

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
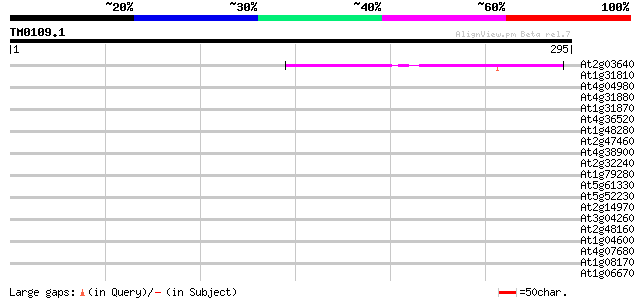
Query= TM0109.1
(295 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g03640 unknown protein 42 4e-04
At1g31810 hypothetical protein 40 0.002
At4g04980 39 0.004
At4g31880 unknown protein 38 0.007
At1g31870 unknown protein 37 0.013
At4g36520 trichohyalin like protein 36 0.028
At1g48280 unknown protein 36 0.028
At2g47460 putative MYB family transcription factor 35 0.037
At4g38900 transcription factor bZIP29 (BZIP29) 35 0.048
At2g32240 putative myosin heavy chain 35 0.048
At1g79280 hypothetical protein 35 0.048
At5g61330 unknown protein 35 0.063
At5g52230 unknown protein 35 0.063
At2g14970 En/Spm-like transposon protein 35 0.063
At3g04260 hypothetical protein 34 0.082
At2g48160 unknown protein 34 0.082
At1g04600 putative myosin heavy chain 34 0.082
At4g07680 hypothetical protein 34 0.11
At1g08170 hypothetical protein 34 0.11
At1g06670 DEIH-box RNA/DNA helicase 34 0.11
>At2g03640 unknown protein
Length = 422
Score = 42.0 bits (97), Expect = 4e-04
Identities = 36/149 (24%), Positives = 65/149 (43%), Gaps = 11/149 (7%)
Query: 146 HVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGDDALA 205
+V+ E TK+V E ++TE NES+ + E ++T +++ G A
Sbjct: 131 YVSDEFVEPEATKEVEESQSTNAITEPANESVEAVIVPTEAKTTVTKPASAIPNGH---A 187
Query: 206 SPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDAL---ASPPPQP 262
P + K+V ++ ++A K E +S V SL+ L ASP +
Sbjct: 188 KVPEE-----KVVNENSSLPKAAEAKLQEEVPKKSFALIVQSLAQSAGTLQVKASPVKRK 242
Query: 263 KTRRKLAAPQPAAPQPAVRRSARIKTTPK 291
+ +AAP+ AP P ++++ P+
Sbjct: 243 PVEKPVAAPERKAPSPIRKQASAESIKPQ 271
>At1g31810 hypothetical protein
Length = 1201
Score = 39.7 bits (91), Expect = 0.002
Identities = 27/94 (28%), Positives = 37/94 (38%), Gaps = 9/94 (9%)
Query: 198 SLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALAS 257
S G+ A PPP P P P R K T+ S+ +G +
Sbjct: 630 STGNKRQAQPPPPP--------PPPPPTRIPAAKCAPPPPPPPPTSHSGSIRVGPPSTPP 681
Query: 258 PPPQPKTRRKLA-APQPAAPQPAVRRSARIKTTP 290
PPP P + ++ AP+P AP P S R+ P
Sbjct: 682 PPPPPPPKANISNAPKPPAPPPLPPSSTRLGAPP 715
Score = 30.4 bits (67), Expect = 1.2
Identities = 22/85 (25%), Positives = 30/85 (34%), Gaps = 7/85 (8%)
Query: 207 PPPQPKTRRKLVA------PQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALA-SPP 259
PPP P TR P P S I+ G ++ + +PP
Sbjct: 643 PPPPPPTRIPAAKCAPPPPPPPPTSHSGSIRVGPPSTPPPPPPPPPKANISNAPKPPAPP 702
Query: 260 PQPKTRRKLAAPQPAAPQPAVRRSA 284
P P + +L AP P P P + A
Sbjct: 703 PLPPSSTRLGAPPPPPPPPLSKTPA 727
>At4g04980
Length = 681
Score = 38.5 bits (88), Expect = 0.004
Identities = 44/144 (30%), Positives = 61/144 (41%), Gaps = 20/144 (13%)
Query: 150 ETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGD--DALAS- 206
ETE+E+ T+D E T +E T+ S EDV S T ++S + +L S
Sbjct: 267 ETEHENETEDHSETTTSE--TDSTESSPKEDVPPPPPLTSPQTPSPTVSTFNTKSSLRSQ 324
Query: 207 -PPPQPKTRRKLVAPQPAVRRSARIKTGE---MEKNESLTTDVASLSLGDDALASPPPQP 262
PPP P K AP P S ++GE K S D A ++ +PP P
Sbjct: 325 PPPPPPSPEHKAPAPPPPPPMSKASESGEFCQFSKTHSTNGDNA------PSMPAPPAPP 378
Query: 263 KTRRKLAAPQPAAPQPAVRRSARI 286
+ R L +RRSA+I
Sbjct: 379 GSGRSL-----KKATSKLRRSAQI 397
>At4g31880 unknown protein
Length = 873
Score = 37.7 bits (86), Expect = 0.007
Identities = 44/179 (24%), Positives = 78/179 (42%), Gaps = 26/179 (14%)
Query: 138 TENKSLMEHVAAETENESLT---------KDVPEETENESLTED--------VNESLTED 180
TE K +EH E ENES + D+ EETE L + V+ S+T
Sbjct: 349 TEEKPDVEHQIEEKENESSSVKQADLSKDSDIKEETEPAELLDSKDVLTSPPVDSSVTAA 408
Query: 181 V-GEMEKNESLTTDVASLSLGDDA--LASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEK 237
E EKN+S+ + S + GD+ ++SP + + V + A ++ T E++
Sbjct: 409 TSSENEKNKSVQI-LPSKTSGDETANVSSPSMAEELPEQSVPKKTANQKKKESSTEEVKP 467
Query: 238 NESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAP--QPAVRRSARIKTTPKKFK 294
+ S+ T+ S + + P K+ +K+A+ P P+ + ++ K + K
Sbjct: 468 SASIATEEVS---EEPNTSEPQVTKKSGKKVASSSKTKPTVPPSKKSTSETKVAKQSEK 523
>At1g31870 unknown protein
Length = 561
Score = 37.0 bits (84), Expect = 0.013
Identities = 45/182 (24%), Positives = 69/182 (37%), Gaps = 36/182 (19%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDV-PEETENESLTEDVNESLTED-- 180
E++ K K+ S+ E + ++ E+ K V PEE ENE + + + ED
Sbjct: 21 EKKKKKKKQKKPSKPEPRGVL----VVDEDPVWQKQVDPEEDENEDDSAEETPLVDEDIE 76
Query: 181 VGEMEKNESLTTDVASLSLGDDALA---------------SPPPQPKTRRKLVAPQPAVR 225
V M + E + A ++ +D SPP + +TR +P+P R
Sbjct: 77 VKRMRRLEEIKARRAHNAIAEDGSGWVTLPLNREDTQSNISPPRRQRTRNDSPSPEPGPR 136
Query: 226 RSA------------RIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQP 273
RS R K E +SL D SPP + K R +P+P
Sbjct: 137 RSVADRVDTDMSPPRRRKRHNSPSPEPNRKHTKPVSLDSD--MSPPRKRKARNDSPSPEP 194
Query: 274 AA 275
A
Sbjct: 195 EA 196
>At4g36520 trichohyalin like protein
Length = 1432
Score = 35.8 bits (81), Expect = 0.028
Identities = 38/168 (22%), Positives = 66/168 (38%), Gaps = 20/168 (11%)
Query: 92 KNEIKTGFAKQHNFKGCVPVFEIIPCECCVTAEEENK-SLKENVASETENKSLMEHVAAE 150
+ +IK K N + V E E + ++E + LKE E EN+ + E A E
Sbjct: 674 ERKIKEAREKAENERRAVEAREKAEQERKMKEQQELELQLKEAFEKEEENRRMREAFALE 733
Query: 151 TENESLTKDVPEETENE------------------SLTEDVNESLTEDVGEMEKNESLTT 192
E E K+ E+ ENE +L ++ E ++ E E+NE
Sbjct: 734 QEKERRIKEAREKEENERRIKEAREKAELEQRLKATLEQEEKERQIKERQEREENERRAK 793
Query: 193 DVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTG-EMEKNE 239
+V + + L Q + R+L + +++ E+E+ E
Sbjct: 794 EVLEQAENERKLKEALEQKENERRLKETREKEENKKKLREAIELEEKE 841
Score = 28.1 bits (61), Expect = 5.9
Identities = 15/64 (23%), Positives = 27/64 (41%)
Query: 125 EENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEM 184
EEN + + EN+ ++ + E E K+ E+ ENE + E ++
Sbjct: 645 EENDRRERVAVEKAENEKRLKAALEQEEKERKIKEAREKAENERRAVEAREKAEQERKMK 704
Query: 185 EKNE 188
E+ E
Sbjct: 705 EQQE 708
>At1g48280 unknown protein
Length = 558
Score = 35.8 bits (81), Expect = 0.028
Identities = 34/154 (22%), Positives = 64/154 (41%), Gaps = 16/154 (10%)
Query: 125 EENKSLKENVASETENKSLMEH-VAAETENESLTK-DVPEETENESLTEDVNESLTEDVG 182
E ++ NV E N+ L + V+AE + SL+ D P + S +D+ + +
Sbjct: 145 ELEEARNSNVELELNNRKLSQDLVSAEAKISSLSSNDKPAKEHQNSRFKDIQRLIASKLE 204
Query: 183 EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLT 242
+ + + + + + LS + + PP P + LV+P ++ G+ ++N S
Sbjct: 205 QPKVKKEVAVESSRLSPPSPSPSRLPPTPPLPKFLVSPASSL--------GKRDENSS-- 254
Query: 243 TDVASLSLGDDALASPPPQPKTRRKLAAPQPAAP 276
+ PPP P+ K A Q + P
Sbjct: 255 ----PFAPPTPPPPPPPPPPRPLAKAARAQKSPP 284
>At2g47460 putative MYB family transcription factor
Length = 371
Score = 35.4 bits (80), Expect = 0.037
Identities = 28/117 (23%), Positives = 49/117 (40%), Gaps = 4/117 (3%)
Query: 133 NVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTT 192
N+ E E + H + +P T+NE + N L+ + + S++
Sbjct: 69 NITPEEEELVVKLHSTLGNRWSLIAGHLPGRTDNE-IKNYWNSHLSRKLHNFIRKPSISQ 127
Query: 193 DVASLSLGDDALASPPPQPKT---RRKLVAPQPAVRRSARIKTGEMEKNESLTTDVA 246
DV+++ + + + A PPPQ K R A +P + R+ KT + DVA
Sbjct: 128 DVSAVIMTNASSAPPPPQAKRRLGRTSRSAMKPKIHRTKTRKTKKTSAPPEPNADVA 184
Score = 30.4 bits (67), Expect = 1.2
Identities = 16/52 (30%), Positives = 30/52 (56%), Gaps = 2/52 (3%)
Query: 237 KNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKT 288
+ S++ DV+++ + + + A PPPQ K R+L +A +P + R+ KT
Sbjct: 121 RKPSISQDVSAVIMTNASSAPPPPQAK--RRLGRTSRSAMKPKIHRTKTRKT 170
>At4g38900 transcription factor bZIP29 (BZIP29)
Length = 547
Score = 35.0 bits (79), Expect = 0.048
Identities = 44/180 (24%), Positives = 69/180 (37%), Gaps = 32/180 (17%)
Query: 127 NKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEM-E 185
N S ++ + EN+ ME A S TK +TE ES VNES ++ E
Sbjct: 244 NSSEADDSKNGNENRDDMESSRA-----SGTKTNGSDTEGES--SSVNESANNNMNSSGE 296
Query: 186 KNESLTTDVAS----------------------LSLGDDALASPPPQPKTRRKLVAPQPA 223
K ES+ A LS GD++L PPP P + + V+P +
Sbjct: 297 KRESVKRRAAGGDIAPTTRHYRSVSVDSCFMEKLSFGDESL-KPPPSPGSMSRKVSPTNS 355
Query: 224 V-RRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRR 282
V S + E E ++ + D PK +++ A + +A + R+
Sbjct: 356 VDGNSGAAFSIEFNNGEFTAAEMKKIMANDKLAEMAMSDPKRVKRILANRQSAARSKERK 415
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 35.0 bits (79), Expect = 0.048
Identities = 34/122 (27%), Positives = 57/122 (45%), Gaps = 7/122 (5%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENE-SLTEDVNESLTEDVG 182
+E N+ + EN E KS +AA E +L+K ETE + S TE + + LT+++
Sbjct: 264 KELNEKMSENEKVEAALKSSAGELAAVQEELALSKSRLLETEQKVSSTEALIDELTQELE 323
Query: 183 EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLT 242
+ + +ES + S+ DA Q K + ++ Q + + E E ESL+
Sbjct: 324 QKKASESRFKEELSVLQDLDA------QTKGLQAKLSEQEGINSKLAEELKEKELLESLS 377
Query: 243 TD 244
D
Sbjct: 378 KD 379
>At1g79280 hypothetical protein
Length = 2111
Score = 35.0 bits (79), Expect = 0.048
Identities = 45/188 (23%), Positives = 74/188 (38%), Gaps = 28/188 (14%)
Query: 118 ECCVTAEEENKSLK-ENVASETENKSLMEHVAAETENES---------LTKDVPEETENE 167
+ T + +N+ + EN +TE E+V A+ +NE+ T+ +P E E+
Sbjct: 1832 DATTTTDGDNEETEAENAEEKTE-----EYVEAQQDNEADEPVEESPTETETIPTEEESR 1886
Query: 168 SLTEDVN-ESLTEDVGEMEKNESLTTDVASLSLGDDA---LASPPPQPKTRRKLVAPQPA 223
TE+ N E LT+ + E+ E + L G D + SP + L P
Sbjct: 1887 DQTEEENQEPLTDMESDKEEGELDLDTLEDLEEGTDVASMMRSPEKEEVQPETLATP--- 1943
Query: 224 VRRSARIKTGEMEKNESLTTDVA--SLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVR 281
+ +R++T E ++ T V G DA P A Q AP+ ++
Sbjct: 1944 TQSPSRMETAMEEAETTIETPVEDDKTDEGGDAAEEAADIPNN----ANDQQEAPETDIK 1999
Query: 282 RSARIKTT 289
TT
Sbjct: 2000 PETSAATT 2007
>At5g61330 unknown protein
Length = 436
Score = 34.7 bits (78), Expect = 0.063
Identities = 20/62 (32%), Positives = 38/62 (61%), Gaps = 1/62 (1%)
Query: 128 KSLKENVASETENKSLMEHVAAETENE-SLTKDVPEETENESLTEDVNESLTEDVGEMEK 186
+S + + SE+E+ S E++ AE++NE D E+ E +S+ +D ES +D G+ E+
Sbjct: 7 RSKRARLDSESEDISDQENLKAESDNEDDQLPDGIEDDEVDSMEDDEGESEEDDEGDTEE 66
Query: 187 NE 188
++
Sbjct: 67 DD 68
>At5g52230 unknown protein
Length = 746
Score = 34.7 bits (78), Expect = 0.063
Identities = 47/171 (27%), Positives = 66/171 (38%), Gaps = 23/171 (13%)
Query: 122 TAEEENKSLKENVASETENKSLMEHVAAETENESLTKD--VPEETENESLTEDVNESLTE 179
T EE LK N++S KS + V + ++ K+ + EE E +S + + S E
Sbjct: 197 TTEEVVVDLKRNLSSSNA-KSEKDSVNSSVRSQKPKKEAVMKEEEEQDSSEKRITRSKVE 255
Query: 180 DVGEMEKNESLTTDVA---SLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEME 236
EK L+ VA S L L P P+ KTR K+ P E
Sbjct: 256 -----EKKNELSNSVARRTSKRLAGIEL-EPTPELKTRAKVQRIVPL--------DDEPT 301
Query: 237 KNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIK 287
T V + DD P P+ KTR K+ P +P + R K
Sbjct: 302 PELKTRTKVQRVVPPDD---EPTPELKTRTKIQRIVPPDDEPTLELKTRTK 349
>At2g14970 En/Spm-like transposon protein
Length = 771
Score = 34.7 bits (78), Expect = 0.063
Identities = 37/111 (33%), Positives = 54/111 (48%), Gaps = 16/111 (14%)
Query: 126 ENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVN---ESLTEDV- 181
EN E++ ETE ++ E + ETE E+ +D+ ETE E+ ED+N E+ ED+
Sbjct: 95 ENIEEVEDIDGETEKEA--EDINGETEKEA--EDINGETEKEA-DEDINGETENEAEDIN 149
Query: 182 GEMEKNESLTT---DVASLSL-GDDALASPPPQPK---TRRKLVAPQPAVR 225
GE E L L L GD+ + P + + TR K +A P R
Sbjct: 150 GETENEAELQAAEEPEGELELSGDEDVVQPKTKRQRGPTRMKDIAKDPNAR 200
Score = 28.1 bits (61), Expect = 5.9
Identities = 22/58 (37%), Positives = 30/58 (50%), Gaps = 6/58 (10%)
Query: 125 EENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENE---SLTEDVNESLTE 179
E K E++ ETEN++ E + ETENE+ EE E E S EDV + T+
Sbjct: 128 ETEKEADEDINGETENEA--EDINGETENEA-ELQAAEEPEGELELSGDEDVVQPKTK 182
>At3g04260 hypothetical protein
Length = 913
Score = 34.3 bits (77), Expect = 0.082
Identities = 26/121 (21%), Positives = 52/121 (42%), Gaps = 7/121 (5%)
Query: 134 VASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTD 193
+ + E+K E V E+E +D+ +E +NE ++ + ED E E+ E + +
Sbjct: 624 IETSVESKETTESVVTG-ESEKAIEDISKEADNEEDDDEEEQEGDEDDDENEEEEVVVPE 682
Query: 194 VASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDD 253
+ + G+D + + K +++ Q +T + K S ++L DD
Sbjct: 683 TENRAEGEDLVKNKAADAKKHLQMIGVQLLKESDEANRTKKRGKRAS------RMTLEDD 736
Query: 254 A 254
A
Sbjct: 737 A 737
>At2g48160 unknown protein
Length = 1366
Score = 34.3 bits (77), Expect = 0.082
Identities = 30/97 (30%), Positives = 48/97 (48%), Gaps = 9/97 (9%)
Query: 121 VTAEEENKSLKENVASETENKS--LMEHVAAETENESLTKDVPEETENESLTEDVNESLT 178
VT E E++ L+ENV+S T + ++E V E E E + P TEN + T+ + +
Sbjct: 1056 VTPEHESRILEENVSSSTAERHTLILEDVDGELEMEDVAP--PWGTENCTHTDQADNTKV 1113
Query: 179 ED--VGEMEKNESLTTDVASLSLGDDAL--ASPPPQP 211
+ +G+ + T +SL L +SPPP P
Sbjct: 1114 SNCQLGQQHR-PVFGTSHQHMSLSSPPLPSSSPPPPP 1149
>At1g04600 putative myosin heavy chain
Length = 1730
Score = 34.3 bits (77), Expect = 0.082
Identities = 41/131 (31%), Positives = 54/131 (40%), Gaps = 16/131 (12%)
Query: 72 EKVNWAEAIYDDLHVSFNKLKNEIKTGFAK--------QHNFKGCVPVFEIIPCECCVTA 123
E N A + L S + L+N+I K + K VPV I +
Sbjct: 994 EMTNDLAAENEQLKESVSSLQNKIDESERKYEEISKISEERIKDEVPV---IDQSAIIKL 1050
Query: 124 EEENKSLKENVASETENKSLMEHVAAETE---NESLTKDVPEETENESLTEDVNESLTED 180
E EN+ LK V+S E ++ ET E L +DV + E S E NE L
Sbjct: 1051 ETENQKLKALVSSMEEKIDELDRKHDETSPNITEKLKEDVSFDYEIVSNLEAENERLKAL 1110
Query: 181 VGEMEK--NES 189
VG +EK NES
Sbjct: 1111 VGSLEKKINES 1121
Score = 28.9 bits (63), Expect = 3.5
Identities = 44/190 (23%), Positives = 73/190 (38%), Gaps = 23/190 (12%)
Query: 124 EEENKSLKENVAS------ETENKSLMEHVAAE--TENESLTKDVPEETENESLTEDVNE 175
E EN+ LK V S E+ N S E + + ESLT+D + E D N+
Sbjct: 1101 EAENERLKALVGSLEKKINESGNNSTDEQEEGKYILKEESLTEDASIDNERVKKLADENK 1160
Query: 176 SLTEDVGEMEKNESLTT---DVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKT 232
L + V +EK T + AS + + + + Q + + ++T
Sbjct: 1161 DLNDLVSSLEKKIDETEKKYEEASRLCEERLKQALDAETGLIDLKTSMQRLEEKVSDMET 1220
Query: 233 GEMEKNESLTTDVASLSLGDD-ALASPPPQPKTRRKLAAPQPAAPQPAVR------RSAR 285
E + + + AS + + PP + +P AP P+ R R +R
Sbjct: 1221 AEQIRRQQALVNSASRRMSPQVSFTGAPPLENGHQ-----EPLAPIPSRRFGTESFRRSR 1275
Query: 286 IKTTPKKFKD 295
I+ P +F D
Sbjct: 1276 IERQPHEFVD 1285
>At4g07680 hypothetical protein
Length = 684
Score = 33.9 bits (76), Expect = 0.11
Identities = 40/199 (20%), Positives = 82/199 (41%), Gaps = 34/199 (17%)
Query: 122 TAEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESL----------TE 171
T+E++ K +K+ A +E+ + + V+ + +SL KD ++ + ++ E
Sbjct: 44 TSEKKKKLVKDKEAEVSESPAKKQKVSQSEDVDSLEKDAEKKKKKKNKKKEVAVESDEDE 103
Query: 172 DVNESLTEDVGEMEKNESLTTDVAS-LSLGDDALASPPPQPKTRRKLVAPQPAVRRSARI 230
DV + GE+E + +++ S L+ D + + + +R ++ Q R A+
Sbjct: 104 DVRSREQDGQGELESLVDVVSNLISPLNSRFDTVDTSIKEMSSRLDVIGGQIESRVEAKF 163
Query: 231 --KTGEMEKN--------ESLTTDVASLSLGDDALASPPPQ-------------PKTRRK 267
+ G +E + +++ +S + D LA P PQ P +K
Sbjct: 164 EGRFGSIENDVKQIKEQLKAIADSKSSSYIRDIFLAKPQPQTQEDNPKAQTQKTPNVPKK 223
Query: 268 LAAPQPAAPQPAVRRSARI 286
Q A P P + A +
Sbjct: 224 TTNNQSATPSPPPSKQADV 242
>At1g08170 hypothetical protein
Length = 243
Score = 33.9 bits (76), Expect = 0.11
Identities = 33/128 (25%), Positives = 53/128 (40%), Gaps = 19/128 (14%)
Query: 131 KENVASETENKSLMEHVAAETENESLTKDVPEETENESLTED------VNESLTE----- 179
K V S T+ K ++E T E V ET N+ T+D V E++T
Sbjct: 5 KPKVVSVTKKKKVVEETIKVTVTEEGDPCVITETANDQETQDLTFSIPVGENVTTVEIPV 64
Query: 180 ---DVGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARI-----K 231
D + E++TT + D++ PP P R +PQP +++ K
Sbjct: 65 EVPDERSLPVGENVTTVKIPVDDRDESSPQPPETPVEVRDEPSPQPPETPASKSEGTLKK 124
Query: 232 TGEMEKNE 239
T ++EK +
Sbjct: 125 TDKVEKKQ 132
>At1g06670 DEIH-box RNA/DNA helicase
Length = 1576
Score = 33.9 bits (76), Expect = 0.11
Identities = 33/145 (22%), Positives = 64/145 (43%), Gaps = 19/145 (13%)
Query: 124 EEENKSLKENVASETE--NKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDV 181
EE +++EN S+ N+ + +A+ + ++ ++ P + N + + N + + D+
Sbjct: 1230 EENFGNMEENPPSDLAIGNEQTLPKLASNLDMGNMEENTPSDLANGNEKTEPNSANSMDL 1289
Query: 182 GEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESL 241
G ME+N + D A + +PK+ KL V + + G NE
Sbjct: 1290 GNMEEN----------TPSDLANGNKKKEPKSVSKLDLGSEKVSIPSNLVNG----NEQH 1335
Query: 242 TTDVASLSLGDDALASPPPQPKTRR 266
++A G+DA A+ P+ K R
Sbjct: 1336 DLNIAP---GEDASAAKQPEKKRSR 1357
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.309 0.127 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,892,487
Number of Sequences: 26719
Number of extensions: 314807
Number of successful extensions: 1969
Number of sequences better than 10.0: 162
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 136
Number of HSP's that attempted gapping in prelim test: 1684
Number of HSP's gapped (non-prelim): 355
length of query: 295
length of database: 11,318,596
effective HSP length: 99
effective length of query: 196
effective length of database: 8,673,415
effective search space: 1699989340
effective search space used: 1699989340
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0109.1


