

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
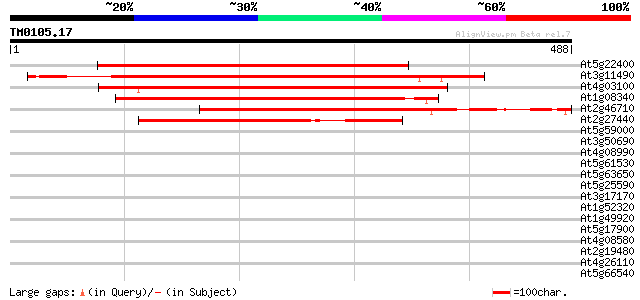
Query= TM0105.17
(488 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g22400 rac GTPase activating protein 387 e-108
At3g11490 putative rac GTPase activating protein 384 e-107
At4g03100 rac GTPase activating protein like 383 e-106
At1g08340 unknown protein 343 9e-95
At2g46710 putative rac GTPase activating protein 340 8e-94
At2g27440 putative rac GTPase activating protein 275 5e-74
At5g59000 unknown protein 41 0.001
At3g50690 putative protein 40 0.003
At4g08990 Met2-type cytosine DNA-methyltransferase-like protein 40 0.004
At5g61530 unknown protein 39 0.005
At5g63650 serine/threonine-protein kinase 39 0.006
At5g25590 unknown protein (At5g25590) 39 0.006
At3g17170 unknown protein 39 0.006
At1g52320 bZIP protein, putative 39 0.006
At1g49920 hypothetical protein 39 0.008
At5g17900 microfibril-associated protein - like 38 0.011
At4g08580 putative microfibril-associated protein 38 0.011
At2g19480 putative nucleosome assembly protein 38 0.011
At4g26110 nucleosome assembly protein I-like protein 38 0.014
At5g66540 unknown protein 37 0.019
>At5g22400 rac GTPase activating protein
Length = 466
Score = 387 bits (995), Expect = e-108
Identities = 186/271 (68%), Positives = 229/271 (83%)
Query: 77 GQSHNQNQNQFAILDILVAALKKSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGF 136
G+ + ++Q ++L +LVA ++SL++C +R ++ S++I WPT VRHV+HVTFDRFNGF
Sbjct: 78 GEEDDGGEDQISLLALLVAIFRRSLISCKSNRRELCSMEIGWPTNVRHVAHVTFDRFNGF 137
Query: 137 LGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKA 196
LGLP E +PEVP++ PSASA VFGVS +SMQ SYD RGN VPTILL+MQN LYS+GGL+A
Sbjct: 138 LGLPVEFEPEVPRRAPSASATVFGVSTESMQLSYDSRGNCVPTILLLMQNCLYSQGGLQA 197
Query: 197 EGIFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMH 256
EGIFR+ A+NS+EE VR QLNRG +P ++VHCL+GLIKAWFRELPT VLDSL+PEQVM
Sbjct: 198 EGIFRLTAENSEEEAVREQLNRGFIPERIDVHCLAGLIKAWFRELPTSVLDSLSPEQVMQ 257
Query: 257 CNSEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDP 316
C +EE+ LV+LLP TEAALLDWAINLMADVV++E NKMN+RNIAMVFAPNMTQM DP
Sbjct: 258 CQTEEENVELVRLLPPTEAALLDWAINLMADVVQYEHLNKMNSRNIAMVFAPNMTQMDDP 317
Query: 317 LTALIHAVQVMNFLKTLILKTLRERDESMAK 347
LTAL++AVQVMNFLKTLI KTLRER +S+ +
Sbjct: 318 LTALMYAVQVMNFLKTLIEKTLRERQDSVVE 348
>At3g11490 putative rac GTPase activating protein
Length = 435
Score = 384 bits (987), Expect = e-107
Identities = 211/408 (51%), Positives = 271/408 (65%), Gaps = 49/408 (12%)
Query: 16 NPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKGNW 75
N P SPF N+ E++E++EE E+E E +
Sbjct: 32 NRDPHSPF--NISRREEEEEEEERSEKERERFELS------------------------- 64
Query: 76 RGQSHNQNQNQFAILDILVAALKKSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNG 135
+ L+ILV+A+++S++ V ED+ S++I PT+VRHV+HVTFDRF+G
Sbjct: 65 ------------SALEILVSAIRRSVIGGCVGEEDLCSMEIGVPTDVRHVAHVTFDRFHG 112
Query: 136 FLGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLK 195
FLGLP E +PEVP++ PSASA VFGVS +SMQ SYD RGN VPTILLMMQ+ LYS GGL+
Sbjct: 113 FLGLPVEFEPEVPRRAPSASATVFGVSTESMQLSYDTRGNIVPTILLMMQSHLYSRGGLR 172
Query: 196 AEGIFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVM 255
EGIFRIN +N QEE++R +LN+G++P ++VHCL+ LIKAWFRELP+GVLDSL+PEQVM
Sbjct: 173 VEGIFRINGENGQEEYIREELNKGIIPDNIDVHCLASLIKAWFRELPSGVLDSLSPEQVM 232
Query: 256 HCNSEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVD 315
SE++C LV+LLPSTEA+LLDWAINLMADVVE EQ NKMNARNIAMVFAPNMTQM+D
Sbjct: 233 ESESEDECVELVRLLPSTEASLLDWAINLMADVVEMEQLNKMNARNIAMVFAPNMTQMLD 292
Query: 316 PLTALIHAVQVMNFLKTLILKTLRERDESMAKARQLSSLL------NSPSCKGDSHPFKD 369
PLTAL++AVQVMNFLKTLI+KTL++R ES K S+ + S + H K
Sbjct: 293 PLTALMYAVQVMNFLKTLIVKTLKDRKESRDKLVPASNPSPRDHNGDQSSSRQLLHLMKA 352
Query: 370 NREES----SAQPVDTCATMPPDKSEFSRMEWCVDEKVWSSEEKGTGG 413
N+EE+ A+ D + ++ E + VD K S +GG
Sbjct: 353 NKEETLDNFEAEMKDKEESADEEEEECAESVELVDIKKSSLVNNSSGG 400
>At4g03100 rac GTPase activating protein like
Length = 430
Score = 383 bits (984), Expect = e-106
Identities = 195/317 (61%), Positives = 243/317 (76%), Gaps = 13/317 (4%)
Query: 78 QSHNQNQNQFAILDILVAALKKSLVTCSVD-RED-----------VSSLDISWPTEVRHV 125
+ QNQ Q ++++ L+ AL+KS+V+C VD R+D V ++I WPT VRH+
Sbjct: 30 EEEEQNQQQLSLVEFLLTALRKSVVSCRVDNRQDDGGVGGGISSAVHHMEIGWPTNVRHI 89
Query: 126 SHVTFDRFNGFLGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQ 185
+HVTFDRF+GFLGLP ELQ E+P +VPSAS VFGVSA+SMQCSYDE+GNSVPTILL+MQ
Sbjct: 90 THVTFDRFHGFLGLPHELQVEIPCRVPSASVSVFGVSAESMQCSYDEKGNSVPTILLLMQ 149
Query: 186 NRLYSEGGLKAEGIFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGV 245
RLYS+ GLKAEGIFRIN +NSQEE VR QLNRG+VP ++VHCL+GLIKAWFRELP+GV
Sbjct: 150 ERLYSQQGLKAEGIFRINPENSQEEHVRDQLNRGIVPENIDVHCLAGLIKAWFRELPSGV 209
Query: 246 LDSLTPEQVMHCNSEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMV 305
LD L+PE+V++CN+E++ L+K L TE+ALL+WA++LMADVVE E+ NKMNARNIAMV
Sbjct: 210 LDGLSPEEVLNCNTEDESVELIKQLKPTESALLNWAVDLMADVVEEEESNKMNARNIAMV 269
Query: 306 FAPNMTQMVDPLTALIHAVQVMNFLKTLILKTLRERDESMAKARQLSSLLNSPS-CKGDS 364
FAPNMTQM DPLTAL+HAVQVMN LKTLI KTL ER+E+ + S +S S DS
Sbjct: 270 FAPNMTQMTDPLTALMHAVQVMNLLKTLITKTLAEREENATGSEGYSPSHSSNSQTDSDS 329
Query: 365 HPFKDNREESSAQPVDT 381
+D +Q D+
Sbjct: 330 DNAQDMEVSCESQATDS 346
>At1g08340 unknown protein
Length = 404
Score = 343 bits (881), Expect = 9e-95
Identities = 171/283 (60%), Positives = 217/283 (76%), Gaps = 9/283 (3%)
Query: 93 LVAALKKSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQKVP 152
+V L + T V+ D ++DI PT +RHV+HVTFDRF+GFLGLP+E +P+VP+K P
Sbjct: 53 VVLELARCFSTAEVEDNDRPAMDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAP 112
Query: 153 SASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFV 212
SASA VFGVS +SMQ SYD RGN VP ILL++Q+RLY +GGL+AEG+FRI +NS+EEFV
Sbjct: 113 SASATVFGVSTESMQLSYDSRGNCVPVILLLLQSRLYDQGGLQAEGVFRITGENSEEEFV 172
Query: 213 RCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPS 272
R QLN+G++P G++VHCL+GLIKAWFRELP GVLD L EQVM C S+ED +V+LLP
Sbjct: 173 REQLNKGIIPDGIDVHCLAGLIKAWFRELPRGVLDPLPSEQVMQCESDEDFVKVVRLLPQ 232
Query: 273 TEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKT 332
TEA+LL+WAINLMADV++ E NKMN+RN+A+VFAPNM+QM DPLTAL++AVQVM LK+
Sbjct: 233 TEASLLNWAINLMADVIQFEHVNKMNSRNLALVFAPNMSQMADPLTALMYAVQVMKLLKS 292
Query: 333 LILKTLRERDESMAKARQLSSLLNSPSCK--GDSHPFKDNREE 373
L KT+RER+ S SS+++ K D KDN EE
Sbjct: 293 LTEKTVREREAS-------SSVVDRRCSKEAEDGEKEKDNEEE 328
Score = 38.5 bits (88), Expect = 0.008
Identities = 19/48 (39%), Positives = 29/48 (59%)
Query: 6 RSKSCGLVEFNPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
R S +V+ S + KD E++E+DEEEEEEEE+ D+E++
Sbjct: 301 REASSSVVDRRCSKEAEDGEKEKDNEEEEEDEEEEEEEEDEDEDEEEE 348
Score = 32.0 bits (71), Expect = 0.79
Identities = 13/28 (46%), Positives = 23/28 (81%), Gaps = 2/28 (7%)
Query: 26 NVKDEEDDEDDEEEEEEEEELYNDDEDD 53
N ++EED+E++EEEE+E+E+ ++E D
Sbjct: 325 NEEEEEDEEEEEEEEDEDED--EEEEGD 350
Score = 28.5 bits (62), Expect = 8.7
Identities = 10/26 (38%), Positives = 21/26 (80%)
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++++DE+EEEE + +Y E++
Sbjct: 334 EEEEEEDEDEDEEEEGDGVYIIKEEE 359
>At2g46710 putative rac GTPase activating protein
Length = 301
Score = 340 bits (873), Expect = 8e-94
Identities = 190/331 (57%), Positives = 223/331 (66%), Gaps = 46/331 (13%)
Query: 166 MQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLNRGLVPRGV 225
MQCSYD+RGNSVPTILL MQ RLY+EGGLKAEGIFRIN DN +EE VR QLN G+VPRG+
Sbjct: 1 MQCSYDDRGNSVPTILLRMQKRLYTEGGLKAEGIFRINPDNGKEEHVRRQLNCGVVPRGI 60
Query: 226 EVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPSTEAALLDWAINLM 285
+VHCL+GLIKAWFRELPTGVLD LTPEQVM CN+EEDC+ LV LLP E+A+LDWAI LM
Sbjct: 61 DVHCLAGLIKAWFRELPTGVLDVLTPEQVMRCNTEEDCSRLVILLPPVESAILDWAIGLM 120
Query: 286 ADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTLRERDESM 345
ADVVEHEQFNKMNARN+AMVFAPNMTQM DPLTALIHAVQVMNFLKTLIL L+ER+ +
Sbjct: 121 ADVVEHEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILMNLKERENAD 180
Query: 346 AKARQLSSLLNSPSCKGDSH------PFKDNREESSAQPVDTCATMPPDKSEFSRMEWCV 399
AKAR L + PS + +S P K N V T + D
Sbjct: 181 AKARWLKKQTSDPSEEWESQHSEILSPEKPNNNNPKFLRVATLCRLEADN---------- 230
Query: 400 DEKVWSSEEKGTGGGALESVSGGSSPSRYESGPLESRYRGIYDSEHWLRLRKGVRRLCQH 459
+E+ W+ +++ G L++ SG + GP V+RLC+H
Sbjct: 231 EEEFWNIKKRNDHEGVLDTSSGNGN-----IGP--------------------VQRLCKH 265
Query: 460 PVFQLSKSTKKRADLGIVNTREG--GGEAWA 488
P+FQLSKSTKK + N EG G EAW+
Sbjct: 266 PLFQLSKSTKKAF---VSNRDEGRKGREAWS 293
>At2g27440 putative rac GTPase activating protein
Length = 368
Score = 275 bits (702), Expect = 5e-74
Identities = 137/229 (59%), Positives = 172/229 (74%), Gaps = 24/229 (10%)
Query: 113 SLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDE 172
++DIS PT + HV+HVT+DRF+GFLGLP+E +P+VP+K PSASA VFGVS +SMQ SYD
Sbjct: 86 AMDISRPTNISHVAHVTYDRFDGFLGLPSEFEPDVPKKPPSASATVFGVSTESMQLSYDS 145
Query: 173 RGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSG 232
RGN VPTIL ++Q+RLY +GGL+ EGIFRI DNS+EEF+R +LN+G++P G+++HCL+G
Sbjct: 146 RGNCVPTILTLLQSRLYDQGGLQVEGIFRITGDNSEEEFIREELNKGVLPEGIDIHCLAG 205
Query: 233 LIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPSTEAALLDWAINLMADVVEHE 292
LIKAWFRELP GVLDSL +QVM C S ED VK+ E
Sbjct: 206 LIKAWFRELPKGVLDSLPSQQVMQCESGED---FVKVF---------------------E 241
Query: 293 QFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTLRER 341
NKM +RN+A+VFAPNM+QM DPLTAL++AVQVMN L+ L KTLRER
Sbjct: 242 VVNKMTSRNLALVFAPNMSQMADPLTALMYAVQVMNLLRNLTDKTLRER 290
>At5g59000 unknown protein
Length = 231
Score = 41.2 bits (95), Expect = 0.001
Identities = 15/32 (46%), Positives = 26/32 (80%)
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDYWSNPV 59
++EEDD DD+E+EE+EEE ++E++Y+ P+
Sbjct: 99 EEEEDDSDDDEDEEDEEEEEEEEEEEYYGLPL 130
Score = 28.5 bits (62), Expect = 8.7
Identities = 11/23 (47%), Positives = 19/23 (81%)
Query: 31 EDDEDDEEEEEEEEELYNDDEDD 53
E +++ E+E+EEEE+ +DDED+
Sbjct: 89 ETNKEAEDEDEEEEDDSDDDEDE 111
>At3g50690 putative protein
Length = 447
Score = 40.0 bits (92), Expect = 0.003
Identities = 18/34 (52%), Positives = 23/34 (66%)
Query: 26 NVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPV 59
N + E DDEDDEE+EE+EEE DE+D S +
Sbjct: 159 NERPESDDEDDEEDEEDEEEEEEGDEEDPGSGEI 192
Score = 35.0 bits (79), Expect = 0.093
Identities = 15/25 (60%), Positives = 20/25 (80%)
Query: 26 NVKDEEDDEDDEEEEEEEEELYNDD 50
N +D DDE+D+EE+EEEEE+ N D
Sbjct: 288 NEEDGVDDEEDDEEDEEEEEVDNAD 312
Score = 31.6 bits (70), Expect = 1.0
Identities = 18/51 (35%), Positives = 28/51 (54%), Gaps = 4/51 (7%)
Query: 25 HNVKDEEDDED----DEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKD 71
H ++D E++ED +E++EE+EEE D+ D ST + AG D
Sbjct: 281 HEIEDSENEEDGVDDEEDDEEDEEEEEVDNADRGLGGSGSTSRLMNAGEID 331
Score = 30.4 bits (67), Expect = 2.3
Identities = 22/66 (33%), Positives = 34/66 (51%), Gaps = 13/66 (19%)
Query: 1 MTRL--FRSKSCGLV-----------EFNPSPPSPFFHNVKDEEDDEDDEEEEEEEEELY 47
+TRL +RS+ GL+ E N P S + +DEED+E++EE +EE+
Sbjct: 131 VTRLKDYRSRVFGLIKTLKYLDKTDAEGNERPESDDEDDEEDEEDEEEEEEGDEEDPGSG 190
Query: 48 NDDEDD 53
D D+
Sbjct: 191 EIDGDE 196
>At4g08990 Met2-type cytosine DNA-methyltransferase-like protein
Length = 1512
Score = 39.7 bits (91), Expect = 0.004
Identities = 21/65 (32%), Positives = 37/65 (56%), Gaps = 4/65 (6%)
Query: 15 FNPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKD-KG 73
++P PS H V++EE +ED+EE+E EE+++ +E+ TP + S+D +
Sbjct: 626 YSPEVPSEAIHEVEEEEIEEDEEEDENEEDDI---EEEAVEVQKSHTPKKSRGNSEDMEI 682
Query: 74 NWRGQ 78
W G+
Sbjct: 683 KWNGE 687
>At5g61530 unknown protein
Length = 376
Score = 39.3 bits (90), Expect = 0.005
Identities = 37/159 (23%), Positives = 68/159 (42%), Gaps = 6/159 (3%)
Query: 154 ASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVR 213
AS+ VFGV+ + + E +P IL+ + L G L + +F+ D + +
Sbjct: 132 ASSDVFGVAIE-ITVQRQESSRPIPLILVKCADYLILTG-LNSPNLFKAEGDRKLIQQLV 189
Query: 214 CQLN---RGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSE-EDCTNLVKL 269
N R +P GV ++ L+K + LPT + ++ S ++
Sbjct: 190 SAYNQDPRASIPEGVNPVDVAALLKYYLASLPTPLTTFELYNEIKDARSSIHRMRQSLQK 249
Query: 270 LPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAP 308
L + L++ L+ V + NKM++ ++AM AP
Sbjct: 250 LSNVNYNTLEFITALLLRVSQKSLLNKMDSHSLAMEMAP 288
>At5g63650 serine/threonine-protein kinase
Length = 360
Score = 38.9 bits (89), Expect = 0.006
Identities = 17/38 (44%), Positives = 25/38 (65%)
Query: 16 NPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
NP+P S D+E+D +DE EEEEEEE ++E++
Sbjct: 301 NPAPSSNAVKGFDDDEEDVEDEVEEEEEEEEEEEEEEE 338
>At5g25590 unknown protein (At5g25590)
Length = 775
Score = 38.9 bits (89), Expect = 0.006
Identities = 21/43 (48%), Positives = 28/43 (64%), Gaps = 1/43 (2%)
Query: 4 LFRSKS-CGLVEFNPSPPSPFFHNVKDEEDDEDDEEEEEEEEE 45
++R KS G V P +P ++EED+E+DEEEEEEEEE
Sbjct: 246 IYRKKSGSGKVVEEMEPKTPEKVEEEEEEDEEEDEEEEEEEEE 288
>At3g17170 unknown protein
Length = 314
Score = 38.9 bits (89), Expect = 0.006
Identities = 20/38 (52%), Positives = 28/38 (73%), Gaps = 3/38 (7%)
Query: 19 PPSPFFHNVK--DEEDDEDDEEE-EEEEEELYNDDEDD 53
PP P FH+V+ DE D+D+EEE EE+E+E +DE+D
Sbjct: 220 PPPPEFHSVRAGDEYYDDDEEEEIEEDEDEGEGEDEED 257
>At1g52320 bZIP protein, putative
Length = 798
Score = 38.9 bits (89), Expect = 0.006
Identities = 16/29 (55%), Positives = 22/29 (75%)
Query: 17 PSPPSPFFHNVKDEEDDEDDEEEEEEEEE 45
P PP P V ++++DE++EEEEEEEEE
Sbjct: 266 PVPPQPHSPVVTEDDEDEEEEEEEEEEEE 294
>At1g49920 hypothetical protein
Length = 785
Score = 38.5 bits (88), Expect = 0.008
Identities = 13/27 (48%), Positives = 23/27 (85%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYW 55
DEE+++DD +++EE+++ +DDEDD W
Sbjct: 759 DEEEEDDDVDDDEEDDDDVDDDEDDEW 785
Score = 35.0 bits (79), Expect = 0.093
Identities = 12/24 (50%), Positives = 22/24 (91%)
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
EED++ D++EEEE++++ +D+EDD
Sbjct: 751 EEDEDGDDDEEEEDDDVDDDEEDD 774
>At5g17900 microfibril-associated protein - like
Length = 435
Score = 38.1 bits (87), Expect = 0.011
Identities = 18/43 (41%), Positives = 24/43 (54%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKD 71
+EED+ +EEEEEEE E D EDD + P+ P +D
Sbjct: 150 EEEDEIQEEEEEEEESEYETDSEDDMPGIAMIKPVFVPKAERD 192
>At4g08580 putative microfibril-associated protein
Length = 435
Score = 38.1 bits (87), Expect = 0.011
Identities = 18/43 (41%), Positives = 24/43 (54%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKD 71
+EED+ +EEEEEEE E D EDD + P+ P +D
Sbjct: 150 EEEDEIQEEEEEEEESEYETDSEDDMPGIALIKPVFVPKAERD 192
>At2g19480 putative nucleosome assembly protein
Length = 379
Score = 38.1 bits (87), Expect = 0.011
Identities = 19/45 (42%), Positives = 29/45 (64%), Gaps = 2/45 (4%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKG 73
DEED+EDDE++EEE++E +DDE++ + + AG K G
Sbjct: 318 DEEDEEDDEDDEEEDDE--DDDEEEEADQGKKSKKKSSAGHKKAG 360
Score = 38.1 bits (87), Expect = 0.011
Identities = 16/25 (64%), Positives = 22/25 (88%), Gaps = 2/25 (8%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE+DDE+DEE++E++EE DDEDD
Sbjct: 314 DEDDDEEDEEDDEDDEE--EDDEDD 336
Score = 33.9 bits (76), Expect = 0.21
Identities = 11/25 (44%), Positives = 23/25 (92%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
D+E+DE+D+E++EEE++ +D+E++
Sbjct: 317 DDEEDEEDDEDDEEEDDEDDDEEEE 341
Score = 33.5 bits (75), Expect = 0.27
Identities = 12/25 (48%), Positives = 21/25 (84%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE D++DDEE+EE++E+ +D++D
Sbjct: 311 DEIDEDDDEEDEEDDEDDEEEDDED 335
Score = 33.1 bits (74), Expect = 0.36
Identities = 10/28 (35%), Positives = 25/28 (88%)
Query: 26 NVKDEEDDEDDEEEEEEEEELYNDDEDD 53
+++D++D+ D++++EE+EE+ +D+E+D
Sbjct: 305 DIEDDDDEIDEDDDEEDEEDDEDDEEED 332
Score = 32.7 bits (73), Expect = 0.46
Identities = 10/24 (41%), Positives = 22/24 (91%)
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
+EDD++++EE++E++E +D++DD
Sbjct: 314 DEDDDEEDEEDDEDDEEEDDEDDD 337
>At4g26110 nucleosome assembly protein I-like protein
Length = 372
Score = 37.7 bits (86), Expect = 0.014
Identities = 22/46 (47%), Positives = 29/46 (62%), Gaps = 2/46 (4%)
Query: 29 DEEDDED-DEEEEEEEEELYNDDEDDYWSNPVSTPLI-NPAGSKDK 72
DEEDD D DE+EE+EE+E +DDED+ S P I N G + +
Sbjct: 310 DEEDDIDEDEDEEDEEDEEDDDDEDEEESKTKKKPSIGNKKGGRSQ 355
Score = 35.4 bits (80), Expect = 0.072
Identities = 13/27 (48%), Positives = 22/27 (81%)
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDD 53
+ D+E+D+ DE+E+EE+EE DD+D+
Sbjct: 307 IDDDEEDDIDEDEDEEDEEDEEDDDDE 333
Score = 31.6 bits (70), Expect = 1.0
Identities = 14/44 (31%), Positives = 26/44 (58%)
Query: 30 EEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKG 73
++D+EDD +E+E+EE+ ++++DD S P+ KG
Sbjct: 308 DDDEEDDIDEDEDEEDEEDEEDDDDEDEEESKTKKKPSIGNKKG 351
Score = 31.2 bits (69), Expect = 1.3
Identities = 14/45 (31%), Positives = 26/45 (57%)
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKG 73
D+++++D +E+E+EE+E +D+DD T G+K G
Sbjct: 308 DDDEEDDIDEDEDEEDEEDEEDDDDEDEEESKTKKKPSIGNKKGG 352
Score = 31.2 bits (69), Expect = 1.3
Identities = 11/18 (61%), Positives = 17/18 (94%)
Query: 28 KDEEDDEDDEEEEEEEEE 45
+DEED+EDD++E+EEE +
Sbjct: 322 EDEEDEEDDDDEDEEESK 339
Score = 30.8 bits (68), Expect = 1.8
Identities = 10/18 (55%), Positives = 18/18 (99%)
Query: 28 KDEEDDEDDEEEEEEEEE 45
+DEED+ED+E++++E+EE
Sbjct: 319 EDEEDEEDEEDDDDEDEE 336
>At5g66540 unknown protein
Length = 524
Score = 37.4 bits (85), Expect = 0.019
Identities = 15/25 (60%), Positives = 22/25 (88%)
Query: 28 KDEEDDEDDEEEEEEEEELYNDDED 52
+DEE++E+DEEEEEEEEE +++D
Sbjct: 139 EDEEEEEEDEEEEEEEEEEEEEEKD 163
Score = 35.4 bits (80), Expect = 0.072
Identities = 14/26 (53%), Positives = 22/26 (83%)
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE+DE++EEEEEEEEE D +++
Sbjct: 142 EEEEEDEEEEEEEEEEEEEEKDGDNE 167
Score = 34.3 bits (77), Expect = 0.16
Identities = 22/78 (28%), Positives = 40/78 (51%), Gaps = 5/78 (6%)
Query: 13 VEFNPSPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDD-EDDYWSNPVSTPLINPAGSKD 71
+E N S +DEE++E++EEEEEEE++ N+ ED ++ + +++
Sbjct: 131 IESNDSEGEDEEEEEEDEEEEEEEEEEEEEEKDGDNEGIEDKFFKIKELEEFLEEGEAEE 190
Query: 72 KG----NWRGQSHNQNQN 85
G N +G + + QN
Sbjct: 191 YGIDHKNKKGVAQRKKQN 208
Score = 31.2 bits (69), Expect = 1.3
Identities = 11/19 (57%), Positives = 17/19 (88%)
Query: 26 NVKDEEDDEDDEEEEEEEE 44
N+ D+ED+EDD++EEE+ E
Sbjct: 208 NLSDDEDEEDDDDEEEDVE 226
Score = 29.3 bits (64), Expect = 5.1
Identities = 12/25 (48%), Positives = 18/25 (72%)
Query: 28 KDEEDDEDDEEEEEEEEELYNDDED 52
+DEEDD+D+EE+ E + DDE+
Sbjct: 213 EDEEDDDDEEEDVEFDAFAGGDDEE 237
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,331,016
Number of Sequences: 26719
Number of extensions: 613376
Number of successful extensions: 11001
Number of sequences better than 10.0: 336
Number of HSP's better than 10.0 without gapping: 270
Number of HSP's successfully gapped in prelim test: 72
Number of HSP's that attempted gapping in prelim test: 5797
Number of HSP's gapped (non-prelim): 2306
length of query: 488
length of database: 11,318,596
effective HSP length: 103
effective length of query: 385
effective length of database: 8,566,539
effective search space: 3298117515
effective search space used: 3298117515
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0105.17


