

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.13
(1262 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
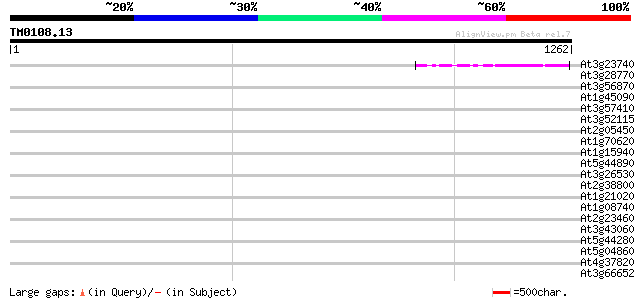
Score E
Sequences producing significant alignments: (bits) Value
At3g23740 unknown protein 152 2e-36
At3g28770 hypothetical protein 42 0.002
At3g56870 hypothetical protein 42 0.003
At1g45090 hypothetical protein 37 0.055
At3g57410 villin 3 37 0.094
At3g52115 gamma response I protein 35 0.21
At2g05450 hypothetical protein 35 0.21
At1g70620 unknown protein 34 0.46
At1g15940 T24D18.4 34 0.46
At5g44890 putative protein 33 0.79
At3g26530 hypothetical protein 33 0.79
At2g38800 unknown protein 33 0.79
At1g21020 hypothetical protein 33 0.79
At1g08740 hypothetical protein 33 0.79
At2g23460 extra-large GTP-binding protein (XLG1) 33 1.0
At3g43060 putative protein 33 1.4
At5g44280 putative protein 32 1.8
At5g04860 unknown protein 32 1.8
At4g37820 unknown protein 32 1.8
At3g66652 hypothetical protein 32 1.8
>At3g23740 unknown protein
Length = 542
Score = 152 bits (383), Expect = 2e-36
Identities = 117/350 (33%), Positives = 174/350 (49%), Gaps = 42/350 (12%)
Query: 914 RIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPISDSDLRATS 973
R K K GS Y+R+LP L + ++ N P QK ++ IS L + S
Sbjct: 216 RGKLFKTPGSVNYRRMLPYLKDIQEDN---------PYPQKNTEEV----ISSPMLNSES 262
Query: 974 GGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGFQSSNSKCKVMQMCDEQ 1033
+ V + T SGT + ++ P R P + K + +
Sbjct: 263 DNEGTQEVVTSNVTRESGTSSDE--------NEEPLPCERVPVNLEQSDPDKEQETQIKH 314
Query: 1034 VLLNGLCKPESSSGTPISVRRIDSPSTPLRPIVNEVMAREEATFALSESLINSKVKGNCS 1093
V+ + E++ G+ I + P+V + E + AL + +++ V G +
Sbjct: 315 VIPD----TENNLGSEIPLSS---------PLVGSRSSSEVNSSALHNTFVDNLV-GEEN 360
Query: 1094 FSISPDDKEKLS----EAHGCCLSLSQLELKEKLEVPA-VDFKKGILKRNPRGCRGLCTC 1148
+ + + K+S EAH + ++ L P+ KGILKR+ RGCRG+C+C
Sbjct: 361 MNGAEITEAKISAEELEAHSSDATAELVDPSVILATPSSFSPSKGILKRSMRGCRGICSC 420
Query: 1149 LNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPIDSVNDNPGFDRDQV 1208
LNC+SFRLHAERAFE+SRNQL D E + DL+ E+SHLRD+L + + + P + Q
Sbjct: 421 LNCSSFRLHAERAFEFSRNQLQDTEVMVLDLVGEISHLRDLLEKYNSADHSEP--YKSQA 478
Query: 1209 KEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKII 1258
EA ++A A +LAK RL M DDL IH RI N QR RV+FA ++ EK I
Sbjct: 479 GEASKRACEAAELAKSRLHQMNDDLQIHYRIPNEQRARVKFAHYIHEKTI 528
Score = 37.0 bits (84), Expect = 0.072
Identities = 44/154 (28%), Positives = 62/154 (39%), Gaps = 54/154 (35%)
Query: 493 DDFILTTPPDAKI------YDNSAVNVDRS----KSV--------------PQDMKRLKP 528
DDF TTPPD+++ + S VN + KSV P KRL P
Sbjct: 90 DDFAQTTPPDSELLAISEEINGSVVNKSDTNLWRKSVLLPCSRPKIFKNTGPFSYKRLLP 149
Query: 529 MDLRTNTQESFSKGVCLKADS------------------------RKDSLLKSKSVRRPH 564
++ + + S C K+ S R S LKS P+
Sbjct: 150 YLMQASDDGTSSSSRCSKSLSQNITKPVSQSMDSVYDKDSTGSFCRDTSPLKSVIASTPN 209
Query: 565 -----LPGKLFKTLGSVS-KRFLPFLMELAKDDP 592
GKLFKT GSV+ +R LP+L ++ +D+P
Sbjct: 210 KNAAFSRGKLFKTPGSVNYRRMLPYLKDIQEDNP 243
>At3g28770 hypothetical protein
Length = 2081
Score = 42.4 bits (98), Expect = 0.002
Identities = 190/1086 (17%), Positives = 384/1086 (34%), Gaps = 148/1086 (13%)
Query: 74 EKREPPEEDDGDTLLVKYLR----KKAKRDDDGGDLPRLLTKDLRARRVYSPQSSARASL 129
EKRE + ++G+++ + L KK +DD+ L +R + + +
Sbjct: 584 EKREETQGNNGESVKNENLENKEDKKELKDDESVGAKTNNETSLEEKR----EQTQKGHD 639
Query: 130 DLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSSAEKGGL 189
+ I+S D K G + S++EK VHV D + N + + K
Sbjct: 640 NSINSKIVDNK----GGNADSNKEKE-VHVGD---STNDNNMESK--------------- 676
Query: 190 CVDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDEEIVVTT 249
+D +V V + G S + + N DS + K+ E + + D++ V
Sbjct: 677 --EDTKSEVEVKKNDGSSEKGEEGKENNKDSMEDKKLENKES-----QTDSKDDKSVDDK 729
Query: 250 PPDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFST 309
+A++ G G+ ++D + GK + S E K +K N++ +++K +
Sbjct: 730 QEEAQIYG--------GESKDD---KSVEAKGKKKESKEN---KKTKTNENRVRNKEENV 775
Query: 310 AANHSFDLDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGG 369
N + G + ++ + V+ + T D ++ G +K++K
Sbjct: 776 QGNKKESEKVEKGEKKESK-DAKSVETKDNKKLSSTENRDE--AKERSGEDNKEDKEESK 832
Query: 370 QDFQLPLSSQSE----------KPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNER 419
+ ++E K +LK + E +K S + E ++ N++
Sbjct: 833 DYQSVEAKEKNENGGVDTNVGNKEDSKDLKDDRSV--EVKANKEESMKKKREE-VQRNDK 889
Query: 420 EVSGECLSVPSGDDKCFNESKCDPAADSDGVKVVGNALHNDDFMQCSIHKNNLDYAKVTN 479
+ E + D + + K GN N D + S + D K
Sbjct: 890 SSTKEVRDFANNMDIDVQKGSGESVKYKKDEKKEGNKEENKDTINTSSKQKGKDKKKKKK 949
Query: 480 DSGCTSV-------------QLGVLNDDFILTTPPD-AKIYDNSAVNVDRSKSVPQDMKR 525
+S +++ +L D+ TT + +K+ + + N ++ +S K
Sbjct: 950 ESKNSNMKKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDSASKN 1009
Query: 526 LKPMDL---RTNTQESFSKGVCLKADSRKDSL----LKSKSVRRPHLPGKLFKTLGSVSK 578
+ + ++ T+E K D +++ KSK + K K +
Sbjct: 1010 REKKEYEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKE 1069
Query: 579 RFLPFLMELAKDDPGISKFGHGTPKKDEADMHGKRFELTLSSQPEEASIDGHKIEKGPIH 638
+ + K + + + KK+E K+ E + S + EE D K+E
Sbjct: 1070 KKESENHKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRKKEEDKKDMEKLE----- 1124
Query: 639 GTVESNALGNILFNSANELSNGNLPKLTSSQDLPELPMQSDVKEVVQECLSAPSVEEPIE 698
+ N+ N NE KL + + +++ K +E S+ S + ++
Sbjct: 1125 ---DQNS--NKKKEDKNEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQKNEVD 1179
Query: 699 KAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGH---DVDIKHMQNSISSLQERGPSP 755
K S + ++ ++ + K+ N+ ++ + + +E+
Sbjct: 1180 KKEKKSSKDQQKKKEKEMKES----EEKKLKKNEEDRKKQTSVEENKKQKETKKEKNKPK 1235
Query: 756 KDQSMLYVNVGVSNQTLEHHSSEVKGFSVNNGEVKQLLNMEKRKYVSRYPSEGQS--SSQ 813
D+ G +++E S E + + + + K + + + S+ S SQ
Sbjct: 1236 DDKKNTTKQSGGKKESMESESKEAENQQKSQATTQADSDESKNEILMQADSQADSHSDSQ 1295
Query: 814 LDTDMQDLE---------ENERSNGAIRKSDDAVVLNHVGVSNEVPRPPPENIISKTQDM 864
D+D E +R+N RK +V N + + P++ K
Sbjct: 1296 ADSDESKNEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKD--DKKNTT 1353
Query: 865 AAHGGNKEGSVQKGIAYGSKGNNGRKGCYAYEIENASESKITPVLNRCPRIKSLKHAGSP 924
GG KE + SK ++ A ++ ESK ++ + S + +
Sbjct: 1354 KQSGGKKESMESE-----SKEAENQQKSQATTQADSDESKNEILMQADSQADSHSDSQAD 1408
Query: 925 TYKRLLPLLSNAKKYNSCASVNRHDPKHQ---------KFLDQTPLLPISDSDLRATSGG 975
+ + +L A + N D K Q K + P D G
Sbjct: 1409 SDESKNEILMQADSQATTQRNNEEDRKKQTSVAENKKQKETKEEKNKPKDDKKNTTEQSG 1468
Query: 976 DSNDSVPMEHFTANSGTKQQ--------KELNACELNDDSSSPRSRSPGFQSSNSKCKVM 1027
+S+ E A + K Q + N + DS + + S SK +++
Sbjct: 1469 GKKESMESESKEAENQQKSQATTQGESDESKNEILMQADSQADTHANSQGDSDESKNEIL 1528
Query: 1028 QMCDEQVLLNGLCKPESSSGTPISVRRIDSPS---TPLRPIVNEVMAREEATFALSESLI 1084
D Q + +S + + DS + T NE++ + ++ + ESL
Sbjct: 1529 MQADSQAD----SQTDSDESKNEILMQADSQADSQTDSDESKNEILMQADSQAKIGESLE 1584
Query: 1085 NSKVKG 1090
++KVKG
Sbjct: 1585 DNKVKG 1590
Score = 32.3 bits (72), Expect = 1.8
Identities = 87/421 (20%), Positives = 152/421 (35%), Gaps = 72/421 (17%)
Query: 206 ISSQARDFDRANTDSSDHKQ---FEEGNDGDCSGKINDFDEEIVVTTPPDAEL---LGDS 259
I QA + TDS + K + + D ++ EI++ A++ L D+
Sbjct: 1527 ILMQADSQADSQTDSDESKNEILMQADSQADSQTDSDESKNEILMQADSQAKIGESLEDN 1586
Query: 260 KVDGDEGKGEE-------DLLVR---DAPFTGKA-ENSGEGLSRKDSKRNDSVL-----K 303
KV G E G+E + V+ + GK EN G+ +S ++ ++ +++ K
Sbjct: 1587 KVKGKEDNGDEVGKENSKTIEVKGRHEESKDGKTNENGGKEVSTEEGSKDSNIVERNGGK 1646
Query: 304 SKYFSTAANHSFDLDLYAGARSQT---AFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHC 360
+ +++ G T + G+I +G G N TK+
Sbjct: 1647 EDSIKEGSEDGKTVEINGGEELSTEEGSKDGKIEEGKE----GKEN------STKEGSKD 1696
Query: 361 DKDEKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNERE 420
DK E+GM G++ SS+ K +E E+ TM E GSK + + + + E
Sbjct: 1697 DKIEEGMEGKENSTKESSKDGKINEIHGDKEA-TMEE--GSKDGGTNSTGKDSKDSKSVE 1753
Query: 421 VSGECLSVPSGDDKCFNESKCDPAADSDGVKVVGNALHNDDFMQCSIHKNNLDYAKVT-- 478
++G DD ++SK N D + + K + VT
Sbjct: 1754 ING------VKDDSLKDDSK------------------NGDINEINNGKEDSVKDNVTEI 1789
Query: 479 --NDSGCTSVQLGVLNDDFILTTPPDAKIYDNSAVNVDRSKSVPQDMKRLKPMDLRTNTQ 536
ND+ T+ N D + T K A S + + K ++ N Q
Sbjct: 1790 QGNDNSLTNSTSSEPNGDKLDTNKDSMKNNTMEAQGGSNGDSTNGETEETKESNVSMNNQ 1849
Query: 537 ESFSKGVCLKADSRKDSLLKSKSVRRPHLPGKLFKTLGSVSKRFLPFLMELAKDDPGISK 596
G S ++S+ + + T + SK F+ L + PG +
Sbjct: 1850 NMQDVG------SNENSMNNQTTGTGDDIISTTTDTESNTSKEVTSFISNLEEKSPGTQE 1903
Query: 597 F 597
F
Sbjct: 1904 F 1904
>At3g56870 hypothetical protein
Length = 670
Score = 41.6 bits (96), Expect = 0.003
Identities = 30/128 (23%), Positives = 61/128 (47%), Gaps = 6/128 (4%)
Query: 1136 KRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPID 1195
+RN + + S + +++A +S+ Q+ D + VA L KEL +R + R +
Sbjct: 510 RRNTQAAKAQPFSTGGTSIQGCSQKAIAFSQGQMRDFQNVAARLTKELKSMRQITKRCLQ 569
Query: 1196 SVNDNPGF---DRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQR---PRVRF 1249
+ ++ + D+VK A E+ K+ LS+++ D + C++ ++ R P
Sbjct: 570 AESNTSNMSDCNLDEVKTVIGNAEKTEESCKKWLSIIERDCNRFCKLMSMVREDSPATEN 629
Query: 1250 ADHVQEKI 1257
H ++KI
Sbjct: 630 IVHKKKKI 637
>At1g45090 hypothetical protein
Length = 1210
Score = 37.4 bits (85), Expect = 0.055
Identities = 95/418 (22%), Positives = 149/418 (34%), Gaps = 81/418 (19%)
Query: 2 APIPTLRESASAMEETLISRKR--SNPSSSPPGALTRSRSQLFVHRNRSGQRRPDSGPRQ 59
AP +R SA+ E ++ R+R S+P + ++ S G+ P G +
Sbjct: 382 APKKQVRPSATG-ERVILKRQRRVSDPVENKKRKISHSDDGEGGPSGGDGEGGPSGGDGE 440
Query: 60 GGLYSG--LGGSSGNDEKREPPEEDD-----------GDTLLVKY---LRKKAKRDDDGG 103
GG G GG SG D + P D GD KY LR KR
Sbjct: 441 GGPSGGDGEGGPSGGDGEGGPNGADGEGGPNGADGEVGDEAFDKYAGELRSFFKRQT--- 497
Query: 104 DLPRLLTKDLRARRVYSPQSSARASLDLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDV- 162
+LL K +V+ + RA+L DRK G P S KS + V
Sbjct: 498 ---QLLDKKFEDLKVFV-REEIRAALQ-----SQDRKEEG-----PRSSHKSDSPSEKVE 543
Query: 163 MIAVNSDCTDRKIADLGFGSSAEKGGLCVDDATMKVTVSPDLGI---SSQARDFDRANTD 219
+ + ++++ EK V ++K + P + SS+ + +
Sbjct: 544 RVTKTVEKVNKRVEKAN--KRVEKAAEQVQRKSVKKSTKPRKFVPRRSSRLNNTPKKAAT 601
Query: 220 SSDHKQFEEGNDGD----CSGKINDFDEEIVVTTPPDAELLGDSKVDGDEGKGEEDLLVR 275
S + GNDGD ++D+D E + L G S + +EG E
Sbjct: 602 GSLPVEEVPGNDGDGEVSAKQTVSDYDMENIY-------LGGASSTEEEEGGNSE----- 649
Query: 276 DAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAA-NHSFDLDLYAGARSQTAFPGQIV 334
E S+ +DSV++ + +H + + G + P I+
Sbjct: 650 ----------IEEDSSKLHQNVSDSVMEDNVDRVPSPSHQPEHGIPDGGNFEQLLPDPIL 699
Query: 335 QGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGG---QDFQLPLSSQSEKPSEPELK 389
P E H D +KG+GG ++ P S Q E+P PE+K
Sbjct: 700 SDPALEKMSDD--------VSSSSHQDV-QKGLGGNPEEEVVAPESQQLEEPQSPEVK 748
>At3g57410 villin 3
Length = 965
Score = 36.6 bits (83), Expect = 0.094
Identities = 43/170 (25%), Positives = 69/170 (40%), Gaps = 15/170 (8%)
Query: 967 SDLRATSGGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSP----GFQSSNS 1022
S ++SG S+ S + + G +Q+ E A + +SSP S+SP G S S
Sbjct: 768 SAFNSSSGRTSSPSRDRSN-GSQGGPRQRAEALAALTSAFNSSPSSKSPPRRSGLTSQAS 826
Query: 1023 KCKVMQMCDEQVLLNGLCKPESSSGTPISVRRIDSPSTPLRPIVNEVMAREEATFALSES 1082
+ QVL + + S T S D + +EV A EEAT A E
Sbjct: 827 QRAAAVAALSQVL---TAEKKKSPDTSPSAEAKDEKA------FSEVEATEEATEAKEEE 877
Query: 1083 LINSKVKGNCSFSISPDDKEKLSEAHGCCLSLSQLELKEKLEVPAVDFKK 1132
++ + + P + E G + +L+ K + V +DFK+
Sbjct: 878 EVSPAAEASAE-EAKPKQDDSEVETTGVTFTYERLQAKSEKPVTGIDFKR 926
>At3g52115 gamma response I protein
Length = 588
Score = 35.4 bits (80), Expect = 0.21
Identities = 31/95 (32%), Positives = 42/95 (43%), Gaps = 9/95 (9%)
Query: 867 HGGNKEGSVQKGIAYGSKGNNGRKGCYAYEIENASESKITPVLNRCPRIKSLKHAGSPTY 926
H EG+ + I GS GNN YE E+ ++ TP+ + P ++S A P
Sbjct: 378 HSETAEGADKVRIGTGSSGNN-------YEKESIIKTVQTPITSISPIVRSPGAAKPPQL 430
Query: 927 KRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPL 961
L S + S S HDP H FLD TP+
Sbjct: 431 SGLKRPASIWRDTRSRQSPGGHDP-HDDFLD-TPI 463
>At2g05450 hypothetical protein
Length = 1153
Score = 35.4 bits (80), Expect = 0.21
Identities = 43/195 (22%), Positives = 73/195 (37%), Gaps = 39/195 (20%)
Query: 229 GNDGD----CSGKINDFDEEIVVTTPPDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAE 284
GNDGD ++D+D + + L G S + +EG+ E
Sbjct: 584 GNDGDGEVSAKHTVSDYDMDNIY-------LGGASSTEQEEGRNSE-------------- 622
Query: 285 NSGEGLSRKDSKRNDSVLKSKYFSTAA-NHSFDLDLYAGARSQTAFPGQIVQGPGFEACG 343
E S+ +DSV++ + +H + + G + P I+ P E
Sbjct: 623 -IEEDSSKLHQNVSDSVMEDNVDHVPSPSHQPEQGIPDGGNFEELLPDPILSDPAIEKMS 681
Query: 344 GTNWPDNFVGTKKLGHCDKDEKGMGG---QDFQLPLSSQSEKPSEPELKSESFTMRETNG 400
H D KG+GG +D P S Q E+P PE+K+E+
Sbjct: 682 DD--------VSSSSHQDV-AKGLGGNREEDVVAPESQQLEEPQSPEVKAETICGDGEEA 732
Query: 401 SKLSSSQNLLESPME 415
K + S ++++ +E
Sbjct: 733 LKEAKSPSVVDEALE 747
>At1g70620 unknown protein
Length = 897
Score = 34.3 bits (77), Expect = 0.46
Identities = 61/254 (24%), Positives = 95/254 (37%), Gaps = 46/254 (18%)
Query: 204 LGISSQARDFDRANTD-SSDHKQFEEGNDGDCSGKINDFDEEIVVTTPPDAELLGDSKVD 262
LG++S A D D A+TD +SD E G + G ++ ++ PD E + +K+D
Sbjct: 528 LGLASYASDDDDADTDAASDANADENGVESLGVGSRHNVSQQPSTEKLPDPEAMASAKLD 587
Query: 263 GDEGKGEEDLLVRDAPFTGKAENSG-EGLSRK--DSKRNDSVLKSKYFSTAANHSFDLDL 319
G +GK SG E S+ ++++D +K +A+ D D
Sbjct: 588 PAVGVNAN---------SGKNSKSGLEDYSQMPGSTRKDDEAGSTKISDVSASSGLDDDT 638
Query: 320 YAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPLSSQ 379
+G+R + PD + K D+ G L
Sbjct: 639 -SGSRKE--------------------HPDR-TDSDKDAILDEPHVKNSGVKSDCNLRQD 676
Query: 380 SEKPSEPELKSESFTMR----ETNGSK-LSSSQNLLESPMELNEREVSGECLSVPSGDDK 434
S KP +L E T R ET G K SQN + M+ N+ + + + V S
Sbjct: 677 SNKPYGKDLSDEVSTDRSRIVETKGGKEKGDSQNDSKDRMKENDLKSAEKVKGVES---- 732
Query: 435 CFNESKCDPAADSD 448
N+ DP D
Sbjct: 733 --NKKSTDPHVKKD 744
>At1g15940 T24D18.4
Length = 1012
Score = 34.3 bits (77), Expect = 0.46
Identities = 90/381 (23%), Positives = 149/381 (38%), Gaps = 48/381 (12%)
Query: 355 KKLGHCDKDEKGMGG--QDFQLPLSSQSEK----PSEPELKSESFTMRETNGSKLSSSQN 408
KKL H D +K G D +P SS+S+K + P K + K + +
Sbjct: 522 KKLVHSDAKKKNSEGASMDTPIPQSSKSKKKDSRATTPATKKSEQAPKSHPKMKRIAGEE 581
Query: 409 LLESPMELNEREVSGECLSVPSGDDKCFNESKCDPAADSDGVKVVGNALHND-DFMQCSI 467
+ + EL E E+ G+ ++V DK F E VK + ++D D + ++
Sbjct: 582 VESNTNELGE-ELVGKRVNVWWPLDKKFYEGVIKSYC---RVKKMHQVTYSDGDVEELNL 637
Query: 468 HKNNLDYAKVTNDSGCTSVQLGVLNDDFILTTPPDAKIYDNS-------AVNVDRSKS-- 518
K K+ D S DD + +TP A I + NV+ S S
Sbjct: 638 KKERF---KIIEDKSSASED---KEDDLLESTPLSAFIQREKSKKRKIVSKNVEPSSSPE 691
Query: 519 VPQDMKRLKPMDLRTNT--QESFSKGVCLKADSRKDSLLKSKSVRR-PHLPGKLFKTLGS 575
V M+ +K D T++ Q +KG LKA S + K+++ L G+ KT G
Sbjct: 692 VRSSMQTMKKKDSVTDSIKQTKRTKG-ALKAVSNEPESTTGKNLKSLKKLNGEPDKTRGR 750
Query: 576 VSKRFLPFLMELAKDDPGISKFGHGTPKKDEADMHGKRFELTLSSQPEEASIDGHKIEKG 635
K+ K + + KDE D +L S E ++ H ++
Sbjct: 751 TGKKQKVTQAMHRKIEKDCDE-QEDLETKDEED----SLKLGKESDAEPDRMEDH--QEL 803
Query: 636 PIHGTVESNALGNILFNSANELSNGNLPKLTSSQDLPELPMQSDVKEVVQECLSAPSVEE 695
P + VE+ G E + + ++ + P ++D KE + L P+ E
Sbjct: 804 PENHNVETKTDG-----EEQEAAKEPTAESKTNGEEPNAEPETDGKE--HKSLKEPNAEP 856
Query: 696 PIEKAVVGSKDECSREPNFDL 716
+ G + E ++EPN +L
Sbjct: 857 KSD----GEEQEAAKEPNAEL 873
>At5g44890 putative protein
Length = 1015
Score = 33.5 bits (75), Expect = 0.79
Identities = 26/100 (26%), Positives = 47/100 (47%), Gaps = 9/100 (9%)
Query: 191 VDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDEEIVVTTP 250
VD+++ + +VS D +S +A + ++ +D + E DG I+D +
Sbjct: 527 VDNSSAEKSVSADKHVSPEAVNINKVGGVVTDTTEDESARDG-----IHDIGSSTLPAKE 581
Query: 251 PDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGL 290
D ++ + V GD+ +D ++ DAP G GEGL
Sbjct: 582 LD-DMFAEKSVSGDKTVSHQDNILVDAPGDG---IGGEGL 617
>At3g26530 hypothetical protein
Length = 1015
Score = 33.5 bits (75), Expect = 0.79
Identities = 26/100 (26%), Positives = 47/100 (47%), Gaps = 9/100 (9%)
Query: 191 VDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDEEIVVTTP 250
VD+++ + +VS D +S +A + ++ +D + E DG I+D +
Sbjct: 527 VDNSSAEKSVSADKHVSPEAVNINKVGGVVTDTTEDESARDG-----IHDIGSSTLPAKE 581
Query: 251 PDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGL 290
D ++ + V GD+ +D ++ DAP G GEGL
Sbjct: 582 LD-DMFAEKSVSGDKTVSHQDNILVDAPGDG---IGGEGL 617
>At2g38800 unknown protein
Length = 612
Score = 33.5 bits (75), Expect = 0.79
Identities = 29/108 (26%), Positives = 42/108 (38%), Gaps = 8/108 (7%)
Query: 219 DSSDHKQFEEGNDGDCSGKINDFDEEIVVTTPPDAELLGDSKVDGDEGKGE------EDL 272
D + K+FE GN G C I+ E V P +E D D E E E+
Sbjct: 230 DLEEKKEFENGNGGSCEVDIDSQISETVSEGAPRSETDSDDYSDSAEMVIELKESCLEET 289
Query: 273 LVRDA--PFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFDLD 318
LV D+ KA G+ K+S +++++ N D D
Sbjct: 290 LVDDSENEVQEKANRDGDTYLLKESDLEETLVEDSMNQDEGNRDGDAD 337
>At1g21020 hypothetical protein
Length = 751
Score = 33.5 bits (75), Expect = 0.79
Identities = 26/100 (26%), Positives = 47/100 (47%), Gaps = 9/100 (9%)
Query: 191 VDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDEEIVVTTP 250
VD+++ + +VS D +S +A + ++ +D + E DG I+D +
Sbjct: 527 VDNSSAEKSVSADKHVSPEAVNINKVGGVVTDTTEDESARDG-----IHDIGSSTLPAKE 581
Query: 251 PDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGL 290
D ++ + V GD+ +D ++ DAP G GEGL
Sbjct: 582 LD-DMFAEKSVSGDKTVSHQDNILVDAPGDG---IGGEGL 617
>At1g08740 hypothetical protein
Length = 950
Score = 33.5 bits (75), Expect = 0.79
Identities = 26/100 (26%), Positives = 47/100 (47%), Gaps = 9/100 (9%)
Query: 191 VDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDEEIVVTTP 250
VD+++ + +VS D +S +A + ++ +D + E DG I+D +
Sbjct: 526 VDNSSAEKSVSADKHVSPEAVNINKVGGVVTDTTEDESARDG-----IHDIGSSTLPAKE 580
Query: 251 PDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGL 290
D ++ + V GD+ +D ++ DAP G GEGL
Sbjct: 581 LD-DMFAEKSVSGDKTVSHQDNILVDAPGDG---IGGEGL 616
>At2g23460 extra-large GTP-binding protein (XLG1)
Length = 888
Score = 33.1 bits (74), Expect = 1.0
Identities = 22/83 (26%), Positives = 38/83 (45%), Gaps = 2/83 (2%)
Query: 354 TKKLGHCDKDEKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESP 413
T + H +++E+ GG LSS E ES + E++ + L ES
Sbjct: 97 TSVIEHTEEEEEEEGGDGEDCELSSSGELLLRSCSVKESLDLNESSSNPLVPDWESNESV 156
Query: 414 MELN--EREVSGECLSVPSGDDK 434
+ ++ V+G+C+S +GD K
Sbjct: 157 LSMDYPSSRVTGDCVSETNGDGK 179
>At3g43060 putative protein
Length = 585
Score = 32.7 bits (73), Expect = 1.4
Identities = 38/146 (26%), Positives = 58/146 (39%), Gaps = 31/146 (21%)
Query: 167 NSDCTDRKIADL-----GFGSSAEKGGLCVDDATMKVTVSPDLGISSQARDFDRANTDSS 221
N C +K ADL G G++ +K S D+G S + D+S
Sbjct: 409 NRGCGFQKEADLDDEMAGVGATGKKA-------------SDDIGGSGFGNE--EQEEDAS 453
Query: 222 DHKQFEEGNDGDCSGKINDFDEEIVVTTPPDAELLGD----------SKVDG-DEGKGEE 270
D + +E ND + +G + E++ T AE D KVD DE EE
Sbjct: 454 DEEGAKEENDDEMAGVEEEDAEQVDATDEEGAEGDNDEMAGVEEETSEKVDATDEEGAEE 513
Query: 271 DLLVRDAPFTGKAENSGEGLSRKDSK 296
+ R+ P G ++ EG+ D +
Sbjct: 514 EASNREGPEEGDDASNREGVEEGDDE 539
>At5g44280 putative protein
Length = 486
Score = 32.3 bits (72), Expect = 1.8
Identities = 27/103 (26%), Positives = 47/103 (45%), Gaps = 5/103 (4%)
Query: 202 PDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDEEIVVTTPPDAELLGDSKV 261
PD+ + R A D + + +E +GD K ++ +E+ V P DAE
Sbjct: 14 PDVADQPRDRFNPEATQDLQEKDETKEEKEGDEEVKHDEAEEDQEVVKPNDAE----EDD 69
Query: 262 DGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKS 304
DGD+ + +E+ V +A +AE E ++ + DS +S
Sbjct: 70 DGDDAEEDEEEEV-EAEEDEEAEEEEEEEEEEEEEEEDSKERS 111
>At5g04860 unknown protein
Length = 782
Score = 32.3 bits (72), Expect = 1.8
Identities = 25/103 (24%), Positives = 47/103 (45%), Gaps = 17/103 (16%)
Query: 754 SPKDQSMLYVNVGVSNQTLEHHSSEVKGFSVN----NGEV--KQLLNMEKRKYVSRYPSE 807
+P+D+ ++Y + H S EV V+ G++ K++L+ +KRK R P +
Sbjct: 313 NPEDEDLIYYSHRSPLAETGHCSDEVSNDVVSLEQAKGQMSKKRMLSWKKRKLSFRSPKQ 372
Query: 808 G-----------QSSSQLDTDMQDLEENERSNGAIRKSDDAVV 839
+ +D D + L ++ SN +SDDA++
Sbjct: 373 KGEPLLKKDCLEEGGDDIDFDRRQLSSSDESNSDWYRSDDAIM 415
>At4g37820 unknown protein
Length = 532
Score = 32.3 bits (72), Expect = 1.8
Identities = 35/148 (23%), Positives = 63/148 (41%), Gaps = 7/148 (4%)
Query: 689 SAPSVEEPIEKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNSISSL 748
S+ E I++ + K+E S + + ++ N + I+ ++++ SS
Sbjct: 380 SSSQEENEIKETEIKEKEESSSQEGNENKETEKKSSESQRKENTNSEKKIEQVESTDSSN 439
Query: 749 QERGPSPK-DQSMLYVNVGVSNQTLEHHSS----EVKGFSVNNGEVKQLLNMEKRKYVSR 803
++G K D+S SN+ E SS E K + NGE ++ N +++ +
Sbjct: 440 TQKGDEQKTDESKRESGNDTSNKETEDDSSKTESEKKEENNRNGETEETQNEQEQTKSAL 499
Query: 804 YPSEGQSSSQLDTDMQDLEENERSNGAI 831
S Q TD++ L E SNG I
Sbjct: 500 EISHTQDVKDARTDLETLPET--SNGLI 525
>At3g66652 hypothetical protein
Length = 980
Score = 32.3 bits (72), Expect = 1.8
Identities = 42/177 (23%), Positives = 68/177 (37%), Gaps = 24/177 (13%)
Query: 832 RKSDDAVVLNHVGVSNEVPRPP---PENIISKTQDMAAHG--GNKEGSVQKGIAYGSKGN 886
R D VV+ + V+N+V P PE T + A+ + GS YGS +
Sbjct: 220 RDPDPGVVIQ-IPVTNDVEELPVRTPEKARCITSNEASRSDVSHSYGSKDLNSVYGSPKD 278
Query: 887 NGRKGCYAYEIENASESKITPVLNRCPRIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVN 946
GC + S K P N C R +P+ K +L +K S + +
Sbjct: 279 EAFVGCQEENAGSFSGEKSLPTENCCSR------EATPSDKEML----EKEKEESVCNSD 328
Query: 947 RHDP----KHQKFLDQTPLLPISDSDLRATSGGDSNDSVPMEHFTANSGTKQQKELN 999
DP + D+ L P S S + D ++ ++ +S T Q+E++
Sbjct: 329 ETDPSSVERESSLGDRIRLSPTSSSSVGINEESDDYETESLK----DSATDDQREVS 381
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.131 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,577,825
Number of Sequences: 26719
Number of extensions: 1468287
Number of successful extensions: 3685
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 51
Number of HSP's that attempted gapping in prelim test: 3631
Number of HSP's gapped (non-prelim): 105
length of query: 1262
length of database: 11,318,596
effective HSP length: 111
effective length of query: 1151
effective length of database: 8,352,787
effective search space: 9614057837
effective search space used: 9614057837
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0108.13


