

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094a.6
(990 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
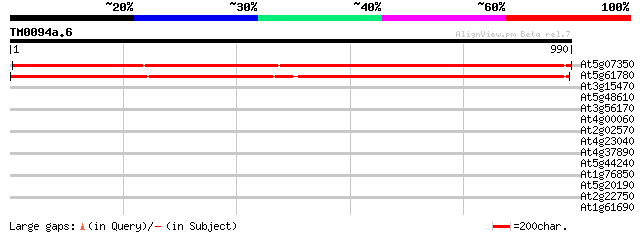
Sequences producing significant alignments: (bits) Value
At5g07350 unknown protein 1431 0.0
At5g61780 100 kDa coactivator - like protein 1406 0.0
At3g15470 unknown protein 38 0.025
At5g48610 putative protein 38 0.032
At3g56170 Ca(2+)-dependent nuclease 35 0.16
At4g00060 unknown protein 33 1.0
At2g02570 unknown protein 32 1.4
At4g23040 unknown protein 31 4.0
At4g37890 unknown protein 30 5.2
At5g44240 ATPase, calcium-transporting 30 6.7
At1g76850 unknown protein 30 6.7
At5g20190 putative protein 30 8.8
At2g22750 putative bHLH transcription factor (bHLH018) 30 8.8
At1g61690 hypothetical protein 30 8.8
>At5g07350 unknown protein
Length = 991
Score = 1431 bits (3703), Expect = 0.0
Identities = 726/995 (72%), Positives = 843/995 (83%), Gaps = 14/995 (1%)
Query: 5 ATGATG-WYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVDEPF 63
ATGA W +GRVKAV SGDCLVI A++ ++ GP PEK+IT +SL+ P++ARRGG+DEPF
Sbjct: 2 ATGAENQWLKGRVKAVTSGDCLVITALSHNRAGPPPEKTITFSSLMAPKMARRGGIDEPF 61
Query: 64 AWESREYLRKLCIGKEVTFRVDYNVASI-NRDFGTVFLGEKNVGVLVVSQGWAKVREQGQ 122
AWES+E+LRKLCIGKEV F+VDY V +I R+FG+VFLG +N+ LVV GWAKVRE GQ
Sbjct: 62 AWESKEFLRKLCIGKEVAFKVDYKVEAIAGREFGSVFLGNENLAKLVVKTGWAKVREPGQ 121
Query: 123 Q-KGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLA 181
Q + +VSPY+ ELL+LEE AKQEG GRWSKVPGAAEASIRNLPPSAIGD++ FDAMGLLA
Sbjct: 122 QNQDKVSPYIKELLQLEELAKQEGYGRWSKVPGAAEASIRNLPPSAIGDSAGFDAMGLLA 181
Query: 182 ANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTAD 241
ANKG PME IVEQVRDGST+RVYLLPEFQFVQVFVAG+Q+P MGRR +VVET D
Sbjct: 182 ANKGKPMEGIVEQVRDGSTIRVYLLPEFQFVQVFVAGVQAPSMGRRTTNGSVVET--VPD 239
Query: 242 ENDGDVPGEPRPPLTSAQRLAVS--SSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVD 299
E +GDV E R PLT+AQRLA S SS E ++DPF +AK+FTE RVL+RDVRIVLEGVD
Sbjct: 240 EPNGDVSAESRGPLTTAQRLAASAASSVEVSSDPFATEAKYFTEHRVLSRDVRIVLEGVD 299
Query: 300 KFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRL 359
KF+NLIGSV+Y DGE+ KDL LELVENG AK+VEWSANMMEEEAK++LK AEL+ KK ++
Sbjct: 300 KFNNLIGSVHYSDGETVKDLGLELVENGLAKFVEWSANMMEEEAKKKLKAAELQCKKDKV 359
Query: 360 RMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCP 419
+MW NYVPPA+NSKAIH+QNFTGKVVEVVSGDC+IVADD++P+GSP AERRV LSSIR P
Sbjct: 360 KMWANYVPPATNSKAIHDQNFTGKVVEVVSGDCLIVADDAVPFGSPAAERRVCLSSIRSP 419
Query: 420 KVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMD 479
K+GNPRR+EKPAPYAREA+EFLR RL+G+QV V+MEYSRK+ DG S AAD R MD
Sbjct: 420 KMGNPRREEKPAPYAREAREFLRQRLIGKQVIVQMEYSRKVTQGDGPTT-SGAAD-RFMD 477
Query: 480 FGSVFLLSATKADSDDT--PSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYY 537
FGSVFL SA KADSD+ P + AGSQP GVN+ ELV+ RGFG V+RHRDFEERSN+Y
Sbjct: 478 FGSVFLPSAAKADSDEVTAPPAAAIAGSQPVGVNIAELVLVRGFGNVVRHRDFEERSNHY 537
Query: 538 DALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVL 597
DALL AE+RAL+G+KGIHSAK+ P MHITDLT ++AKKAKDFLP L R RRIPAVVEYVL
Sbjct: 538 DALLAAEARALAGKKGIHSAKESPAMHITDLTVSAAKKAKDFLPSLQRIRRIPAVVEYVL 597
Query: 598 SGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNG 657
SGHRFKL IPK TCSIAF+ SGVRCPGRGEPYSEEAI++MRR+IMQRDVEIEVETVDR G
Sbjct: 598 SGHRFKLYIPKITCSIAFSFSGVRCPGRGEPYSEEAISVMRRRIMQRDVEIEVETVDRTG 657
Query: 658 TFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVE 717
TFLGS+WESRTNVA LLEAGLAK+QTSFG+DRI E HLL++AE+SAK QKLKIWEN+VE
Sbjct: 658 TFLGSMWESRTNVATVLLEAGLAKMQTSFGADRIAEAHLLEQAERSAKNQKLKIWENYVE 717
Query: 718 GEEVSNGAN--VESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPV 775
GEEVSNG VE++Q+E LKV+VTEVLGGG+FYVQ+ GDQKIASIQ QLA+L++K+AP+
Sbjct: 718 GEEVSNGNTNTVETRQKETLKVVVTEVLGGGRFYVQSAGDQKIASIQNQLASLSIKDAPI 777
Query: 776 LGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQL 835
+G+F+PK+GDIVL F D SW RAM+V PR V+SP + FEVFYIDYGNQE V YS +
Sbjct: 778 IGSFNPKRGDIVLAQFSLDNSWNRAMIVTAPRAAVQSPDEKFEVFYIDYGNQETVPYSAI 837
Query: 836 RPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTS 895
RP+D SVSAAPGLAQLC LAYIK PSLE+DFG EA EYL +TL SGKEF+A +EERDTS
Sbjct: 838 RPIDPSVSAAPGLAQLCRLAYIKVPSLEDDFGPEAGEYLHTVTLGSGKEFKAVIEERDTS 897
Query: 896 GGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQ 955
GGKVKGQGTGT VTL+AVD EISVNAAMLQEG+ARMEKR +W K ++ LD+L+KFQ
Sbjct: 898 GGKVKGQGTGTEFVVTLIAVDDEISVNAAMLQEGIARMEKRQKWGHKGKQAALDALEKFQ 957
Query: 956 DDARKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
++ARK R G+WQYGD+ESDDEDT PARK GRR
Sbjct: 958 EEARKSRIGIWQYGDIESDDEDTG-PARKPAGGRR 991
>At5g61780 100 kDa coactivator - like protein
Length = 985
Score = 1406 bits (3639), Expect = 0.0
Identities = 710/996 (71%), Positives = 833/996 (83%), Gaps = 19/996 (1%)
Query: 1 MASAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVD 60
MA+ A W +GRVKAV SGDCLVI A+ ++ GP PEK+ITL+SL+ P++ARRGG+D
Sbjct: 1 MATGAATENQWLKGRVKAVTSGDCLVITALTHNRAGPPPEKTITLSSLMAPKMARRGGID 60
Query: 61 EPFAWESREYLRKLCIGKEVTFRVDYNVASI-NRDFGTVFLGEKNVGVLVVSQGWAKVRE 119
EPFAWESRE+LRKLCIGKEV F+VDY V +I R+FG+V+LG +N+ LVV GWAKVR
Sbjct: 61 EPFAWESREFLRKLCIGKEVAFKVDYKVEAIAGREFGSVYLGNENLAKLVVQNGWAKVRR 120
Query: 120 QGQQ-KGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMG 178
GQQ + +VSPY+AEL +LEEQA+QEG GRWSKVPGAAEASIRNLPPSA+GD+ NFDAMG
Sbjct: 121 PGQQNQDKVSPYIAELEQLEEQAQQEGFGRWSKVPGAAEASIRNLPPSAVGDSGNFDAMG 180
Query: 179 LLAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRR-AAPETVVETE 237
LLAA+KG PME IVEQVRDGST+RVYLLPEFQFVQVFVAG+Q+P MGRR + E VV+ +
Sbjct: 181 LLAASKGKPMEGIVEQVRDGSTIRVYLLPEFQFVQVFVAGLQAPSMGRRQSTQEAVVDPD 240
Query: 238 LTADENDGDVPGEPRPPLTSAQRLAVS--SSTETAADPFGPDAKFFTEMRVLNRDVRIVL 295
+TA N GD E R PLT+AQRLA S SS E ++DPF +AK+FTE+RVLNRDVRIVL
Sbjct: 241 VTATSN-GDASAETRGPLTTAQRLAASAASSVEVSSDPFAMEAKYFTELRVLNRDVRIVL 299
Query: 296 EGVDKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAK 355
EGVDKF+NLIGSVYY DG++ KDL LELVENG AKYVEWSANM++EEAK++LK EL+ K
Sbjct: 300 EGVDKFNNLIGSVYYSDGDTVKDLGLELVENGLAKYVEWSANMLDEEAKKKLKATELQCK 359
Query: 356 KSRLRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSS 415
K+R++MW NYVPPASNSKAIH+QNFTGKVVEVVSGDC++VADDSIP+GSP+AERRV LSS
Sbjct: 360 KNRVKMWANYVPPASNSKAIHDQNFTGKVVEVVSGDCLVVADDSIPFGSPMAERRVCLSS 419
Query: 416 IRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADS 475
IR PK+GNPRR+EKPAPYAREAKEFLR +L+G +V V+MEYSRKI P DG V + A
Sbjct: 420 IRSPKMGNPRREEKPAPYAREAKEFLRQKLIGMEVIVQMEYSRKISPGDG--VTTSGAGD 477
Query: 476 RVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSN 535
RVMDFGSVFL S TK D+ ++ P G N+ EL++ RG GTV+RHRDFEERSN
Sbjct: 478 RVMDFGSVFLPSPTKGDTAVAAAATP-------GANIAELIISRGLGTVVRHRDFEERSN 530
Query: 536 YYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEY 595
+YDALL AE+RA++G+K IHSAKD P +HI DLT SAKKAKDFLP L R +I AVVEY
Sbjct: 531 HYDALLAAEARAIAGKKNIHSAKDSPALHIADLTVASAKKAKDFLPSLQRINQISAVVEY 590
Query: 596 VLSGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDR 655
VLSGHRFKL IPKE+CSIAFA SGVRCPGRGEPYSEEAIALMRRKIMQRDVEI VE VDR
Sbjct: 591 VLSGHRFKLYIPKESCSIAFAFSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIVVENVDR 650
Query: 656 NGTFLGSLWE--SRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWE 713
GTFLGS+WE S+TN LLEAGLAK+QT FG+DRIPE H+L+ AE+SAK QKLKIWE
Sbjct: 651 TGTFLGSMWEKNSKTNAGTYLLEAGLAKMQTGFGADRIPEAHILEMAERSAKNQKLKIWE 710
Query: 714 NFVEGEEVSNGAN-VESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKE 772
N+VEGEEV NG++ VE++Q+E LKV+VTEVLGGG+FYVQTVGDQK+ASIQ QLAAL+LK+
Sbjct: 711 NYVEGEEVVNGSSKVETRQKETLKVVVTEVLGGGRFYVQTVGDQKVASIQNQLAALSLKD 770
Query: 773 APVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAY 832
AP++G+F+PKKGDIVL F D SW RAM+VN PRG V+SP++ FEVFYIDYGNQE V Y
Sbjct: 771 APIIGSFNPKKGDIVLAQFSLDNSWNRAMIVNGPRGAVQSPEEEFEVFYIDYGNQEIVPY 830
Query: 833 SQLRPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEER 892
S +RP+D SVS+APGLAQLC LAYIK P EEDFG++A EYL +TL SGKEFRA VEER
Sbjct: 831 SAIRPVDPSVSSAPGLAQLCRLAYIKVPGKEEDFGRDAGEYLHTVTLESGKEFRAVVEER 890
Query: 893 DTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQ 952
DTSGGKVKGQGTGT L VTL+AVD EISVNAAMLQEG+ARMEKR RW+ K+++ LD+L+
Sbjct: 891 DTSGGKVKGQGTGTELVVTLIAVDDEISVNAAMLQEGIARMEKRRRWEPKDKQAALDALE 950
Query: 953 KFQDDARKERRGMWQYGDVESDDEDTAPPARKAGAG 988
KFQD+ARK R G+W+YGD++SDDED P RK G G
Sbjct: 951 KFQDEARKSRTGIWEYGDIQSDDEDNV-PVRKPGRG 985
>At3g15470 unknown protein
Length = 883
Score = 38.1 bits (87), Expect = 0.025
Identities = 30/106 (28%), Positives = 48/106 (44%), Gaps = 5/106 (4%)
Query: 862 LEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISV 921
L E E E + L +GKEF + D + KVK GTGT + + + E+ V
Sbjct: 234 LRESVVNEEVEVCTIKNLDNGKEFVVNEIQEDGTWKKVKEVGTGTQMTME----EFEMCV 289
Query: 922 NAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARK-ERRGMW 966
+ + + L R + D+ K DS +D+A K +++G W
Sbjct: 290 GHSPIVQELMRRQNVEDSDKNTSKENEDSGNSNKDNASKSKKKGSW 335
>At5g48610 putative protein
Length = 470
Score = 37.7 bits (86), Expect = 0.032
Identities = 48/182 (26%), Positives = 72/182 (39%), Gaps = 27/182 (14%)
Query: 129 PYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSA-IGDASNFDAMGLLAANKGSP 187
P L +LEE+ K R K+P + SI+NL +G + G L N +P
Sbjct: 254 PKLIRGPKLEEREKDSPDLRNCKLPDVSRTSIKNLHTEGNLGKRKDHMTNGFLYENGTTP 313
Query: 188 -----MEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADE 242
+ A V V +G TL P +V G P+ V E +
Sbjct: 314 HKLQKLSASVPSVENGRTLGAPRTPPMPASEV---------QGTTCKPQ-VKEVRI---- 359
Query: 243 NDGDVPGE-----PRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEG 297
N V GE P PL + ++ V + E +A P PD K+ + +L+ R +L
Sbjct: 360 NGFAVSGEKRKVCPPSPLAATMKVKVKENGEASAKPPHPDLKYLNQ--ILHVPTRELLPE 417
Query: 298 VD 299
+D
Sbjct: 418 ID 419
>At3g56170 Ca(2+)-dependent nuclease
Length = 323
Score = 35.4 bits (80), Expect = 0.16
Identities = 30/119 (25%), Positives = 59/119 (49%), Gaps = 7/119 (5%)
Query: 596 VLSGHRFKLLIPKETCSIAFA--LSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETV 653
+ SG+R +E + F LSG+ P PY +EA + + + + +++ V T
Sbjct: 191 IASGYRMISFQNEEVLAKKFRIRLSGIDSPESKMPYGKEAHDELLKMVEGKCLKVLVYTE 250
Query: 654 DRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIW 712
DR G +G ++ + V +L+ GLA ++ D+ E L + E A+++++ +W
Sbjct: 251 DRYGRCVGDIYCNGKFVQEVMLKKGLAWHYVAY--DKRAE---LAKWENEARQKRVGLW 304
>At4g00060 unknown protein
Length = 1344
Score = 32.7 bits (73), Expect = 1.0
Identities = 22/90 (24%), Positives = 39/90 (42%), Gaps = 7/90 (7%)
Query: 397 DDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEY 456
DD+ SP + L S + G R +KP P A+ K + G++ ++E+
Sbjct: 270 DDNTHKCSPSSSSNQKLGSTNRKQKGKTRNMKKPTPEAKSDKNVNLSTKNGKKDQAKLEF 329
Query: 457 SR-------KIVPTDGSAVPSPAADSRVMD 479
++ K VPT + + P A + M+
Sbjct: 330 NKSREAIECKKVPTASTMINDPEASAATME 359
>At2g02570 unknown protein
Length = 300
Score = 32.3 bits (72), Expect = 1.4
Identities = 38/156 (24%), Positives = 69/156 (43%), Gaps = 20/156 (12%)
Query: 705 KKQKLKIWENFVEGEEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIAS---- 760
K+Q ++ + E S A++E + +EV+ + EVL K ++ D +++
Sbjct: 21 KEQLEQVRQLLSEDPRNSEYADMEKELKEVI-ALTEEVLATAKQNEISLSDAGVSAEATP 79
Query: 761 ----IQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMV-VNTPRGPVESPKD 815
++ L+ P+ P G V F D WY A + +T G
Sbjct: 80 GSPDLEGAWEKTGLRNDPIHEGKFPV-GTKVQAVFSDDGEWYDATIEAHTANG------- 131
Query: 816 IFEVFYIDYGNQEQVAYSQLRPLDQ-SVSAAPGLAQ 850
+ V Y ++GN+E+V +RP++Q ++ A LAQ
Sbjct: 132 -YFVAYDEWGNKEEVDPDNVRPIEQNAIVEAERLAQ 166
>At4g23040 unknown protein
Length = 525
Score = 30.8 bits (68), Expect = 4.0
Identities = 22/58 (37%), Positives = 31/58 (52%), Gaps = 3/58 (5%)
Query: 225 GRRAAPETVVETELTADENDGDVPGEPRPPLTSAQ-RLAVSSSTETAAD--PFGPDAK 279
GR AAP ++ E + D++D D E PL S + R AVS S + D P P+A+
Sbjct: 245 GRFAAPSSLSEDDDDDDDDDPDYVEEEEEPLVSHRPRRAVSGSRSSLNDDLPRSPEAE 302
>At4g37890 unknown protein
Length = 739
Score = 30.4 bits (67), Expect = 5.2
Identities = 23/73 (31%), Positives = 36/73 (48%), Gaps = 12/73 (16%)
Query: 229 APETVVETELTADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGPDAKF-------- 280
+P +L D + P R PL+ L+VSSS+ T + P P A F
Sbjct: 75 SPSLPASPKLQCDTSGDVTPTRNRSPLSF---LSVSSSSSTPSSPKSP-ASFSLLKSKLC 130
Query: 281 FTEMRVLNRDVRI 293
FT++R+LNR+ ++
Sbjct: 131 FTKVRILNRNNQV 143
>At5g44240 ATPase, calcium-transporting
Length = 1078
Score = 30.0 bits (66), Expect = 6.7
Identities = 10/24 (41%), Positives = 19/24 (78%)
Query: 319 LALELVENGYAKYVEWSANMMEEE 342
++L+LV+ YAK++EW M+++E
Sbjct: 290 VSLDLVKGLYAKFIEWDVEMIDQE 313
>At1g76850 unknown protein
Length = 1090
Score = 30.0 bits (66), Expect = 6.7
Identities = 35/140 (25%), Positives = 55/140 (39%), Gaps = 26/140 (18%)
Query: 827 QEQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFR 886
++ VA +P Q AA S A ++ PS++ED E + L++SSG +
Sbjct: 37 RKPVANLVQQPRQQKPVAAAAAPPKKSAAAVRKPSMDEDEESE----VELLSISSGDD-- 90
Query: 887 AQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKE--- 943
+E GG G G G + + ++G AR E WD E
Sbjct: 91 -DLEREREIGGSSGGAGRGR---------------GSDVREKGRARKEDDGAWDGGEPDC 134
Query: 944 -RKVGLDSLQKFQDDARKER 962
++V L + D R+ R
Sbjct: 135 WKRVNEAELARRVRDMRESR 154
>At5g20190 putative protein
Length = 290
Score = 29.6 bits (65), Expect = 8.8
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 1/44 (2%)
Query: 324 VENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVP 367
V+ YA+++ W A EEE K ELE + SR+ +T P
Sbjct: 243 VQASYARFL-WDAEEEEEEEKEERHEEELEHQTSRMNFFTGPSP 285
>At2g22750 putative bHLH transcription factor (bHLH018)
Length = 304
Score = 29.6 bits (65), Expect = 8.8
Identities = 18/71 (25%), Positives = 37/71 (51%), Gaps = 3/71 (4%)
Query: 328 YAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPASNSKAIHNQNFTGKVVEV 387
+ KY++ S EE+ K + + + KKS L + N+ P +S+S + + + + E+
Sbjct: 167 HIKYLQESVKEYEEQKKEKTMESVVLVKKSSLVLDENHQPSSSSSSDGNRNSSSSNLPEI 226
Query: 388 ---VSGDCIIV 395
VSG +++
Sbjct: 227 EVRVSGKDVLI 237
>At1g61690 hypothetical protein
Length = 576
Score = 29.6 bits (65), Expect = 8.8
Identities = 29/111 (26%), Positives = 45/111 (40%), Gaps = 8/111 (7%)
Query: 405 PLAERRVNLSSIRCPKVGNPRR----DEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKI 460
P A + N S PK + R ++ + R+A + R + ++ E E ++ +
Sbjct: 399 PPAREKENSPSSSAPKAMSGRDRFKLQQESLSHKRQAMKLRREGKM-QEAEAEFEIAKTL 457
Query: 461 ---VPTDGSAVPSPAADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPT 508
+ S+ P P D V DF LLSA KA D P + P T
Sbjct: 458 EAQLEDSTSSKPEPVDDVAVEDFLDPQLLSALKAIGLDNPVNPPPVSKTDT 508
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.133 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,982,015
Number of Sequences: 26719
Number of extensions: 966585
Number of successful extensions: 2007
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1963
Number of HSP's gapped (non-prelim): 16
length of query: 990
length of database: 11,318,596
effective HSP length: 109
effective length of query: 881
effective length of database: 8,406,225
effective search space: 7405884225
effective search space used: 7405884225
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0094a.6


