

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.14
(487 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
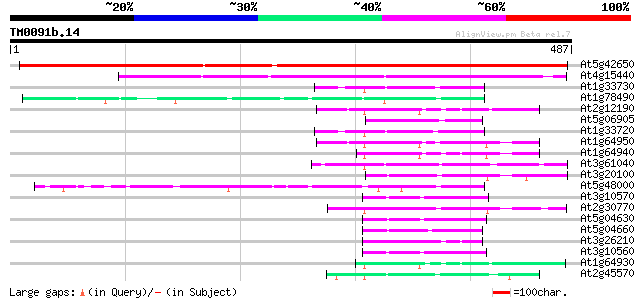
Score E
Sequences producing significant alignments: (bits) Value
At5g42650 allene oxide synthase (emb|CAA73184.1) 473 e-133
At4g15440 hydroperoxide lyase (HPL1) 273 2e-73
At1g33730 cytochrome P450, putative 59 5e-09
At1g78490 unknown protein 56 4e-08
At2g12190 pseudogene 55 9e-08
At5g06905 cytochrome P450-like protein 55 1e-07
At1g33720 cytochrome P450, putative 54 1e-07
At1g64950 unknown protein 52 7e-07
At1g64940 hypothetical protein 52 7e-07
At3g61040 cytochrome P450 monooxygenase-like protein 51 1e-06
At3g20100 cytochrome P450, putative 51 1e-06
At5g48000 cytochrome P450-like protein 51 2e-06
At3g10570 putative cytochrome P450 51 2e-06
At2g30770 putative cytochrome P450 51 2e-06
At5g04630 cytochrome P450 - like protein 50 2e-06
At5g04660 cytochrom P450 - like protein 50 3e-06
At3g26210 cytochrome P450 like protein 50 4e-06
At3g10560 putative cytochrome P450 49 5e-06
At1g64930 hypothetical protein 49 5e-06
At2g45570 cytochrome P450 (CYP76C2) 49 6e-06
>At5g42650 allene oxide synthase (emb|CAA73184.1)
Length = 518
Score = 473 bits (1217), Expect = e-133
Identities = 236/477 (49%), Positives = 323/477 (67%), Gaps = 5/477 (1%)
Query: 9 TKNSSSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIRTNMPPG 68
T+ S +LP++ IPG+YGLP G + DR DYFY+QG ++FF +R+ KYNSTV R NMPPG
Sbjct: 45 TRTGSKDLPIRNIPGNYGLPIVGPIKDRWDYFYDQGAEEFFKSRIRKYNSTVYRVNMPPG 104
Query: 69 PGISSDSKVIALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPSH 128
I+ + +V+ALLD SFP+LFD KVEK+D+ GT+MPST TGGYR ++ D +EP H
Sbjct: 105 AFIAENPQVVALLDGKSFPVLFDVDKVEKKDLFTGTYMPSTELTGGYRILSYLDPSEPKH 164
Query: 129 KLLKTFFMQVLSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFNFLFL 188
+ LK +L S N P + T S+ F L+ +L+ K GKA F S FNFL
Sbjct: 165 EKLKNLLFFLLKSSRNRIFPEFQATYSELFDSLEKELSLK-GKADFGGSSDGTAFNFLAR 223
Query: 189 LLTDKHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPAWTVKW 248
+P++T + PGL+T W+ L PL ++GLP++ E+ LI T P VK
Sbjct: 224 AFYGTNPADTKLKADAPGLITKWVLFNLHPLLSIGLPRVI---EEPLIHTFSLPPALVKS 280
Query: 249 SYKKLCEGFASAGASLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIGLA 308
Y++L E F + +L EA+++GI R+EA HNL+F FN +GG+ FP ++K IG A
Sbjct: 281 DYQRLYEFFLESAGEILVEADKLGISREEATHNLLFATCFNTWGGMKILFPNMVKRIGRA 340
Query: 309 GPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVV 368
G ++H +LAEEIR+V++ + GE+++ A++KM LTKSVVYE LR EP V QY +A++D+V
Sbjct: 341 GHQVHNRLAEEIRSVIKSNGGELTMGAIEKMELTKSVVYECLRFEPPVTAQYGRAKKDLV 400
Query: 369 VESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVG-EGEKLLKYVYWSNGRE 427
+ESHDAAF++K GEM++GYQP AT+DPK+F+ +EFV RFVG EGEKLL++V WSNG E
Sbjct: 401 IESHDAAFKVKAGEMLYGYQPLATRDPKIFDRADEFVPERFVGEEGEKLLRHVLWSNGPE 460
Query: 428 TEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKAT 484
TE PTV NKQC K+ VVL+ RL+V+E F RYD+F E LG +V SL KA+
Sbjct: 461 TETPTVGNKQCAGKDFVVLVARLFVIEIFRRYDSFDIEVGTSPLGSSVNFSSLRKAS 517
>At4g15440 hydroperoxide lyase (HPL1)
Length = 384
Score = 273 bits (697), Expect = 2e-73
Identities = 160/392 (40%), Positives = 226/392 (56%), Gaps = 15/392 (3%)
Query: 95 VEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPSHKLLKTFFMQVLSSKHNTFLPLLRTTL 154
V+KRDVL G F PS GF GG R + DTTEP H +K F M+ L +L LR+ L
Sbjct: 4 VDKRDVLIGDFRPSLGFYGGVRVGVYLDTTEPKHAKIKGFAMETLKRSSKVWLQELRSNL 63
Query: 155 SDHFSDLDSQLAGKSGKASFNTSISAATFNFLFLLLTDKHPSETV-IGDKGPGLVTAWLG 213
+ + ++S+++ K+G AS+ + F+FL L S + I + G + WL
Sbjct: 64 NIFWGTIESEIS-KNGAASYIFPLQRCIFSFLCASLAGVDASVSPDIAENGWKTINTWLA 122
Query: 214 AQLAPLATLGLPKIFNYAEDFLIRTIPFPAWTVKWSYKKLCEGF-ASAGASLLDEAERVG 272
Q+ P A LG+ + E+ L+ T P+P+ + +YKKL +AG L E G
Sbjct: 123 LQVIPTAKLGV--VPQPLEEILLHTWPYPSLLIAGNYKKLYNFIDENAGDCLRLGQEEFG 180
Query: 273 IKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIGLAGPELHAKLAEEIRTVVRESNGEVS 332
+ RDEA NL+F+LGFNA+GG + P LI I L ++ E+R V S +++
Sbjct: 181 LTRDEAIQNLLFVLGFNAYGGFSVFLPSLIGRITGDNSGLQERIRTEVRRVCG-SGSDLN 239
Query: 333 LSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIKKGEMIFGYQPFAT 392
+++M L KSVVYE LR P VP Q+A+AR+D + SHDA FE+KKGE++ GYQP
Sbjct: 240 FKTVNEMELVKSVVYETLRFSPPVPLQFARARKDFQISSHDAVFEVKKGELLCGYQPLVM 299
Query: 393 KDPKVFENPEEFVGHRFVGE-GEKLLKYVYWSNGRETEEPTVDNKQCPAKNLVVLMCRLY 451
+D VF+ PEEF R+VGE G +LL Y+YWSNG +T P+ NKQC AK++V L L
Sbjct: 300 RDANVFDEPEEFKPDRYVGETGSELLNYLYWSNGPQTGTPSASNKQCAAKDIVTLTASLL 359
Query: 452 VVEFFLRYDTFTFEFKPVVLGPNVTIESLTKA 483
V + FLRYDT T G + +I+++ KA
Sbjct: 360 VADLFLRYDTIT--------GDSGSIKAVVKA 383
>At1g33730 cytochrome P450, putative
Length = 372
Score = 59.3 bits (142), Expect = 5e-09
Identities = 45/152 (29%), Positives = 72/152 (46%), Gaps = 14/152 (9%)
Query: 265 LDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIR 321
L + + I DE EH L+ M F T+ ++W L P+ K+ +EI
Sbjct: 156 LQQGDESEINIDEIEHLLLDM-----FLAGTDTNSSTVEWAMTELLGNPKTMTKVQDEIN 210
Query: 322 TVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQY-AKAREDVVVESHDAAFEIKK 380
V+R+ NG+V S + K+ ++V+ E R+ PA P+ KA DV + F + K
Sbjct: 211 RVIRQ-NGDVQESHISKLPYLQAVIKETFRLHPAAPFLLPRKAERDVDI----LGFHVPK 265
Query: 381 GEMIFGYQPFATKDPKVFENPEEFVGHRFVGE 412
+ +DP V+ENP +F RF+G+
Sbjct: 266 DSHVLVNVWAIGRDPNVWENPTQFEPERFLGK 297
>At1g78490 unknown protein
Length = 479
Score = 56.2 bits (134), Expect = 4e-08
Identities = 97/412 (23%), Positives = 165/412 (39%), Gaps = 42/412 (10%)
Query: 12 SSSNLPLKPIPGSYGLPFFGAMHDRHDYFYNQGRDKFFATRVEKYNSTVIRTNMPPGPGI 71
S+ P K PGS G P G D +G F R+ +Y + RTN+ +
Sbjct: 27 SNPKCPGKLPPGSMGFPIIGETLDFFKPCGVEGIPTFVKKRMIRYGP-LFRTNIFGSKTV 85
Query: 72 -SSDSKVIALL---DSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPS 127
S+D VI + ++ SF + + + V K D F+
Sbjct: 86 VSTDPDVIHQIFRQENTSFELGYPDIFV-KVFGKDNLFLKEVFI---------------- 128
Query: 128 HKLLKTFFMQVLSSK--HNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFNF 185
HK L+ MQ+L S+ T L + DH + SQ + K N ++ T
Sbjct: 129 HKYLQKITMQILGSEGLKQTMLGNMDKATRDHIRSIASQGSFNVRKEVENLVVAYMTPKL 188
Query: 186 LFLLLTDKHPSETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPAWT 245
+ L K +++ + D W + L + K E+ + +
Sbjct: 189 ISNL---KPETQSKLIDNLNAFNLDWFKSFLRLSTWKAVTKALKSREEAI--QVMKDVLM 243
Query: 246 VKWSYKKLCEGFASAGASLLDEAERVGIKRDEAEH-NLVFMLGFNAFGGLTNQFPILIKW 304
++ ++ E F + +LL+E E+ G D+ NL+F+L F G ++ + +K+
Sbjct: 244 MRKETREKQEDFLN---TLLEELEKDGSFFDQGSAINLIFLLAFALREGTSSCTALAVKF 300
Query: 305 IGLAGPELHAKLAEEIRTVV---RESNGEVSLSAL-DKMTLTKSVVYEVLRIEPAVPYQY 360
I P++ A+L E + +V ++ VS MT T V EVLR+ P +
Sbjct: 301 IS-KDPKVLAELKREHKAIVDNRKDKEAGVSWEEYRHNMTFTNMVSNEVLRLANTTPLLF 359
Query: 361 AKAREDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE 412
KA +DV ++ + I G ++ DP ++ENP EF R+ G+
Sbjct: 360 RKAVQDVEIK----GYTIPAGWIVAVAPSAVHFDPAIYENPFEFNPWRWEGK 407
>At2g12190 pseudogene
Length = 512
Score = 55.1 bits (131), Expect = 9e-08
Identities = 58/201 (28%), Positives = 84/201 (40%), Gaps = 17/201 (8%)
Query: 267 EAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIRTV 323
E E KR E +V + GG T+ ++WI + PE+ +L EEI++V
Sbjct: 287 ELELPDEKRKLNEDEIVSLCSEFLNGG-TDTTATALQWIMANLVKNPEIQKRLYEEIKSV 345
Query: 324 VRESNGEVSLSALDKMTLTKSVVYEVLRIEP----AVPYQYAKAREDVVVESHDAAFEIK 379
V E EV KM K+VV E LR P +P+ ED V+ + K
Sbjct: 346 VGEEAKEVEEEDAQKMPYLKAVVMEGLRRHPPGHFVLPH---SVTEDTVLGGYKVP---K 399
Query: 380 KGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYWSNGRETEEPTVDNKQCP 439
KG + F +DP V+E P F RF+GE E + + S G + + CP
Sbjct: 400 KGTINFMVAEIG-RDPMVWEEPMAFKPERFMGEEEAV--DITGSRGIKMMPFGAGRRICP 456
Query: 440 AKNLVVLMCRLYVVEFFLRYD 460
L +L YV ++
Sbjct: 457 GIGLAMLHLEYYVANMVREFE 477
>At5g06905 cytochrome P450-like protein
Length = 528
Score = 54.7 bits (130), Expect = 1e-07
Identities = 28/101 (27%), Positives = 55/101 (53%), Gaps = 4/101 (3%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
P++ AK+ +EI++VV +N + S L K+ ++ + E LR+ P P ++ D+ +
Sbjct: 328 PDIFAKIRDEIKSVVGTTNRLIKESDLQKLPYLQAAIKETLRLHPVGPLLRRESNTDMKI 387
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFV 410
+D +K G IF +DP +++P++F+ RF+
Sbjct: 388 NGYD----VKSGTKIFINAYGIMRDPTTYKDPDKFMPERFL 424
>At1g33720 cytochrome P450, putative
Length = 511
Score = 54.3 bits (129), Expect = 1e-07
Identities = 43/151 (28%), Positives = 67/151 (43%), Gaps = 12/151 (7%)
Query: 265 LDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIR 321
L + + I DE EH L+ M F T+ ++W L P+ K+ +EI
Sbjct: 288 LQQGDETEINIDEIEHLLLDM-----FVAGTDTNSSTVEWAMAELLGNPKTMTKVQDEIN 342
Query: 322 TVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIKKG 381
V+ + NG+ S + K+ K+VV E R+ PA P+ + E V F + K
Sbjct: 343 HVIGQ-NGDFQESDISKLPYLKAVVKETFRLHPAAPFLLQRKAETNV---EILGFTVLKD 398
Query: 382 EMIFGYQPFATKDPKVFENPEEFVGHRFVGE 412
+ +DP V+ENP F RF+G+
Sbjct: 399 SQVLVNVWAIGRDPLVWENPTHFEPERFLGK 429
>At1g64950 unknown protein
Length = 510
Score = 52.0 bits (123), Expect = 7e-07
Identities = 56/206 (27%), Positives = 85/206 (41%), Gaps = 29/206 (14%)
Query: 267 EAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIRTV 323
E E KR E +V + GG T+ ++WI + P++ +L EEI++V
Sbjct: 287 ELELPDEKRKLNEDEIVSLCSEFLNGG-TDTTATALQWIMANLVKNPDIQKRLYEEIKSV 345
Query: 324 VRESNGEVSLSALDKMTLTKSVVYEVLRIEP----AVPYQYAKAREDVVVESHDAAFEIK 379
V E EV KM ++VV E LR P +P+ ED V+ +++
Sbjct: 346 VGEEANEVEEEDAQKMPYLEAVVMEGLRRHPPGHFVLPH---SVTEDTVL----GGYKVP 398
Query: 380 KGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE-----GEKLLKYVYWSNGRETEEPTVD 434
K I +DPKV+E P F RF+ E G + +K + + GR
Sbjct: 399 KNGTINFMVAEIGRDPKVWEEPMAFKPERFMEEAVDITGSRGIKMMPFGAGR-------- 450
Query: 435 NKQCPAKNLVVLMCRLYVVEFFLRYD 460
+ CP L +L YV +D
Sbjct: 451 -RICPGIGLAMLHLEYYVANMVREFD 475
>At1g64940 hypothetical protein
Length = 511
Score = 52.0 bits (123), Expect = 7e-07
Identities = 49/171 (28%), Positives = 74/171 (42%), Gaps = 28/171 (16%)
Query: 302 IKWIG---LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEP---- 354
++WI + PE+ +L EEI+++V E EV KM K+VV E LR P
Sbjct: 322 LQWIMANLVKNPEIQRRLYEEIKSIVGEEAKEVEEQDAQKMPYLKAVVMEGLRRHPPGHF 381
Query: 355 AVPYQYAKAREDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE-- 412
+P+ ED V+ + KKG + F +DPKV+E P F RF+ E
Sbjct: 382 VLPH---SVTEDTVLGGYKVP---KKGTINFMVAEIG-RDPKVWEEPMAFKPERFMEEAV 434
Query: 413 ---GEKLLKYVYWSNGRETEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYD 460
G + +K + + GR + CP L +L YV ++
Sbjct: 435 DITGSRGIKMMPFGAGR---------RICPGIGLAMLHLEYYVANMVREFE 476
>At3g61040 cytochrome P450 monooxygenase-like protein
Length = 498
Score = 51.2 bits (121), Expect = 1e-06
Identities = 57/225 (25%), Positives = 93/225 (41%), Gaps = 17/225 (7%)
Query: 263 SLLDEAERVGIKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEE 319
SLLD A + K E + N + L + F + ++W L P++ K+ EE
Sbjct: 272 SLLDIAHK---KESELDDNNIKHLLLDLFLAGVDTSSSAVEWAMAELLRNPKMIVKVQEE 328
Query: 320 IRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVVESHDAAFEIK 379
IR V+ G V + K+ ++VV E LR+ P P+ + E V+ + F I
Sbjct: 329 IRQVIG-LKGTVQDLDIVKLPYLQAVVKESLRLHPPAPFLVPRKSESDDVQIFE--FLIP 385
Query: 380 KGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEKLLKYVYWSNGRETEEPTVDNKQCP 439
K + +DP V++NP +F RF+G G + N E + CP
Sbjct: 386 KNTQVLVNVWAIGRDPNVWKNPTQFEPERFLGRGIDVK-----GNHFELIPFGAGRRICP 440
Query: 440 AKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKAT 484
L + L + +D +E++ V+ NV + AT
Sbjct: 441 GMPLAFRIMHLVLASLLYGFD---WEYQNGVVPENVDMNEAFGAT 482
>At3g20100 cytochrome P450, putative
Length = 513
Score = 51.2 bits (121), Expect = 1e-06
Identities = 48/188 (25%), Positives = 80/188 (42%), Gaps = 27/188 (14%)
Query: 310 PELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKAREDVVV 369
P + ++ EEI +VV +S + + L K+ ++VV E LR+ P P + +E V
Sbjct: 331 PNIFVRIREEIDSVVGKSR-LIQETDLPKLPYLQAVVKEGLRLHPPTPLMVREFQEGCKV 389
Query: 370 ESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEG---------EKLLKYV 420
+ F I + +DP V+E+PEEF RF+ E+ LKY+
Sbjct: 390 KG----FYIPASTTLVVNGYAVMRDPNVWEDPEEFKPERFLASSRLMQEDEIREQALKYI 445
Query: 421 YWSNGRETEEPTVDNKQCPAKNLVVLM----CRLYVVEFFLRYDTFTFEFKPVVLGPNVT 476
+ +GR + CP N+ + + V F R + + K + G N+T
Sbjct: 446 AFGSGR---------RGCPGANVAYIFVGTAIGMMVQCFDWRINGEKVDMKEAIGGLNLT 496
Query: 477 IESLTKAT 484
+ K T
Sbjct: 497 LAHPLKCT 504
>At5g48000 cytochrome P450-like protein
Length = 477
Score = 50.8 bits (120), Expect = 2e-06
Identities = 90/406 (22%), Positives = 170/406 (41%), Gaps = 50/406 (12%)
Query: 22 PGSYGLPFFGAMHDRHDYFYNQGR---DKFFATRVEKYNSTVIRTNMPPGPGISSDSKVI 78
PGS GLP G + D+F G F R+ KY + RTN+ S++ V+
Sbjct: 37 PGSMGLPIIG---ETCDFFEPHGLYEISPFVKKRMLKYGP-LFRTNI-----FGSNTVVL 87
Query: 79 ALLDSASFPILFDNSKVEKRDVLDGTFMPSTGFTGGYRACAFQDTTEPSHKLLKTFFMQV 138
D I+F+ + E + + F F + HK +K +Q
Sbjct: 88 TEPD-----IIFEVFRQENKSFV---FSYPEAFVKPFGKENVFLKHGNIHKHVKQISLQH 139
Query: 139 LSSKHNTFLPLLRTTLSDHFSDLDSQLAGKSGKASFNTSISAATFNFLFL---LLTDKHP 195
L S+ L + + + L K+ + SF+ + + L ++++ P
Sbjct: 140 LGSE-----ALKKKMIGEIDRVTYEHLRSKANQGSFDAKEAVESVIMAHLTPKIISNLKP 194
Query: 196 -SETVIGDKGPGLVTAWLGAQLAPLATLGLPKIFNYAEDFLIRTIPFPAWTVKWSYKKLC 254
++ + D L + W + L + + K+F A + ++ I +T + + +++C
Sbjct: 195 ETQATLVDNIMALGSEWFQSPLKLTTLISIYKVF-IARRYALQVIK-DVFTRRKASREMC 252
Query: 255 EGFASAGASLLDEAERVG-IKRDEAEHNLVFMLGFNAFGGLTNQFPILIKWIGLAGPELH 313
F ++++E E+ I +E+ NL+F + A ++ + IK++ E H
Sbjct: 253 GDFLD---TMVEEGEKEDVIFNEESAINLIFAILVVAKESTSSVTSLAIKFLA----ENH 305
Query: 314 AKLAE---EIRTVVRESNGEVSLSALDK----MTLTKSVVYEVLRIEPAVPYQYAKARED 366
LAE E +++ NG+ + + ++ MT T V+ E LR+ P Y KA D
Sbjct: 306 KALAELKREHAAILQNRNGKGAGVSWEEYRHQMTFTNMVINETLRMANMAPIMYRKAVND 365
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGE 412
V ++ + I G ++ P + ++ENP EF R+ G+
Sbjct: 366 VEIK----GYTIPAGWIVAVIPPAVHFNDAIYENPLEFNPWRWEGK 407
>At3g10570 putative cytochrome P450
Length = 513
Score = 50.8 bits (120), Expect = 2e-06
Identities = 31/109 (28%), Positives = 56/109 (50%), Gaps = 5/109 (4%)
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKARED 366
+ PE+ ++L +EI++ V + EV +DKM ++VV E+LR P Y
Sbjct: 333 IVNPEIQSRLYDEIKSTVGDR--EVEEKDVDKMVFLQAVVKEILRKHPPT---YFTLTHS 387
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGEK 415
V + A +++ G + Y P +DPK++ +P++F RF+ E+
Sbjct: 388 VTEPTTVAGYDVPVGINVEFYLPGINEDPKLWSDPKKFNPDRFISGKEE 436
>At2g30770 putative cytochrome P450
Length = 497
Score = 50.8 bits (120), Expect = 2e-06
Identities = 53/216 (24%), Positives = 92/216 (42%), Gaps = 28/216 (12%)
Query: 277 EAEHNLVFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIRTVVRESNGEVSL 333
+ + N + + + F G T+ L++W + P+ KL +EIR+ +R +
Sbjct: 282 QVQRNDIKFMILDMFIGGTSTTSTLLEWTMTELIRSPKSMKKLQDEIRSTIRPHGSYIKE 341
Query: 334 SALDKMTLTKSVVYEVLRIEPAVPYQYAK-AREDVVVESHDAAFEIKKGEMIFGYQPFAT 392
++ M K+V+ EVLR+ P++P + EDV V+ + I G +
Sbjct: 342 KEVENMKYLKAVIKEVLRLHPSLPMILPRLLSEDVKVK----GYNIAAGTEVIINAWAIQ 397
Query: 393 KDPKVF-ENPEEFVGHRFVGEG----EKLLKYVYWSNGRETEEPTVDNKQCPAKNLVVLM 447
+D ++ + EEF R + G K L Y+ + +GR + CP NL + +
Sbjct: 398 RDTAIWGPDAEEFKPERHLDSGLDYHGKNLNYIPFGSGR---------RICPGINLALGL 448
Query: 448 CRLYVVEFFLRYDTFTFEFKPVVLGPNVTIESLTKA 483
+ V R+D V GPN LT+A
Sbjct: 449 AEVTVANLVGRFDW------RVEAGPNGDQPDLTEA 478
>At5g04630 cytochrome P450 - like protein
Length = 509
Score = 50.4 bits (119), Expect = 2e-06
Identities = 31/108 (28%), Positives = 56/108 (51%), Gaps = 4/108 (3%)
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKARED 366
++ P++ ++L +EI++ V + V L+KM ++ V E+LR P Y
Sbjct: 328 ISNPKIQSRLYDEIKSTVGDDR-TVEEKDLNKMVFLQAFVKELLRRHPPT---YFTLTHG 383
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGE 414
V ++ A ++I G + Y P ++DPK++ PE+F RF+ GE
Sbjct: 384 VTEPTNLAGYDIPVGANVEFYLPGISEDPKIWSKPEKFDPDRFITGGE 431
>At5g04660 cytochrom P450 - like protein
Length = 512
Score = 50.1 bits (118), Expect = 3e-06
Identities = 30/104 (28%), Positives = 56/104 (53%), Gaps = 4/104 (3%)
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKARED 366
+A PE+ ++L +EI++ V + V +DKM ++ V E+LR P + A
Sbjct: 331 IANPEIQSRLYDEIKSTVGDDR-RVDEKDVDKMVFLQAFVKELLRKHPPTYFSLTHA--- 386
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFV 410
V+ + A ++I G + Y P ++DP+++ NP++F RF+
Sbjct: 387 VMETTTLAGYDIPAGVNVEVYLPGISEDPRIWNNPKKFDPDRFM 430
>At3g26210 cytochrome P450 like protein
Length = 501
Score = 49.7 bits (117), Expect = 4e-06
Identities = 29/105 (27%), Positives = 54/105 (50%), Gaps = 5/105 (4%)
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAK-ARE 365
+ P + K+ +E+RTV+ E ++ L+++ K V+ E R+ PA P + A
Sbjct: 320 IRNPRVMKKVQDEVRTVLGEKRDRITEQDLNQLNYFKLVIKETFRLHPAAPLLLPREAMA 379
Query: 366 DVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFV 410
+ ++ +D +K +++ +DP ++ENPEEF RFV
Sbjct: 380 KIKIQGYDIP---EKTQIMVNVYAIG-RDPDLWENPEEFKPERFV 420
>At3g10560 putative cytochrome P450
Length = 514
Score = 49.3 bits (116), Expect = 5e-06
Identities = 33/108 (30%), Positives = 55/108 (50%), Gaps = 5/108 (4%)
Query: 307 LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEPAVPYQYAKARED 366
+A PE+ ++L +EI++ V + V +DKM L ++VV E+LR P Y
Sbjct: 333 IANPEIQSRLYDEIKSTVGDR--AVDERDVDKMVLLQAVVKEILRRHPPT---YFTLSHG 387
Query: 367 VVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEGE 414
V + + + I G I Y P ++DPK++ P++F RF+ E
Sbjct: 388 VTEPTTLSGYNIPVGVNIEFYLPGISEDPKIWSEPKKFDPDRFLSGRE 435
>At1g64930 hypothetical protein
Length = 511
Score = 49.3 bits (116), Expect = 5e-06
Identities = 53/189 (28%), Positives = 75/189 (39%), Gaps = 16/189 (8%)
Query: 301 LIKWIG---LAGPELHAKLAEEIRTVVRESNGEVSLSALDKMTLTKSVVYEVLRIEP--- 354
+++WI + E+ +L EEI VV E V KM K+VV E LR P
Sbjct: 319 VLQWIMANLVKNQEIQERLYEEITNVVGEEAKVVEEKDTQKMPYLKAVVMEALRRHPPGN 378
Query: 355 -AVPYQYAKAREDVVVESHDAAFEIKKGEMIFGYQPFATKDPKVFENPEEFVGHRFVGEG 413
+P+ ED V+ + KKG + F +DPKV+E P F RF+GE
Sbjct: 379 TVLPHSVT---EDTVLGGYKVP---KKGTINFLVAEIG-RDPKVWEEPMAFKPERFMGEE 431
Query: 414 EKLLKYVYWSNGRETEEPTVDNKQCPAKNLVVLMCRLYVVEFFLRYDTFTFEFKPVVLGP 473
E + + S G + + CP L +L YV + E V L
Sbjct: 432 EAV--DITGSRGIKMMPFGAGRRICPGIGLAMLHLEYYVANMVREFQWKEVEGHEVDLTE 489
Query: 474 NVTIESLTK 482
V + K
Sbjct: 490 KVEFTVIMK 498
>At2g45570 cytochrome P450 (CYP76C2)
Length = 512
Score = 48.9 bits (115), Expect = 6e-06
Identities = 54/193 (27%), Positives = 76/193 (38%), Gaps = 20/193 (10%)
Query: 276 DEAEHNL--VFMLGFNAFGGLTNQFPILIKWIG---LAGPELHAKLAEEIRTVVRESNGE 330
DEAE N + L + FG T+ ++W L PE K EI V+ + G
Sbjct: 293 DEAELNTNDIVHLLLDLFGAGTDTNSSTVEWAMAELLRNPETMVKAQAEIDCVIGQK-GV 351
Query: 331 VSLSALDKMTLTKSVVYEVLRIEPAVPYQYA-KAREDVVVESHDAAFEIKKGEMIFGYQP 389
V S + + ++VV E R+ PA P KA DV V F + K +F
Sbjct: 352 VEESDISALPYLQAVVKETFRLHPAAPLLVPRKAESDVEV----LGFMVPKDTQVFVNVW 407
Query: 390 FATKDPKVFENPEEFVGHRFVGEGEKLLKYVYWSNGRETEEPT--VDNKQCPAKNLVVLM 447
+DP V+EN F RF+G+ L GR+ E + CP L V
Sbjct: 408 AIGRDPNVWENSSRFKPERFLGKDIDL-------RGRDYELTPFGAGRRICPGLPLAVKT 460
Query: 448 CRLYVVEFFLRYD 460
L + +D
Sbjct: 461 VPLMLASLLYSFD 473
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,239,895
Number of Sequences: 26719
Number of extensions: 499029
Number of successful extensions: 1306
Number of sequences better than 10.0: 175
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 129
Number of HSP's that attempted gapping in prelim test: 1176
Number of HSP's gapped (non-prelim): 201
length of query: 487
length of database: 11,318,596
effective HSP length: 103
effective length of query: 384
effective length of database: 8,566,539
effective search space: 3289550976
effective search space used: 3289550976
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0091b.14


