

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0102b.4
(278 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
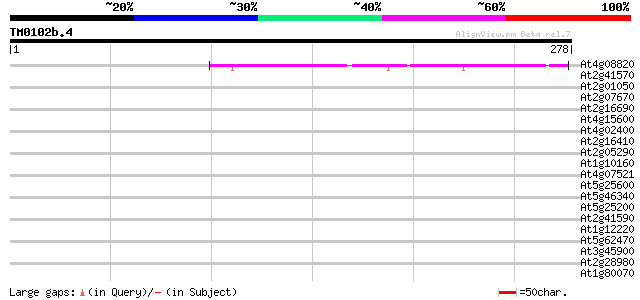
Score E
Sequences producing significant alignments: (bits) Value
At4g08820 putative protein 42 3e-04
At2g41570 putative Ta11-like non-LTR retroelement protein 36 0.020
At2g01050 hypothetical protein 34 0.076
At2g07670 putative non-LTR retrolelement reverse transcriptase 33 0.13
At2g16690 putative Ta11-like non-LTR retroelement protein 32 0.38
At4g15600 hypothetical protein 32 0.49
At4g02400 32 0.49
At2g16410 pseudogene 30 1.1
At2g05290 putative Ta11-like non-LTR retroelement protein 30 1.4
At1g10160 putative reverse transcriptase 30 1.4
At4g07521 putative protein 30 1.9
At5g25600 putative protein 29 2.4
At5g46340 unknown protein 28 4.2
At5g25200 putative protein 28 4.2
At2g41590 putative Ta11-like non-LTR retroelement protein 28 4.2
At1g12220 NBS/LRR disease resistance protein, putative 28 4.2
At5g62470 MYB96 transcription factor-like protein 28 5.4
At3g45900 unknown protein 28 5.4
At2g28980 putative non-LTR retroelement reverse transcriptase 28 5.4
At1g80070 hypothetical protein 28 5.4
>At4g08820 putative protein
Length = 404
Score = 42.4 bits (98), Expect = 3e-04
Identities = 44/195 (22%), Positives = 83/195 (42%), Gaps = 21/195 (10%)
Query: 100 LKPWSSSFAP-----INRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
++ WS +F P + WVR T IP++ + + + +G ++K+D +T F
Sbjct: 94 VQDWSPNFDPLRDDIVTTPVWVRLTNIPVNYYHRCLLEEIARGLGKLLKVDLNTITFGRG 153
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTISI--IEEGEALLHPNCCCREESSVEDDVFS 212
F+R+ + L +LI G Y ++ +EEG ++ + R E + VF+
Sbjct: 154 RFARVCIEVNLAK--PLKGTVLINGDRYFVAYEEVEEGFTVVRQS-GIRVEKPAQKMVFA 210
Query: 213 GWSLNGVTVTP----------ELSESGGEDDDGLWPVETKSQHDLQGSPNHSLGDNKNPR 262
S G + E+S G ++ + E L+GS + KN R
Sbjct: 211 AGSSGGRSNRSLRELPKNQGVEISNRFGGLEEDMVSAEIGEVAILEGSNKENKYKGKNMR 270
Query: 263 SNTGAVNRLLSSRLN 277
+G V ++ +++N
Sbjct: 271 KESG-VTQVKEAQVN 284
>At2g41570 putative Ta11-like non-LTR retroelement protein
Length = 418
Score = 36.2 bits (82), Expect = 0.020
Identities = 33/154 (21%), Positives = 66/154 (42%), Gaps = 12/154 (7%)
Query: 67 LRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKPWSSS-------FAPINRVTWVRCT 119
L +R E +L+E+L+ + E+ ++ W F P+ W+R
Sbjct: 72 LSQERFQFFFKSEDDLLEILKTGVWTQDEWCVVMERWIEKSTEEYLMFLPV----WMRLR 127
Query: 120 GIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARKILIKG 179
IP++ +T D K + VG V+K++ D + ++ R+ V N + ++I +
Sbjct: 128 NIPVNYYTQDTIKKIASCVGKVLKVELDLEKSQAQDYIRVQVIIDVRNPLRNFKEIQLP- 186
Query: 180 TTYTISIIEEGEALLHPNCCCREESSVEDDVFSG 213
T +S+ + E + C+ + + D SG
Sbjct: 187 TGEIVSVTFDYERIRKRCFLCQRLTHEKGDCDSG 220
>At2g01050 hypothetical protein
Length = 515
Score = 34.3 bits (77), Expect = 0.076
Identities = 25/104 (24%), Positives = 45/104 (43%), Gaps = 9/104 (8%)
Query: 100 LKPWSSSFAP-----INRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
++ WSS F P + WVR + IP + + + +G +K+D +T F
Sbjct: 147 VQDWSSRFDPLRDDIVTTPVWVRLSNIPYNYYHRCLLMEIARGLGRPLKVDMNTINFDKG 206
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNC 198
F+R+ + L +LI G Y ++ EG + + +C
Sbjct: 207 RFARVCIEVNLAK--PLKGTVLINGDRYFVAY--EGLSKICSSC 246
>At2g07670 putative non-LTR retrolelement reverse transcriptase
Length = 913
Score = 33.5 bits (75), Expect = 0.13
Identities = 26/104 (25%), Positives = 44/104 (42%), Gaps = 9/104 (8%)
Query: 100 LKPWSSSFAPI-NRVT----WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
++ WS F P+ + +T WVR IPL ++ + +G +K+D T
Sbjct: 122 VQAWSPEFDPLRDEITTTPIWVRLMNIPLSLYHTSILMGIAGSLGKPVKVDMTTLHVERA 181
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNC 198
F+R+ + L +L+ G Y +S EG A + C
Sbjct: 182 RFARMCIEVDLAK--PLKGTLLLNGERYFVSY--EGLANICSRC 221
>At2g16690 putative Ta11-like non-LTR retroelement protein
Length = 240
Score = 32.0 bits (71), Expect = 0.38
Identities = 14/63 (22%), Positives = 32/63 (50%)
Query: 115 WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARK 174
W+R IP++ +T + K + VG V+K++ D + ++ R+ V N + ++
Sbjct: 123 WIRLRNIPVNYYTEETIKKIAGCVGQVVKVELDLEKSQAQDYVRVQVNLDVRNPLRNSKS 182
Query: 175 ILI 177
+ +
Sbjct: 183 VQV 185
>At4g15600 hypothetical protein
Length = 655
Score = 31.6 bits (70), Expect = 0.49
Identities = 22/91 (24%), Positives = 38/91 (41%), Gaps = 7/91 (7%)
Query: 100 LKPWSSSFAPINRV-----TWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
++ WS F P+ V WVR +P+ + + + +G IK+D T
Sbjct: 162 VQAWSPEFNPLRDVIETTPVWVRVANLPVTFYHNEILLGIAAGLGKPIKVDLTTLRKERG 221
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTIS 185
F+R+ V N L +++ G Y +S
Sbjct: 222 RFARVCVEVNLKN--PLKGTLVVNGERYFVS 250
>At4g02400
Length = 870
Score = 31.6 bits (70), Expect = 0.49
Identities = 19/60 (31%), Positives = 30/60 (49%), Gaps = 4/60 (6%)
Query: 212 SGWSLNGVTVTPELSESGGEDDDGLWPVETKSQHDLQGSPNHSLG---DNKNPRSNTGAV 268
SG + G T E + +D D + + S +D++G N+ LG D +P NTGA+
Sbjct: 515 SGRRVFGATSKVEAPKESKKDSDNFYD-NSDSDNDMEGIENNDLGAVGDTASPARNTGAI 573
>At2g16410 pseudogene
Length = 204
Score = 30.4 bits (67), Expect = 1.1
Identities = 19/82 (23%), Positives = 34/82 (41%), Gaps = 3/82 (3%)
Query: 67 LRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKPWSSSFA---PINRVTWVRCTGIPL 123
L D R E++L ++L S++ L+ W S P W++ GIP
Sbjct: 83 LGDSRFQFFFESETDLQKVLNKRPCHFSKWSFALERWKSHIGISFPDTMTFWIKTEGIPT 142
Query: 124 HMWTMDCFKHLLLQVGDVIKID 145
W + ++ +G V ++D
Sbjct: 143 EFWEEEVLRNFGASIGAVRRVD 164
>At2g05290 putative Ta11-like non-LTR retroelement protein
Length = 383
Score = 30.0 bits (66), Expect = 1.4
Identities = 22/98 (22%), Positives = 43/98 (43%), Gaps = 3/98 (3%)
Query: 67 LRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKPWSSSFAP---INRVTWVRCTGIPL 123
L DR E +L E+L+ + ++ ++ W P + + W+R IP+
Sbjct: 74 LSRDRFQFIFKYEEDLQEVLKTGVWTQDDWGVVMERWVEDLPPNYLMFLLIWLRLRNIPV 133
Query: 124 HMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLV 161
+ +T K + VG VI+ D +E ++ R+ +
Sbjct: 134 NHYTQATIKSIAKCVGQVIEFPFDENEAQSKDYVRIRI 171
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 30.0 bits (66), Expect = 1.4
Identities = 12/25 (48%), Positives = 18/25 (72%)
Query: 77 HQESNLVELLQGSKGWRSEFVDNLK 101
HQ +NL++ L+G G+R E VD +K
Sbjct: 377 HQATNLIKFLRGDDGFRVENVDQIK 401
>At4g07521 putative protein
Length = 223
Score = 29.6 bits (65), Expect = 1.9
Identities = 24/101 (23%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 92 WRSEFVDNL--KPWSSSFAPINRV-----TWVRCTGIPLHMWTMDCFKHLLLQVGDVIKI 144
WR F NL + W F P+ V W+R T +P++ + + +G K+
Sbjct: 27 WRV-FGSNLTVQAWLPEFDPLRDVIATTPVWIRLTNVPVNFYHPSILMSIAGGLGKPFKV 85
Query: 145 DEDTSEFVVLEFSRLLVRTTTLNFITLARKILIKGTTYTIS 185
D T F+R+ V L I++ G Y ++
Sbjct: 86 DTMTLNHERARFARVCVEMNLAK--PLKGTIMVNGDRYYVA 124
>At5g25600 putative protein
Length = 331
Score = 29.3 bits (64), Expect = 2.4
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 1/61 (1%)
Query: 115 WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARK 174
WV+ GIPL++ TM + + +G V +D D + F R+ VR + + + R+
Sbjct: 124 WVQVRGIPLYVSTM-TVRFIASTLGPVTGLDFDEETSTQIAFIRVKVRISITDRMRFFRR 182
Query: 175 I 175
+
Sbjct: 183 V 183
>At5g46340 unknown protein
Length = 540
Score = 28.5 bits (62), Expect = 4.2
Identities = 17/77 (22%), Positives = 38/77 (49%), Gaps = 8/77 (10%)
Query: 135 LLQVGDVIKIDEDTSEFVV--------LEFSRLLVRTTTLNFITLARKILIKGTTYTISI 186
L+++GDV + D+D ++ + ++F +RT + F+++ L++ ++
Sbjct: 48 LVELGDVAEKDDDKADLLEGGLARSPSVKFHNSSIRTNIIRFLSMEDSFLLEHRATLRAM 107
Query: 187 IEEGEALLHPNCCCREE 203
E G L++ C R E
Sbjct: 108 SEFGAILIYFYICDRTE 124
>At5g25200 putative protein
Length = 367
Score = 28.5 bits (62), Expect = 4.2
Identities = 25/87 (28%), Positives = 37/87 (41%), Gaps = 5/87 (5%)
Query: 115 WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVR---TTTLNFITL 171
WV+ GIPL + + + +G++I +D + + F R+ VR T L F
Sbjct: 124 WVQIRGIPLPYVSEETVLEIAQDLGEIISLDFHEATSPQIAFIRVRVRFGITDRLRF--F 181
Query: 172 ARKILIKGTTYTISIIEEGEALLHPNC 198
R I G T TI E L +C
Sbjct: 182 QRIIFDSGETATIRFQYERLRRLCSSC 208
>At2g41590 putative Ta11-like non-LTR retroelement protein
Length = 367
Score = 28.5 bits (62), Expect = 4.2
Identities = 20/73 (27%), Positives = 34/73 (46%), Gaps = 5/73 (6%)
Query: 115 WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVR---TTTLNFITL 171
WV+ GIPL + + + +G+V+ +D + + + + R+ VR T L F
Sbjct: 124 WVQIRGIPLPYVSEETVMEIAQDLGEVLMLDYHDTTSIQIAYIRVRVRFGITDRLRF--F 181
Query: 172 ARKILIKGTTYTI 184
R + G T TI
Sbjct: 182 QRIVFDSGETATI 194
>At1g12220 NBS/LRR disease resistance protein, putative
Length = 889
Score = 28.5 bits (62), Expect = 4.2
Identities = 36/149 (24%), Positives = 63/149 (42%), Gaps = 23/149 (15%)
Query: 36 SLVREMKNLELI--------------PSLREAFMVEGHITIQVRYLRDDRV-LLTNHQES 80
SLV+E++ LE + P L +VE + +YL+++ V +LT
Sbjct: 650 SLVKELQLLEHLEVITLDISSSLVAEPLLCSQRLVECIKEVDFKYLKEESVRVLTLPTMG 709
Query: 81 NLVELLQGSKGWRSEFVD--------NLKPWSSSFAPINRVTWVRCTGIPLHMWTMDCFK 132
NL +L G R ++ N P + F+ ++RV +C G+ W +
Sbjct: 710 NLRKLGIKRCGMREIKIERTTSSSSRNKSPTTPCFSNLSRVFIAKCHGLKDLTWLLFAPN 769
Query: 133 HLLLQVGDVIKIDEDTSEFVVLEFSRLLV 161
L+VG ++++ SE E S +V
Sbjct: 770 LTFLEVGFSKEVEDIISEEKAEEHSATIV 798
>At5g62470 MYB96 transcription factor-like protein
Length = 352
Score = 28.1 bits (61), Expect = 5.4
Identities = 13/44 (29%), Positives = 26/44 (58%)
Query: 207 EDDVFSGWSLNGVTVTPELSESGGEDDDGLWPVETKSQHDLQGS 250
++D+ + W+ + +++ESG ED+DG+ T SQ + Q +
Sbjct: 101 DNDIKNYWNTHLKKKLKKINESGEEDNDGVSSSNTSSQKNHQST 144
>At3g45900 unknown protein
Length = 389
Score = 28.1 bits (61), Expect = 5.4
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 1/61 (1%)
Query: 25 PEIPKEEWMLRSLVREMKNLELIPSLREAFMVEGHITIQVRYLRDDRVLLTNHQESNLVE 84
PE +E +RS V E K + + L ++F+V ++ + +L +D + L S + E
Sbjct: 4 PEEEEERLRMRSFVEEEKEHDELKKLAKSFLVLS-FSVMLAHLPNDAISLVPRLTSQVTE 62
Query: 85 L 85
L
Sbjct: 63 L 63
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 28.1 bits (61), Expect = 5.4
Identities = 33/157 (21%), Positives = 62/157 (39%), Gaps = 10/157 (6%)
Query: 115 WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARK 174
W+ +P M+T L +G+ K+ DT E +++ V + ++
Sbjct: 158 WITIKNVPRSMFTWKGLSFLASPIGEPKKLHPDTVLCNSFEEAKVFVEADLTQ--EMPKQ 215
Query: 175 ILIKGTTYTISIIEEGEALLHPNC-CCREESSVEDDVFSGWSLNGVTVTPELSESGGEDD 233
K T +++E L P C C + +++ + S N ++ E+ E +
Sbjct: 216 FRFKSETGVDAMVEYKYPWLPPRCSSCSKWGHIQEVCLTRPSPNQLSTPTEI-----ETE 270
Query: 234 DGLWP--VETKSQHDLQGSPNHSLGDNKNPRSNTGAV 268
D P ++ K L SP+ +L N S+T V
Sbjct: 271 DKTEPPLMKEKPLEILSKSPSATLTKTLNGDSHTQKV 307
>At1g80070 hypothetical protein
Length = 2382
Score = 28.1 bits (61), Expect = 5.4
Identities = 18/57 (31%), Positives = 30/57 (52%), Gaps = 10/57 (17%)
Query: 58 GHITI---QVRYLRDDRVLLTN------HQESNLV-ELLQGSKGWRSEFVDNLKPWS 104
GHI I +RY + V +T+ H+E L+ L + + W SEF+D+ + W+
Sbjct: 1378 GHILIPQSDLRYSKQTDVGVTHFRSGMSHEEDQLIPNLYRYIQPWESEFIDSQRVWA 1434
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,469,498
Number of Sequences: 26719
Number of extensions: 276882
Number of successful extensions: 770
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 759
Number of HSP's gapped (non-prelim): 31
length of query: 278
length of database: 11,318,596
effective HSP length: 98
effective length of query: 180
effective length of database: 8,700,134
effective search space: 1566024120
effective search space used: 1566024120
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0102b.4


