

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.9
(742 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
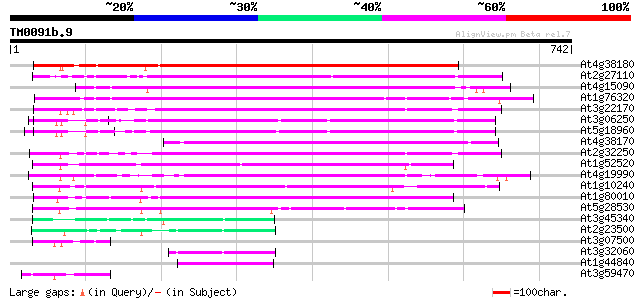
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 472 e-133
At2g27110 Mutator-like transposase 380 e-105
At4g15090 unknown protein 353 1e-97
At1g76320 putative phytochrome A signaling protein 349 3e-96
At3g22170 far-red impaired response protein, putative 339 4e-93
At3g06250 unknown protein 320 2e-87
At5g18960 FAR1 - like protein 317 1e-86
At4g38170 hypothetical protein 315 8e-86
At2g32250 Mutator-like transposase 313 2e-85
At1g52520 F6D8.26 292 5e-79
At4g19990 putative protein 281 9e-76
At1g10240 unknown protein 275 7e-74
At1g80010 hypothetical protein 273 2e-73
At5g28530 far-red impaired response protein (FAR1) - like 225 8e-59
At3g45340 putative protein 67 5e-11
At2g23500 Mutator-like transposase 59 8e-09
At3g07500 hypothetical protein 56 6e-08
At3g32060 hypothetical protein 54 3e-07
At1g44840 hypothetical protein 54 3e-07
At3g59470 unknown protein 53 7e-07
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 472 bits (1215), Expect = e-133
Identities = 232/575 (40%), Positives = 358/575 (61%), Gaps = 20/575 (3%)
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR---AVL----VCCNEGQHKVKISRTE 84
G+EFES E + FYNS+A++ GF RV SS+ R A++ VC EG + RT+
Sbjct: 76 GLEFESEEAAKAFYNSYARRIGFSTRVSSSRRSRRDGAIIQRQFVCAKEGFRNMNEKRTK 135
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCH 144
+ + KR + R GC+ASL V + KW++ F DHNH +V P V +R H
Sbjct: 136 D-----REIKRPRTITRVGCKASLSVKMQDS-GKWLVSGFVKDHNHELVPPDQVHCLRSH 189
Query: 145 KKMSVAAKSLVEKFEEEGI-PTGKVAAIFNDGDST----FTNRDCWNYIRNVRRKNLDVG 199
+++S AK+L++ + G+ P ++A+ + FT DC NY+RN R+K+++ G
Sbjct: 190 RQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRNYMRNNRQKSIE-G 248
Query: 200 DAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKT 259
+ Q + DY ++ +N NFFY++Q ED + N+FW D ++ + + FGD +TFDTTY++
Sbjct: 249 EIQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDFTHFGDTVTFDTTYRS 308
Query: 260 NKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDL 319
N+Y +PFAPF G+N+H Q IL+GCA + +E+EASFVWLF T L AM P+SI TD D
Sbjct: 309 NRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAAMSAHPPVSITTDHDA 368
Query: 320 AMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEE 379
++AA+ VFP +RHR C WHI+KK EKL+H++ K F+ D +C+ + S++ FE
Sbjct: 369 VIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHPSFESDFHKCVNLTESVEDFERC 428
Query: 380 WMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAST 439
W ++ +Y L+++EWLQ +Y R W+P+Y R TFFA M+ T RS+SIN++FD +++AST
Sbjct: 429 WFSLLDKYELRDHEWLQAIYSDRRQWVPVYLRDTFFADMSLTHRSDSINSYFDGYINAST 488
Query: 440 TLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGEL 499
L +F +EKA+ESRLE E K +Y++ + +L T S +E+ A+ +YTR +F +F+ EL
Sbjct: 489 NLSQFFKLYEKALESRLEKEVKADYDTMNSPPVLKTPSPMEKQASELYTRKLFMRFQEEL 548
Query: 500 EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHI 559
F K DG YQV+ +A V ++ A C +++EF GI+CRHI
Sbjct: 549 VGTLTFMASKADDDGDLVTYQVAKYGEAHKAHFVKFNVLEMRANCSCQMFEFSGIICRHI 608
Query: 560 LVIFQAKGVVQIPSHFIMERWTKDANRG-LEDTYN 593
L +F+ ++ +P ++I++RWT++A + D YN
Sbjct: 609 LAVFRVTNLLTLPPYYILKRWTRNAKSSVIFDDYN 643
>At2g27110 Mutator-like transposase
Length = 851
Score = 380 bits (976), Expect = e-105
Identities = 210/622 (33%), Positives = 345/622 (54%), Gaps = 29/622 (4%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDST 90
+GMEF S ++ + FY+ ++++ GF +SK L+ +G V R S+
Sbjct: 51 VGMEFNSEKEAKSFYDEYSRQLGF-----TSK-----LLPRTDGSVSV---REFVCSSSS 97
Query: 91 NQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVA 150
++KR+ S C+A + + KW++ F +H H + S + +R + + +
Sbjct: 98 KRSKRRLS---ESCDAMVRIEL-QGHEKWVVTKFVKEHTHGLASSNMLHCLRPRRHFANS 153
Query: 151 AKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKR 210
KS + E +P+G + + D +S R N K DA + +Y KR
Sbjct: 154 EKSSYQ--EGVNVPSGMMY-VSMDANS----RGARNASMATNTKRTIGRDAHNLLEYFKR 206
Query: 211 KQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFV 270
Q EN FFYA+Q DED++M N+FW D+RSR++Y FGD +T DT Y+ N++ +PFAPF
Sbjct: 207 MQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRVPFAPFT 266
Query: 271 GLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFP 330
G+N+H Q+IL+GCAL+ DES+ SF+WLFKT L AM + P+S++TDQD A++ A +VFP
Sbjct: 267 GVNHHGQAILFGCALILDESDTSFIWLFKTFLTAMRDQPPVSLVTDQDRAIQIAAGQVFP 326
Query: 331 ESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIMVEYNLK 390
+RH + W ++++ EKLAH+ F+ ++ CI + +I+ FE W ++ +Y+L
Sbjct: 327 GARHCINKWDVLREGQEKLAHVCLAYPSFQVELYNCINFTETIEEFESSWSSVIDKYDLG 386
Query: 391 ENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEK 450
+EWL LY R W+P+Y R +FFA + +Q +FFD +V+ TTL F +E+
Sbjct: 387 RHEWLNSLYNARAQWVPVYFRDSFFAAVFPSQGYS--GSFFDGYVNQQTTLPMFFRLYER 444
Query: 451 AVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKI 510
A+ES E E + + ++ + +L T S +E AA+++TR IF KF+ EL + T ++I
Sbjct: 445 AMESWFEMEIEADLDTVNTPPVLKTPSPMENQAANLFTRKIFGKFQEELVETFAHTANRI 504
Query: 511 RRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQ 570
DG ++V+N + + V A C +++E GILCRH+L +F ++
Sbjct: 505 EDDGTTSTFRVANFENDNKAYIVTFCYPEMRANCSCQMFEHSGILCRHVLTVFTVTNILT 564
Query: 571 IPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAY 630
+P H+I+ RWT++A +E D + E I R H+ REA A+ + EAY
Sbjct: 565 LPPHYILRRWTRNAKSMVE---LDEHVSENGHDSSIHRYNHLCREAIKYAEEGAITAEAY 621
Query: 631 NFIISEMKQTHKSAAAMKTSEG 652
N + ++++ K + ++ G
Sbjct: 622 NIALGQLREGGKKVSVVRKRIG 643
>At4g15090 unknown protein
Length = 768
Score = 353 bits (907), Expect = 1e-97
Identities = 198/587 (33%), Positives = 319/587 (53%), Gaps = 26/587 (4%)
Query: 88 DSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKM 147
+S+ + R+ + ++ C+AS+ V R KW+I F DHNH ++ P + R + +
Sbjct: 50 ESSGSSSRRSTVKKTDCKASMHVKR-RPDGKWIIHEFVKDHNHELL-PALAYHFRIQRNV 107
Query: 148 SVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTN------RDCWNYIRNVRRKNLDVGDA 201
+A K+ ++ T K+ + + N D + + R L+ GD+
Sbjct: 108 KLAEKNNIDILHAVSERTKKMYVEMSRQSGGYKNIGSLLQTDVSSQVDKGRYLALEEGDS 167
Query: 202 QAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNK 261
Q + +Y KR + EN FFYAI +ED R+ NLFW DA+SR Y F DV++FDTTY
Sbjct: 168 QVLLEYFKRIKKENPKFFYAIDLNEDQRLRNLFWADAKSRDDYLSFNDVVSFDTTYVKFN 227
Query: 262 YSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAM 321
+P A F+G+N+HSQ +L GCAL+ DES +FVWL KT L+AMGG+ P I+TDQD +
Sbjct: 228 DKLPLALFIGVNHHSQPMLLGCALVADESMETFVWLIKTWLRAMGGRAPKVILTDQDKFL 287
Query: 322 KAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWM 381
+A++++ P +RH LWH+++K PE +H+ + F +CI S + F+ W
Sbjct: 288 MSAVSELLPNTRHCFALWHVLEKIPEYFSHVMKRHENFLLKFNKCIFRSWTDDEFDMRWW 347
Query: 382 RIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTL 441
+++ ++ L+ +EWL L++ R+ W+P + F AGM+T+QRSES+N+FFD ++ TL
Sbjct: 348 KMVSQFGLENDEWLLWLHEHRQKWVPTFMSDVFLAGMSTSQRSESVNSFFDKYIHKKITL 407
Query: 442 QEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEK 501
+EF+ ++ +++R E E ++++ HK L + S E+ A+ YT IF KF+ E+
Sbjct: 408 KEFLRQYGVILQNRYEEESVADFDTCHKQPALKSPSPWEKQMATTYTHTIFKKFQVEVLG 467
Query: 502 INRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILV 561
+ K + D ++V +C + D F V C ++E+ G LCRH L+
Sbjct: 468 VVACHPRKEKEDENMATFRVQDC-EKDDDFLVTWSKTKSELCCFCRMFEYKGFLCRHALM 526
Query: 562 IFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQRE---ASF 618
I Q G IP +I++RWTKDA G+ GE +D + + VQR S
Sbjct: 527 ILQMCGFASIPPQYILKRWTKDAKSGVL-------AGEGADQI----QTRVQRYNDLCSR 575
Query: 619 LADLAEE---SEEAYNFIISEMKQTHKSAAAMKTSEGVVLLESSEKN 662
+L+EE SEE YN + + +T K+ M + + +S+ N
Sbjct: 576 ATELSEEGCVSEENYNIALRTLVETLKNCVDMNNARNNITESNSQLN 622
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 349 bits (896), Expect = 3e-96
Identities = 208/671 (30%), Positives = 346/671 (50%), Gaps = 24/671 (3%)
Query: 33 MEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDSTNQ 92
MEFE+ E FY +AK GFG SS+ RA + ++ ++ D+ N
Sbjct: 1 MEFETHEDAYLFYKDYAKSVGFGTAKLSSRRSRASKEFIDAKFSCIRYGSKQQSDDAINP 60
Query: 93 TKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVAAK 152
++ + GC+AS+ V R KW + SF +HNH ++ P+ Y R H+ +
Sbjct: 61 R----ASPKIGCKASMHVKR-RPDGKWYVYSFVKEHNHDLL-PEQAHYFRSHRNTELVKS 114
Query: 153 SLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQ 212
+ ++ P + D F + N RR LD GDA+ + ++ R Q
Sbjct: 115 NDSRLRRKKNTPLTDCKHLSAYHDLDFIDGYMRNQHDKGRRLVLDTGDAEILLEFLMRMQ 174
Query: 213 VENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGL 272
EN FF+A+ ED + N+FWVDA+ Y+ F DV++F+T+Y +KY +P FVG+
Sbjct: 175 EENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKVPLVLFVGV 234
Query: 273 NNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFPES 332
N+H Q +L GC LL D++ ++VWL ++ L AMGG+KP ++TDQ+ A+KAA+A V PE+
Sbjct: 235 NHHVQPVLLGCGLLADDTVYTYVWLMQSWLVAMGGQKPKVMLTDQNNAIKAAIAAVLPET 294
Query: 333 RHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIMVEYNLKEN 392
RH CLWH++ + P L + Q F + + +CI S S + F+ W++++ +++L++
Sbjct: 295 RHCYCLWHVLDQLPRNLDYWSMWQDTFMKKLFKCIYRSWSEEEFDRRWLKLIDKFHLRDV 354
Query: 393 EWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEKAV 452
W++ LY+ R+ W P + R FAG++ RSES+N+ FD +V T+L+EF+ + +
Sbjct: 355 PWMRSLYEERKFWAPTFMRGITFAGLSMRCRSESVNSLFDRYVHPETSLKEFLEGYGLML 414
Query: 453 ESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKIRR 512
E R E E K ++++ H++ L + S E+ +Y+ IF +F +LE + H +
Sbjct: 415 EDRYEEEAKADFDAWHEAPELKSPSPFEKQMLLVYSHEIFRRF--QLEVLGAAACHLTKE 472
Query: 513 DGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIP 572
Y V + +D + VD D C +E+ G LCRH +V+ Q GV IP
Sbjct: 473 SEEGTTYSVKD-FDDEQKYLVDWDEFKSDIYCSCRSFEYKGYLCRHAIVVLQMSGVFTIP 531
Query: 573 SHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAYNF 632
+++++RWT +A R + +L + + I R + R A L + S+E+Y+
Sbjct: 532 INYVLQRWT-NAARNRHQISRNLELVQSN----IRRFNDLCRRAIILGEEGSLSQESYDI 586
Query: 633 IISEMKQTHKSAAA--------MKTSEGVVLLESSEKNVNQVCS-SEQVSEPPQLTIGN- 682
+ MK+ K A + E + + NQ S S Q+ P + GN
Sbjct: 587 AMFAMKEAFKQCAVTINTIKHPARCEEAAIQAGDPVQEENQYGSTSTQIGPEPNIHAGNV 646
Query: 683 PHVSQTKGRKK 693
P ++T+ K+
Sbjct: 647 PWQAETRREKR 657
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 339 bits (869), Expect = 4e-93
Identities = 202/633 (31%), Positives = 331/633 (51%), Gaps = 42/633 (6%)
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR-------AVLVCCNEG---QHKVKIS 81
GMEFES + FY +++ GF +++S+ + A C G ++ +
Sbjct: 73 GMEFESHGEAYSFYQEYSRAMGFNTAIQNSRRSKTTREFIDAKFACSRYGTKREYDKSFN 132
Query: 82 RT---EEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSV 138
R + QD N R+ + ++ C+AS+ V R KW+I SF +HNH ++ ++V
Sbjct: 133 RPRARQSKQDPENMAGRR-TCAKTDCKASMHVKR-RPDGKWVIHSFVREHNHELLPAQAV 190
Query: 139 SYMRCHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDV 198
S + K + AK E V ++ +D S+F R +++
Sbjct: 191 SE-QTRKIYAAMAKQFAEY--------KTVISLKSDSKSSFEKG---------RTLSVET 232
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYK 258
GD + + D+ R Q NSNFFYA+ +D R+ N+FWVDA+SR +Y F DV++ DTTY
Sbjct: 233 GDFKILLDFLSRMQSLNSNFFYAVDLGDDQRVKNVFWVDAKSRHNYGSFCDVVSLDTTYV 292
Query: 259 TNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQD 318
NKY MP A FVG+N H Q ++ GCAL+ DES A++ WL +T L+A+GG+ P +IT+ D
Sbjct: 293 RNKYKMPLAIFVGVNQHYQYMVLGCALISDESAATYSWLMETWLRAIGGQAPKVLITELD 352
Query: 319 LAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEE 378
+ M + + ++FP +RH L LWH++ K E L + + F ++CI S + F
Sbjct: 353 VVMNSIVPEIFPNTRHCLFLWHVLMKVSENLGQVVKQHDNFMPKFEKCIYKSGKDEDFAR 412
Query: 379 EWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAS 438
+W + + + LK+++W+ LY+ R+ W P Y AGM+T+QR++SINAFFD ++
Sbjct: 413 KWYKNLARFGLKDDQWMISLYEDRKKWAPTYMTDVLLAGMSTSQRADSINAFFDKYMHKK 472
Query: 439 TTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGE 498
T++QEFV ++ ++ R E E K + E +K + + S E+ + +YT +F KF+ E
Sbjct: 473 TSVQEFVKVYDTVLQDRCEEEAKADSEMWNKQPAMKSPSPFEKSVSEVYTPAVFKKFQIE 532
Query: 499 LEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRH 558
+ + + RD ++V + ++ F V + C L+E+ G LCRH
Sbjct: 533 VLGAIACSPREENRDATCSTFRVQD-FENNQDFMVTWNQTKAEVSCICRLFEYKGYLCRH 591
Query: 559 ILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSD-TLKILRRVHVQREAS 617
L + Q + IPS +I++RWTKDA + + GE ++LR + A
Sbjct: 592 TLNVLQCCHLSSIPSQYILKRWTKDAK-------SRHFSGEPQQLQTRLLRYNDLCERAL 644
Query: 618 FLADLAEESEEAYNFIISEMKQTHKSAAAMKTS 650
L + A S+E+YN ++ + A + TS
Sbjct: 645 KLNEEASLSQESYNIAFLAIEGAIGNCAGINTS 677
>At3g06250 unknown protein
Length = 764
Score = 320 bits (819), Expect = 2e-87
Identities = 194/619 (31%), Positives = 312/619 (50%), Gaps = 61/619 (9%)
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRS---SKPKRAV----LVCCNEG-QHKVKISRT 83
G+EF S + +FY ++A+ GF VR+ SK ++ VC EG QH
Sbjct: 193 GLEFNSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSKEGFQHPS----- 247
Query: 84 EEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRC 143
R GC A + + R + W++ N DHNH + K + M
Sbjct: 248 -----------------RMGCGAYMRIKRQDSGG-WIVDRLNKDHNHDLEPGKKNAGM-- 287
Query: 144 HKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQA 203
KK+ T V + D N D N+I + R +
Sbjct: 288 -KKI-----------------TDDVTGGLDSVDLIELN-DLSNHISSTRENTIGKEWYPV 328
Query: 204 VFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYS 263
+ DY + KQ E+ FFYAI+ D + +++FW D+RSR + FGD + FDT+Y+ YS
Sbjct: 329 LLDYFQSKQAEDMGFFYAIELDSNGSCMSIFWADSRSRFACSQFGDAVVFDTSYRKGDYS 388
Query: 264 MPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKA 323
+PFA F+G N+H Q +L G AL+ DES+ +F WLF+T L+AM G++P S++ DQDL ++
Sbjct: 389 VPFATFIGFNHHRQPVLLGGALVADESKEAFSWLFQTWLRAMSGRRPRSMVADQDLPIQQ 448
Query: 324 ALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRI 383
A+A+VFP + HR W I K E L ++ FK + ++C+ S + F+ W +
Sbjct: 449 AVAQVFPGTHHRFSAWQIRSKERENLRSFPNE---FKYEYEKCLYQSQTTVEFDTMWSSL 505
Query: 384 MVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQE 443
+ +Y L++N WL+ +Y+ RE W+P Y R++FF G++ + + F+ + +++ T+L+E
Sbjct: 506 VNKYGLRDNMWLREIYEKREKWVPAYLRASFFGGIHV---DGTFDPFYGTSLNSLTSLRE 562
Query: 444 FVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKIN 503
F+ ++E+ +E R E ERKE++ S + L T +EE +YT IF F+ EL +
Sbjct: 563 FISRYEQGLEQRREEERKEDFNSYNLQPFLQTKEPVEEQCRRLYTLTIFRIFQSELAQSY 622
Query: 504 RFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIF 563
+ K +G + V C + + V + A C +++E+ G+LCRHIL +F
Sbjct: 623 NYLGLKTYEEGAISRFLVRKCGNENEKHAVTFSASNLNASCSCQMFEYEGLLCRHILKVF 682
Query: 564 QAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLA 623
+ ++PS +I+ RWTK+A G D + G S LK L ++ AS +
Sbjct: 683 NLLDIRELPSRYILHRWTKNAEFGF---VRDVESGVTSQDLKALMIWSLREAASKYIEFG 739
Query: 624 EESEEAYNFIISEMKQTHK 642
S E Y M++ K
Sbjct: 740 TSSLEKYKLAYEIMREGGK 758
Score = 48.5 bits (114), Expect = 1e-05
Identities = 39/115 (33%), Positives = 57/115 (48%), Gaps = 31/115 (26%)
Query: 25 GNSKLE--IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISR 82
G+S LE +G+EF++ E+ R++YNS+A + GF VR GQ + SR
Sbjct: 22 GDSGLEPYVGLEFDTAEEARDYYNSYATRTGFKVRT---------------GQ--LYRSR 64
Query: 83 TEEIQDSTNQTKR-KCS------TIRSGCEASLIVSRGTTKSKWMIKSFNNDHNH 130
T D T ++R CS R+GC A + V R T KW++ +HNH
Sbjct: 65 T----DGTVSSRRFVCSKEGFQLNSRTGCPAFIRVQRRDT-GKWVLDQIQKEHNH 114
>At5g18960 FAR1 - like protein
Length = 788
Score = 317 bits (813), Expect = 1e-86
Identities = 193/620 (31%), Positives = 318/620 (51%), Gaps = 60/620 (9%)
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRS---SKPKRAV----LVCCNEG-QHKVKISRT 83
G+EF S + +FY ++A+ GF VR+ SK ++ VC EG QH
Sbjct: 214 GLEFGSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSREGFQHPS----- 268
Query: 84 EEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRC 143
R GC A + + R + W++ N DHNH + K
Sbjct: 269 -----------------RMGCGAYMRIKRQDSGG-WIVDRLNKDHNHDLEPGKKND---- 306
Query: 144 HKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQA 203
+ ++K ++G TG + ++ + F N N+I+ R +
Sbjct: 307 ---------AGMKKIPDDG--TGGLDSVDLIELNDFGN----NHIKKTRENRIGKEWYPL 351
Query: 204 VFDYCKRKQVENSNFFYAIQCD-EDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKY 262
+ DY + +Q E+ FFYA++ D + +++FW D+R+R + FGD + FDT+Y+ Y
Sbjct: 352 LLDYFQSRQTEDMGFFYAVELDVNNGSCMSIFWADSRARFACSQFGDSVVFDTSYRKGSY 411
Query: 263 SMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMK 322
S+PFA +G N+H Q +L GCA++ DES+ +F+WLF+T L+AM G++P SI+ DQDL ++
Sbjct: 412 SVPFATIIGFNHHRQPVLLGCAMVADESKEAFLWLFQTWLRAMSGRRPRSIVADQDLPIQ 471
Query: 323 AALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMR 382
AL +VFP + HR W I +K E L S FK + ++CI + +I F+ W
Sbjct: 472 QALVQVFPGAHHRYSAWQIREKERENLIPF---PSEFKYEYEKCIYQTQTIVEFDSVWSA 528
Query: 383 IMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQ 442
++ +Y L+++ WL+ +Y+ RE+W+P Y R++FFAG+ + +I FF + +DA T L+
Sbjct: 529 LINKYGLRDDVWLREIYEQRENWVPAYLRASFFAGIPI---NGTIEPFFGASLDALTPLR 585
Query: 443 EFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKI 502
EF+ ++E+A+E R E ERKE++ S + L T +EE +YT +F F+ EL +
Sbjct: 586 EFISRYEQALEQRREEERKEDFNSYNLQPFLQTKEPVEEQCRRLYTLTVFRIFQNELVQS 645
Query: 503 NRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVI 562
+ K +G + V C + + V + + C +++E G+LCRHIL +
Sbjct: 646 YNYLCLKTYEEGAISRFLVRKCGNESEKHAVTFSASNLNSSCSCQMFEHEGLLCRHILKV 705
Query: 563 FQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADL 622
F + ++PS +I+ RWTK+A G D + G + LK L ++ AS +
Sbjct: 706 FNLLDIRELPSRYILHRWTKNAEFGF---VRDMESGVSAQDLKALMVWSLREAASKYIEF 762
Query: 623 AEESEEAYNFIISEMKQTHK 642
S E Y M++ K
Sbjct: 763 GTSSLEKYKLAYEIMREGGK 782
Score = 49.7 bits (117), Expect = 6e-06
Identities = 42/128 (32%), Positives = 61/128 (46%), Gaps = 31/128 (24%)
Query: 20 TSDIGGNSKLE--IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHK 77
+ D G S +E +G+EF++ E+ REFYN++A + GF VR GQ
Sbjct: 32 SEDEEGGSGVEPYVGLEFDTAEEAREFYNAYAARTGFKVRT---------------GQ-- 74
Query: 78 VKISRTEEIQDSTNQTKR-KCS------TIRSGCEASLIVSRGTTKSKWMIKSFNNDHNH 130
+ SRT D T ++R CS R+GC A + V R T KW++ +HNH
Sbjct: 75 LYRSRT----DGTVSSRRFVCSKEGFQLNSRTGCTAFIRVQRRDT-GKWVLDQIQKEHNH 129
Query: 131 VMVSPKSV 138
+ SV
Sbjct: 130 ELGGEGSV 137
>At4g38170 hypothetical protein
Length = 531
Score = 315 bits (806), Expect = 8e-86
Identities = 162/445 (36%), Positives = 259/445 (57%), Gaps = 9/445 (2%)
Query: 204 VFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNK-Y 262
V +Y KR+Q+EN F YAI+ D N+FW D RL+Y FGD + FDTTY+ K Y
Sbjct: 7 VLNYLKRRQLENPGFLYAIEDD----CGNVFWADPTCRLNYTYFGDTLVFDTTYRRGKRY 62
Query: 263 SMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMK 322
+PFA F G N+H Q +L+GCAL+ +ESE+SF WLF+T LQAM P SI + D ++
Sbjct: 63 QVPFAAFTGFNHHGQPVLFGCALILNESESSFAWLFQTWLQAMSAPPPPSITVEPDRLIQ 122
Query: 323 AALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMR 382
A+++VF ++R R I ++ EKLAH++ F+ + C+ + + FE W
Sbjct: 123 VAVSRVFSQTRLRFSQPLIFEETEEKLAHVFQAHPTFESEFINCVTETETAAEFEASWDS 182
Query: 383 IMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQ 442
I+ Y +++N+WLQ +Y R+ W+ ++ R TF+ ++T + S +N+FF FVDASTT+Q
Sbjct: 183 IVRRYYMEDNDWLQSIYNARQQWVRVFIRDTFYGELSTNEGSSILNSFFQGFVDASTTMQ 242
Query: 443 EFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKI 502
+ ++EKA++S E E K +YE+ + + ++ T S +E+ AAS+YTR F KF+ E +
Sbjct: 243 MLIKQYEKAIDSWREKELKADYEATNSTPVMKTPSPMEKQAASLYTRAAFIKFQEEFVET 302
Query: 503 NRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVI 562
+ I G Y+V+ + TV D A C +++E+ GI+CRHIL +
Sbjct: 303 LAIPANIISDSGTHTTYRVAKFGEVHKGHTVSFDSLEVKANCSCQMFEYSGIICRHILAV 362
Query: 563 FQAKGVVQIPSHFIMERWTKDAN-RGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLAD 621
F AK V+ +PS +++ RWTK+A RG E+ ++ ++S L +++EA+ +
Sbjct: 363 FSAKNVLALPSRYLLRRWTKEAKIRGTEEQPEFSNGCQESLNLCF---NSLRQEATKYVE 419
Query: 622 LAEESEEAYNFIISEMKQTHKSAAA 646
+S + Y + + + K AA
Sbjct: 420 EGAKSIQIYKVAMDALDEAAKKVAA 444
>At2g32250 Mutator-like transposase
Length = 684
Score = 313 bits (802), Expect = 2e-85
Identities = 185/633 (29%), Positives = 320/633 (50%), Gaps = 64/633 (10%)
Query: 27 SKLEIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR-------AVLVCCNEGQHKVK 79
S+L GM+FES E FY +A+ GFG+ +++S+ + + C G + K
Sbjct: 36 SELRNGMDFESKEAAYYFYREYARSVGFGITIKASRRSKRSGKFIDVKIACSRFGTKREK 95
Query: 80 ISRTEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVS 139
+T R C ++GC+A L + R + KW+I +F +HNH +
Sbjct: 96 ---------ATAINPRSCP--KTGCKAGLHMKRKEDE-KWVIYNFVKEHNHEI------- 136
Query: 140 YMRCHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVG 199
C V+ + + P G +A I+ + L+
Sbjct: 137 ---CPDDFYVSVRG-------KNKPAGALA------------------IKKGLQLALEEE 168
Query: 200 DAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKT 259
D + + ++ Q + FFYA+ D D R+ N+FW+DA+++ Y F DV+ FDT Y
Sbjct: 169 DLKLLLEHFMEMQDKQPGFFYAVDFDSDKRVRNVFWLDAKAKHDYCSFSDVVLFDTFYVR 228
Query: 260 NKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDL 319
N Y +PFAPF+G+++H Q +L GCAL+ + SE+++ WLF+T L+A+GG+ P +ITDQD
Sbjct: 229 NGYRIPFAPFIGVSHHRQYVLLGCALIGEVSESTYSWLFRTWLKAVGGQAPGVMITDQDK 288
Query: 320 AMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEE 379
+ + +VFP+ RH CLW ++ K E L + F C+ S + + FE
Sbjct: 289 LLSDIVVEVFPDVRHIFCLWSVLSKISEMLNPFVSQDDGFMESFGNCVASSWTDEHFERR 348
Query: 380 WMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAST 439
W ++ ++ L ENEW+Q L++ R+ W+P Y AG++ +RS SI + FD ++++
Sbjct: 349 WSNMIGKFELNENEWVQLLFRDRKKWVPHYFHGICLAGLSGPERSGSIASHFDKYMNSEA 408
Query: 440 TLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGEL 499
T ++F + K ++ R + E K++ E + K L + E+ + IYT F KF+ E+
Sbjct: 409 TFKDFFELYMKFLQYRCDVEAKDDLEYQSKQPTLRSSLAFEKQLSLIYTDAAFKKFQAEV 468
Query: 500 EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHI 559
+ K R DG ++++ + ++ R F V ++ + A C L+E+ G LC+H
Sbjct: 469 PGVVSCQLQKEREDGTTAIFRIED-FEERQNFFVALNNELLDACCSCHLFEYQGFLCKHA 527
Query: 560 LVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTL--KILRRVHVQREAS 617
+++ Q+ V ++PS +I++RW+K N N D +K T+ ++ R + R
Sbjct: 528 ILVLQSADVSRVPSQYILKRWSKKGN-------NKEDKNDKCATIDNRMARFDDLCRRFV 580
Query: 618 FLADLAEESEEAYNFIISEMKQTHKSAAAMKTS 650
L +A S+EA + +++T K +M S
Sbjct: 581 KLGVVASLSDEACKTALKLLEETVKHCVSMDNS 613
>At1g52520 F6D8.26
Length = 703
Score = 292 bits (747), Expect = 5e-79
Identities = 180/574 (31%), Positives = 294/574 (50%), Gaps = 39/574 (6%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR-------AVLVCCNEGQHKVKISRT 83
+GMEFES + +YN +A + GF VRV++S KR AVL C ++G ++
Sbjct: 87 VGMEFESYDDAYNYYNCYASEVGFRVRVKNSWFKRRSKEKYGAVLCCSSQGFKRI----- 141
Query: 84 EEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRC 143
N R R+GC A +I R +W + DHNH++ S R
Sbjct: 142 -------NDVNRVRKETRTGCPA-MIRMRQVDSKRWRVVEVTLDHNHLLGCKLYKSVKRK 193
Query: 144 HKKMS--VAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDA 201
K +S V+ ++ + + G N ++ N+ N + NL GD+
Sbjct: 194 RKCVSSPVSDAKTIKLYRACVVDNGS-----NVNPNSTLNKKFQNSTGSPDLLNLKRGDS 248
Query: 202 QAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNK 261
A+++Y R Q+ N NFFY + +++ ++ N+FW DA S++S FGDVI D++Y + K
Sbjct: 249 AAIYNYFCRMQLTNPNFFYLMDVNDEGQLRNVFWADAFSKVSCSYFGDVIFIDSSYISGK 308
Query: 262 YSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAM 321
+ +P F G+N+H ++ L C L E+ S+ WL K L M + P +I+TD+ +
Sbjct: 309 FEIPLVTFTGVNHHGKTTLLSCGFLAGETMESYHWLLKVWLSVM-KRSPQTIVTDRCKPL 367
Query: 322 KAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWM 381
+AA+++VFP S R L HI++K PEKL + H ++ + + + + FE W
Sbjct: 368 EAAISQVFPRSHQRFSLTHIMRKIPEKLGGL-HNYDAVRKAFTKAVYETLKVVEFEAAWG 426
Query: 382 RIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTL 441
++ + + ENEWL+ LY+ R W P+Y + TFFAG+ E++ FF+ +V T L
Sbjct: 427 FMVHNFGVIENEWLRSLYEERAKWAPVYLKDTFFAGIAAAHPGETLKPFFERYVHKQTPL 486
Query: 442 QEFVLKFEKAVESRLEAERKENYESRH-KSRILSTGSKLEEHAASIYTRNIFAKFKGELE 500
+EF+ K+E A++ + E + ES+ + L T E + IYTR++F KF+ E+E
Sbjct: 487 KEFLDKYELALQKKHREETLSDIESQTLNTAELKTKCSFETQLSRIYTRDMFKKFQIEVE 546
Query: 501 KI-NRFTRHKIRRDGPKYVYQV-------SNCYDARDTFTVDIDLDSQIAKCGYELYEFM 552
++ + F+ ++ DGP ++ V S+ + RD F V + +C + F
Sbjct: 547 EMYSCFSTTQVHVDGPFVIFLVKERVRGESSRREIRD-FEVLYNRSVGEVRCICSCFNFY 605
Query: 553 GILCRHILVIFQAKGVVQIPSHFIMERWTKDANR 586
G LCRH L + GV +IP +I+ RW KD R
Sbjct: 606 GYLCRHALCVLNFNGVEEIPLRYILPRWRKDYKR 639
>At4g19990 putative protein
Length = 672
Score = 281 bits (719), Expect = 9e-76
Identities = 196/693 (28%), Positives = 335/693 (48%), Gaps = 99/693 (14%)
Query: 26 NSKLEIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR-------AVLVCCNEGQHKV 78
N +++ G EFES E+ EFY +A GF +++S+ R A VC G K
Sbjct: 18 NLEIDEGREFESKEEAFEFYKEYANSVGFTTIIKASRRSRMTGKFIDAKFVCTRYGSKKE 77
Query: 79 KISR---TE--EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMV 133
I T+ I + + + S+ ++ C+A L V R +W+++S +HNH +
Sbjct: 78 DIDTGLGTDGFNIPQARKRGRINRSSSKTDCKAFLHVKR-RQDGRWVVRSLVKEHNHEIF 136
Query: 134 SPKSVSYMRCHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRR 193
+ ++ S ++ + +EK AI ++ V+
Sbjct: 137 TGQADSLRE------LSGRRKLEKLN---------GAI----------------VKEVKS 165
Query: 194 KNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITF 253
+ L+ GD + + NFF +Q + N+FWVDA+ R Y F DV++
Sbjct: 166 RKLEDGDVERLL-----------NFFTDMQS-----LRNIFWVDAKGRFDYTCFSDVVSI 209
Query: 254 DTTYKTNKYSMPFAPFVGLNNHSQSILYGCALL-QDESEASFVWLFKTLLQAMGGKKPIS 312
DTT+ N+Y +P F G+N+H Q +L G LL DES++ FVWLF+ L+AM G +P
Sbjct: 210 DTTFIKNEYKLPLVAFTGVNHHGQFLLLGFGLLLTDESKSGFVWLFRAWLKAMHGCRPRV 269
Query: 313 IITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHS 372
I+T D +K A+ +VFP SRH +W + + PEKL H+ + ++ I GS
Sbjct: 270 ILTKHDQMLKEAVLEVFPSSRHCFYMWDTLGQMPEKLGHVIRLEKKLVDEINDAIYGSCQ 329
Query: 373 IQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFD 432
+ FE+ W ++ +++++N WLQ LY+ RE W+P+Y + AGM T QRS+S+N+ D
Sbjct: 330 SEDFEKNWWEVVDRFHMRDNVWLQSLYEDREYWVPVYMKDVSLAGMCTAQRSDSVNSGLD 389
Query: 433 SFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIF 492
++ TT + F+ +++K ++ R E E K E+ +K L + S + A +YTR +F
Sbjct: 390 KYIQRKTTFKAFLEQYKKMIQERYEEEEKSEIETLYKQPGLKSPSPFGKQMAEVYTREMF 449
Query: 493 AKFKGE-LEKINRFTRHKIRRDG-PKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYE 550
KF+ E L + + + DG K ++V + Y+ +F V + +S C
Sbjct: 450 KKFQVEVLGGVACHPKKESEEDGVNKRTFRVQD-YEQNRSFVVVWNSESSEVVCS----- 503
Query: 551 FMGILCRHILVIFQAKGVVQIPSHFIMERWTKDA-NRGLEDTYNDNDLGEKSDTLK--IL 607
CR +F+ KG + IPS ++++RWTKDA +R + ++ + K+ K L
Sbjct: 504 -----CR----LFELKGELSIPSQYVLKRWTKDAKSREVMESDQTDVESTKAQRYKDLCL 554
Query: 608 RRVHVQREASFLADLAEESEEAYNFIISEMKQTHKS-------AAAMKTSEGVVL----L 656
R + + EAS SEE+YN +++ + + + ++ SE V +
Sbjct: 555 RSLKLSEEASL-------SEESYNAVVNVLNEALRKWENKSNLIQNLEESESVTAQDLPI 607
Query: 657 ESSEKNVNQVCSSEQVSEPPQLTIGNPHVSQTK 689
+ N + + V++ Q+ + VS K
Sbjct: 608 HEEQNNTYDMNKDDNVADTGQVLFSSQLVSWNK 640
>At1g10240 unknown protein
Length = 680
Score = 275 bits (703), Expect = 7e-74
Identities = 185/648 (28%), Positives = 308/648 (46%), Gaps = 57/648 (8%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPK--------RAVLVCCNEGQHKVKISR 82
+G F + + EFY++FAK+ GF +R ++ K R VC G +K
Sbjct: 50 LGQIFLTHDTAYEFYSTFAKRCGFSIRRHRTEGKDGVGKGLTRRYFVCHRAGNTPIKT-- 107
Query: 83 TEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTT--KSKWMIKSFNNDHNHVMVSPKSVSY 140
+ + Q R+ S R GC+A L +S+ T ++W + F N HNH ++ P V +
Sbjct: 108 ---LSEGKPQRNRRSS--RCGCQAYLRISKLTELGSTEWRVTGFANHHNHELLEPNQVRF 162
Query: 141 MRCHKKMSVAAKSLVEKFEEEGIPTGKVAAIFN------DGDSTFTNRDCWNYIRNVRRK 194
+ ++ +S A KS + F + GI ++ + G FT +D N +++ K
Sbjct: 163 LPAYRSISDADKSRILMFSKTGISVQQMMRLLELEKCVEPGFLPFTEKDVRNLLQSF--K 220
Query: 195 NLDVGDAQAVF-DYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITF 253
LD D F C+ + ++ NF + D + ++ N+ W A S SY+LFGD + F
Sbjct: 221 KLDPEDENIDFLRMCQSIKEKDPNFKFEFTLDANDKLENIAWSYASSIQSYELFGDAVVF 280
Query: 254 DTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISI 313
DTT++ + MP +VG+NN+ +GC LL+DE+ S+ W + M GK P +I
Sbjct: 281 DTTHRLSAVEMPLGIWVGVNNYGVPCFFGCVLLRDENLRSWSWALQAFTGFMNGKAPQTI 340
Query: 314 ITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKL-AHIYHKQSIFKRDMKRCIRGSHS 372
+TD ++ +K A+A P ++H LC+W ++ KFP A + + + +K + R + S
Sbjct: 341 LTDHNMCLKEAIAGEMPATKHALCIWMVVGKFPSWFNAGLGERYNDWKAEFYR-LYHLES 399
Query: 373 IQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFD 432
++ FE W ++ + L N + LY R W Y RS F AGM T RS++INAF
Sbjct: 400 VEEFELGWRDMVNSFGLHTNRHINNLYASRSLWSLPYLRSHFLAGMTLTGRSKAINAFIQ 459
Query: 433 SFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIF 492
F+ A T L FV + V+ + +A ++ + ++ L TG+ +E HAAS+ T F
Sbjct: 460 RFLSAQTRLAHFVEQVAVVVDFKDQATEQQTMQQNLQNISLKTGAPMESHAASVLTPFAF 519
Query: 493 AKFKGELEKINRF----------TRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIA 542
+K + +L + RH + DG + VY V I
Sbjct: 520 SKLQEQLVLAAHYASFQMDEGYLVRHHTKLDGGRKVYWVP---------------QEGII 564
Query: 543 KCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDN--DLGEK 600
C +L+EF G LCRH L + Q+P ++ RW + + T+ N D GE+
Sbjct: 565 SCSCQLFEFSGFLCRHALRVLSTGNCFQVPDRYLPLRWRR-ISTSFSKTFRSNAEDHGER 623
Query: 601 SDTLKILRRVHVQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMK 648
L+ L V A L +E+ + ++S +++ S+ A++
Sbjct: 624 VQLLQNLVSTLVSESAKSKERLDIATEQT-SILLSRIREQPVSSLAIR 670
>At1g80010 hypothetical protein
Length = 696
Score = 273 bits (699), Expect = 2e-73
Identities = 176/584 (30%), Positives = 294/584 (50%), Gaps = 48/584 (8%)
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRSS------KPKRAVLVCCNEGQHKVKISRTEE 85
GMEFES + FYNS+A++ GF +RV+SS K KR ++CCN K+
Sbjct: 69 GMEFESYDDAYSFYNSYARELGFAIRVKSSWTKRNSKEKRGAVLCCNCQGFKL------- 121
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHK 145
++D+ ++ K R+GC+A +I R +W + DHNH P+ + HK
Sbjct: 122 LKDAHSRRKET----RTGCQA-MIRLRLIHFDRWKVDQVKLDHNHSF-DPQRAHNSKSHK 175
Query: 146 KMSVAAKSLVEKFEEEGIPTGKVAAI-----------------FNDGDSTFTNRDCWNYI 188
K S +A S K E P +V I + G+++ + D +
Sbjct: 176 KSSSSA-SPATKTNPEPPPHVQVRTIKLYRTLALDTPPALGTSLSSGETSDLSLD---HF 231
Query: 189 RNVRRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFG 248
++ RR L G +A+ D+ + Q+ + NF Y + +D + N+FW+DAR+R +Y FG
Sbjct: 232 QSSRRLELR-GGFRALQDFFFQIQLSSPNFLYLMDLADDGSLRNVFWIDARARAAYSHFG 290
Query: 249 DVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGK 308
DV+ FDTT +N Y +P FVG+N+H +IL GC LL D+S ++VWLF+ L M G+
Sbjct: 291 DVLLFDTTCLSNAYELPLVAFVGINHHGDTILLGCGLLADQSFETYVWLFRAWLTCMLGR 350
Query: 309 KPISIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIR 368
P IT+Q AM+ A+++VFP + HRL L H++ + + + +F + R +
Sbjct: 351 PPQIFITEQCKAMRTAVSEVFPRAHHRLSLTHVLHNICQSVVQL-QDSDLFPMALNRVVY 409
Query: 369 GSHSIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESIN 428
G ++ FE W +++ + + NE ++ +++ RE W P+Y + TF AG T
Sbjct: 410 GCLKVEEFETAWEEMIIRFGMTNNETIRDMFQDRELWAPVYLKDTFLAGALTFPLGNVAA 469
Query: 429 AF-FDSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIY 487
F F +V +T+L+EF+ +E ++ + E + ES L T E A ++
Sbjct: 470 PFIFSGYVHENTSLREFLEGYESFLDKKYTREALCDSESLKLIPKLKTTHPYESQMAKVF 529
Query: 488 TRNIFAKFKGELEKINR-FTRHKIRRDG--PKYVYQVSNCYDARDTFTV-DIDLDSQI-A 542
T IF +F+ E+ ++ F ++ +G YV + RD + + +Q+
Sbjct: 530 TMEIFRRFQDEVSAMSSCFGVTQVHSNGSASSYVVKEREGDKVRDFEVIYETSAAAQVRC 589
Query: 543 KCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANR 586
C + F G CRH+L++ G+ ++P +I++RW KD R
Sbjct: 590 FCVCGGFSFNGYQCRHVLLLLSHNGLQEVPPQYILQRWRKDVKR 633
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 225 bits (573), Expect = 8e-59
Identities = 149/598 (24%), Positives = 278/598 (45%), Gaps = 44/598 (7%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVR-VRSSKPK-----RAVLVCCNEGQHKVKISRTE 84
+G F + ++ E+Y++FA+K+GF +R RS++ + R VC G ++ +
Sbjct: 72 VGQIFTTDDEAFEYYSTFARKSGFSIRKARSTESQNLGVYRRDFVCYRSGFNQPR----- 126
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTK--SKWMIKSFNNDHNHVMVSPKSVSYMR 142
+ + + R+ ++R GC+ L +++ S W + F+N HNH ++ V +
Sbjct: 127 --KKANVEHPRERKSVRCGCDGKLYLTKEVVDGVSHWYVSQFSNVHNHELLEDDQVRLLP 184
Query: 143 CHKKMSVAAKSLVEKFEEEGIPTGKVAAIFN------DGDSTFTNRDCWNYIRNVRRKNL 196
++K+ + + + + G P ++ + G F +D N++R ++
Sbjct: 185 AYRKIQQSDQERILLLSKAGFPVNRIVKLLELEKGVVSGQLPFIEKDVRNFVRACKKSVQ 244
Query: 197 D---------VGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLF 247
+ D + + CK + +F Y DE+ ++ N+ W S Y LF
Sbjct: 245 ENDAFMTEKRESDTLELLECCKGLAERDMDFVYDCTSDENQKVENIAWAYGDSVRGYSLF 304
Query: 248 GDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGG 307
GDV+ FDT+Y++ Y + F G++N+ +++L GC LLQDES SF W +T ++ M G
Sbjct: 305 GDVVVFDTSYRSVPYGLLLGVFFGIDNNGKAMLLGCVLLQDESCRSFTWALQTFVRFMRG 364
Query: 308 KKPISIITDQDLAMKAALAKVFPESRHRLCLWHIIKK----FPEKLAHIYHKQSIFKRDM 363
+ P +I+TD D +K A+ + P + H + + HI+ K F + L Y + F+
Sbjct: 365 RHPQTILTDIDTGLKDAIGREMPNTNHVVFMSHIVSKLASWFSQTLGSHYEE---FRAGF 421
Query: 364 KRCIRGSHSIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQR 423
R ++ FE++W ++ + L + LY R SW+P R F A T++
Sbjct: 422 DMLCRAG-NVDEFEQQWDLLVTRFGLVPDRHAALLYSCRASWLPCCIREHFVAQTMTSEF 480
Query: 424 SESINAFFDSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHA 483
+ SI++F VD +T +Q +L E A++ A + R L T +E+HA
Sbjct: 481 NLSIDSFLKRVVDGATCMQ--LLLEESALQVSAAASLAKQILPRFTYPSLKTCMPMEDHA 538
Query: 484 ASIYTRNIFAKFKGELEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAK 543
I T F+ + E+ ++ ++ +GP V+ V + +++ +
Sbjct: 539 RGILTPYAFSVLQNEMVLSVQYAVAEM-ANGPFIVHHYKK---MEGECCVIWNPENEEIQ 594
Query: 544 CGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKS 601
C + +E GILCRH L + K IP + + RW +++ + N +G+ S
Sbjct: 595 CSCKEFEHSGILCRHTLRVLTVKNCFHIPEQYFLLRWRQESPHVATENQNGQGIGDDS 652
>At3g45340 putative protein
Length = 693
Score = 66.6 bits (161), Expect = 5e-11
Identities = 74/330 (22%), Positives = 123/330 (36%), Gaps = 43/330 (13%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDST 90
+G EF S E V + N +K+ FGV+ I+ T
Sbjct: 190 VGQEFRSKEAVWDLINRASKEEVFGVKT---------------------------IKSDT 222
Query: 91 NQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVA 150
+ +CS GC+ L V++ + W +K H P++ Y + +
Sbjct: 223 GRLMLECSQASKGCDWYLRVTKTSETDFWCVKKHTQIHK-CSRCPEATRYEKQKGTPRLV 281
Query: 151 AKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDA--------Q 202
A L E + + T K I + DC +Y +R K V D +
Sbjct: 282 ADVLHEDYPGL-LNTPKPKQIMSIVKGRL-GLDC-SYSTTLRGKKKHVSDVSGSPETSYK 338
Query: 203 AVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKY 262
+F Y + N ++ D D R LF S ++ VI D T+ Y
Sbjct: 339 LLFAYLHILRSVNHGTITEVELDADDRFKFLFIALGASIEGFKAMRKVIVIDATFLKTIY 398
Query: 263 S--MPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLA 320
+ A N+H I +G ++ E+ AS+ W F+ L + + I D +
Sbjct: 399 GGMLVIATAQDPNHHHYPIAFG--VIDSENHASWNWFFRKLNSIIPDDPELVFIRDIHGS 456
Query: 321 MKAALAKVFPESRHRLCLWHIIKKFPEKLA 350
+ +A VFP++ H C+WH+ + + LA
Sbjct: 457 IIKGVADVFPKASHGHCVWHLSQNIKKMLA 486
>At2g23500 Mutator-like transposase
Length = 784
Score = 59.3 bits (142), Expect = 8e-09
Identities = 67/332 (20%), Positives = 129/332 (38%), Gaps = 42/332 (12%)
Query: 29 LEIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVC----CNEGQHKVKISRTE 84
L + EF + + E + A N FG ++ S +R VL C C+ IS T+
Sbjct: 199 LTLRQEFPNKAALHEVVDRAAFANSFGYVIKKSDKERYVLKCAKESCSWRLRASNISTTD 258
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCH 144
C+ + G + L +G + ++ + +DH + S +
Sbjct: 259 IFSIRRYNKMHSCTRLSKG-SSRLRKRKGNPQ---LVAALLHDHFPGQLETPVPSII--- 311
Query: 145 KKMSVAAKSLVEKFEEEGIPTGKVAAIFN---DGDSTFTNRDCWNYIRNVRRKNLDVGDA 201
M + L K GK AI++ + ++ + +C+ Y+
Sbjct: 312 --MELVQTKLGVKVSYSTALRGKYHAIYDLKGSPEESYKDINCYLYM------------- 356
Query: 202 QAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNK 261
K+V + Y ++ DE+ + +F S +++ V+ D T+ N
Sbjct: 357 --------LKKVNDGTVTY-LKLDENDKFQYVFVALGASIEGFRVMRKVLIVDATHLKNG 407
Query: 262 YS--MPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDL 319
Y + FA N H I + A+L E++AS+ W F+ L + + +TD++
Sbjct: 408 YGGVLVFASAQDPNRHHYIIAF--AVLDGENDASWEWFFEKLKTVVPDTSELVFMTDRNA 465
Query: 320 AMKAALAKVFPESRHRLCLWHIIKKFPEKLAH 351
++ A+ V+ + H C+WH+ + H
Sbjct: 466 SLIKAIRNVYTAAHHGYCIWHLSQNVKGHATH 497
>At3g07500 hypothetical protein
Length = 214
Score = 56.2 bits (134), Expect = 6e-08
Identities = 36/110 (32%), Positives = 57/110 (51%), Gaps = 16/110 (14%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRV----RSSKPKRAV---LVCCNEGQHKVKISRT 83
IGMEFES E + FY+++A GF +RV RS + V LVC EG + + R+
Sbjct: 43 IGMEFESEEAAKSFYDNYATCMGFVMRVDAFRRSMRDGTVVWRRLVCNKEGFRRSRPRRS 102
Query: 84 EEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMV 133
E +++ + R GC+A ++V R W++ F +HNH ++
Sbjct: 103 E--------SRKPRAITREGCKALIVVKR-EKSGTWLVTKFEKEHNHPLL 143
>At3g32060 hypothetical protein
Length = 487
Score = 53.9 bits (128), Expect = 3e-07
Identities = 34/143 (23%), Positives = 66/143 (45%), Gaps = 5/143 (3%)
Query: 211 KQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYS--MPFAP 268
K+V + Y ++ DE+ + +F S +++ V+ D T+ N Y + FA
Sbjct: 228 KKVNDGTVTY-LKLDENDKFQYVFVALGASIEGFRVMRKVLIVDATHLKNGYGGVLVFAS 286
Query: 269 FVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKV 328
N H I + A+L E++AS+ W F+ L + + +TD++ ++ A+ V
Sbjct: 287 AQDPNRHHYIIAF--AVLDGENDASWEWFFEKLKTVVPDTSELVFMTDRNASLIKAIRNV 344
Query: 329 FPESRHRLCLWHIIKKFPEKLAH 351
+ + H C+WH+ + H
Sbjct: 345 YTAAHHGYCIWHLSQNVKGHATH 367
>At1g44840 hypothetical protein
Length = 926
Score = 53.9 bits (128), Expect = 3e-07
Identities = 27/127 (21%), Positives = 60/127 (46%)
Query: 222 IQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILY 281
++ + R + +F S ++ ++ D T+ KY G + + Q
Sbjct: 546 VEDESKERFLYMFLAFGASIAGFRHLRRILVVDGTHLKGKYKGVLLTSSGQDANFQVYPL 605
Query: 282 GCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFPESRHRLCLWHI 341
G A++ E++ S+ W F L + + K ++I++D+ ++ A+ +VFP++ H C+ H+
Sbjct: 606 GFAVVDSENDESWTWFFTKLERIIADSKTLTILSDRHSSILVAVKRVFPQANHGACIIHL 665
Query: 342 IKKFPEK 348
+ K
Sbjct: 666 CRNIQTK 672
>At3g59470 unknown protein
Length = 251
Score = 52.8 bits (125), Expect = 7e-07
Identities = 35/125 (28%), Positives = 57/125 (45%), Gaps = 18/125 (14%)
Query: 16 SILDTSDIGGNSKLEIGMEFESIEKVREFYNSFAKKNGFGVRV-------RSSKPKRAVL 68
++ D ++ G+ +G EFES FYN++A K GF +RV P L
Sbjct: 58 AVADMTEAQGDEPY-VGQEFESEAAAHGFYNAYATKVGFVIRVSKLSRSRHDGSPIGRQL 116
Query: 69 VCCNEGQHKVKISRTEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDH 128
VC EG + ++ R+ + R GC+A +++ R KW+I F +H
Sbjct: 117 VCNKEGY---------RLPSKRDKVIRQRAETRVGCKAMILI-RKENSGKWVITKFVKEH 166
Query: 129 NHVMV 133
NH ++
Sbjct: 167 NHSLM 171
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,299,609
Number of Sequences: 26719
Number of extensions: 690246
Number of successful extensions: 2350
Number of sequences better than 10.0: 133
Number of HSP's better than 10.0 without gapping: 83
Number of HSP's successfully gapped in prelim test: 50
Number of HSP's that attempted gapping in prelim test: 2138
Number of HSP's gapped (non-prelim): 195
length of query: 742
length of database: 11,318,596
effective HSP length: 107
effective length of query: 635
effective length of database: 8,459,663
effective search space: 5371886005
effective search space used: 5371886005
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0091b.9


