYVSACEAMWRIKGYPIHYRQTLVTKLTFHEKGKQPIYVKEGET 793
Query: 729 IEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQ-GESIG 787
E VL R + F AW + N + +L Y + P + ++ K K++ RK G +G
Sbjct: 794 AESVLYRVNDDETQFTAWFELNKRDPEAAKLLYEQIPNFYTWNGKDKNFRRRKMPGFVVG 853
Query: 788 RMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFID 847
R++ VPP + Y+LR+L+N RG ++D++++ VH T+R AC ALGLL DD+++I+
Sbjct: 854 RINHVPPKIDDAYHLRILINNIRGPKGFDDIKTIEGVVHKTYRDACYALGLLDDDKDYIN 913
Query: 848 GILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNNRGMC 906
GI + Y R LFV L+ S+ +P VW TW +L++ D QRK ++ C
Sbjct: 914 GIEEANFWCFHKYVRKLFVIMLIFESLSSPAVVWEHTWKILSE----DFQRKVRDKLKC 968
Score = 112 bits (281), Expect = 1e-24
Identities = 53/115 (46%), Positives = 74/115 (64%)
Query: 6 QKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTSIG 65
++ S S CCM+G+I+LP L + PE + L + +++F N+R YN +F+FTS+G
Sbjct: 118 KEHGSSTTSRCCMQGQIVLPMLKESPEYMWWLFTSDHPDAKNFRANVRPYNMLFSFTSLG 177
Query: 66 GKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRM 120
GKVD + GRGP F + G+NYH I +L P GD KF QLY+ DT NEV NR+
Sbjct: 178 GKVDRSVKKGRGPSMFALQGENYHLIDALKPKPGDYAKFQQLYVMDTDNEVDNRI 232
>At2g05080 putative helicase
Length = 1219
Score = 365 bits (937), Expect = e-100
Identities = 180/424 (42%), Positives = 267/424 (62%), Gaps = 28/424 (6%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYTIEFQKRGLPHAHIL+WL ++ KL ID I AE+PD P LFE + MVHG
Sbjct: 264 MYTIEFQKRGLPHAHILIWLDSKCKLTRAEHIDKAISAEIPDKLKDPELFEVIKEMMVHG 323
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG+ + PCM+NG+C+KF+PK + T D +G+P+YRRR ++ DN +V
Sbjct: 324 PCGVVNPKCPCMENGKCSKFYPKDHVPKTIIDKEGFPIYRRRRIDDFVQKKDFKCDNRYV 383
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDK------- 671
+PYN L ++Y+AHIN+E+CN+S S+KY+FKYI+KG DRVT+ + S +++K
Sbjct: 384 IPYNRSLSLRYRAHINVEWCNQSGSVKYIFKYIHKGPDRVTVVVGSSLNSKNKEKGKQKV 443
Query: 672 --------PVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLF 723
P + +E++ +F+CRY+S CEA WRI +Y IH++ V KL++HLP +Q + F
Sbjct: 444 NADTDGSEPKKKNEVEDFFNCRYVSACEAAWRILKYPIHYRSTSVMKLSFHLPGEQYIYF 503
Query: 724 KENEAIEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQG 783
K +E +E VL + + S+ +A R+LTY P F YDPK K ++ RK+G
Sbjct: 504 KGDEEVETVLNKADLDGSIQIA-----------RKLTYPNIPTRFTYDPKEKKFNLRKKG 552
Query: 784 ESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDR 843
+IGR+++VP + YYLR+LLNV G +E+L++VN ++ ++ ACEALGLL +D+
Sbjct: 553 FAIGRINYVPRDIEDGYYLRILLNVVPGPRSFEELKTVNGVLYKEWKDACEALGLLDNDQ 612
Query: 844 EFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNNR 903
E+ID + + + G Y R LFV L +++ +P +VW TW L++ I + ++ N
Sbjct: 613 EYIDDLKRTSFWSSGWYLRQLFVIML--DALISPENVWAATWQHLSEDIQNEKKKYFNRP 670
Query: 904 GMCL 907
CL
Sbjct: 671 VTCL 674
>At1g52960 hypothetical protein
Length = 924
Score = 358 bits (919), Expect = 1e-98
Identities = 176/386 (45%), Positives = 243/386 (62%), Gaps = 17/386 (4%)
Query: 523 KLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPCGLSKRESPCMKNGRCAKFFPKK 582
KL++ D +I AE+PD K P L+ V + M+HGPCG+ SPCM+NG+C K+FPK
Sbjct: 6 KLSTAEDTDKIITAEIPDKKKKPGLYAVVKDCMIHGPCGVGHPNSPCMENGKCKKYFPKS 65
Query: 583 YADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSN 642
Y+D T D+DG+PVYRRR+TG+ + G DN +V+PYN K+ ++YQAHIN+E CN+S
Sbjct: 66 YSDTTKVDNDGFPVYRRRDTGIYVEKNGFQCDNRYVIPYNEKVSLRYQAHINVELCNQSG 125
Query: 643 SIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIH 702
SIKYLFKY++KG DRVT+++ K E DE+K YFDCRY+S CEA+WRIF++ IH
Sbjct: 126 SIKYLFKYVHKGHDRVTVTVEPNDQDTAKK-EKDEVKDYFDCRYVSACEAMWRIFKFPIH 184
Query: 703 HKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIVKNSMFLAWMDANCK---------- 752
++ PV KL +H KQ V +K E E V+ R + + FLAW N K
Sbjct: 185 YRTTPVVKLFFHEEGKQPVYYKPGETTESVMDRLSSEATQFLAWFQLNKKPPSRTIRANA 244
Query: 753 ------YVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGELYYLRMLL 806
+L + E P F ++ K K + R++G +IGR++FVP + YYLR+LL
Sbjct: 245 KKLPKAAPDPTKLLFEEIPNHFTWNSKEKKFMIRERGFAIGRINFVPRTIEDAYYLRILL 304
Query: 807 NVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGGAYTRALFV 866
N++RG T Y+DL++V VH +FR A ALGLL DD+E+I+GI D Y R LFV
Sbjct: 305 NIKRGVTSYKDLKTVKGVVHKSFRDAVFALGLLDDDKEYINGIKDAKFWCSAKYVRRLFV 364
Query: 867 SFLLPNSMCNPLHVWNETWHVLADGI 892
LL S+ P VW+ETW +L++ I
Sbjct: 365 IMLLSESLTKPEMVWDETWRILSEDI 390
>At1g35940 hypothetical protein
Length = 1678
Score = 351 bits (900), Expect = 2e-96
Identities = 180/412 (43%), Positives = 252/412 (60%), Gaps = 43/412 (10%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHILL++ A+ KL + ID +I AE+PD + P L+E + N M+HG
Sbjct: 788 MYTVEFQKRGLPHAHILLFMHAKSKLPTADDIDKLISAEIPDKEKEPDLYEVIKNSMIHG 847
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM +G C+K +PKK+ D+T SDGYP+YRRR T + G+ DN +V
Sbjct: 848 PCGSANVNSPCMVDGECSKLYPKKHQDITKIGSDGYPIYRRRKTDDYVEKGGIKCDNRYV 907
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTT----QDKPVE 674
VPYN KL ++Y AHIN+E+CN++ SIKYLFKYINKG D+V + + T + P +
Sbjct: 908 VPYNKKLSLRYNAHINVEWCNQNGSIKYLFKYINKGPDKVVFIVEPTQQTTAGDSETPQQ 967
Query: 675 D--------DEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKEN 726
+ +EIK +FDCRY+S EAVWRIF+Y I H PVQKL++H+ KQ F
Sbjct: 968 EQGSAEKKKNEIKDWFDCRYVSASEAVWRIFKYPIQHISTPVQKLSFHVEGKQPAYFDPK 1027
Query: 727 EAIEDVLRRNIVKNSMFLAWMDANCKYVHG------RQLTYVEFPELFVYDPKSKSWHPR 780
IEDVL R +S F+AW+ N + G R+ Y E P F +D ++KS+ R
Sbjct: 1028 SNIEDVLERVANVDSQFMAWLTLNRRNAVGKNGKRARECLYAEIPAYFTWDGENKSFKKR 1087
Query: 781 KQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLK 840
+G SIGR+ +V + Y+LR+LLN+ RG T Y ++++ + V+ TF+ AC A G+L
Sbjct: 1088 TRGFSIGRIHYVSRKMEDEYFLRVLLNIVRGPTSYAEIKTYDGVVYKTFKEACFARGILD 1147
Query: 841 DDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
DD+ FIDG+++ HV ++TWH+LA+ I
Sbjct: 1148 DDQVFIDGLVEAT-------------------------HVRSQTWHLLAEDI 1174
Score = 120 bits (301), Expect = 6e-27
Identities = 57/116 (49%), Positives = 75/116 (64%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K + + F++CC +G + LP+L + P LL NLL S+H+ DN R +N +FA TS
Sbjct: 350 KKETNKESVFTLCCGEGSVKLPFLKESPHLLKNLLSGNHPLSKHYRDNARIFNMVFAMTS 409
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
GGKVD + GRGP F + G NYH IGSL T GD K++QLYI DT+NEV+NR
Sbjct: 410 FGGKVDKSMPKGRGPAMFRLQGGNYHLIGSLKLTPGDYAKYSQLYIIDTENEVENR 465
>At1g54430 hypothetical protein
Length = 1639
Score = 346 bits (887), Expect = 6e-95
Identities = 175/403 (43%), Positives = 251/403 (61%), Gaps = 11/403 (2%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YTIEFQKRGLPHAHILLWL + K + ID I AE+P P V +M+HG
Sbjct: 699 VYTIEFQKRGLPHAHILLWLQGDLKKPTPNDIDKYISAEIPVKDKDPEGHTLVEQHMMHG 758
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM+ G C+K FP+++ + T + G+ +YRRRN + LDN FV
Sbjct: 759 PCGKDRPSSPCMEKGICSKKFPREFVNHTKMNESGFILYRRRNDQRYVLKGQTRLDNRFV 818
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTT---------Q 669
VP+N ++L KY+AHIN+E+CNKS++IKYLFKYI KGVD+ T + G + +
Sbjct: 819 VPHNLEILKKYKAHINVEWCNKSSAIKYLFKYITKGVDKATFIIQKGNSVNGQGSGNGFE 878
Query: 670 DKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAI 729
+KP +EI +Y DCRYLS CEA+WRIF ++IHH PPVQ+L HLP +Q +F+E E +
Sbjct: 879 EKP--RNEINEYLDCRYLSACEAMWRIFMFNIHHHNPPVQRLPLHLPGEQSTIFEEEENL 936
Query: 730 EDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRM 789
E+V R + +M + + N R+L YV+ P +FV+D +K + RKQ E+IGR+
Sbjct: 937 ENVEYRYGHERTMLTEYFELNKICEDARKLKYVQVPTMFVWDSTNKMYTRRKQRENIGRI 996
Query: 790 SFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGI 849
+ P G+LYYLR+LLN +G T ++ L++V VH++F+ AC GLL D+E+ D +
Sbjct: 997 VNILPTAGDLYYLRILLNKVKGATSFDYLKTVGGVVHESFKAACHTRGLLDGDKEWHDAM 1056
Query: 850 LDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
+ A + R+LFV L+ + PL +W+ W +AD +
Sbjct: 1057 DEAAQWSTSYLLRSLFVLILIYCEVSEPLKLWSHCWESMADDV 1099
Score = 95.9 bits (237), Expect = 1e-19
Identities = 50/99 (50%), Positives = 65/99 (65%), Gaps = 4/99 (4%)
Query: 23 MLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGPPQFV 82
+LP P++L LL T S H+ NIR+YNS+ AFTS+G ++D + GP F
Sbjct: 319 VLPPEPQPPQMLKKLL----TESPHYQRNIRTYNSILAFTSMGAQIDKNVMHKGGPFTFR 374
Query: 83 ISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMS 121
I GQN+H++GSLVP EG PK QLYI+DT NEV NR+S
Sbjct: 375 IHGQNHHKLGSLVPEEGKPPKILQLYIFDTANEVHNRIS 413
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 323 bits (829), Expect = 3e-88
Identities = 166/401 (41%), Positives = 235/401 (58%), Gaps = 37/401 (9%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHI++W+ YK ++ +D +I AE+PD + +P L++ VS M+HG
Sbjct: 405 MYTVEFQKRGLPHAHIIVWMDPRYKFHTADHVDKIIFAEIPDKEKHPELYQAVSECMIHG 464
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PC L SPCM+NG+C+K++PK + + T+ D GYP+YRRR++G + DN +V
Sbjct: 465 PCRLVNPNSPCMENGKCSKYYPKNHVENTSLDYKGYPIYRRRDSGRFIEKNKYQCDNWYV 524
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
VPYN LL KY+AHIN+E+CN+S SIKYLFKY+NKG DRVT + + G D P E +++
Sbjct: 525 VPYNDVLLRKYRAHINVEWCNQSVSIKYLFKYVNKGPDRVTQN-NVGEINND-PQERNQV 582
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
+ YFDCR Y IH++ V KLT+H KQ V KE E E VL R
Sbjct: 583 QDYFDCR------------GYPIHYRQTSVTKLTFHEKGKQSVYVKEGETAESVLYRVNN 630
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
+ F+AW + N + +L Y + P + ++ VPP +
Sbjct: 631 DETQFIAWFELNKRDPEAAKLLYEQIPNFYT-------------------INHVPPKIDD 671
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGG 858
Y+LR+L+N R ++D+++V VH T+R AC ALGLL DD+E+I GI +
Sbjct: 672 AYHLRILINNIRAPKGFDDIKTVEGVVHKTYRDACYALGLLDDDKEYIHGIEEANFWCSP 731
Query: 859 AYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRK 899
Y R FV L+ S+ +P+ VW TW +L + D QRK
Sbjct: 732 KYVRKSFVIMLISESLSSPVVVWEHTWKILFE----DFQRK 768
Score = 37.0 bits (84), Expect = 0.080
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLL 38
+ +KSA+P ++ CCM+G+I+LP L + PE + LL
Sbjct: 83 RRKKSANPVYTGCCMQGQIVLPMLKESPEYMWWLL 117
>At2g07620 putative helicase
Length = 1241
Score = 300 bits (768), Expect = 4e-81
Identities = 145/342 (42%), Positives = 214/342 (62%), Gaps = 13/342 (3%)
Query: 501 TIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPC 560
T+ FQKRGLPHAHILL++ K + ID+++ AE+PD P L+E V++ M+HGPC
Sbjct: 406 TVMFQKRGLPHAHILLFMDKSCKFPTSDDIDNILSAEIPDKAKDPKLYEVVNDCMIHGPC 465
Query: 561 GLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVP 620
G + +ESPC+ +G+C+KFFPKK + TT DGYP+YRRR + + G+ DN +VVP
Sbjct: 466 GAANKESPCIVDGKCSKFFPKKLVEQTTVGKDGYPIYRRRESEHFVEKGGIKCDNTYVVP 525
Query: 621 YNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPV------- 673
YN L ++Y+AHIN+E+C +S SIKYLFKYINKG DRV + + T + +
Sbjct: 526 YNRMLSLRYRAHINVEWCKQSGSIKYLFKYINKGQDRVAIVVEPKDKTSNMVLFSGSQKL 585
Query: 674 ------EDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENE 727
+D EIK YFDCRY+S EAVWRIF++ I ++ PV KL+YHLP KQ + F++ +
Sbjct: 586 LVAVIDDDKEIKDYFDCRYVSASEAVWRIFKFPIQYRTTPVMKLSYHLPGKQPLCFEDTQ 645
Query: 728 AIEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIG 787
I+++ + ++ MF+ ++ N + RQ Y E P F +D ++K W R++G IG
Sbjct: 646 NIDELSEKKANEDFMFIGFLKLNQECEFARQFIYTEIPPYFTWDGQNKQWKLRERGFYIG 705
Query: 788 RMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTF 829
RM++ YY+++LL + G ED+R+ + V F
Sbjct: 706 RMNYASIKMEPEYYMKILLGIVCGPKSDEDIRTYKDVVRRKF 747
Score = 137 bits (346), Expect = 3e-32
Identities = 63/119 (52%), Positives = 84/119 (69%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K +KS P FS+CC++G + LP+L + PEL+ LL D RHF +NIR+YN +F+ TS
Sbjct: 10 KRRKSKKPKFSLCCLQGSVKLPFLTESPELIRELLSCDDALRRHFRENIRAYNMLFSMTS 69
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMSH 122
+GGKVD G+GP QF + G NYH IGS++P EGD KF+QLYI DT NEV+NR ++
Sbjct: 70 LGGKVDRSNPQGKGPNQFQLHGANYHLIGSMLPGEGDYAKFSQLYIVDTGNEVENRSTN 128
>At3g32320 hypothetical protein
Length = 494
Score = 243 bits (621), Expect = 4e-64
Identities = 124/306 (40%), Positives = 180/306 (58%), Gaps = 22/306 (7%)
Query: 517 WLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPCGLSKRESPCMKNGRCA 576
+L A YK + +D +I AE+PD + P ++ VS ++HGPCGL SPCMKNG+C+
Sbjct: 163 YLDAIYKFPTADHVDMIIFAEIPDKEKDPEQYQAVSECIIHGPCGLVNPNSPCMKNGKCS 222
Query: 577 KFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVPYNPKLLMKYQAHINIE 636
K++PK + + T+ D++GYP+YRRR+TG + DN +VVPYN LL KY+AHIN+E
Sbjct: 223 KYYPKNHVENTSLDNEGYPIYRRRDTGRFIKKNKYQCDNWYVVPYNDVLLRKYKAHINVE 282
Query: 637 YCNKSNSIKYLFKYINKGVDRVTLSMSSGR-----------TTQDKPVEDDEIKQYFDCR 685
+CN+S S+KYLFKY+NKG DRVT+S+ R T + E ++++ YFDC
Sbjct: 283 WCNQSVSVKYLFKYVNKGPDRVTVSVEPHRKEVVSEQNNVGETNNDQQERNQVQDYFDC- 341
Query: 686 YLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIVKNSMFLA 745
RI Y IH++ + KLT+H KQ V KE E E VL R + F A
Sbjct: 342 ---------RIRGYPIHYRQTSITKLTFHEKGKQPVYVKEGETAESVLYRVNNDETQFTA 392
Query: 746 WMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQ-GESIGRMSFVPPGTGELYYLRM 804
W + N + +L Y + P + ++ K K + RK G +GR++ VPP + Y+LR+
Sbjct: 393 WFELNKRDPEAAKLLYEQIPNFYTWNGKDKDFRGRKMPGFVVGRINHVPPKIDDAYHLRI 452
Query: 805 LLNVQR 810
L+N R
Sbjct: 453 LINNMR 458
>At4g07800 hypothetical protein
Length = 448
Score = 234 bits (598), Expect = 2e-61
Identities = 133/327 (40%), Positives = 188/327 (56%), Gaps = 30/327 (9%)
Query: 503 EFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPCGL 562
+F+KRGLP A ILL++ A KL D + L+E + N M+HG CG
Sbjct: 133 KFKKRGLPRARILLFMEANRKLPI-----------ADDIETEQELYELIKNSMIHGLCGS 181
Query: 563 SKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVPYN 622
+ S CM +G+C+K +PKK+ ++T +DGYPVYRRR + GV DN +VVPYN
Sbjct: 182 ANTNSLCMVDGQCSKLYPKKHQELTKVGADGYPVYRRRPIDGCIEKGGVKCDNMYVVPYN 241
Query: 623 PKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTL--------SMSSGRTTQDKPV- 673
+ +KYQAHIN+E+CN++ SIKYLFKYINKG DRV + S+ T D+ V
Sbjct: 242 -RFSLKYQAHINVEWCNQNGSIKYLFKYINKGPDRVVFIVELVKEETNSNTTTLGDETVT 300
Query: 674 ---EDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIE 730
+ D IK +FD Y+S EA+WRIF++ I H+ PVQKL++H+ KQ F +
Sbjct: 301 TKKKKDGIKDWFDYIYVSASEAIWRIFKFPIQHRSTPVQKLSFHVEGKQPAYFDAKAKMV 360
Query: 731 DVLRRNIVKNSMFLAWMDANCKYVHG------RQLTYVEFPELFVYDPKSKSWHPRKQGE 784
DVL R ++S F+ W+ N K V G R Y E P F +D ++K + R +G
Sbjct: 361 DVLERVSNEDSQFMVWLTLNKKNVVGKNGKRARNCLYAEIPTYFTWDGENKPFKKRTRGF 420
Query: 785 SIGRMSFVPPGTGELYYLRMLLNVQRG 811
+GR+++V + YL +LLN+ RG
Sbjct: 421 FLGRINYVLRKMEDENYLIVLLNIVRG 447
>At5g37110 putative helicase
Length = 1307
Score = 233 bits (594), Expect = 6e-61
Identities = 139/396 (35%), Positives = 206/396 (51%), Gaps = 85/396 (21%)
Query: 523 KLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPCGLSKRESPCMKNGRCAKFFPKK 582
KL ID VI AE+PD P LFE + MVHGPCG+ + PCM+NG+
Sbjct: 520 KLTKAEHIDKVISAEIPDKLKDPELFEVIKESMVHGPCGVVNPKCPCMENGK-------- 571
Query: 583 YADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSN 642
R T ++ DN +V+PYN L ++Y+AHIN+E+CN+S
Sbjct: 572 -----------------RRTDDFVEKKDFKCDNRYVIPYNRSLSLRYRAHINVEWCNQSG 614
Query: 643 SIKYLFKYINKGVDRVTLSMSSGRTTQDKP---------------VEDDEIKQYFDCRYL 687
S+KY+FKYI+KG DRVT+ + S +++K + +E++ YF+CR
Sbjct: 615 SVKYIFKYIHKGPDRVTVVVESSLNSKNKENGKQKDNADTDGSETKKKNEVEDYFNCR-- 672
Query: 688 SPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIVKNSMFLAWM 747
IE VL R+ + SMFLAW
Sbjct: 673 ----------------------------------------RIETVLNRSDLDGSMFLAWF 692
Query: 748 DANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGELYYLRMLLN 807
+ N R+LTY + P F YD K K ++ RK+G +IGR+++VP + YYLR+LLN
Sbjct: 693 ELNKVSKIARKLTYADIPTRFTYDSKEKKFNLRKKGFAIGRINYVPRDIEDGYYLRILLN 752
Query: 808 VQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGGAYTRALFVS 867
VQ G +E+LR+VN+ ++ ++ ACEALGLL +D+E+ID + + + G Y R LFV
Sbjct: 753 VQPGPRCFEELRTVNDVLYKEWKDACEALGLLDNDQEYIDDLKRTSFWSSGGYLRQLFVI 812
Query: 868 FLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNNR 903
L +++ +P +VW TW L++ I + +RK+ NR
Sbjct: 813 ML--DALISPENVWAATWQHLSEDIQ-NNKRKYFNR 845
Score = 119 bits (297), Expect = 2e-26
Identities = 59/119 (49%), Positives = 75/119 (62%)
Query: 1 RSVKAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFA 60
R K K+ F+ CC++G++ LP+L + PELL L N D SRHF +NIR+YN +F
Sbjct: 117 RIEKTVKNKKSKFTSCCLQGQVKLPFLKNSPELLYALPTNDDEISRHFRENIRAYNMIFY 176
Query: 61 FTSIGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
FTS+GG + + GP F I +NYHRIGSL P PKF QLYI DT+NEV NR
Sbjct: 177 FTSLGGDTENSVRASGGPQMFQIHRENYHRIGSLKPDNDIPPKFMQLYIVDTENEVDNR 235
>At2g14470 pseudogene
Length = 1265
Score = 230 bits (587), Expect = 4e-60
Identities = 137/397 (34%), Positives = 196/397 (48%), Gaps = 99/397 (24%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHILL++ A+ KL + ID +I AE+PD + P L+E + N M+HG
Sbjct: 447 MYTVEFQKRGLPHAHILLFMHAKSKLPTSDDIDKLISAEIPDKEKEPELYEVIKNSMIHG 506
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + +SPCM +G C+K +PKK+ D+T SDGYP+YRRR + G+ DN +V
Sbjct: 507 PCGSANVKSPCMVDGECSKLYPKKHQDITKVGSDGYPIYRRRKIDDYVEKGGIKCDNRYV 566
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
+PYN K ++Y AHIN+E+CN+++SIKYLFKYINKG D+V + TQ D E
Sbjct: 567 MPYNKKFSLRYNAHINVEWCNQNDSIKYLFKYINKGPDKVIFIVEP---TQQATAGDSET 623
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVL--RRN 736
Q ++Q K+ I+D RRN
Sbjct: 624 PQ------------------------------------QEQRSAEKKKNEIKDWFDCRRN 647
Query: 737 IV-KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPG 795
V KN R+ Y E P F +D ++K++ R +G SIGR+ +V
Sbjct: 648 AVGKNGK------------RARECLYAEIPAYFTWDGENKAFKKRTRGFSIGRIHYVSRK 695
Query: 796 TGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATL 855
+ Y+LR+LLN+ ++
Sbjct: 696 MEDDYFLRVLLNI---------------------------------------------SV 710
Query: 856 AGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
++ F LL +S+ P HVW++TWH+LA+ I
Sbjct: 711 LFWRLSQEFFAMLLLSDSLSRPAHVWSQTWHILAEDI 747
Score = 127 bits (318), Expect = 6e-29
Identities = 58/116 (50%), Positives = 78/116 (67%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K + F++CC +G + LP+L + P+LL NLL S+H+ DN R++N +FA TS
Sbjct: 9 KKETKKESGFTLCCGEGSVKLPFLKESPDLLKNLLSGNHPLSKHYRDNARTFNMVFAVTS 68
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
+GGKVD + GRGP F + G NYH IGSL PT GD K++QLYI DT+NEV+NR
Sbjct: 69 LGGKVDKSMPKGRGPAMFRLQGGNYHLIGSLKPTPGDYAKYSQLYIVDTENEVENR 124
>At3g42320 putative protein
Length = 541
Score = 203 bits (517), Expect = 5e-52
Identities = 128/353 (36%), Positives = 176/353 (49%), Gaps = 88/353 (24%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYTIEFQKRGL HAHILL+L A KL + D VI AE+PD K L+ V + M+HG
Sbjct: 220 MYTIEFQKRGLLHAHILLFLHATNKLYTAEDTDKVITAEIPDKKKKLELYVLVKDCMIHG 279
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG+ SPCM NG+C K+F K Y D T D+DG+PVYRR NTG+
Sbjct: 280 PCGVGHPNSPCMVNGQCKKYFSKSYYDTTKVDNDGFPVYRRHNTGI-------------- 325
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
Y K+ IKYLFKY++K DRVT+++ K E DE+
Sbjct: 326 ------------------YAEKNGLIKYLFKYVHKEHDRVTVTIEPNNQDTTKK-EKDEL 366
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
+K PP +++ H K N+
Sbjct: 367 ------------------------NKKPPSRRV--HASAK-----------------NLP 383
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
K + F +L E P F ++ K K + +++G +IGR +FVP +
Sbjct: 384 KAAPF------------PTKLLVEEIPNHFTWNSKEKKFMIKERGFAIGRTNFVPHTIED 431
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILD 851
YYLR+LLN++RG T Y+DL++V +H++FR A ALGLL DD+E+++G D
Sbjct: 432 AYYLRILLNIKRGVTSYKDLKTVKGVIHESFRDAVFALGLLHDDKEYMNGFKD 484
Score = 107 bits (267), Expect = 5e-23
Identities = 54/99 (54%), Positives = 67/99 (67%)
Query: 21 KIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGPPQ 80
+I+LP L D PE L +LL + D S+HF +NIR+YN++F+FTSIG KVD L G P
Sbjct: 6 QIVLPSLKDSPEFLWHLLTSDDELSKHFRENIRAYNTLFSFTSIGSKVDHFLPKGPQPNM 65
Query: 81 FVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
F I G+NYH IG+L P KF QLYI DT+NEV NR
Sbjct: 66 FAIQGENYHLIGALKPKSSAKAKFQQLYIADTENEVNNR 104
>At5g32630 putative protein
Length = 856
Score = 172 bits (436), Expect = 1e-42
Identities = 119/353 (33%), Positives = 166/353 (46%), Gaps = 76/353 (21%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
M+T+EFQKRGL HA LL++ A+ KL + ID +I AE+PD P +E + N M+HG
Sbjct: 275 MHTVEFQKRGLLHAPTLLFMDAKSKLPTTDDIDKLISAEIPDKDKEPEFYEVIKNSMIHG 334
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM D Y + G DN +V
Sbjct: 335 PCGAANMNSPCMVE-------------------DDY-----------IEKGGFKCDNSYV 364
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
VPYN KL ++YQAHIN+E +SN IK TL T + K DEI
Sbjct: 365 VPYNQKLSLRYQAHINVEC--QSNRIKQ---------KAATLGEPPNSTEKKK----DEI 409
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
K +FDC L + Y +L++H KQ F N IEDVL R
Sbjct: 410 KDWFDCSCLEDLQISTSASIYS-------SSRLSFHCEWKQPAYFDPNAIIEDVLERISN 462
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
++SMF+AW+ N G K G S+GR+++ +
Sbjct: 463 EDSMFMAWLTLNRNNDVG------------------------KNGFSLGRINYGARKMED 498
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILD 851
YYL++LLN+ RG ED+++ N ++ +F+ AC A G+L DD+ IDG+L+
Sbjct: 499 EYYLQVLLNIVRGPMSCEDIKTFNGVLYPSFKEACFARGILDDDQVNIDGLLE 551
Score = 65.9 bits (159), Expect = 2e-10
Identities = 30/57 (52%), Positives = 40/57 (69%)
Query: 63 SIGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
S+GG+VD + G+GP F + NYH IGSL P GD K++QLYI DT+N+V+NR
Sbjct: 3 SLGGRVDNSMPKGKGPNMFRLQEGNYHLIGSLKPKPGDYAKYSQLYIVDTENKVENR 59
>At1g64410 unknown protein
Length = 753
Score = 149 bits (376), Expect = 1e-35
Identities = 72/161 (44%), Positives = 100/161 (61%), Gaps = 1/161 (0%)
Query: 498 GMYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVH 557
GMYTIEFQKRGLPHAHILL++ KL++ D VI AE+PD K P L+ V + M+H
Sbjct: 178 GMYTIEFQKRGLPHAHILLFMHPTSKLSTAEDTDKVITAEIPDKKKKPELYAVVKDCMIH 237
Query: 558 GPCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGF 617
GPCG+ SPCM+NG+C K+FPK Y+D T D+DG+PVYRRR+TG+ + G D
Sbjct: 238 GPCGVGHPNSPCMENGKCKKYFPKSYSDTTKVDNDGFPVYRRRDTGIYVEKNGHDRVTVT 297
Query: 618 VVPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRV 658
V P + K + + +Y + S K++ + + R+
Sbjct: 298 VEPNDQDTAKKEKDEVK-DYFDCSKEKKFMIRERGFAIGRI 337
Score = 106 bits (264), Expect = 1e-22
Identities = 56/127 (44%), Positives = 81/127 (63%), Gaps = 4/127 (3%)
Query: 773 KSKSWHPRKQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGA 832
K K + R++G +IGR++FVP + YYLR+LLN++RG T +DL++V V+ +FR A
Sbjct: 321 KEKKFMIRERGFAIGRINFVPRTIEDAYYLRILLNIKRGVTSSKDLKTVKAVVYKSFRDA 380
Query: 833 CEALGLLKDDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
ALGLL DD+E+I+ I D Y R LFV LL S+ P VW+ETW +L++
Sbjct: 381 VFALGLLDDDKEYINEIKDANFWCSAKYVRRLFVIMLLSESLTKPEMVWDETWKILSE-- 438
Query: 893 ACDLQRK 899
D++RK
Sbjct: 439 --DIERK 443
Score = 79.7 bits (195), Expect = 1e-14
Identities = 43/87 (49%), Positives = 54/87 (61%), Gaps = 7/87 (8%)
Query: 38 LRNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPT 97
L++ D ++HF +NIR+YN +F+FTSIGGKVD L GRGP F I G+L P
Sbjct: 65 LQSDDELAKHFRENIRAYNMLFSFTSIGGKVDHCLPKGRGPNMFAIQ-------GALKPK 117
Query: 98 EGDNPKFAQLYIYDTKNEVQNRMSHFR 124
KF QLYI DT+NEV NR + R
Sbjct: 118 SVAKAKFQQLYIVDTENEVNNRYNIMR 144
>At5g28670 putative protein
Length = 647
Score = 125 bits (313), Expect = 2e-28
Identities = 70/211 (33%), Positives = 111/211 (52%), Gaps = 25/211 (11%)
Query: 543 LYPSLFECVSNYMVHGPCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNT 602
+ PS F Y++ P L CM+N +C+KF+PK + + T+ D++GY +YRR +T
Sbjct: 281 IIPSSFTGGPAYILVNPNSL------CMENDKCSKFYPKNHVENTSLDNEGYQIYRRIDT 334
Query: 603 GVSTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSM 662
G + DN +V+PYN L KY+AHIN+E+CN+S +KYLFKY+NK +DRVT+S+
Sbjct: 335 GRFIEKNKFQYDNWYVIPYNDVFLRKYRAHINVEWCNQSVFVKYLFKYVNKDLDRVTVSI 394
Query: 663 SSGR-----------TTQDKPVEDDEIKQYFDCRYLSPCE-AVW-------RIFQYDIHH 703
R T P E +++ Y+DC + W + F+ D H
Sbjct: 395 KPHRKEDVTEQNNVGETNKDPQERNQVHDYYDCSEHDDTQFMAWLELNKPLQSFEVDEEH 454
Query: 704 KWPPVQKLTYHLPRKQVVLFKENEAIEDVLR 734
+ P LT +++ + + IE ++R
Sbjct: 455 ETSPRYILTRIQISDDILIPEGDNPIESIIR 485
Score = 81.3 bits (199), Expect = 4e-15
Identities = 39/76 (51%), Positives = 50/76 (65%)
Query: 45 SRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKF 104
++ F NIR YN +F+ TSIGGKV + RGP F + G+NYH IG+L P D KF
Sbjct: 177 AKTFRANIRPYNMLFSVTSIGGKVYRSVKKRRGPSMFALQGENYHLIGALKPKADDYAKF 236
Query: 105 AQLYIYDTKNEVQNRM 120
QLYI DT+NEV N++
Sbjct: 237 QQLYIMDTENEVDNQI 252
>At3g42420 putative protein
Length = 1018
Score = 121 bits (303), Expect = 3e-27
Identities = 50/112 (44%), Positives = 76/112 (67%)
Query: 530 IDSVICAEVPDPKLYPSLFECVSNYMVHGPCGLSKRESPCMKNGRCAKFFPKKYADVTTF 589
ID +I AE+PD + P L+E + M+HGPCG + + SPCM +G+C+K +PKK+ ++T
Sbjct: 422 IDKMISAEIPDKEKEPELYEVIKKCMIHGPCGAANKNSPCMVDGKCSKNYPKKFEEITKV 481
Query: 590 DSDGYPVYRRRNTGVSTNRRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKS 641
DG+PVYRR+ + + G DN +VV YN L ++Y AHIN+E+CN++
Sbjct: 482 GKDGFPVYRRKQSHNYVEKSGFRCDNRYVVSYNKILSVRYGAHINVEWCNQN 533
Score = 48.9 bits (115), Expect = 2e-05
Identities = 31/88 (35%), Positives = 43/88 (48%), Gaps = 11/88 (12%)
Query: 29 DCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGPPQFVISGQNY 88
D EL+ LL + D + R++ + YN +FA TSIGGKVD + G+GP F + G
Sbjct: 318 DSIELVKELLTDDDAQGRNYRKHAWIYNMVFAMTSIGGKVDKSMPKGQGPIMFRLQG--- 374
Query: 89 HRIGSLVPTEGDNPKFAQLYIYDTKNEV 116
G K + YI T +EV
Sbjct: 375 --------AIGRKEKDQRTYIRPTTSEV 394
Score = 40.4 bits (93), Expect = 0.007
Identities = 21/76 (27%), Positives = 36/76 (46%)
Query: 809 QRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGGAYTRALFVSF 868
Q G T + +++ N V+ T++ A G+L DD+ +ID ++D + G + R F
Sbjct: 532 QNGPTCFAHIKTYNGVVYPTYKTVSFARGILDDDQVYIDSLVDASQFCFGNFLRNFFAML 591
Query: 869 LLPNSMCNPLHVWNET 884
LL + N T
Sbjct: 592 LLSELELTEAEIKNYT 607
>At3g31980 hypothetical protein
Length = 1099
Score = 117 bits (292), Expect = 6e-26
Identities = 73/249 (29%), Positives = 117/249 (46%), Gaps = 50/249 (20%)
Query: 659 TLSMSSGRTTQDKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRK 718
T +M +G T+ K DEIK YFDCR
Sbjct: 353 TSNMQTGSETRSKEKSTDEIKDYFDCR--------------------------------- 379
Query: 719 QVVLFKENEAIEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWH 778
K NE +SMF+A++ N + RQ TY E P+ F +D ++K W
Sbjct: 380 -----KANE------------DSMFMAFLKLNQECEFARQFTYTEIPQYFTWDGQNKQWK 422
Query: 779 PRKQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGL 838
R++G IGRM++ YY+R+LL + G T ED+R+ + V++T++ AC A G+
Sbjct: 423 LRERGFCIGRMNYASIKMDPEYYMRILLGIVCGPTSDEDIRTYKDVVYETYKEACLARGI 482
Query: 839 LKDDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQR 898
L DD+ +ID I++ + G + R LF LL + P +VW + +L + I ++
Sbjct: 483 LTDDQAYIDTIVEGSLYFFGDHLRNLFSMMLLDKCLARPEYVWEKCSRILIEDIETKKRK 542
Query: 899 KHNNRGMCL 907
+++N + L
Sbjct: 543 QYDNPDLVL 551
Score = 92.0 bits (227), Expect = 2e-18
Identities = 40/76 (52%), Positives = 55/76 (71%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHILL++ K + ID++I AE+PD P L+E V + M+HG
Sbjct: 265 MYTVEFQKRGLPHAHILLFMDKSCKFPTSDDIDNIISAEIPDKSKDPKLYEVVKDCMIHG 324
Query: 559 PCGLSKRESPCMKNGR 574
PCG + +ESPC+ +G+
Sbjct: 325 PCGAANKESPCIVDGQ 340
>At4g03690 hypothetical protein
Length = 570
Score = 107 bits (266), Expect = 6e-23
Identities = 45/94 (47%), Positives = 65/94 (68%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+ FQKRGLPHAHI+LW+ YK ++ +D +I AE+ D L++ VS M+HG
Sbjct: 1 MYTVGFQKRGLPHAHIILWMDPRYKFPTVDDVDKIIFAEILDKVKDSELYQVVSECMIHG 60
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSD 592
PCGL SPCM+NG+ +KF+PK + + T+ D++
Sbjct: 61 PCGLVNPNSPCMENGKYSKFYPKNHVENTSLDNE 94
Score = 46.2 bits (108), Expect = 1e-04
Identities = 24/67 (35%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Query: 837 GLLKDDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDL 896
GLL DD+E+I GI + Y LFV L+ + +P VW TW +L++ D
Sbjct: 113 GLLDDDKEYIHGIEEANFWCSPKYVCKLFVIMLISERLSSPAVVWEHTWKILSE----DF 168
Query: 897 QRKHNNR 903
+RK N+
Sbjct: 169 KRKLRNQ 175
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.353 0.158 0.566
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,963,059
Number of Sequences: 26719
Number of extensions: 1112413
Number of successful extensions: 5147
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 5030
Number of HSP's gapped (non-prelim): 73
length of query: 1408
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1296
effective length of database: 8,326,068
effective search space: 10790584128
effective search space used: 10790584128
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0110a.10


