

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0101.12
(310 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
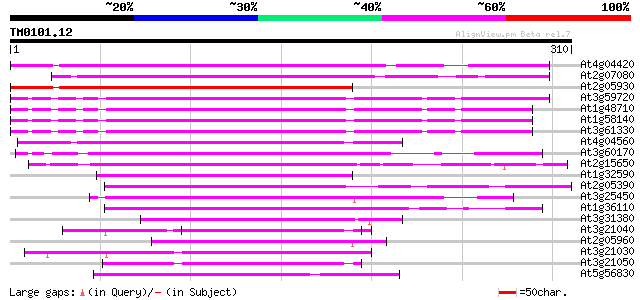
Score E
Sequences producing significant alignments: (bits) Value
At4g04420 putative transposon protein 180 8e-46
At2g07080 putative gag-protease polyprotein 163 1e-40
At2g05930 copia-like retroelement pol polyprotein 137 6e-33
At3g59720 copia-type reverse transcriptase-like protein 123 1e-28
At1g48710 hypothetical protein 122 2e-28
At1g58140 hypothetical protein 122 2e-28
At3g61330 copia-type polyprotein 120 7e-28
At4g04560 putative transposon protein 120 9e-28
At3g60170 putative protein 110 7e-25
At2g15650 putative retroelement pol polyprotein 100 2e-21
At1g32590 hypothetical protein, 5' partial 97 9e-21
At2g05390 putative retroelement pol polyprotein 87 2e-17
At3g25450 hypothetical protein 86 2e-17
At1g36110 hypothetical protein 84 7e-17
At3g31380 hypothetical protein 73 2e-13
At3g21040 hypothetical protein 73 2e-13
At2g05960 putative retroelement pol polyprotein 70 1e-12
At3g21030 hypothetical protein 67 2e-11
At3g21050 hypothetical protein 62 3e-10
At5g56830 unknown protein 57 1e-08
>At4g04420 putative transposon protein
Length = 1008
Score = 180 bits (457), Expect = 8e-46
Identities = 95/298 (31%), Positives = 170/298 (56%), Gaps = 21/298 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A ++ + + W A GWK PV +G K +D+ AEE + NS+A++
Sbjct: 28 MRALIRGLGKEAWIATSIGWKAPV---IKGEDGEDVLKTKDQWNDAEEAKAKANSRALSL 84
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
IFN V++N F+ I C AK+AW+ L +EGT+ V+ SR+ ML ++FENL+M E E I
Sbjct: 85 IFNFVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLSMEETENIE 144
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
E+ ++ +++ + LG+ D+KLV+K+LR + S+F K A+ + D S+ +E++G
Sbjct: 145 EFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTDSIDFEEVVG 204
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
LQ Y+L++ +K +A ++ + +E + L + ++++A+ F+RA+R+++++
Sbjct: 205 MLQAYELEITSGKGGYSKGLALAASAKK-----NEIQELKDTMSMMAKDFSRAMRRVEKK 259
Query: 241 TRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
+Q + D+S + +QCHEC+GYGHI++EC + +K K
Sbjct: 260 -------------GFGRNQGTDRYRDRSSKRDEIQCHECQGYGHIKAECPSLKRKDLK 304
>At2g07080 putative gag-protease polyprotein
Length = 627
Score = 163 bits (412), Expect = 1e-40
Identities = 91/275 (33%), Positives = 154/275 (55%), Gaps = 20/275 (7%)
Query: 24 VKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFNAVDKNMFRLINTCTVAKDAW 83
++ Q +G T KP+ T+ E+ S N++AM AIFN VD++ F+LI C AK AW
Sbjct: 32 IRGQEDGFKIT---KPKANWTAEEKLQSKFNARAMKAIFNGVDEDEFKLIQGCKSAKQAW 88
Query: 84 EILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHMRVRDLSNASFALGEPMSDE 143
+ L+ +HEGT+ V+ +RL + T+FE L M DE+I ++ ++ L+N + +G+ D+
Sbjct: 89 DTLQKSHEGTSSVKRTRLDHIATQFEYLKMEPDEKIVKFSSKISALANEAEVMGKTYKDQ 148
Query: 144 KLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFV 203
KLV+K+LR + KFA + A + + +L+G L+ ++K + K +K+IAF
Sbjct: 149 KLVKKLLRCLPPKFAAHKAVMRVAGNTDKISFVDLVGMLKLEEMKADQDKVKPSKNIAF- 207
Query: 204 SNTTEGDDADSESENLSEALALLARKFNRALRKIDRRTRPNVQDIKSDNSKSYNSQRKVK 263
D + + + + +ALLAR F +AL+++ +I + S+ S+
Sbjct: 208 ----NADQGSEQFQEIKDGMALLARNFGKALKRV---------EIDGERSRGRFSR---S 251
Query: 264 EEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
E D + K +QC+EC G+GHI+ EC +K+ K
Sbjct: 252 ENDDLRKKKEIQCYECGGFGHIKPECPITKRKEMK 286
>At2g05930 copia-like retroelement pol polyprotein
Length = 916
Score = 137 bits (346), Expect = 6e-33
Identities = 70/189 (37%), Positives = 115/189 (60%), Gaps = 3/189 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A ++ + + W A GWK PV +G K ED+ AEE + NS+A++
Sbjct: 40 MRALIRGLGKEAWIATSIGWKAPV---IKGEDGEDVLKTEDQWNDAEEAKATANSRALSL 96
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
IFN+V++N F+ I C AK+AW+ L +EGT+ V+ SR+ ML ++FENLTM E E I
Sbjct: 97 IFNSVNQNQFKQIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLTMEETENIE 156
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
E+ ++ +++ + LG+ D+KLV+K+LR + S+F K A+ + D +S+ +E++G
Sbjct: 157 EFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTNSIDFEEVVG 216
Query: 181 SLQTYKLKL 189
Q Y+L++
Sbjct: 217 MFQAYELEI 225
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 123 bits (308), Expect = 1e-28
Identities = 95/300 (31%), Positives = 152/300 (50%), Gaps = 23/300 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK + V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSECATYLKKQKK 298
RP+ + N + NS R + K KS +K V+C+ C +GH SEC K+ K
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFK 303
>At1g48710 hypothetical protein
Length = 1352
Score = 122 bits (307), Expect = 2e-28
Identities = 94/291 (32%), Positives = 149/291 (50%), Gaps = 23/291 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK I V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIIEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSEC 289
RP+ + N + NS R + K KS +K V+C+ C +GH SEC
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
>At1g58140 hypothetical protein
Length = 1320
Score = 122 bits (306), Expect = 2e-28
Identities = 93/291 (31%), Positives = 149/291 (50%), Gaps = 23/291 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK + V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSEC 289
RP+ + N + NS R + K KS +K V+C+ C +GH SEC
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
>At3g61330 copia-type polyprotein
Length = 1352
Score = 120 bits (302), Expect = 7e-28
Identities = 93/291 (31%), Positives = 148/291 (49%), Gaps = 23/291 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIAEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSEC 289
RP+ + N + NS R + K KS +K V+C+ C +GH SEC
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
>At4g04560 putative transposon protein
Length = 590
Score = 120 bits (301), Expect = 9e-28
Identities = 68/213 (31%), Positives = 113/213 (52%), Gaps = 7/213 (3%)
Query: 5 LKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFNA 64
++SID+ W AV GW P A+ + K E + E+ A+ NS+A++ IF +
Sbjct: 364 IQSIDMDAWFAVEDGWMPPTTKDAKRDIVS---KSRTEWIADEKTAANHNSQALSVIFGS 420
Query: 65 VDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHM 124
+ +N F + C AK+ WEIL+ + E T V+ +RL ML + FENLTM +E + +++
Sbjct: 421 LLRNKFTQVQGCLSAKEVWEILQVSFECTNNVKRTRLDMLASEFENLTMEAEESVDDFNG 480
Query: 125 RVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQT 184
++ ++ + LG+ D+K+V+K LRS+ KF AI+ + + LK D+++G +Q
Sbjct: 481 KLSSITQEAVVLGKTYKDKKMVKKFLRSLPDKFQSHKSAIDVSLNSDQLKFDQVVGMMQA 540
Query: 185 YKLKLGEKPEKKTKSIAFVSNTTEGDDADSESE 217
Y E+ S A E DD E +
Sbjct: 541 Y----DTDKEEILNSYATYFGAIEDDDHTVEED 569
>At3g60170 putative protein
Length = 1339
Score = 110 bits (276), Expect = 7e-25
Identities = 84/293 (28%), Positives = 137/293 (46%), Gaps = 53/293 (18%)
Query: 4 FLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFN 63
FL+S ++ W+ V +G + A G+T E + SA EEA L++ K N +F
Sbjct: 29 FLRSREL--WRLVEEG----IPAIVVGTTPVSEAQ-----RSAVEEAKLKDLKVKNFLFQ 77
Query: 64 AVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYH 123
A+D+ + I + +K WE +K ++G+T+V+ ++LQ L FE L M E E+I +
Sbjct: 78 AIDREILETILDKSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAMKEGEKIDTFL 137
Query: 124 MRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQ 183
R + N GE M +V KILRS+T KF V +IEE+ D+S+L +DEL GSL
Sbjct: 138 GRTLTVVNKMKTNGEVMEQSTIVSKILRSLTPKFNYVVCSIEESNDLSTLSIDELHGSLL 197
Query: 184 TYKLKL-GEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRRTR 242
++ +L G E++ + ++G R + R
Sbjct: 198 VHEQRLNGHVQEEQALKVTHEERPSQGRG-----------------------RGVFR--- 231
Query: 243 PNVQDIKSDNSKSYNSQRKVKEEDKSGQNKG-VQCHECEGYGHIRSECATYLK 294
S+ + + +SG N+ V+C++C GH + EC + K
Sbjct: 232 --------------GSRGRGRGRGRSGTNRAIVECYKCHNLGHFQYECPEWEK 270
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 99.8 bits (247), Expect = 2e-21
Identities = 80/301 (26%), Positives = 148/301 (48%), Gaps = 32/301 (10%)
Query: 11 KTWKAVVKGWKHPVK-AQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFNAVDKNM 69
K W V +G PV+ QAE + T K + EEA ++ A+ + AV +
Sbjct: 32 KLWSVVEEGV--PVEPVQAEETPETARAK------TLREEAVTNDTMALQILQTAVTDQI 83
Query: 70 FRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHMRVRDL 129
F I + +K+AW++LK ++G+ +VR+ +LQ L +ENL M +++ I + ++ L
Sbjct: 84 FSRIAAASSSKEAWDVLKDEYQGSPQVRLVKLQSLRREYENLKMYDNDNIKTFTDKLIVL 143
Query: 130 SNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQTYKLKL 189
GE ++ +L++KIL S+ +KF V +E+ +D+ +L + EL+G L+ + ++
Sbjct: 144 EIQLTYHGEKKTNTQLIQKILISLPAKFDSIVSVLEQTRDLDALTMSELLGILKAQEARV 203
Query: 190 GEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRRTRPNVQDIK 249
+ E+ TK AF ++G ++ + +N + + + + K
Sbjct: 204 TAR-EESTKEGAFYVR-SKGRESGFKQDNTNNRV---------------NQDKKWCGFHK 246
Query: 250 SDNSKSYNSQRKVKEEDKSGQNK--GVQCHECEGYGHIRSECATYLKKQKKGMVVTWSDE 307
S + K K +D G+NK ++C++C GH +EC + K K+ VT +E
Sbjct: 247 SSKHTEEECREKPKNDD-HGKNKRSNIKCYKCGKIGHYANECRS---KNKERAHVTLEEE 302
Query: 308 D 308
D
Sbjct: 303 D 303
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 97.4 bits (241), Expect = 9e-21
Identities = 54/141 (38%), Positives = 81/141 (57%)
Query: 49 EASLENSKAMNAIFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRF 108
E ++++ K N +F ++DK + + I +KD WE +K ++G RV+ ++LQ L F
Sbjct: 20 EKTVKDHKVKNYLFASIDKTILKTILQKETSKDLWESMKRKYQGNDRVQSAQLQRLRRSF 79
Query: 109 ENLTMTEDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQ 168
E L M E I+ Y RV +++N LGE M D K+V KILR++ KF V AIEE+
Sbjct: 80 EVLEMKIGETITGYFSRVMEITNDMRNLGEDMPDSKVVEKILRTLVEKFTYVVCAIEESN 139
Query: 169 DISSLKVDELIGSLQTYKLKL 189
+I L VD L SL ++ L
Sbjct: 140 NIKELTVDGLQSSLMVHEQNL 160
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 86.7 bits (213), Expect = 2e-17
Identities = 65/260 (25%), Positives = 117/260 (45%), Gaps = 35/260 (13%)
Query: 53 ENSKAMNAIFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLT 112
+N A +F +V ++ + +K WE +KT + G RV+ ++LQ L F+ L
Sbjct: 23 KNDMARALLFQSVPESTILQVGKHKTSKAMWEAIKTRNLGAERVKEAKLQTLMAEFDRLN 82
Query: 113 MTEDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVI-AIEEAQDIS 171
M ++E I E+ R+ ++S S +LGE + + K+V+K L+S+ K + +I A+E+ D++
Sbjct: 83 MKDNETIDEFVGRISEISTKSESLGEEIEESKIVKKFLKSLPRKKYIHIIAALEQILDLN 142
Query: 172 SLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFN 231
+ ++++G ++TY+ + DD+ E L A N
Sbjct: 143 TTGFEDIVGRMKTYE-----------------DRVCDEDDSPEEQGKLMYA--------N 177
Query: 232 RALRKIDRRTRPNVQDIKSDNSK-SYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECA 290
R R + S + Y Q++ K + V C+ C+ GH SEC
Sbjct: 178 SESSYDTRGGRGRGRGRSSGRGRGGYGYQQRDKSK--------VICYRCDKTGHYASECL 229
Query: 291 TYLKKQKKGMVVTWSDEDSE 310
L K K ++ED +
Sbjct: 230 DRLLKLIKAQEQQQNNEDDD 249
>At3g25450 hypothetical protein
Length = 1343
Score = 86.3 bits (212), Expect = 2e-17
Identities = 57/238 (23%), Positives = 117/238 (48%), Gaps = 25/238 (10%)
Query: 45 SAEEEASLENSKAMNAIFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQML 104
SA+EE +N A +F ++ +++ + + WE +K+ + G RV+ +RLQ L
Sbjct: 54 SADEE---KNDMARALLFQSIPESLILQVGKQKTSSAVWEAIKSRNLGAERVKEARLQTL 110
Query: 105 TTRFENLTMTEDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSV-TSKFAMKVIA 163
F+ L M + E I +Y R+ +++ + ALGE + + K+V+K L+S+ K+ V A
Sbjct: 111 MAEFDKLKMKDSETIDDYVGRISEITTKAAALGEDIEESKIVKKFLKSLPRKKYIHIVAA 170
Query: 164 IEEAQDISSLKVDELIGSLQTYKLKL---GEKPEKKTKSIAFVSNTTEGDDADSESENLS 220
+E+ D+ + +++ G ++TY+ ++ + E + K + V D + E E ++
Sbjct: 171 LEQVLDLKTTTFEDIAGRIKTYEDRVWDDDDSHEDQGKLMTEVEEEVVDDLEEEEEEVIN 230
Query: 221 EALALLARKFNRALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHE 278
+ + + +R L+ I + ++K KEED + + + + HE
Sbjct: 231 KEIKAKSHVIDRLLKLIRLQ------------------EQKEKEEDDTHEAESLMMHE 270
>At1g36110 hypothetical protein
Length = 745
Score = 84.3 bits (207), Expect = 7e-17
Identities = 60/243 (24%), Positives = 111/243 (44%), Gaps = 33/243 (13%)
Query: 53 ENSKAMNAIFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLT 112
+N+ A IF ++ +++ + T AK W+ +KT + G RV+ +RLQ L FE +
Sbjct: 56 KNTMARGLIFQSIPESLTLQVGTLATAKLVWDSIKTRYVGADRVKEARLQTLMAEFEKMK 115
Query: 113 MTEDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIA-IEEAQDIS 171
M E E+I + R+ +L+ S ALG + KLV+K L ++ + + +IA +E+ D++
Sbjct: 116 MKESEKIDVFAGRLAELATRSDALGSNIETSKLVKKFLNALPLRKYIHIIASLEQVLDLN 175
Query: 172 SLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFN 231
+ ++++G ++ Y+ ++ + E++ + T+ D+ S + FN
Sbjct: 176 NTSFEDIVGRIKVYEERVWDGEEQEDDQGKLMYANTDTQDSWYASRGRGQ-----GGGFN 230
Query: 232 RALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECAT 291
R R +R D SK V C+ C+ GH S C
Sbjct: 231 GRGRGRGRGSR--------DTSK-------------------VTCYRCDKLGHYASNCPD 263
Query: 292 YLK 294
+K
Sbjct: 264 SVK 266
>At3g31380 hypothetical protein
Length = 262
Score = 73.2 bits (178), Expect = 2e-13
Identities = 41/150 (27%), Positives = 84/150 (55%), Gaps = 6/150 (4%)
Query: 73 INTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHMRVRDLSNA 132
+ AK+ W+ +KT H G RV+ +R+Q L FE + M E E+I ++ R+ +LS
Sbjct: 18 VGNLQTAKEVWDSIKTRHVGAERVKEARVQTLMADFEKMKMKEAEKIDDFAGRLSELSTK 77
Query: 133 SFALGEPMSDEKLVRKILRSVTSKFAMKVIA-IEEAQDISSLKVDELIGSLQTYKLKLGE 191
S LG + KLV+K L S+ K + +IA +E+ D+++ ++++G ++ + ++ +
Sbjct: 78 SAPLGVNIEVPKLVKKFLNSLPRKKYIHIIASLEQVLDLNNTTFEDIVGCMKVNEERVYD 137
Query: 192 KPEKKT----KSIAFVSNTTEGDDADSESE 217
PE++T + + S+ ++ + D + +
Sbjct: 138 -PEEETNEDQNKLMYTSSDSKSGNRDYQGD 166
>At3g21040 hypothetical protein
Length = 534
Score = 72.8 bits (177), Expect = 2e-13
Identities = 46/177 (25%), Positives = 86/177 (47%), Gaps = 9/177 (5%)
Query: 30 GSTSTPEEKPEDELTSAEEEAS------LENSKAMNAIFNAVDKNMFRLINTCTVAKDAW 83
G + P PE T E+ + +E++KA+ + +A+ ++FR + +K+ W
Sbjct: 38 GVSPNPTTDPEVAATIKVEDLTQWRNRMIEDNKALKIMQSALPDSVFRKTISVASSKELW 97
Query: 84 EILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHMRVRDLSNASFALGEPMSDE 143
++LK +G +L+ L + ENLTM E ER+ Y RV + F G P+SDE
Sbjct: 98 DLLK---KGNDNEEAKKLRRLEKQLENLTMYEGERMKSYLKRVEKIIEEFFVWGNPISDE 154
Query: 144 KLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSI 200
K++ K+L S+ + + ++E + L +L+ + + + P++ K I
Sbjct: 155 KVIAKLLTSLPRPYDDSITVLKEFMTLPDLTHRDLLKAFEMFGSNPKTMPQELMKFI 211
Score = 48.1 bits (113), Expect = 6e-06
Identities = 28/99 (28%), Positives = 51/99 (51%), Gaps = 4/99 (4%)
Query: 96 VRMSRLQMLTTRFENLTMTEDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTS 155
++ ++L L +FE L M + E + Y RV D A G P+SD+ ++ K+L S++
Sbjct: 233 IKDAKLLRLEKQFEKLMMYDGESMDSYLERVLDTIEQFRASGNPLSDDDVIAKLLTSLSW 292
Query: 156 KFAMKVIAIEEAQDISSLKVDELIGSLQTYKLKLGEKPE 194
+ + +EE ++ + +LIG L+ + G PE
Sbjct: 293 PYDDAIPVLEELMNLPDMTPHDLIGVLEMF----GSHPE 327
>At2g05960 putative retroelement pol polyprotein
Length = 1200
Score = 70.1 bits (170), Expect = 1e-12
Identities = 40/138 (28%), Positives = 79/138 (56%), Gaps = 8/138 (5%)
Query: 79 AKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHMRVRDLSNASFALGE 138
+K W+ +++ + G RV+ ++L+ L F+ L M ++E I E R+ ++S S +LGE
Sbjct: 43 SKAVWDKIQSRNLGAERVKEAKLKTLMAEFDKLKMKDNETIDECAGRLSEISTKSTSLGE 102
Query: 139 PMSDEKLVRKILRSV-TSKFAMKVIAIEEAQDISSLKVDELIGSLQTYKLK-------LG 190
+ + K+V+K L+S+ T K+ V A+E+ D+ + +++G ++TY+ K L
Sbjct: 103 DIEETKVVKKFLKSLPTKKYIHIVAALEQVLDLKNTTFKDIVGRIKTYEDKIWVLITCLK 162
Query: 191 EKPEKKTKSIAFVSNTTE 208
++ E++ KS+ V E
Sbjct: 163 KEAEEEEKSVVGVEAEEE 180
>At3g21030 hypothetical protein
Length = 411
Score = 66.6 bits (161), Expect = 2e-11
Identities = 47/201 (23%), Positives = 95/201 (46%), Gaps = 13/201 (6%)
Query: 9 DIKTWKAVVKG-------WKHPVKAQAEGSTSTPEEKPEDELTSAEEEASL--ENSKAMN 59
D + W +VK W + T+ PE + + +L +++KA+
Sbjct: 14 DYEDWSPMVKTRLVENGVWDVVQNGVSPNPTTNPELAATIKAVDLAQWRNLVVKDNKALK 73
Query: 60 AIFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERI 119
+ +++ ++FR + AK+ W++LK ++ + ++L+ L +FENL M E ER+
Sbjct: 74 IMQSSLPDSVFRKTISIASAKELWDLLKKGND----TKEAKLRRLEKQFENLMMYEGERM 129
Query: 120 SEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELI 179
Y R+ + F LG P+SD+K++ K+L S+ + V ++E + L +L+
Sbjct: 130 KLYLKRLEKIIEELFVLGNPISDDKVIAKLLTSLPRSYDDSVPVLKEFMTLPELTHRDLL 189
Query: 180 GSLQTYKLKLGEKPEKKTKSI 200
+ + + P++ K I
Sbjct: 190 KAFEMFGSDPKTMPQELMKFI 210
>At3g21050 hypothetical protein
Length = 420
Score = 62.4 bits (150), Expect = 3e-10
Identities = 35/143 (24%), Positives = 79/143 (54%), Gaps = 8/143 (5%)
Query: 52 LENSKAMNAIFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENL 111
++++KA+ + +++ ++FR + AK+ W++LK ++ + ++L+ L +FE L
Sbjct: 66 IKDNKALKIMQSSLPDSVFRKTISIASAKELWDLLKKGND----TKEAKLRRLEKQFEKL 121
Query: 112 TMTEDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDIS 171
M E E + Y RV +++ LG P+SD+K++ K+L S++ + + ++E +
Sbjct: 122 MMYEGEPMDLYLKRVEEITERFEVLGNPISDDKVITKLLTSLSWPYDDSIPVLKEFMTLP 181
Query: 172 SLKVDELIGSLQTYKLKLGEKPE 194
L + +L+ + + + G PE
Sbjct: 182 DLTLRDLLKAFELF----GSHPE 200
>At5g56830 unknown protein
Length = 242
Score = 57.0 bits (136), Expect = 1e-08
Identities = 41/169 (24%), Positives = 80/169 (47%), Gaps = 5/169 (2%)
Query: 47 EEEASLENSKAMNAIFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTT 106
E A +E + ++ I + + +F I T +K+AWEIL G + +L L
Sbjct: 3 ETMARMERCEWLSLIQSGLSTEIFSWIAYATSSKEAWEILMVRSVGGPIFQARKLLDLKK 62
Query: 107 RFENLTMTEDERISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEE 166
+ N TM++ E + E+ ++ +L + G + ++ LV+KIL S+ +F ++ + E
Sbjct: 63 EYNNTTMSDAETVHEFTGKLLELVFRMKSCGWKVEEKILVKKILSSLPPRFDKVLVEVAE 122
Query: 167 AQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSE 215
L V+ +I L ++ + E +S+ N E +A++E
Sbjct: 123 -----KLSVNVIIDCLLVHEYNMRPDQESFQESVIQSVNVEESFEAEAE 166
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.126 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,538,511
Number of Sequences: 26719
Number of extensions: 264891
Number of successful extensions: 1246
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 50
Number of HSP's successfully gapped in prelim test: 70
Number of HSP's that attempted gapping in prelim test: 1127
Number of HSP's gapped (non-prelim): 171
length of query: 310
length of database: 11,318,596
effective HSP length: 99
effective length of query: 211
effective length of database: 8,673,415
effective search space: 1830090565
effective search space used: 1830090565
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0101.12


