

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.11
(245 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
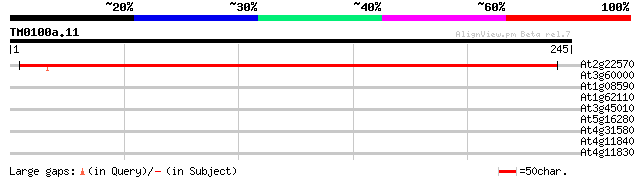
Score E
Sequences producing significant alignments: (bits) Value
At2g22570 unknown protein 324 3e-89
At3g60000 putative protein 30 1.5
At1g08590 putative receptor kinase, CLV1 28 4.5
At1g62110 hypothetical protein 28 5.8
At3g45010 carboxypeptidase precursor-like protein 27 7.6
At5g16280 putative protein 27 10.0
At4g31580 splicing factor 9G8-like SR protein / SRZ-22 27 10.0
At4g11840 putative phospholipase D-gamma 27 10.0
At4g11830 phospholipase D-gamma-2 27 10.0
>At2g22570 unknown protein
Length = 244
Score = 324 bits (830), Expect = 3e-89
Identities = 145/236 (61%), Positives = 194/236 (81%), Gaps = 1/236 (0%)
Query: 5 IIELLKNEIPL-EQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINES 63
I + LK +IP+ E+E ++L D+ GLV+VD++NGFCT+G+GN+AP + N QIS+M+ ES
Sbjct: 7 IFDQLKKQIPVDEEEPLILNRDSSVGLVIVDVVNGFCTIGSGNMAPTKHNEQISKMVEES 66
Query: 64 ARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRR 123
A+LAR FC++ PV+AF+DSHHP+ PE PYPPHCI GT+ES LVPAL+WLE+E TLRR
Sbjct: 67 AKLAREFCDRKWPVLAFIDSHHPDIPERPYPPHCIIGTEESELVPALKWLESEDCATLRR 126
Query: 124 KECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKP 183
K+C +G++GS E DGSNVFVDWVK+ +I+ ++VVGICTDICV DFV + +SA+N G L P
Sbjct: 127 KDCINGFVGSMESDGSNVFVDWVKEKQIKVIVVVGICTDICVFDFVATALSARNHGVLSP 186
Query: 184 LENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLF 239
+E+VVVYS GCATFD+PL VA++ KGA AHPQE +HH+GLYMAK RGA++ +++ F
Sbjct: 187 VEDVVVYSRGCATFDLPLHVAKDIKGAQAHPQELMHHVGLYMAKGRGAQVVSKISF 242
>At3g60000 putative protein
Length = 451
Score = 29.6 bits (65), Expect = 1.5
Identities = 20/70 (28%), Positives = 36/70 (50%), Gaps = 5/70 (7%)
Query: 2 VSQIIELLKN---EIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISE 58
VSQ I+ N +PL+ + +V ++ +GLV I T GN P+ +++
Sbjct: 342 VSQTIQAFSNASLRLPLDGDIMVDSKQLGDGLVAASKIVDGITQNVGNYMPKA--KEMES 399
Query: 59 MINESARLAR 68
+++E R+AR
Sbjct: 400 LLSELTRVAR 409
>At1g08590 putative receptor kinase, CLV1
Length = 1029
Score = 28.1 bits (61), Expect = 4.5
Identities = 18/84 (21%), Positives = 39/84 (46%), Gaps = 5/84 (5%)
Query: 81 LDSHHPNKPEDP---YPPHCIAGTDESNLVPALRWLENETNVTLRRKECFDGYIGSA--E 135
++ HH + E+ + + G N+V L ++ NE V + + +G +G+A
Sbjct: 756 IEDHHQEEDEEDDILREVNLLGGLRHRNIVKILGYVHNEREVMMVYEYMPNGNLGTALHS 815
Query: 136 EDGSNVFVDWVKKNKIRTLLVVGI 159
+D + DW+ + + +V G+
Sbjct: 816 KDEKFLLRDWLSRYNVAVGVVQGL 839
>At1g62110 hypothetical protein
Length = 595
Score = 27.7 bits (60), Expect = 5.8
Identities = 15/40 (37%), Positives = 24/40 (59%), Gaps = 1/40 (2%)
Query: 52 SNRQISEMINESARLARLFCEKNLPV-MAFLDSHHPNKPE 90
+N QIS +I + RL + EK+L + + FL+S + PE
Sbjct: 97 TNSQISSIITDYPRLLLIDAEKSLDIKLQFLESRGASSPE 136
>At3g45010 carboxypeptidase precursor-like protein
Length = 510
Score = 27.3 bits (59), Expect = 7.6
Identities = 12/28 (42%), Positives = 14/28 (49%)
Query: 197 FDVPLEVARNTKGALAHPQEFLHHMGLY 224
FD+P V R G Q+F HH G Y
Sbjct: 79 FDLPAAVDRRGSGGSPSVQDFGHHAGYY 106
>At5g16280 putative protein
Length = 1265
Score = 26.9 bits (58), Expect = 10.0
Identities = 37/159 (23%), Positives = 60/159 (37%), Gaps = 26/159 (16%)
Query: 19 CVVLAEDTV--NGLVLVDIINGFCTVGAGNLAPRESNRQI-------------SEMINES 63
C L EDT NGL V+ + FC ++ R S+ Q+ S++ +
Sbjct: 25 CTPLVEDTFLRNGLSFVETLKPFCNFSNIDVPVRTSSDQLYRLKKFTLRLFYASDIKQPN 84
Query: 64 ARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPA-LRWLENETNVTL- 121
+A+ E+ + + + DP I ES + P+ R+ E TL
Sbjct: 85 VEVAKQRLERVITQAG--EKDFQDLKSDPPQITDILSNPESEIAPSWFRYYNKELIRTLS 142
Query: 122 -RRKECFDG------YIGSAEEDGSNVFVDWVKKNKIRT 153
E FD + S +E+ FVD N++ T
Sbjct: 143 FSDHEAFDHPVACLLVVSSKDEEPIKKFVDLFNSNRLPT 181
>At4g31580 splicing factor 9G8-like SR protein / SRZ-22
Length = 200
Score = 26.9 bits (58), Expect = 10.0
Identities = 14/50 (28%), Positives = 22/50 (44%)
Query: 42 VGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDSHHPNKPED 91
V GNL PR + R++ + + ++ + P AFLD P D
Sbjct: 4 VYVGNLDPRVTERELEDEFRAFGVVRSVWVARRPPGYAFLDFEDPRDARD 53
>At4g11840 putative phospholipase D-gamma
Length = 866
Score = 26.9 bits (58), Expect = 10.0
Identities = 13/44 (29%), Positives = 20/44 (44%)
Query: 169 VCSTMSAKNRGFLKPLENVVVYSHGCATFDVPLEVARNTKGALA 212
+C K F+K E +Y+H T V E A+N + +A
Sbjct: 352 LCPRYGGKGHSFIKKSEVETIYTHHQKTMIVDAEAAQNRRKIVA 395
>At4g11830 phospholipase D-gamma-2
Length = 824
Score = 26.9 bits (58), Expect = 10.0
Identities = 13/44 (29%), Positives = 20/44 (44%)
Query: 169 VCSTMSAKNRGFLKPLENVVVYSHGCATFDVPLEVARNTKGALA 212
+C K F+K E +Y+H T V E A+N + +A
Sbjct: 311 LCPRYGGKGHSFIKKSEVETIYTHHQKTMIVDAEAAQNRRKIVA 354
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,644,953
Number of Sequences: 26719
Number of extensions: 237902
Number of successful extensions: 541
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 536
Number of HSP's gapped (non-prelim): 9
length of query: 245
length of database: 11,318,596
effective HSP length: 97
effective length of query: 148
effective length of database: 8,726,853
effective search space: 1291574244
effective search space used: 1291574244
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0100a.11


