

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.3
(244 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
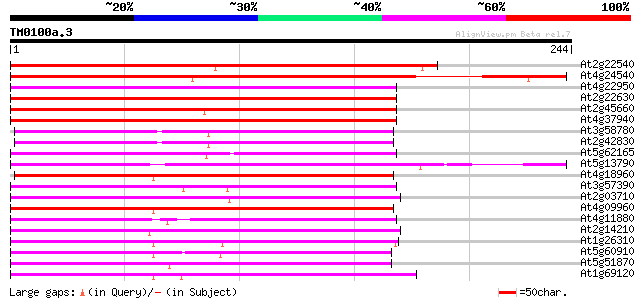
Score E
Sequences producing significant alignments: (bits) Value
At2g22540 putative MADS-box protein 170 6e-43
At4g24540 MADS-box protein AGL24 169 1e-42
At4g22950 putative MADS Box / AGL protein 119 1e-27
At2g22630 MADS-box protein AGL17 (AGL17) 118 3e-27
At2g45660 MADS-box protein (AGL20) 117 7e-27
At4g37940 MADS-box protein AGL17 -like protein 116 9e-27
At3g58780 shatterproof 1 (SHP1)/ agamous -like 1 (AGL1) 114 5e-26
At2g42830 MADS box transcription factor (AGL5) 113 8e-26
At5g62165 unknown protein 111 3e-25
At5g13790 floral homeotic protein AGL15 (sp|Q38847) 111 3e-25
At4g18960 floral homeotic protein agamous 111 4e-25
At3g57390 MADS transcription factor-like protein 110 5e-25
At2g03710 MADS-box protein (AGL3) 110 7e-25
At4g09960 MADS-box protein AGL11 110 9e-25
At4g11880 MADS-box protein AGL14 109 2e-24
At2g14210 pseudogene 108 2e-24
At1g26310 cauliflower (CAL) 108 2e-24
At5g60910 floral homeotic protein AGL8 108 3e-24
At5g51870 MADS box transcription factor-like 108 3e-24
At1g69120 unknown protein 108 3e-24
>At2g22540 putative MADS-box protein
Length = 210
Score = 170 bits (431), Expect = 6e-43
Identities = 91/188 (48%), Positives = 132/188 (69%), Gaps = 2/188 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQI+KIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLFE+ SS
Sbjct: 1 MAREKIQIRKIDNATARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + S +K PS +Q E+ + + K++ +K+H LRQ+ GE+LQGL
Sbjct: 61 SMKEVLERHNLQSKNLEKLDQPSLELQLVENSDHARMSKEIADKSHRLRQMRGEELQGLD 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLF- 178
+ +LQ+LE+ L+ L V K +K M EIS L++K ++L++EN++L+Q + L LL
Sbjct: 121 IEELQQLEKALETGLTRVIETKSDKIMSEISELQKKGMQLMDENKRLRQQVCVLPSLLIT 180
Query: 179 NEYLCSII 186
N +L S I
Sbjct: 181 NPFLLSTI 188
>At4g24540 MADS-box protein AGL24
Length = 220
Score = 169 bits (428), Expect = 1e-42
Identities = 103/244 (42%), Positives = 153/244 (62%), Gaps = 30/244 (12%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+G+FKKA ELS LCDAD+ALI+FSAT KLFE++SS
Sbjct: 1 MAREKIRIKKIDNITARQVTFSKRRRGIFKKADELSVLCDADVALIIFSATGKLFEFSSS 60
Query: 61 SRGGLLPLPDANSAT*DK-TTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
+L +++ +K PPS ++ E+ + L K+VE+KT +LR+L GEDL GL
Sbjct: 61 RMRDILGRYSLHASNINKLMDPPSTHLRLENCNLSRLSKEVEDKTKQLRKLRGEDLDGLN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
L +LQ+LE++L+ L+ VS K E M +I +L+++ EL++EN++L+ L +L +
Sbjct: 121 LEELQRLEKLLESGLSRVSEKKGECVMSQIFSLEKRGSELVDENKRLRDKLETLERAK-- 178
Query: 180 EYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISS-SSYLLEEDGSDTSL 238
+ +L + + +S+ + +SS S ED SDTSL
Sbjct: 179 --------------------------LTTLKEALETESVTTNVSSYDSGTPLEDDSDTSL 212
Query: 239 KLGL 242
KLGL
Sbjct: 213 KLGL 216
>At4g22950 putative MADS Box / AGL protein
Length = 219
Score = 119 bits (299), Expect = 1e-27
Identities = 66/168 (39%), Positives = 99/168 (58%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + ++K+I+N +SRQVTFSKRR GL KKA ELS LCDA++AL++FS +KL+E++SS
Sbjct: 1 MVRGKTEMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALVIFSPRSKLYEFSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S + + Q D L KK+E+ R+L GE + ++
Sbjct: 61 SIAATIERYQRRIKEIGNNHKRNDNSQQARDETSGLTKKIEQLEISKRKLLGEGIDACSI 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+LQ+LE L RSL+ + K + +EI LK +E L++EN+ LK+
Sbjct: 121 EELQQLENQLDRSLSRIRAKKYQLLREEIEKLKAEERNLVKENKDLKE 168
>At2g22630 MADS-box protein AGL17 (AGL17)
Length = 227
Score = 118 bits (295), Expect = 3e-27
Identities = 66/168 (39%), Positives = 102/168 (60%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I I+KID+ +SRQVTFSKRRKGL KKA+EL+ LCDA++ LI+FS T+KL+++ASS
Sbjct: 1 MGRGKIVIQKIDDSTSRQVTFSKRRKGLIKKAKELAILCDAEVCLIIFSNTDKLYDFASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S + + + P+ ++F +TLR+++ RQL G +L GL++
Sbjct: 61 SVKSTIERFNTAKMEEQELMNPASEVKFWQREAETLRQELHSLQENYRQLTGVELNGLSV 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+LQ +E L+ SL + +++ EI L RK + EN +L +
Sbjct: 121 KELQNIESQLEMSLRGIRMKREQILTNEIKELTRKRNLVHHENLELSR 168
>At2g45660 MADS-box protein (AGL20)
Length = 214
Score = 117 bits (292), Expect = 7e-27
Identities = 67/169 (39%), Positives = 103/169 (60%), Gaps = 1/169 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + Q+K+I+N +SRQVTFSKRR GL KKA ELS LCDA+++LI+FS KL+E+ASS
Sbjct: 1 MVRGKTQMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVSLIIFSPKGKLYEFASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPS-PTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
+ + ++ T P S MQ + KK+E+ R+L GE + +
Sbjct: 61 NMQDTIDRYLRHTKDRVSTKPVSEENMQHLKYEAANMMKKIEQLEASKRKLLGEGIGTCS 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+ +LQ++E+ L++S+ + K + F ++I LK+KE L EN+KL +
Sbjct: 121 IEELQQIEQQLEKSVKCIRARKTQVFKEQIEQLKQKEKALAAENEKLSE 169
>At4g37940 MADS-box protein AGL17 -like protein
Length = 228
Score = 116 bits (291), Expect = 9e-27
Identities = 64/168 (38%), Positives = 103/168 (61%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I I++ID+ +SRQVTFSKRRKGL KKA+EL+ LCDA++ LI+FS+T KL+++ASS
Sbjct: 1 MGRGKIVIQRIDDSTSRQVTFSKRRKGLIKKAKELAILCDAEVGLIIFSSTGKLYDFASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S ++ + + + P+ ++F LR+++ RQ+ GE L GL++
Sbjct: 61 SMKSVIDRYNKSKIEQQQLLNPASEVKFWQREAAVLRQELHALQENHRQMMGEQLNGLSV 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
++L LE ++ SL + K++ QEI L +K + +EN L +
Sbjct: 121 NELNSLENQIEISLRGIRMRKEQLLTQEIQELSQKRNLIHQENLDLSR 168
>At3g58780 shatterproof 1 (SHP1)/ agamous -like 1 (AGL1)
Length = 248
Score = 114 bits (285), Expect = 5e-26
Identities = 64/168 (38%), Positives = 100/168 (59%), Gaps = 5/168 (2%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSSR 62
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++AL++FS +L+EYA++S
Sbjct: 18 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVIFSTRGRLYEYANNSV 77
Query: 63 GGLLPLPDANSAT*DKTTPPSPT---MQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
G + A D PPS T Q+ LR+++ + + R + GE L L
Sbjct: 78 RG--TIERYKKACSDAVNPPSVTEANTQYYQQEASKLRRQIRDIQNSNRHIVGESLGSLN 135
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+L+ LE L++ ++ V K+E + EI ++++E+EL N L+
Sbjct: 136 FKELKNLEGRLEKGISRVRSKKNELLVAEIEYMQKREMELQHNNMYLR 183
>At2g42830 MADS box transcription factor (AGL5)
Length = 246
Score = 113 bits (283), Expect = 8e-26
Identities = 63/168 (37%), Positives = 100/168 (59%), Gaps = 5/168 (2%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSSR 62
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++AL++FS +L+EYA++S
Sbjct: 18 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVIFSTRGRLYEYANNSV 77
Query: 63 GGLLPLPDANSAT*DKTTPPSPT---MQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
G + A D PP+ T Q+ LR+++ + + R + GE L L
Sbjct: 78 RG--TIERYKKACSDAVNPPTITEANTQYYQQEASKLRRQIRDIQNLNRHILGESLGSLN 135
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+L+ LE L++ ++ V K E + EI ++++E+EL +N L+
Sbjct: 136 FKELKNLESRLEKGISRVRSKKHEMLVAEIEYMQKREIELQNDNMYLR 183
>At5g62165 unknown protein
Length = 210
Score = 111 bits (278), Expect = 3e-25
Identities = 65/170 (38%), Positives = 102/170 (59%), Gaps = 3/170 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I++KKI+N +SRQVTFSKRR GL KKA ELS LCDA ++LI+FS +L+E++SS
Sbjct: 1 MVRGKIEMKKIENATSRQVTFSKRRNGLLKKAYELSVLCDAQLSLIIFSQRGRLYEFSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSP--TMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGL 118
+ + + + S Q + +++ + K+E R+L G+ +
Sbjct: 61 DMQKTIERYRKYTKDHETSNHDSQIHLQQLKQEASHMI-TKIELLEFHKRKLLGQGIASC 119
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+L +LQ+++ L+RSL V K + F +++ LK KE +L+EEN KL Q
Sbjct: 120 SLEELQEIDSQLQRSLGKVRERKAQLFKEQLEKLKAKEKQLLEENVKLHQ 169
>At5g13790 floral homeotic protein AGL15 (sp|Q38847)
Length = 268
Score = 111 bits (278), Expect = 3e-25
Identities = 80/244 (32%), Positives = 125/244 (50%), Gaps = 31/244 (12%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N +SRQVTFSKRR GL KKA+ELS LCDA++A+IVFS + KLFEY+S+
Sbjct: 1 MGRGKIEIKRIENANSRQVTFSKRRSGLLKKARELSVLCDAEVAVIVFSKSGKLFEYSST 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
+ S + + + + + D L+ ++ + + QL G+ L LT
Sbjct: 61 G------MKQTLSRYGNHQSSSASKAEEDCAEVDILKDQLSKLQEKHLQLQGKGLNPLTF 114
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLL--F 178
+LQ LE+ L +L +V K+ ++ + KE EN+ L++ + L L F
Sbjct: 115 KELQSLEQQLYHALITVRERKERLLTNQLEESRLKEQRAELENETLRRQVQELRSFLPSF 174
Query: 179 NEYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSL 238
Y+ S I + ID ++ L+ + S L+ SDT+L
Sbjct: 175 THYVPSYIKCF-AIDPKNALINH----------------------DSKCSLQNTDSDTTL 211
Query: 239 KLGL 242
+LGL
Sbjct: 212 QLGL 215
>At4g18960 floral homeotic protein agamous
Length = 284
Score = 111 bits (277), Expect = 4e-25
Identities = 63/166 (37%), Positives = 103/166 (61%), Gaps = 1/166 (0%)
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSS- 61
R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS+ +L+EY+++S
Sbjct: 51 RGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSSRGRLYEYSNNSV 110
Query: 62 RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLH 121
+G + A S + + Q+ + LR+++ + RQL GE + ++
Sbjct: 111 KGTIERYKKAISDNSNTGSVAEINAQYYQQESAKLRQQIISIQNSNRQLMGETIGSMSPK 170
Query: 122 QLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+L+ LE L+RS+ + K+E EI ++++EV+L +NQ L+
Sbjct: 171 ELRNLEGRLERSITRIRSKKNELLFSEIDYMQKREVDLHNDNQILR 216
>At3g57390 MADS transcription factor-like protein
Length = 256
Score = 110 bits (276), Expect = 5e-25
Identities = 68/176 (38%), Positives = 105/176 (59%), Gaps = 8/176 (4%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R RI+IKKI+NI+SRQVTFSKRR GL KKA+ELS LCDA++ALI+FS+T K+++++S
Sbjct: 1 MGRGRIEIKKIENINSRQVTFSKRRNGLIKKAKELSILCDAEVALIIFSSTGKIYDFSSV 60
Query: 61 SRGGLLPLPDANSA-T*DKTTPPSPTMQFESDSN-------DTLRKKVEEKTHELRQLNG 112
+L +A T K + S N D+++ ++E + +L G
Sbjct: 61 CMEQILSRYGYTTASTEHKQQREHQLLICASHGNEAVLRNDDSMKGELERLQLAIERLKG 120
Query: 113 EDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
++L+G++ L LE L SL SV K + + +I + +E + +EENQ L++
Sbjct: 121 KELEGMSFPDLISLENQLNESLHSVKDQKTQILLNQIERSRIQEKKALEENQILRK 176
>At2g03710 MADS-box protein (AGL3)
Length = 257
Score = 110 bits (275), Expect = 7e-25
Identities = 63/172 (36%), Positives = 96/172 (55%), Gaps = 2/172 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R ++++K+I+N +RQVTF+KRR GL KKA ELS LCDA+IAL++FS KL+E+ SS
Sbjct: 1 MGRGKVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEIALLIFSNRGKLYEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSND--TLRKKVEEKTHELRQLNGEDLQGL 118
G + + P + D L+ +VE H R L GE+L +
Sbjct: 61 PSGMARTVDKYRKHSYATMDPNQSAKDLQDKYQDYLKLKSRVEILQHSQRHLLGEELSEM 120
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVL 170
+++L+ LE + SL + K + ++S LK KE L+E N+ L++ L
Sbjct: 121 DVNELEHLERQVDASLRQIRSTKARSMLDQLSDLKTKEEMLLETNRDLRRKL 172
>At4g09960 MADS-box protein AGL11
Length = 230
Score = 110 bits (274), Expect = 9e-25
Identities = 61/168 (36%), Positives = 104/168 (61%), Gaps = 1/168 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS +L+EYA++
Sbjct: 1 MGRGKIEIKRIENSTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYANN 60
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
+ R + A S + + +T + + LR++++ + R L G+ L L+
Sbjct: 61 NIRSTIERYKKACSDSTNTSTVQEINAAYYQQESAKLRQQIQTIQNSNRNLMGDSLSSLS 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLK 167
+ +L+++E L+++++ + K E + EI +++E+EL EN L+
Sbjct: 121 VKELKQVENRLEKAISRIRSKKHELLLVEIENAQKREIELDNENIYLR 168
>At4g11880 MADS-box protein AGL14
Length = 221
Score = 109 bits (272), Expect = 2e-24
Identities = 69/177 (38%), Positives = 97/177 (53%), Gaps = 17/177 (9%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R + ++K+I+N +SRQVTFSKRR GL KKA ELS LCDA++ALI+FS KL+E++SS
Sbjct: 1 MVRGKTEMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALIIFSPRGKLYEFSSS 60
Query: 61 SRGGLLP---------LPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLN 111
S +P + D S + Q D L +K+E R++
Sbjct: 61 SS---IPKTVERYQKRIQDLGS-----NHKRNDNSQQSKDETYGLARKIEHLEISTRKMM 112
Query: 112 GEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
GE L ++ +LQ+LE L RSL + K + +E LK KE LI EN+ L +
Sbjct: 113 GEGLDASSIEELQQLENQLDRSLMKIRAKKYQLLREETEKLKEKERNLIAENKMLME 169
>At2g14210 pseudogene
Length = 234
Score = 108 bits (271), Expect = 2e-24
Identities = 66/171 (38%), Positives = 103/171 (59%), Gaps = 1/171 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYAS- 59
M R +I I++IDN +SRQVTFSKRR GL KKA+ELS LCDA++ +I+FS+T KL++YAS
Sbjct: 1 MGRGKIVIRRIDNSTSRQVTFSKRRSGLLKKAKELSILCDAEVGVIIFSSTGKLYDYASN 60
Query: 60 SSRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
SS ++ + + + ++F +L+++++ R+L GE+L G+
Sbjct: 61 SSMKTIIERYNRVKEEQHQLLNHASEIKFWQREVASLQQQLQYLQECHRKLVGEELSGMN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVL 170
+ LQ LE+ L SL V KD+ EI L RK + +EN +L+ ++
Sbjct: 121 ANDLQNLEDQLVTSLKGVRLKKDQLMTNEIRELNRKGQIIQKENHELQNIV 171
>At1g26310 cauliflower (CAL)
Length = 255
Score = 108 bits (271), Expect = 2e-24
Identities = 68/174 (39%), Positives = 104/174 (59%), Gaps = 5/174 (2%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTFSKRR GL KKAQE+S LCDA+++LIVFS KLFEY+S
Sbjct: 1 MGRGRVELKRIENKINRQVTFSKRRTGLLKKAQEISVLCDAEVSLIVFSHKGKLFEYSSE 60
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFESD---SNDTLRKKVEEKTHELRQLNGEDLQ 116
S +L + S + P + +++ L+ K+E R GE+L+
Sbjct: 61 SCMEKVLERYERYSYAERQLIAPDSHVNAQTNWSMEYSRLKAKIELLERNQRHYLGEELE 120
Query: 117 GLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL-KQV 169
++L LQ LE+ L+ +L + K++ + ++ L+RKE E+ EEN L KQ+
Sbjct: 121 PMSLKDLQNLEQQLETALKHIRSRKNQLMNESLNHLQRKEKEIQEENSMLTKQI 174
>At5g60910 floral homeotic protein AGL8
Length = 242
Score = 108 bits (270), Expect = 3e-24
Identities = 69/169 (40%), Positives = 98/169 (57%), Gaps = 4/169 (2%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+Q+K+I+N +RQVTFSKRR GL KKA E+S LCDA++ALIVFS+ KLFEY++
Sbjct: 1 MGRGRVQLKRIENKINRQVTFSKRRSGLLKKAHEISVLCDAEVALIVFSSKGKLFEYSTD 60
Query: 61 S-RGGLLPLPDANSAT*DKTTPPSPTMQFES--DSNDTLRKKVEEKTHELRQLNGEDLQG 117
S +L D + DK Q E+ + L+ +VE R GEDL
Sbjct: 61 SCMERILERYDRYLYS-DKQLVGRDVSQSENWVLEHAKLKARVEVLEKNKRNFMGEDLDS 119
Query: 118 LTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL 166
L+L +LQ LE L ++ S+ K++ + IS L++K+ L + N L
Sbjct: 120 LSLKELQSLEHQLDAAIKSIRSRKNQAMFESISALQKKDKALQDHNNSL 168
>At5g51870 MADS box transcription factor-like
Length = 207
Score = 108 bits (270), Expect = 3e-24
Identities = 65/168 (38%), Positives = 100/168 (58%), Gaps = 2/168 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IKKI+N++SRQVTFSKRR GLFKKA ELS LCDA +A IVFS + +L EY+SS
Sbjct: 1 MVRGKIEIKKIENVTSRQVTFSKRRSGLFKKAHELSVLCDAQVAAIVFSQSGRLHEYSSS 60
Query: 61 SRGGLLPL--PDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGL 118
++ +N+ + +Q D + KK++ R+L G+ L
Sbjct: 61 QMEKIIDRYGKFSNAFYVAERPQVERYLQELKMEIDRMVKKIDLLEVHHRKLLGQGLDSC 120
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKL 166
++ +LQ+++ +++SL V K E + ++ LK KE EL+ E ++L
Sbjct: 121 SVTELQEIDTQIEKSLRIVRSRKAELYADQLKKLKEKERELLNERKRL 168
>At1g69120 unknown protein
Length = 256
Score = 108 bits (270), Expect = 3e-24
Identities = 69/179 (38%), Positives = 102/179 (56%), Gaps = 2/179 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+Q+K+I+N +RQVTFSKRR GL KKA E+S LCDA++AL+VFS KLFEY++
Sbjct: 1 MGRGRVQLKRIENKINRQVTFSKRRAGLLKKAHEISVLCDAEVALVVFSHKGKLFEYSTD 60
Query: 61 S-RGGLLPLPDANS-AT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGL 118
S +L + S A P S S + L+ K+E R GEDLQ +
Sbjct: 61 SCMEKILERYERYSYAERQLIAPESDVNTNWSMEYNRLKAKIELLERNQRHYLGEDLQAM 120
Query: 119 TLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLL 177
+ +LQ LE+ L +L + K++ + I+ L++KE + E+N L + + K+L
Sbjct: 121 SPKELQNLEQQLDTALKHIRTRKNQLMYESINELQKKEKAIQEQNSMLSKQIKEREKIL 179
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.136 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,897,584
Number of Sequences: 26719
Number of extensions: 196793
Number of successful extensions: 1077
Number of sequences better than 10.0: 168
Number of HSP's better than 10.0 without gapping: 115
Number of HSP's successfully gapped in prelim test: 53
Number of HSP's that attempted gapping in prelim test: 919
Number of HSP's gapped (non-prelim): 189
length of query: 244
length of database: 11,318,596
effective HSP length: 97
effective length of query: 147
effective length of database: 8,726,853
effective search space: 1282847391
effective search space used: 1282847391
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0100a.3


