

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.3
(169 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
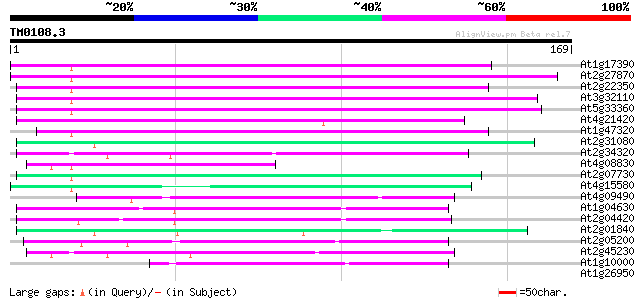
Score E
Sequences producing significant alignments: (bits) Value
At1g17390 hypothetical protein 74 5e-14
At2g27870 putative non-LTR retroelement reverse transcriptase 71 2e-13
At2g22350 putative non-LTR retroelement reverse transcriptase 69 2e-12
At3g32110 non-LTR reverse transcriptase, putative 62 2e-10
At5g33360 putative protein 57 6e-09
At4g21420 hypothetical protein 55 2e-08
At1g47320 hypothetical protein 55 2e-08
At2g31080 putative non-LTR retroelement reverse transcriptase 52 1e-07
At2g34320 putative non-LTR retroelement reverse transcriptase 48 3e-06
At4g08830 putative protein 46 8e-06
At2g07730 putative non-LTR retroelement reverse transcriptase 45 1e-05
At4g15580 splicing factor like protein 44 5e-05
At4g09490 putative proteins 44 5e-05
At1g04630 unknown protein 44 5e-05
At2g04420 putative non-LTR retrolelement reverse transcriptase 43 7e-05
At2g01840 putative non-LTR retroelement reverse transcriptase 43 9e-05
At2g05200 putative non-LTR retroelement reverse transcriptase 42 2e-04
At2g45230 putative non-LTR retroelement reverse transcriptase 40 5e-04
At1g10000 putative reverse transcriptase 40 6e-04
At1g26950 hypothetical protein 38 0.003
>At1g17390 hypothetical protein
Length = 322
Score = 73.6 bits (179), Expect = 5e-14
Identities = 50/146 (34%), Positives = 69/146 (47%), Gaps = 1/146 (0%)
Query: 1 LCWLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAI 59
+ W P VG +K N DG S R A GGV+RD DGNW GF + + LAELW
Sbjct: 102 IAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICSAPLAELWGA 161
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
+++A +RG + + +E DS + + P S +VR + V +
Sbjct: 162 YYGLNIAWERGVTQLEMEIDSEMVVGFLRTGIDDSHPLSFLVRLCHGLLSKDWSVRISHV 221
Query: 120 PREANMVADCLAQNAHVFNFGVHLLH 145
REAN +AD LA A G HL +
Sbjct: 222 YREANRLADGLANYAFFLPLGFHLFN 247
>At2g27870 putative non-LTR retroelement reverse transcriptase
Length = 314
Score = 71.2 bits (173), Expect = 2e-13
Identities = 53/166 (31%), Positives = 72/166 (42%), Gaps = 1/166 (0%)
Query: 1 LCWLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAI 59
+ W P G WK N DG S A GGVLRD +G W GF + + LAELW +
Sbjct: 144 IAWSKPEEGWWKLNTDGASRGNPGLASAGGVLRDEEGAWRGGFALNIGVCSAPLAELWGV 203
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
+ +A +R + + IE DS + + P S +VR I +V +
Sbjct: 204 YYGLYIAWERRVTRLEIEVDSEIVVGFLKIGINEVHPLSFLVRLCHDFISRDWRVRISHV 263
Query: 120 PREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSMTVVAS 165
REAN +AD LA A G H L R++L D+ S
Sbjct: 264 YREANRLADGLANYAFSLPLGFHSLSLVPDSLRFILLDDTSGATVS 309
>At2g22350 putative non-LTR retroelement reverse transcriptase
Length = 321
Score = 68.6 bits (166), Expect = 2e-12
Identities = 49/143 (34%), Positives = 68/143 (47%), Gaps = 1/143 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W+ P G K N DG S A GGVLRD +G W+ GF + + LAELW +
Sbjct: 153 WVSPEDGWVKLNTDGASRGNPGFATAGGVLRDHNGAWIGGFAVNIGVCSAPLAELWGVYY 212
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A RG + +E DS +T+ P S ++R + G V + R
Sbjct: 213 GLFIAWGRGARRVELEVDSKMVVGFLTTGIADSHPLSFLLRLCYDFLSKGWIVRISHVYR 272
Query: 122 EANMVADCLAQNAHVFNFGVHLL 144
EAN +AD LA A + G+HLL
Sbjct: 273 EANRLADGLANYAFSLSLGLHLL 295
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 61.6 bits (148), Expect = 2e-10
Identities = 46/158 (29%), Positives = 70/158 (44%), Gaps = 1/158 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W P G +K N DG S A GGVLR+ G W GF + + LAELW +
Sbjct: 1743 WTPPMEGWYKINTDGASRGNPGLASAGGVLRNSAGAWCGGFAVNIGRCSAPLAELWGVYY 1802
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A + + + +E DS + + P S +VR + V + R
Sbjct: 1803 GLYMAWAKQLTHLELEVDSEVVVGFLKTGIGETHPLSFLVRLCHNFLSKDWTVRISHVYR 1862
Query: 122 EANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDS 159
EAN +AD LA +A + G+H+ + LL +D+
Sbjct: 1863 EANSLADGLANHAFSLSLGLHVFDEIPISLVMLLSEDN 1900
>At5g33360 putative protein
Length = 306
Score = 56.6 bits (135), Expect = 6e-09
Identities = 47/159 (29%), Positives = 65/159 (40%), Gaps = 1/159 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W P +G K N DG S A GG LR+ G W GF + + LAELW +
Sbjct: 138 WSKPSLGWCKLNTDGASHGNPGLAIAGGALRNEYGEWCFGFALNIGRCSAPLAELWGVYY 197
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A RG + + +E DS + + P S +VR + V + R
Sbjct: 198 GLFMAWDRGITRLELEVDSEMVVGFLRTGIGSSHPLSFLVRMCHGFLSRDWIVRIGHVYR 257
Query: 122 EANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSM 160
EAN +AD LA A G H P +L D +
Sbjct: 258 EANRLADELANYAFDLPLGYHGFASPPSSLDSILRDDEL 296
>At4g21420 hypothetical protein
Length = 229
Score = 54.7 bits (130), Expect = 2e-08
Identities = 37/136 (27%), Positives = 65/136 (47%), Gaps = 1/136 (0%)
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTA 62
W P G K N DGS + +A GGV R+ ++ G+ + + +AE A++
Sbjct: 63 WKKPENGRIKLNFDGSRGREGQASIGGVFRNHKAEFLLGYSESIGEATSTMAEFAALKRG 122
Query: 63 VDLAVQRGCSPIIIETDSAEAHAAVTSAQPL-IPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
++LA++ G + + +E D+ ++ L ++ V +I+ +P V + R
Sbjct: 123 LELALENGLTDLWLEGDAKIIMDIISRRGRLRCEKTNKHVNYIKVVMPELNNCVLSHVYR 182
Query: 122 EANMVADCLAQNAHVF 137
E N VAD LA+ H F
Sbjct: 183 EGNRVADKLAKLGHQF 198
>At1g47320 hypothetical protein
Length = 259
Score = 54.7 bits (130), Expect = 2e-08
Identities = 45/137 (32%), Positives = 59/137 (42%), Gaps = 1/137 (0%)
Query: 9 GSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAV 67
G K N DG S A GGVL+D +G W GF + S+ +AELW + LA
Sbjct: 99 GCLKINTDGASRGNPGLATAGGVLQDNEGRWCGGFSLNIGRSSAPMAELWGAYYGLYLAW 158
Query: 68 QRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVA 127
+R S I +E DS + + P S +VR I +V + REAN A
Sbjct: 159 ERKSSHIELEVDSEIVVGFLKTGISDHHPLSFLVRLCHGFISKDWRVRIFHVYREANRFA 218
Query: 128 DCLAQNAHVFNFGVHLL 144
D LA G H +
Sbjct: 219 DGLANYVFSLPLGFHFV 235
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 52.4 bits (124), Expect = 1e-07
Identities = 43/157 (27%), Positives = 63/157 (39%), Gaps = 1/157 (0%)
Query: 3 WLLPPVGSWKFNVDGSVWQTRE-AGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W +P G K DG+ A GG +R+ G W+ GF + S LAELW
Sbjct: 1063 WQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALNIGSCAAPLAELWGAYY 1122
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A +G + ++ D +++ P S +VR + V + R
Sbjct: 1123 GLLIAWDKGFRRVELDLDCKLVVGFLSTGVSNAHPLSFLVRLCQGFFTRDWLVRVSHVYR 1182
Query: 122 EANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQD 158
EAN +AD LA A G+H R LL D
Sbjct: 1183 EANRLADGLANYAFTLPLGLHCFDACPEGVRLLLLAD 1219
>At2g34320 putative non-LTR retroelement reverse transcriptase
Length = 292
Score = 47.8 bits (112), Expect = 3e-06
Identities = 43/139 (30%), Positives = 65/139 (45%), Gaps = 5/139 (3%)
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCG--GVLRDCDGNWVQGFCRRL*-SSNPLLAELWAI 59
W PP K N D + WQ CG +LR+ G + R L + N L AEL A+
Sbjct: 137 WKAPPYQWVKCNTDAT-WQLENPRCGIGWILRNESGGVLWMGARALPRTKNVLEAELEAL 195
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
R AV + II E+D A+A + ++ P + I++ + H +V F+
Sbjct: 196 RWAVLTMSRFNYKRIIFESD-AQALVNLLNSDDFWPTLQPALEDIQQLLHHFEEVKFEFT 254
Query: 120 PREANMVADCLAQNAHVFN 138
PR N VAD +A+ + F+
Sbjct: 255 PRGGNKVADRIARESISFS 273
>At4g08830 putative protein
Length = 947
Score = 46.2 bits (108), Expect = 8e-06
Identities = 30/77 (38%), Positives = 40/77 (50%), Gaps = 2/77 (2%)
Query: 6 PPVGSW-KFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAV 63
PP G W K N DG S A GGVLRD G+W GF + + LAELW + +
Sbjct: 843 PPDGEWVKLNTDGASRGNLGLATTGGVLRDGIGHWCGGFALDIGVCSAPLAELWGVYYGL 902
Query: 64 DLAVQRGCSPIIIETDS 80
+A +R + + +E DS
Sbjct: 903 YMAWERRFTRVELEVDS 919
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 45.4 bits (106), Expect = 1e-05
Identities = 38/141 (26%), Positives = 55/141 (38%), Gaps = 1/141 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W P K DG S A G + + G W+ GF + S + LAELW
Sbjct: 802 WKAPSDRWVKLTTDGASRGHQGLAAASGAILNLQGEWLGGFALNIGSCDAPLAELWGAYY 861
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A +G + + DS +++ P S +VR + V + R
Sbjct: 862 GLLIAWDKGFRRVELNLDSELVVGFLSTGISKAHPLSFLVRLCQGFFTRDWLVRVSHVYR 921
Query: 122 EANMVADCLAQNAHVFNFGVH 142
EAN +AD LA A G H
Sbjct: 922 EANRLADGLANYAFFLPLGFH 942
>At4g15580 splicing factor like protein
Length = 559
Score = 43.5 bits (101), Expect = 5e-05
Identities = 42/140 (30%), Positives = 54/140 (38%), Gaps = 15/140 (10%)
Query: 1 LCWLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAI 59
+ W PP G K + DG S A GGV+RD DG WV GF +L
Sbjct: 26 IAWTKPPEGWVKVSTDGASRGNPGPAAAGGVIRDEDGLWVGGFALQL------------- 72
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
V L Q + +E DS + S P + +VR + V +
Sbjct: 73 -AFVRLRWQSCGGRVELEVDSELVVGFLQSGISEAHPLAFLVRLCHGLLSKDWLVRVSHV 131
Query: 120 PREANMVADCLAQNAHVFNF 139
REAN +AD LA A F
Sbjct: 132 YREANRLADGLANYAFSLQF 151
>At4g09490 putative proteins
Length = 170
Score = 43.5 bits (101), Expect = 5e-05
Identities = 35/117 (29%), Positives = 56/117 (46%), Gaps = 6/117 (5%)
Query: 21 QTREAGCGGVLRDCD---GNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIE 77
+T++ G G V+R+C Q R + PL+AE A+ A+ A G + + +
Sbjct: 42 ETKDVGFGWVIRNCPELTALHYQSAARNV--RLPLMAEAIALFLALQYAQSIGITKLSMA 99
Query: 78 TDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQNA 134
+DS + A+TS P Y I + S+ V F +PR N VAD LA+++
Sbjct: 100 SDSQQLITAITSESPSTEFYGIIFDILNLSLGFA-DVSFSFVPRSENRVADELAKSS 155
>At1g04630 unknown protein
Length = 356
Score = 43.5 bits (101), Expect = 5e-05
Identities = 39/132 (29%), Positives = 57/132 (42%), Gaps = 4/132 (3%)
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL*S--SNPLLAELWAIR 60
W PP+G K N DG+ + G +LRD G ++ G + S +NP+ +EL A+
Sbjct: 195 WTTPPIGWTKCNYDGTYHSNAPSKAGWLLRDDRGTFL-GAAHAIGSITTNPMESELQALV 253
Query: 61 TAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIP 120
A+ RG I E D+ E V ++ I R I + F
Sbjct: 254 MAMQHCWSRGYRKIYFEGDNKEVSEIVNGRSSNFAVFNWI-RDISAWRSKFEECKFTWTR 312
Query: 121 REANMVADCLAQ 132
R +NM AD LA+
Sbjct: 313 RFSNMAADALAK 324
>At2g04420 putative non-LTR retrolelement reverse transcriptase
Length = 221
Score = 43.1 bits (100), Expect = 7e-05
Identities = 38/134 (28%), Positives = 63/134 (46%), Gaps = 5/134 (3%)
Query: 3 WLLPPVGSWKFNVDGSV-WQTREAGCGGVLRDCDGNWVQGFCRRL*S--SNPLLAELWAI 59
W PP+G K N DGS ++T++ G ++RD D + +G + + +N L +EL A+
Sbjct: 63 WEQPPMGWIKCNYDGSFNYRTQQTNSGWLIRD-DKGFYKGAAQAVGGTMNNALESELQAL 121
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
A+ +G +I E DS + + Q ++ I R +V+F
Sbjct: 122 VMAMQHTWSQGYRKVIFEGDSKQVEELLNRKQMHFGAFNWI-REAWSWSKRFEEVIFSWT 180
Query: 120 PREANMVADCLAQN 133
PR N AD LA++
Sbjct: 181 PRTNNQPADMLAKS 194
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 42.7 bits (99), Expect = 9e-05
Identities = 42/158 (26%), Positives = 62/158 (38%), Gaps = 7/158 (4%)
Query: 3 WLLPPVGSWKFNVDGSVWQTRE-AGCGGVLRDCDGNWVQGFCRRL*SS-NPLLAELWAIR 60
W PP G K N D Q R+ G +LRDC+G + C +L S + L AE
Sbjct: 1556 WSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSYSALQAEALGFL 1615
Query: 61 TAVDLAVQRGCSPIIIETDSAEAHAAV--TSAQPLIPPYSEIVRHIRRSIPHGMKVVFDL 118
A+ + RG + E D+ E + T L+ +R +P
Sbjct: 1616 HALQMVWIRGYCYVWFEGDNLELTNLINKTEDHHLLETLLYDIRFWMTKLPFSS---IGY 1672
Query: 119 IPREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLY 156
+ RE N+ AD L + A+ + H P + LY
Sbjct: 1673 VNRERNLAADKLTKYANSMSSLYETFHVPPRWLQLYLY 1710
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 42.0 bits (97), Expect = 2e-04
Identities = 41/135 (30%), Positives = 64/135 (47%), Gaps = 10/135 (7%)
Query: 5 LPPVGSWKFNVDGSVW--QTREAGCGGVLRDC-----DGNWVQGFCRRL*SSNPLLAELW 57
LP + S F + W Q+ AG G V + + CRRL S+ L AE W
Sbjct: 1078 LPVLRSGYFCFVDAAWIAQSSLAGSGWVFQSATALEKETATYSAGCRRLPSA--LSAEAW 1135
Query: 58 AIRTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFD 117
AI++A+ A+Q G + +++ +DS A+TS + Y ++ IR + F+
Sbjct: 1136 AIKSALLHALQLGRTDLMVLSDSKSVVDALTSNISINEIYG-LLMEIRALRVSFHSLCFN 1194
Query: 118 LIPREANMVADCLAQ 132
I R AN +AD A+
Sbjct: 1195 FISRSANAIADATAK 1209
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 40.4 bits (93), Expect = 5e-04
Identities = 41/133 (30%), Positives = 60/133 (44%), Gaps = 6/133 (4%)
Query: 6 PPVGSW-KFNVDGSVWQTREAGCG--GVLRDCDGNWVQGFCRRL*SSNPLL-AELWAIRT 61
PP W K N DG+ W CG VLR+ G + R L S +L E+ A+R
Sbjct: 1217 PPSHGWVKCNTDGA-WSKDLGNCGVGWVLRNHTGRLLWLGLRALPSQQSVLETEVEALRW 1275
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
AV + +I E+DS + + + IP + ++ IR + H +V F R
Sbjct: 1276 AVLSLSRFNYRRVIFESDSQYLVSLIQNEMD-IPSLAPRIQDIRNLLRHFEEVKFQFTRR 1334
Query: 122 EANMVADCLAQNA 134
E N VAD A+ +
Sbjct: 1335 EGNNVADRTARES 1347
>At1g10000 putative reverse transcriptase
Length = 303
Score = 40.0 bits (92), Expect = 6e-04
Identities = 31/90 (34%), Positives = 45/90 (49%), Gaps = 3/90 (3%)
Query: 43 CRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVR 102
CRR S PL AE WAI++A+ A+Q S +++ +DS A+ S L + +V
Sbjct: 198 CRRFPS--PLAAEAWAIKSAMLHALQLERSDLLVLSDSKSIVDALNSNVSLNEIFGLLV- 254
Query: 103 HIRRSIPHGMKVVFDLIPREANMVADCLAQ 132
IR + F IPR N +AD A+
Sbjct: 255 EIRSIRNRFRSISFQFIPRLVNSIADAAAK 284
>At1g26950 hypothetical protein
Length = 158
Score = 37.7 bits (86), Expect = 0.003
Identities = 37/141 (26%), Positives = 56/141 (39%)
Query: 29 GVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHAAVT 88
G LRD G+W F L LAELW + + +A ++G + + +E DS +T
Sbjct: 17 GALRDEYGDWRGSFALNLGRCTAPLAELWGVYYGLVIAWEKGITRLELEVDSKLVAGFLT 76
Query: 89 SAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQNAHVFNFGVHLLHHPL 148
+ S +VR V + REAN AD LA A G L
Sbjct: 77 TGIEDSHLLSFLVRLCYGFSSKDWIVRVSHVYREANRFADELANYAFSLLLGFLSFDSGL 136
Query: 149 MECRYLLYQDSMTVVASLSLP 169
++ +D+ V + +P
Sbjct: 137 PFVDSIMREDAAGVAGARHVP 157
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.141 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,883,160
Number of Sequences: 26719
Number of extensions: 139813
Number of successful extensions: 401
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 354
Number of HSP's gapped (non-prelim): 46
length of query: 169
length of database: 11,318,596
effective HSP length: 92
effective length of query: 77
effective length of database: 8,860,448
effective search space: 682254496
effective search space used: 682254496
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0108.3


